Abstract
Objective
To evaluate the ability of large language models (LLMs) to perform structured information extraction and guideline-based clinical inferences from radiology and pathology reports in real-world oncology.
Methods
We constructed a Question Answering (Q&A) benchmark dataset using 3650 radiological and 588 pathological reports from 1632 patients. The tasks included direct extraction of genomic and histological findings, as well as clinical reasoning tasks, such as Response Evaluation Criteria in Solid Tumors (RECIST)-based tumor response classification and American Joint Committee on Cancer (AJCC)-based tumor-node-metastasis (TNM) staging. We compared the performance of the Gemma family of open-source LLMs (Gemma 4B, a lightweight 4-billion parameter model, and Gemma 12B, a larger 12-billion parameter model) with and without structured reasoning prompts designed according to clinical guidelines.
Results
The 12B model achieved high performance in direct extraction tasks from pathology reports, with F1-score ranging from 92.6 to 93.3 across genomic and histological variables. Furthermore, when guided by structured reasoning prompts, it also showed substantial improvements in reasoning tasks, achieving an F1-score of 81.5 (95% CI: 79.8–83.3) for tumor response, 74.3 (95% CI: 70.8–77.8) for T-stage, 87.1 (95% CI: 85.1–89.0) for N-stage, and 90.8 (95% CI: 89.1–92.2) for M-stage. In contrast, the 4B model showed inconsistent performance and was sometimes degraded under reasoning prompts.
Conclusion
This study shows that LLMs can perform complex guideline-based clinical reasoning using real-world radiology reports. By combining the RECIST/AJCC criteria with structured prompts, we demonstrated how LLMs can move beyond surface-level extraction to support nuanced inference in oncology, with implications for future clinical applications.
Keywords
Introduction
Electronic health records (EHRs) of patients with cancer contain essential information for clinical decision-making, including diagnostic results, genomic tests, pathology reports, treatment plans, and therapeutic responses. 1 However, effectively leveraging this information requires transforming the unstructured text into structured data, a process that relies heavily on the manual efforts of domain experts. In practice, clinicians reportedly spend >90 min/day reviewing clinical documents to understand treatment progression and make informed decisions.2–4
In addition to this time burden, prior work has highlighted broader usability issues. For example, Olakotan et al. conducted a scoping review of 28 studies and concluded that poor EHR usability (such as complex navigation, task-switching, and fragmented information) substantially increases documentation burden and disrupts clinical workflows. 5
A major source of inefficiency lies in the unstructured and non-standardized nature of EHRs. While clinicians record nuanced symptoms, complex diagnoses, and therapeutic contexts in natural language, existing EHR systems have a limited capacity to structurally encode such detailed information. 6 Furthermore, institutional heterogeneity in documentation formats 7 and inter-physician variability in writing styles8,9 make it difficult to consistently store and utilize equivalent information. This lack of standardization not only impedes data utilization but has also been linked to poorer clinical outcomes; key clinical variables are often missing in oncology EHRs and are significantly associated with adverse prognoses.10,11
Recent advances in large language models (LLMs) have shown promise in addressing these challenges.12–14 Johnson et al. demonstrated that Gemma 2 achieved an F1-score of 95.2% for identifying colorectal cancer histopathological diagnoses from pathology reports in the Veterans Health Administration. 15 Huang et al. evaluated ChatGPT-3.5 on more than 1000 lung cancer pathology reports and showed that prompt engineering enabled the model to extract pathological classifications with 89% overall accuracy. 16 Kim et al. compared GPT-3.5 and GPT-4 for extracting demographics, medical history, and test results from case reports, finding GPT-4 highly accurate for sex (95%) and GPT-3.5 better for BMI (78% vs. 57%), with task-specific prompts correcting GPT-4's errors. 17
Beyond information extraction, when integration with techniques such as retrieval-augmented generation (RAG), 18 even smaller-scale LLMs can achieve high performance in document-level information extraction tasks. 19 For instance, Wada et al. demonstrated that RAG markedly improved the performance of a locally deployed Llama 3.2 model for radiology contrast media consultation, eliminating hallucinations (0% vs. 8%) and raising clinical accuracy and safety scores while preserving the privacy advantages of on-premise deployment. 20 These advances suggest new possibilities for understanding complex clinical documentation and inferring patient status and tasks that are challenging using traditional rule-based or simple classification systems.
Additionally, the use of chain-of-thought-based reasoning prompts has been shown to improve prediction accuracy and interpretability by guiding models to follow human-like clinical reasoning steps.
21
However, most prior applications have remained limited to relatively simple information extraction tasks, such as identifying explicit variables within a report. In oncology practice, critical details are often expressed in heterogeneous or implicit ways, and clinically meaningful interpretation requires alignment with guideline-based criteria. Because such inferences directly inform treatment decisions, advancing from extraction to reasoning-oriented approaches is essential. To address this gap, the present study aimed to evaluate whether LLMs can effectively perform information retrieval and clinical reasoning using complex real-world clinical documents of patients with cancer, such as radiology and pathology reports. The evaluation focused on two key aspects—whether LLMs can accurately interpret unstructured clinical text and extract relevant information and whether they can synthesize dispersed, non-explicit data within the document to produce structured outputs. In particular, we analyzed whether LLMs could perform complex reasoning tasks, such as tumor response classification and cancer staging, by providing prompts that incorporated standard clinical criteria, including the Response Evaluation Criteria in Solid Tumors, version 1.1 (RECIST v1.1)
22
and the
Methods
Study design and data collection
This retrospective cohort study utilized EHRs from Ajou University Hospital. We collected 3650 radiology reports from 1632 patients with lung cancer and 588 pathology reports from 208 patients diagnosed between 2012 and 2022. Clinical documents were manually curated and reviewed to extract structured labels, including radiological impressions (tumor response and staging) and pathological findings (
This study was approved by the Institutional Review Board (IRB) of the Ajou University Hospital (AJOUIRB-MDB-2022-249). Furthermore, the requirement for informed consent from all participants was waived by the IRB because of the retrospective nature of this study. All procedures were performed in accordance with the principles of the Declaration of Helsinki. The study followed the STROBE reporting guideline. 24
Question Answering (Q&A) Dataset Construction
Based on the collected data, we designed two tasks—information extraction and clinical reasoning—to evaluate the ability of the LLM to retrieve and reason with clinical content within unstructured documents. These tasks were constructed using the Q&A benchmark dataset. Table 1 summarizes the detailed structure of each task, including the number of questions of each type.
Q&A dataset.

Each clinical document could contain multiple distinct data elements, and thus a single report could generate several questions. For instance, one pathology report might yield separate questions on EGFR mutation status, EGFR mutation subtype, and PD-L1 assay results. Accordingly, the question counts in Table 1 represent not only the number of documents but also the number of extractable labels per document. A random subset of 10 patients was used solely for prompt design and refinement. As this study aimed to evaluate the models in a zero-shot setting under guideline-based instructions, no fine-tuning, train/validation split, or cross-validation was performed.
Annotation process
All clinical documents were independently annotated by two researchers with prior experience in clinical document data management. Because the task involved extracting explicitly documented test results and radiologists’ impressions from the reports, annotation discrepancies were rare. When disagreements occurred, they were easily resolved through simple review. Inter-rater reliability was very high, with a Cohen's kappa of 0.95, indicating almost perfect agreement.
Information extraction dataset
Pathology reports contain complex medical terminology related to genomic findings and histological assessments; therefore, extraction of the information is particularly challenging. This task was designed to evaluate the ability of the LLM to recognize and extract critical structured information, including
Clinical reasoning dataset
To assess whether the LLM could infer clinically relevant findings not explicitly stated in the text, we defined three reasoning tasks. These tasks require contextual understanding and reasoning based on clinical guidelines rather than simple keyword matching.
Tumor response: In total, 2133 radiology reports containing radiologist impressions of tumor size changes, necrosis, or metastatic spread were selected. The LLM was tasked with inferring the tumor response, classified as partial response (PR), stable disease (SD), or progressive disease (PD), based on these descriptions. T-stage: In total, 579 radiology reports with explicit descriptions of primary tumor size, morphology, pleural invasion, or anatomical extent were used to assess the ability of the LLM to infer the T-stage. Reports lacking sufficient detail for reliable classification were excluded. The LLM was tasked with classifying the primary tumor as T1–T4 according to the AJCC 8th edition guidelines. N-stage: Using 1424 computed tomography (CT) or positron emission tomography (PET) radiology reports that included descriptions of lymph node involvement and anatomical location, we evaluated the ability of the LLM to infer the N-stage. M-stage: In total, 962 reports with clearly stated findings on distant metastasis (e.g., brain, liver, bone), meaning that both the primary tumor and metastatic sites were explicitly documented in the text to provide sufficient grounds for inference, were used to assess the ability of the LLM to infer the M-stage.
Unlike simple keyword-based information retrieval, this task assesses the model's ability to comprehensively understand information described in clinical documents and apply clinical standards, such as RECIST and AJCC, to make informed clinical decisions. This represents a core competency that LLMs must possess to serve not merely as document search tools but also as effective clinical decision support systems. The overall flow of data collection and Q&A dataset construction is illustrated in Figure 1.

Question and answer (Q&A) dataset construction workflow. Overview of data collection and Q&A dataset construction from the radiology and pathology reports.
Prompt Design
Information extraction from pathology reports was performed by providing the LLM with both the original report and a prompt with explicit instructions on which clinical information to extract. The model then selects an appropriate output based on the predefined answer format. The structured information extraction process is illustrated in Figure 2.
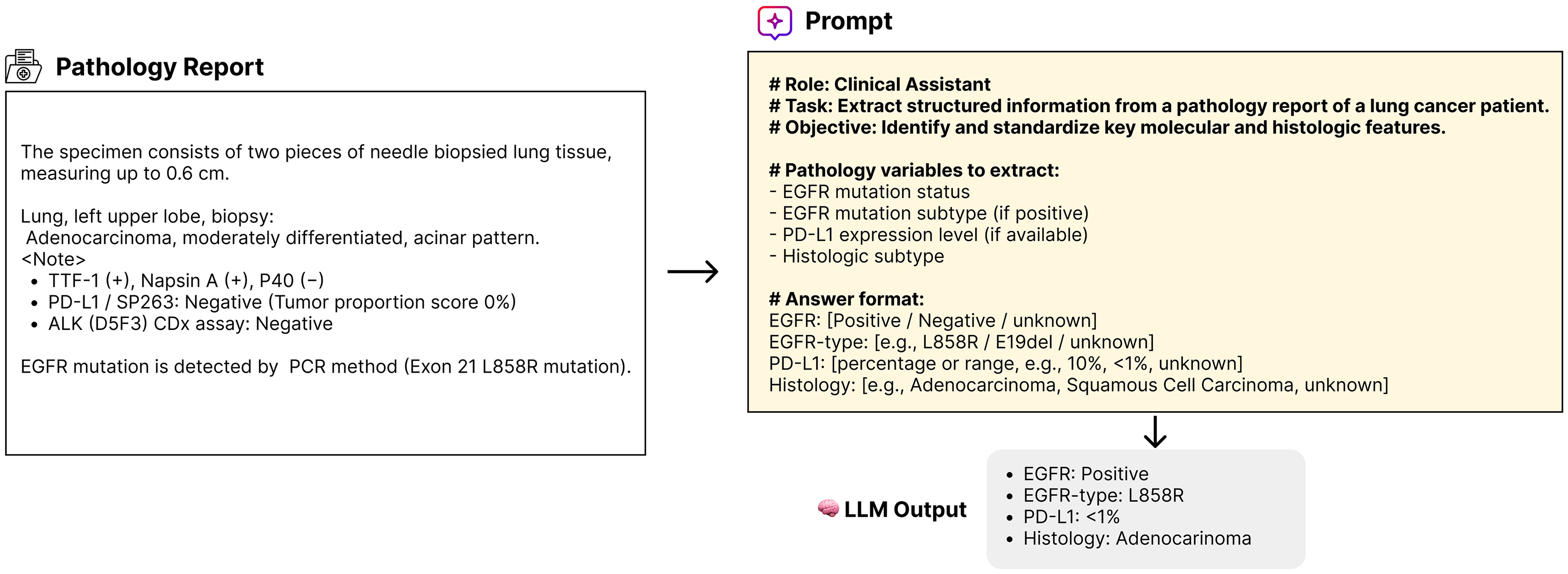
An example of information extraction. Example of extracting
In contrast, clinical reasoning tasks require more than simple information retrieval; they require an assessment of whether the model can comprehend established clinical criteria (e.g., RECIST and AJCC) and make complex, guideline-based decisions accordingly. Domain-specific reasoning prompts are designed to evaluate this capability. These prompts follow a step-by-step structure, guiding the model first to identify key clinical findings within the document and then synthesize them to reach a conclusion based on the relevant clinical standard.
To ensure the prompt's validity and strict adherence to clinical guidelines, the reasoning prompts were developed through an iterative refinement process using data from 10 patients reserved solely for this purpose. This development involved continuously checking LLM outputs against AJCC and RECIST criteria. When the LLM failed to accurately apply a specific step of the guideline, the prompt was explicitly modified and re-instructed at that stage to guarantee consistency and clinical soundness. For example, in the case of N staging, the reasoning prompted the model to (i) determine whether lymph node metastasis was present, (ii) identify the anatomical location of the involved lymph nodes, and (iii) classify the N-stage (N0 to N3) accordingly. For tumor response classification, the prompt included a brief summary of the RECIST v1.1 criteria. For cancer staging, it provided a simplified description of the AJCC 8th edition guidelines. To enable a direct performance comparison, we constructed control prompts that omitted the reasoning structure and instead posed simple direct questions (e.g., “What is the N-stage of this patient?”). The full content and logic of our reasoning prompts are illustrated in Figure 3. The exact prompts used for each task are listed in Supplementary Table 2.

An example of prompt-based clinical reasoning. The examples show how reasoning prompts the large language models (LLMs) to infer the tumor response and N-stage from radiology reports.
For all tasks, we used the Gemma 3 25 family of open-source LLMs, including a lightweight 4-billion parameter variant (4B) and a larger 12-billion parameter variant (12B). Both are decoder-only Transformer architectures; the 4B model employs multi-query attention for efficiency, whereas the 12B model adopts standard multi-head attention to enhance reasoning capacity. Detailed model configurations are provided in the Model configuration and experimental setup section.
Evaluation protocol
Model configuration and experimental setup
The objective of this study was not to compare different LLM architectures but to quantitatively assess how well a single LLM architecture performs both information extraction and guideline-based clinical reasoning using medical documents. We also aimed to analyze the impact of the prompt-reasoning design on model performance. We selected two versions of the Gemma 3 model family with different parameter sizes: the 4B and 12B variants.
To ensure architectural consistency, we adopted the Gemma family of open models, which are decoder-only Transformers trained on trillions of tokens with a 256k vocabulary, rotary positional embeddings, RMSNorm, and GeGLU activations. The 4B variant employs multi-query attention optimized for efficiency, whereas the 12B variant uses standard multi-head attention to improve representational capacity, particularly for complex reasoning tasks.
Implementation environment
All experiments were implemented using the HuggingFace Transformers library (v4.52.4) with PyTorch (v2.6.0 + cu118) and executed on NVIDIA RTX 4090 GPUs (24 GB VRAM) under CUDA 11.8. Each input consisted of a prompt–document pair, and the output was limited to a maximum of 128 tokens. For reproducibility, random seeds were fixed to 42 across all experiments.
Evaluation metrics and statistical analysis
For information extraction tasks, performance was evaluated based on accuracy, measured as an exact match with the ground truth labels. For clinical reasoning tasks, including tumor response classification and N/M staging, performance was assessed using F1-score, precision, and recall. Bootstrap resampling with 1000 iterations was conducted, following a widely adopted practical standard in biomedical statistics that balances computational efficiency and estimate stability,26,27 to assess the statistical significance of performance differences with and without reasoning prompts, as well as between long and short document groups. To account for multiple comparisons across tasks, p-values were further adjusted using the Benjamini–Hochberg procedure to control the false discovery rate (FDR, α=0.05).
Results
Dataset overview
For the information extraction task, we constructed Q&A pairs targeting key variables in the pathology reports, including
Characteristics of the QA benchmark dataset.

Abbreviations: EGFR: epidermal growth factor receptor; PD-L1: programmed death-ligand 1; PR: partial response; SD: stable disease; PD: progressive disease; T: tumor; N: node; M: metastasis; ex20ins: exon 20 insertion; 19del: exon 19 deletion.
Other mutations include variants such as G719X, G719C, T790M, S768I, L861Q.
Information extraction results
The Gemma 4B model demonstrated high performance in extracting structured information from pathology reports:
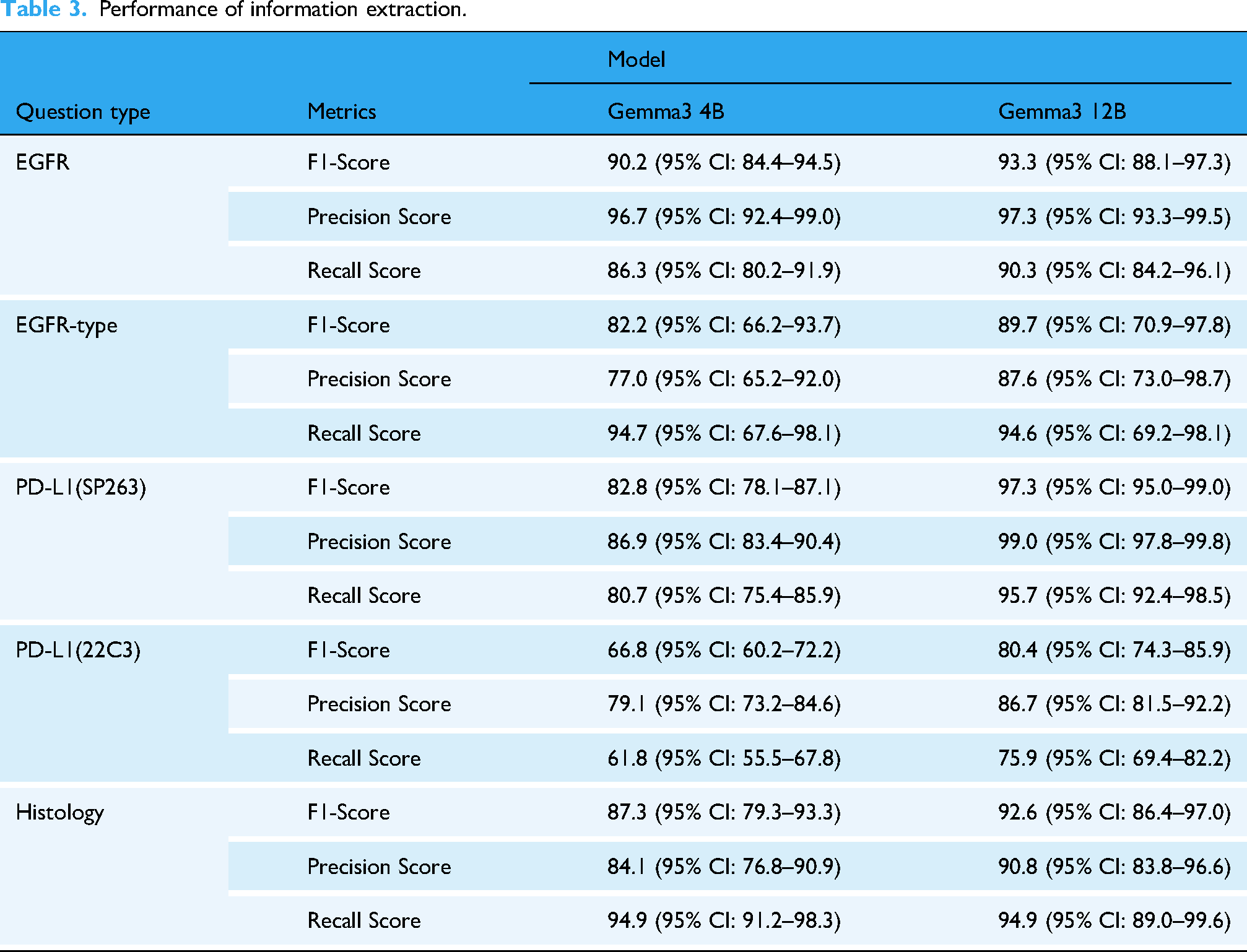
In contrast, the Gemma 12B model achieved improved f1-score for PD-L1 expression (f1-score 97.3% for SP263 and f1-score 80.4% for 22C3), demonstrating better disambiguation between the assays (Table 3). These results suggest that increasing the number of model parameters can enhance the performance of complex biomedical text extraction. The detailed class-wise error distributions are further illustrated in the confusion matrices provided in Supplementary Figure 1.
Performance of information extraction.
Clinical reasoning results
We evaluated the accuracy of the LLMs in inferring clinical judgments based on radiology report impressions. Each model (4B and 12B) was tested both with and without reasoning prompts to compare performance (Figure 4).

Performance comparison by model and prompt. F1, precision, and recall scores for reasoning tasks were compared between the 4B versus 12B large language models (LLMs) and between with and without reasoning.
Tumor response inference
In the tumor response classification, the 4B model achieved an F1-score of 73.5 (95% CI: 71.3–75.2) without prompts and 75.2 (95% CI: 73.2–77.1) with reasoning prompts, showing marginal improvement. However, the 12B model showed a significant gain; its F1-score was 69.4 (95% CI: 66.9–70.9) without prompts, which improved to 81.5 (95% CI: 79.8–83.3) with reasoning prompts.
Staging inference
For T staging, the 4B model achieved an F1-score of 42.3 (95% CI: 37.6–46.4) without reasoning prompts and 64.0 (95% CI: 59.7–67.5) with prompts, while the 12B model improved from 41.7 (95% CI: 38.1–45.5) to 74.3 (95% CI: 70.8–77.8) with prompts. Misclassifications were most frequent in borderline cases where tumor size was near the cut-off between T3 and T4 or when invasion into adjacent structures was ambiguously described. For N staging, the F1-score of the 4B model improved from 54.5 (95% CI: 52.6–56.2) to 77.3 (95% CI: 75.5–79.5) with the addition of reasoning prompts. The 12B model showed a similar trend, reaching a score of 87.1 (95% CI: 85.1–89.0) with prompts. Performance improvements were particularly notable for fine-grained distinctions between N1 and N2, as well as between N2 and N3. In M staging, the 4B model exhibited decreased performance when reasoning prompts were used; the F1-score dropped from 71.9 (95% CI: 69.6–74.4) to 53.6 (95% CI: 50.7–57.1). In contrast, the 12B model improved from 74.4 (95% CI: 71.8–77.3) to 90.8 (95% CI: 89.1–92.2) with reasoning prompts. These results indicate that reasoning prompts are more beneficial in larger models but may hinder performance in smaller models.
Document length analysis
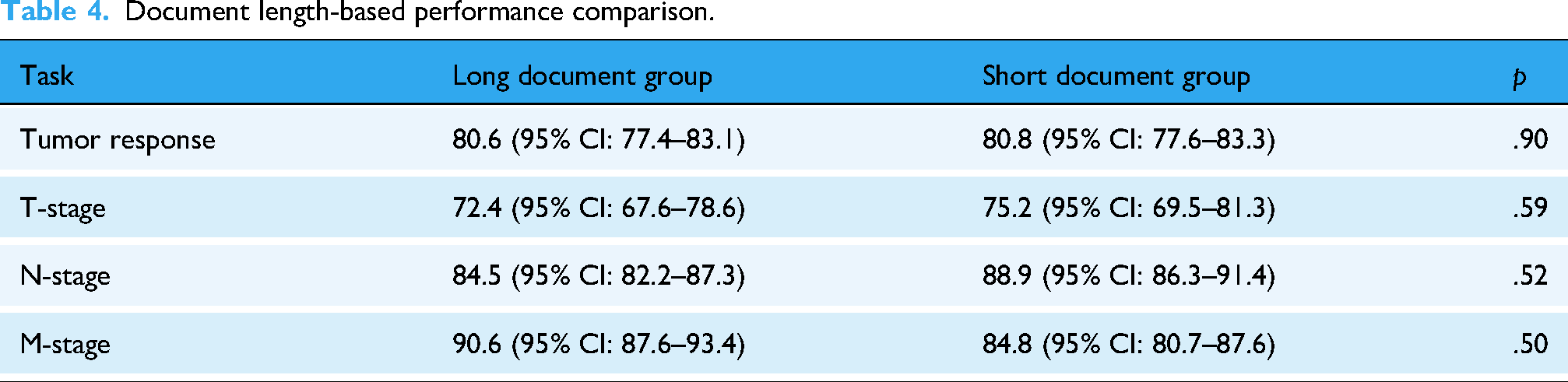
We examined whether the length of radiology reports influenced model performance. For each Q&A dataset, reports were divided into long and short document groups based on the median word count. Comparative analysis across tumor response, T-stage, N-stage, and M-stage based on F1-scores showed no statistically significant differences between the two groups (all
Document length-based performance comparison.

Effect of reasoning prompts on model performance
The application of reasoning prompts led to a statistically significant improvement in performance across all tasks in the 12B model (

Confusion matrices for clinical reasoning tasks. Confusion matrices for tumor response, T-stage, N-stage, and M-stage predictions across models and prompt settings.
Computational benchmarking
Inference time and token usage for each setting are summarized in Table 5, showing that larger models and reasoning prompts require greater computational resources. Specifically, the 4B model was executed on a single NVIDIA RTX 4090 GPU (24 GB memory), whereas the 12B model required two RTX 4090 GPUs due to its higher memory demand.
Inference time.
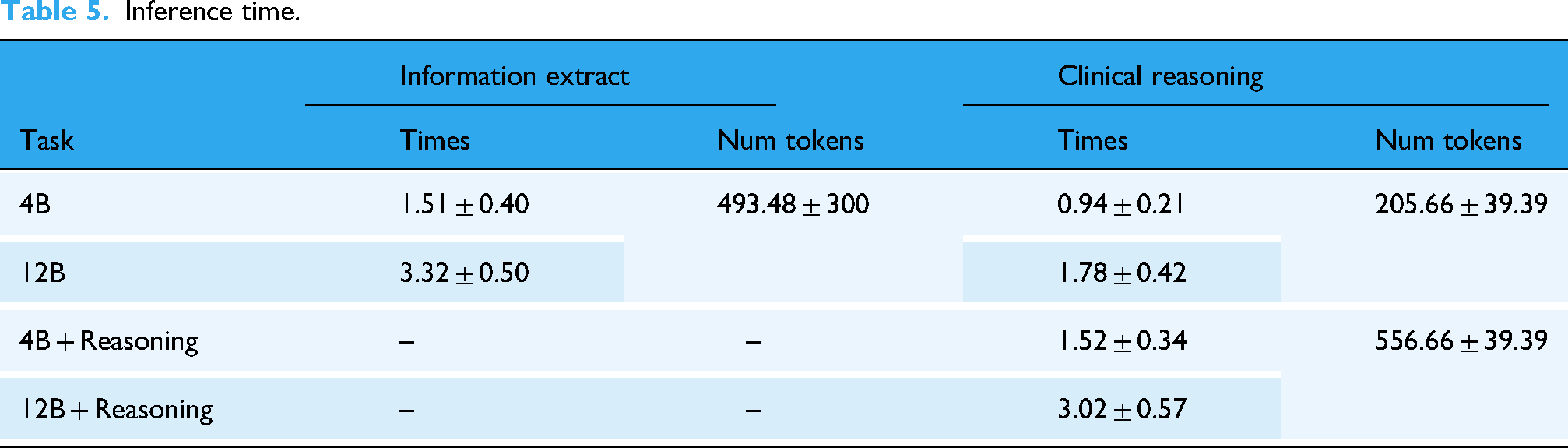
To allow a fair batch-level comparison, we benchmarked on a representative subset of documents whose token counts fell near the corpus median (550 tokens, IQR 495–604). This avoided distortion from extreme-length inputs while reflecting typical cases in our dataset. For each model and batch size (1, 8, 16, 32), we report average latency (sec/doc), peak GPU memory (GB), throughput (docs/hour), and token processing rate (tokens/sec). Detailed results are provided in Supplementary Table 1.
Interpretability in borderline cases
To further assess interpretability, we examined borderline cases where staging was inherently ambiguous, such as tumors with a maximum diameter near the T3/T4 threshold, contact versus true invasion of the mediastinum or chest wall, and lymph node involvement with equivocal laterality or distribution. Figure 6 presents representative examples where the reasoning prompt guided the LLM (Gemma 12B) to apply AJCC 8th edition criteria step by step, producing structured and guideline-based outputs.

Clinical reasoning examples for borderline TNM staging cases. Representative cases illustrating how the LLM applied AJCC guideline criteria step by step to resolve ambiguities in borderline T, N, and M staging.
In the T stage example, a 6.7 cm right upper lobe (RUL) mass contacting the mediastinum and chest wall was staged as T3. The model explicitly reasoned that contact without invasion does not fulfill T4 criteria, demonstrating adherence to guideline logic rather than keyword matching. In the N stage example, hypermetabolic lymph nodes were reported in ipsilateral (N2) and contralateral (N3) stations, including the right paratracheal and left supraclavicular regions. By mapping each nodal station to AJCC definitions, the model correctly determined the final stage as N3. In the M stage example, reports of hepatic masses and osteolytic rib lesions were directly linked to distant metastases, leading to an M1 classification.
Error analysis
Representative error cases were reviewed to clarify the main sources of misclassification.
Information extraction: Most errors came from confusion between unknown and negative labels. Reports with valid answers were sometimes marked as unknown, while unknown labels were misclassified as negative. PD-L1 errors often involved misinterpreting nearby numbers or mixing assay types, such as assigning SP263 values to 22C3 queries when only one assay was reported.
Tumor response: Subtle changes were sometimes interpreted as stable disease rather than progressive disease, while new or ambiguous findings occasionally led to progressive disease calls when the reference was stable disease or partial response.
T staging: Lesions described with borderline language were sometimes overcalled as invasive or assigned to a higher T category despite insufficient evidence. These errors reflected over-interpretation of ambiguous descriptors.
N staging: Equivocal FDG uptake in suspected nodes led to inconsistent classifications, with some cases under-called as negative and others over-called as more advanced nodal involvement.
M staging: Indeterminate findings with mild or nonspecific FDG uptake occasionally caused under-calling of potential metastasis or over-calling of physiologic uptake as metastatic disease.
Overall, most errors stemmed from borderline or ambiguous report language rather than random hallucinations, highlighting the inherent interpretive challenges of clinical text.
Discussion
Principal results
Our study demonstrates that even a medium-sized 12B model, when guided by structured prompts aligned with RECIST and AJCC criteria, maintained robust performance, achieving F1 scores of 81.5 (95% CI, 79.8–83.3) for tumor response, 74.3 (95% CI, 70.8–77.8) for T-stage, 87.1 (95% CI, 85.1–89.0) for N-stage, and 90.8 (95% CI, 89.1–92.2) for M-stage, highlighting the feasibility of accurate and resource-efficient deployment in real-world oncology workflows.
Comparison with prior work
Previous studies applying LLMs to clinical text have largely focused on relatively simple tasks such as named entity recognition (NER) or straightforward information extraction, for example identifying specific entities from patient records. 28 In oncology, some studies attempted to extract metastasis status or cancer stage from pathology reports, but these were primarily based on rule-based approaches or keyword matching.29,30 More recently, LLMs have been used to infer cancer stage even when it was not explicitly mentioned in the document, relying instead on contextual cues. However, such methods often depend on the model's pretrained knowledge rather than a structured reasoning process grounded in clinical guidelines, which limits their clinical reliability. 31 Additionally, very large models such as GPT-4 and Llama-3 70B can achieve near-perfect accuracy (≥97%) for pathology report extraction, but these models pose challenges for deployment in hospitals due to privacy and computational constraints, and smaller models showed marked performance drops in the same studies. 32
Strengths and contributions
This study makes two major contributions to the literature. First, we constructed a Q&A benchmark dataset that encompassed both information extraction and clinical reasoning tasks using authentic clinical documents. Unlike prior NER-focused studies, 33 our benchmark emphasizes higher-order clinical reasoning tasks, such as tumor response classification and tumor-node-metastasis (TNM) staging, thereby providing a more realistic evaluation of the clinical potential of LLMs. Second, we proposed a prompt-based framework that incorporates standard clinical criteria as reasoning instructions. To the best of our knowledge, this is one of the first attempts to directly assess whether LLMs can understand and apply structured medical guidelines rather than relying solely on text patterns. Importantly, this framework does not merely improve raw performance but also enhances interpretability by exposing the reasoning process itself (Figure 6). By aligning model outputs with guideline-based steps, clinicians can treat the LLM's response as a form of structured explanation rather than a black-box answer, which is essential for clinical adoption
Our results showed that both the 4B and 12B models achieved high f1-score in information extraction tasks. The smaller model demonstrated sufficient performance for clearly defined variables, such as
The benefits of reasoning prompts are more pronounced for clinical reasoning tasks. The 4B model frequently misclassified borderline cases, especially between PR and PD, and struggled to integrate anatomical details for accurate N staging. In contrast, the 12B model showed improved precision and consistency when combined with reasoning prompts, particularly for the fine-grained classification of the N-stage (N1–N3). This suggests that reasoning prompts do more than provide task instructions; they serve to guide the model through a structured clinical reasoning process that mirrors human decision-making. Such guided reasoning may offer a foundation for building explainable and trustworthy artificial intelligence (AI) tools for clinical decision support systems. Furthermore, performance was consistent regardless of document length. Comparative analysis between long and short reports across tumor response and TNM staging tasks showed no statistically significant differences (all
These findings demonstrate that LLMs can effectively organize and structure information in unstructured clinical documents, such as radiology and pathology reports when guided by standardized medical criteria. However, the use of reasoning prompts and larger models also increased the inference latency (Table 5). For example, in clinical reasoning tasks, the average inference time increased from 0.94 to 1.78 s for the 4B model and from 1.52 to 3.02 s for the 12B model when reasoning prompts were applied. Although this latency is acceptable for individual patient reviews, it may present a bottleneck in real-time decision support or high-throughput clinical data processing. Therefore, the balance between accuracy and efficiency must be considered when deploying LLMs in a clinical setting. Nevertheless, the information extraction task exhibited clear efficiency gains. For instance, the 12B model processed a pathology report in approximately 3.32 s, which is several dozen times faster than the 1–3 min required for manual extraction by clinicians. 33 Thus, the proposed LLM-based pipeline has practical value in various clinical scenarios. For example, it could enable automatic extraction and structuring of tumor response or staging information immediately after a pathology or radiology report is generated, facilitating faster clinical decisions. Additionally, they can be used to retrospectively process historical free-text reports into high-quality structured datasets for clinical research. By automating repetitive documentation tasks, LLMs can significantly enhance workflow efficiency and support the development of AI-powered clinical decision tools in oncology.
Challenges and requirements for real-world implementation
In particular, clinical deployment may require near-perfect accuracy for certain endpoints. For example, biomarker extraction tasks such as PD-L1 or EGFR mutation status, which directly inform treatment eligibility, may demand ≥99% accuracy, whereas staging tasks may tolerate slightly lower thresholds if subject to clinician oversight. The regulatory landscape must also be considered for clinical deployment. Both Food and Drug Administration (FDA) and European Medicines Agency (EMA) emphasize transparency, explainability, safety, and clinical validity in AI-based clinical decision support systems.34–36 Our approach, which explicitly models reasoning steps aligned with RECIST and AJCC guidelines, contributes to transparency and clinical validity by exposing intermediate reasoning processes. Nevertheless, further work is needed to align outputs with regulatory requirements for explainability, such as generating rationales or saliency maps that can be validated against expert clinical reasoning. In addition, ensuring data privacy is critical; one feasible strategy is to deploy smaller models on-premise within hospital networks to mitigate risks of transmitting sensitive patient information.
Another challenge is the integration of such systems into existing EHR infrastructures. Hospital databases vary widely in structure and reporting conventions, making interoperability a significant barrier. Furthermore, inference scenarios differ between retrospective research, where batch processing is acceptable, and real-time clinical care, where low-latency predictions are essential. These differences imply varying GPU and resource requirements, raising scalability concerns. As summarized in Supplementary Table 1, the 4B model ran efficiently on a single 24 GB GPU with throughput up to ∼2900 documents/hour, whereas the 12B model required two GPUs and exceeded memory limits at batch size 32. This highlights practical trade-offs between accuracy, efficiency, and hardware scalability in real-world deployment. Although our study demonstrated promising performance even with smaller LLMs, future work should evaluate efficiency and scalability under both batch and real-time inference settings to assess feasibility for clinical integration. The ability to accurately structure complex medical information, such as tumor response or cancer staging, according to standards, such as the AJCC or RECIST, can also support the use of LLMs in cases such as multidisciplinary care coordination or clinical trial eligibility screening. Our results showed that LLMs achieved 80–91% concordance with the oncologists’ documented assessments, suggesting that they are already suitable for use as decision-support tools. Such systems could reduce documentation burdens and contribute to improved patient–clinician interactions.
Limitations
This study had several limitations. First, the reasoning prompts were designed specifically for lung cancer, which may restrict their applicability to other malignancies. Generalization to other cancer types would require redesigning prompts in accordance with disease-specific clinical guidelines. Alternatively, the development of a more cancer-agnostic reasoning framework could allow broader applicability without the need for repeated prompt customization. In addition, the reasoning prompts occasionally led to degraded performance in the 4B model, suggesting that further optimization of prompt design will be necessary to ensure consistent effectiveness across different model sizes. Second, the dataset was sourced from a single institution, which may have limited the diversity of document formats and vocabulary; future validation on multi-institutional cohorts with prospective study designs is needed to ensure robustness across heterogeneous clinical environments. In addition, temporal validation was not performed; all reports were evaluated collectively without separating earlier and later cases, which may limit assessment of the model's generalizability to future clinical data. Third, clinical judgments such as staging and tumor response are often made by synthesizing information across multiple reports from different time points. In contrast, this study relied on single-report inferences. Fourth, some subgroups in our dataset were relatively small (e.g., patients with squamous cell carcinoma,
Future work will focus on three main directions. First, incorporating temporal reasoning that integrates longitudinal documents, for example through graph-based modeling of patient trajectories, to more accurately reflect real-world clinical decision-making. Second, designing prompts applicable to various cancer types and verifying whether clinical reasoning can be effectively applied to LLMs across heterogeneous clinical document formats from multiple institutions. Third, evaluating the impact of using LLMs for information retrieval and extraction from clinical documents on reducing clinician workload and improving efficiency in real-world oncology practice. Fourth, expanding the evaluation of reasoning prompts through systematic studies, including testing multiple variations and examining how sensitive model performance is to the ordering and specificity of guideline-based reasoning steps. Such work will help establish more generalizable and robust principles for prompt design in clinical applications of LLMs. Finally, as our framework relied on a generation-based LLM that does not produce probabilistic outputs, we were unable to perform conventional calibration analyses. This limitation highlights the need for future work to incorporate probability-based evaluation in order to better assess prediction reliability and enhance clinical applicability.
Conclusions
In conclusion, this study presents a novel application of LLMs for structured clinical reasoning, utilizing reasoning prompts aligned with standardized guidelines, such as the AJCC and RECIST. Unlike prior studies that treated guidelines as static reference texts, 16 we explicitly modeled guideline-driven reasoning pathways. Furthermore, by employing a smaller LLM that is deployable within a hospital's internal infrastructure, we demonstrated a practical path toward secure and efficient AI systems that are compatible with clinical workflows. This finding supports the feasibility of LLM-based systems for real-world clinical decision support.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251394622 - Supplemental material for Clinical reasoning from real-world oncology reports using large language models
Supplemental material, sj-docx-1-dhj-10.1177_20552076251394622 for Clinical reasoning from real-world oncology reports using large language models by Jun Hyeong Park, Seonhwa Kim and Jaesung Heo in DIGITAL HEALTH
Footnotes
Ethical considerations
This study was approved by the Institutional Review Board (IRB) of Ajou University Hospital (AJOUIRB-MDB-2022-249). Furthermore, the requirement for informed consent from all participants was waived by the IRB due to the retrospective nature of the study. All procedures followed the principles of the Declaration of Helsinki.
Consent to participate
The requirement for informed consent from all participants was waived by the IRB due to the retrospective nature of the study.
Author contributions
Conceptualization: Jun Hyeong Park, Jaesung Heo
Data curation: Jun Hyeong Park, Seonhwa Kim
Formal analysis: Jun Hyeong Park
Investigation: Jun Hyeong Park
Methodology: Jun Hyeong Park, Jaesung Heo
Writing–original draft: Jun Hyeong Park
Writing–review and editing: Jun Hyeong Park, Jaesung Heo.
Supervision: Jaesung Heo
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by grants from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant numbers: RS-2021-KH113822 and RS-2022-KH130307). Additional funding was provided by the National R&D Program for Cancer Control through the National Cancer Center (NCC), funded by the Ministry of Health & Welfare, Republic of Korea (grant number: RS-2025-02214710).
Data availability
The datasets generated and/or analyzed in this study are available from the corresponding author upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
