Abstract
Radiology reporting is narrative, and its content depends on the clinician’s ability to interpret the images accurately. A tertiary hospital, such as anonymous institute, focuses on writing reports narratively as part of training for medical personnel. Nevertheless, free-text reports make it inconvenient to extract information for clinical audits and data mining. Therefore, we aim to convert unstructured breast radiology reports into structured formats using natural language processing (NLP) algorithm. This study used 327 de-identified breast radiology reports from the anonymous institute. The radiologist identified the significant data elements to be extracted. Our NLP algorithm achieved 97% and 94.9% accuracy in training and testing data, respectively. Henceforth, the structured information was used to build the predictive model for predicting the value of the BIRADS category. The model based on random forest generated the highest accuracy of 92%. Our study not only fulfilled the demands of clinicians by enhancing communication between medical personnel, but it also demonstrated the usefulness of mineable structured data in yielding significant insights.
Introduction
Electronic medical records (EMRs) are collection of the patient's medical information stored on a computer system that allow data tracking over time and monitor and improve the quality of care within the practice.1,2 EMRs can come in structured and unstructured data, rich in valuable real-world data on the patient, clinical findings, and genomic information. 3 A radiology report in electronic form is an example of an unstructured EMR.
A breast radiology report contains the interpretation of a breast imaging study prepared by a radiologist, which plays a vital role in communication between radiologists and clinicians.4,5 The free-text radiology report is often associated with excessive variability in language and format when reported by different radiologists. This may decrease clarity and increase the difficulty for clinicians to highlight information from the report. 6 Conversely, structured reporting, an alternative reporting method, helps minimize the variability and maximize the clarity of the report by improving the consistency and communication of information. Additionally, structured reporting has more potential for data analysis by using a machine learning-based techniques compared to unstructured datasets. 7 Machine learning-based analysis requires well-curated input data, which is typically absent from the EMRs. When utilizing the data in the radiology report, the structured and clean data required to derive the machine learning-based predictive models can be extracted by using natural language processing (NLP), thereby minimizing the cost of manually labelling of training data.
Text mining is an artificial intelligence (AI) technique based on NLP to structure the input text, extracting meaningful, non-trivial patterns or knowledge from unstructured texts.8,9 There are two main approaches used in NLP: rule-based and machine learning-based techniques. 10 Previously, researchers have utilized the NLP techniques to extract specific information from the EMRs. However, the majority of clinical NLP mainly focuses on a single task, such as, smoking status and hip fracture classification, 11 thromboembolic disease detection, 12 actionable appendicitis findings extraction, 13 skeletal site-specific fractures identification, 14 ureteric stones identification, 15 stroke detection, 16 section label generation, 17 prostate cancer severity classification, 18 and cancer morphology classification. 19
Our goal is to develop a text mining program based on rule-based NLP techniques to generate structured and summarized breast radiology reports from the anonymous institute. The extracted information in a structured data format allows clinicians to audit and secondary use in machine learning data analysis, for example, predicting Breast Imaging Reporting and Data System (BIRADS) category value using the extracted response for mammogram findings.
Method
NLP algorithm to extract important variables from breast radiology report
Study setting
The Medical Research Ethics Committee (MREC) has approved this study to develop a point-of-care data capture for the Institutional Breast Cancer Registry. The dataset consists of 327 de-identified breast radiology reports from the years 2015 to 2019. These reports were written by various radiologists from the anonymous institute and were randomly retrieved from the radiology information system (RIS) in the Department of Radiology. The random retrieval of the radiology reports was performed by the radiologist involved in the study with the aim of minimizing dataset bias. The third-party RIS implemented in the anonymous institute restricts access for non-medical staff and does not allow direct export of breast radiology reports. Therefore, the radiology reports were manually copied from the RIS to another editable format by the radiologist, which increased the difficulty in retrieving the radiology reports within the time constraints.
The structure for mammography or ultrasound reports, the lexicon for breast imaging findings, and the final assessment category are based on the 5th edition of the American College of Radiology (ACR) BIRADS classification system. 20 However, each breast radiology report may only contain either mammogram or ultrasound examination findings.
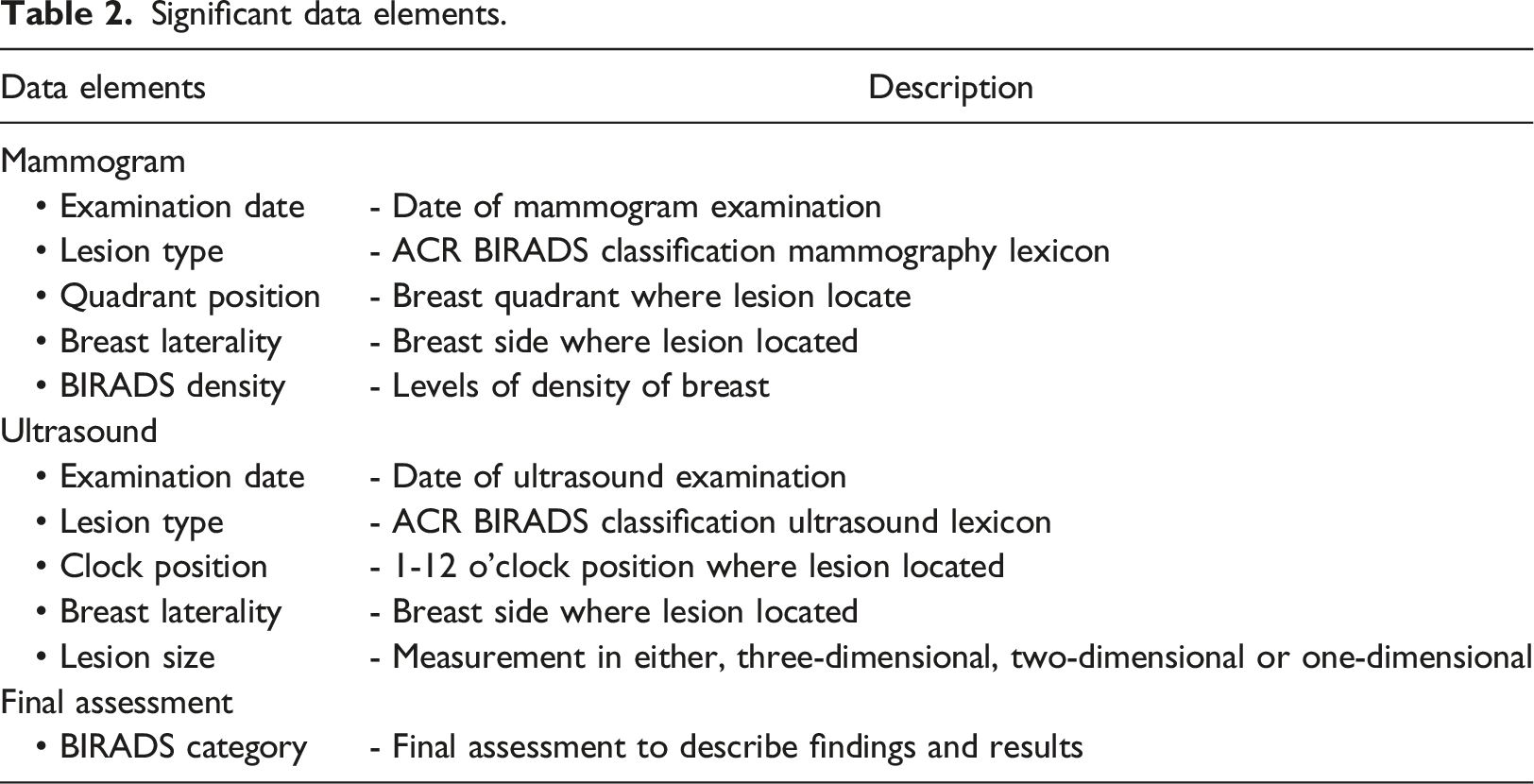
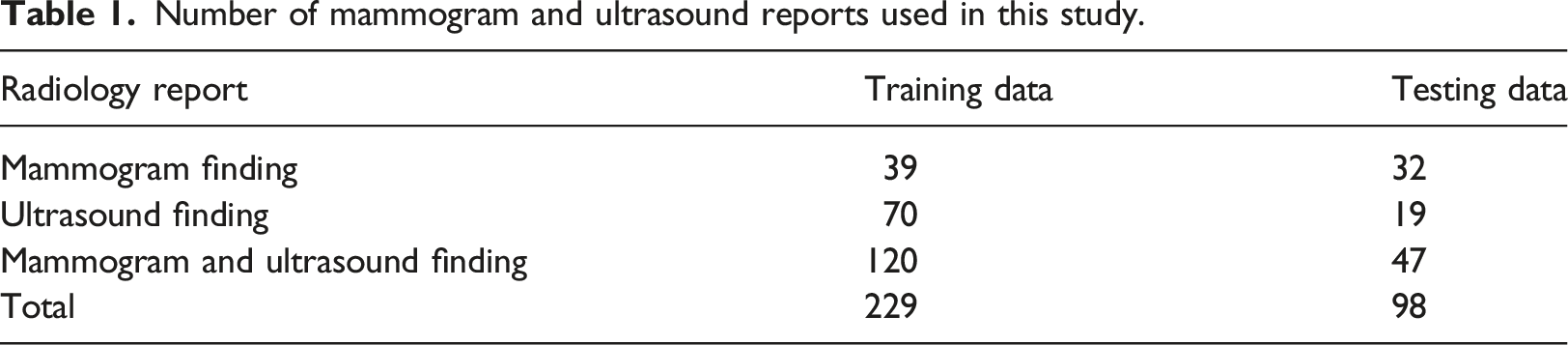
During algorithm development, the radiology reports were randomly divided into 229 training data and 98 radiology reports testing data by using the randomized tools provided by Microsoft Excel. The proposed workflow is illustrated in Figure 1. Table 1 shows the number of mammogram and ultrasound reports involved in this study. Table 2 illustrates the significant data elements identified from the breast radiology report by the radiologists in the anonymous institute. Overview of study. Number of mammogram and ultrasound reports used in this study. Significant data elements.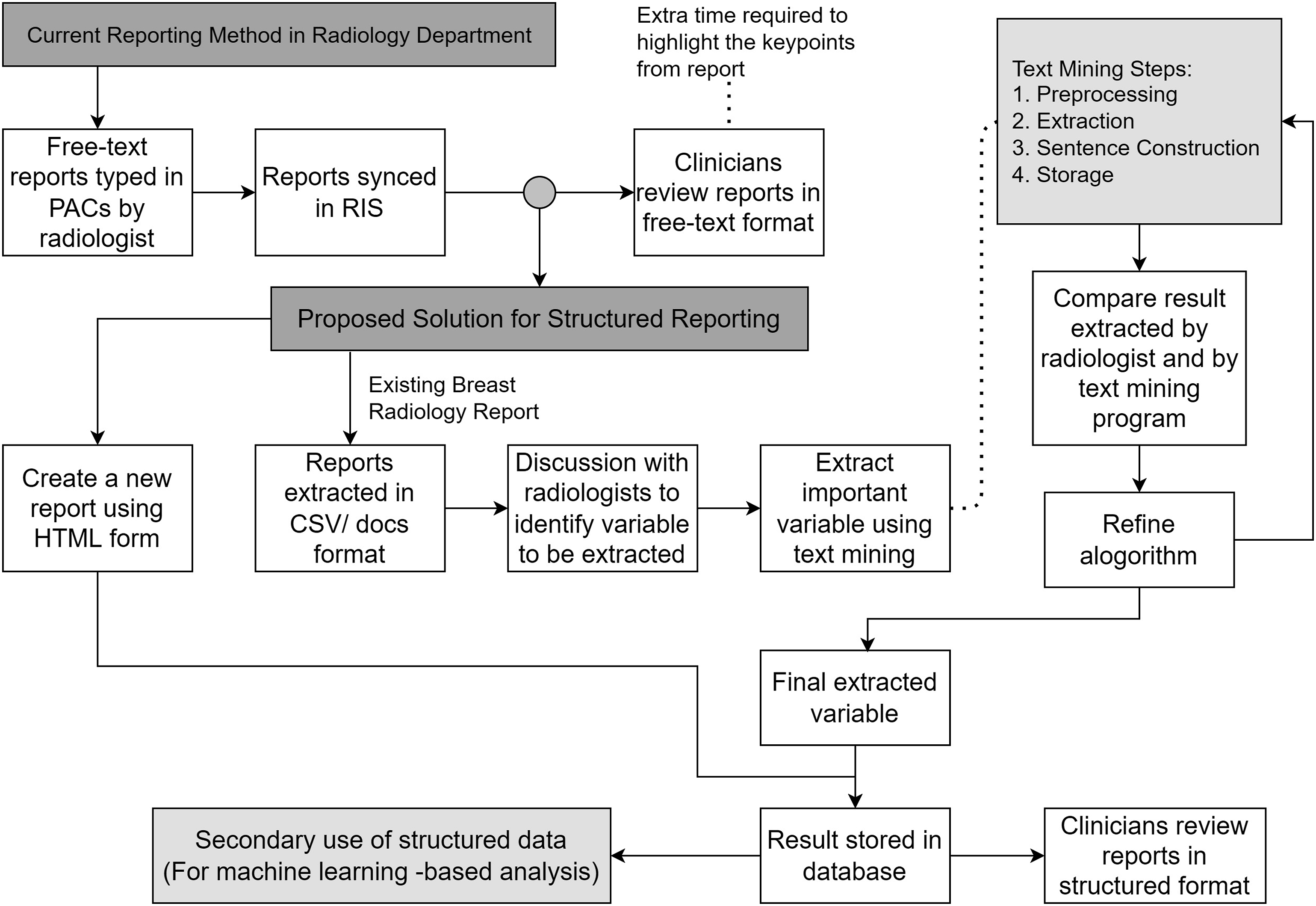

Rule-based NLP algorithm
R programming language version 3.6.1 was used to create the proposed algorithm.
21
Based on training dataset, a set of rules was developed for identifying the desired data elements, guided by the radiologist’s suggestions. The NLP algorithm underwent refinement after radiologist verified the correctness of response extracted by the NLP algorithm. Figure 2 shows the overall algorithm design. Algorithm design.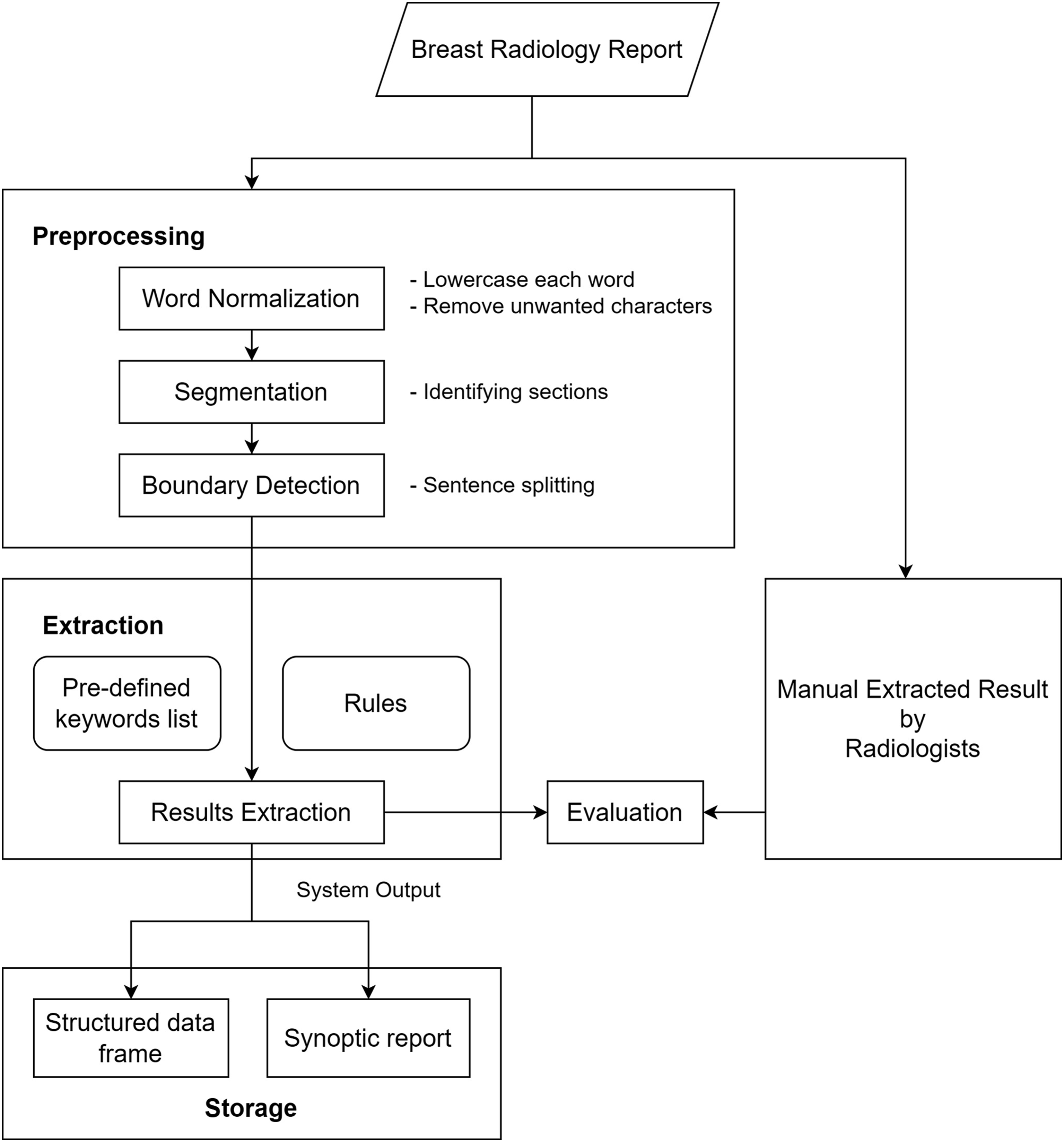
In the first preprocessing step, all characters were converted into lowercase to prevent case-sensitive problems in the subsequent extraction process. Unnecessary characters generated during the exportation of the report from the working platform were removed. The Roman numerals previously used to denote BIRADS category value were replaced with Arabic numerals to ensure format uniformity. Therefore, the report was divided into mammogram and ultrasound sections. The segmentation step was conducted by recognizing the occurrence of the terms “mammogram” and “ultrasound” as the keyword. The successive preprocessing steps enabled the use of a subset of sections with specific rules to maximize efficiency. Boundary detection was performed to further split the report into sub-rows coinciding with each new sentence. This approach allowed semantic analysis to be performed on each sentence individually.
For each mammogram finding, the extracted response included lesion type, breast quadrant, and breast laterality. The size and position of the lesion were not often reported in mammogram section, as mammography could not be precisely determined them.22,23 The presence of at least one of the following data elements in a sentence such as breast quadrant, breast laterality, or a term that describing the lesion location is mandatory as a keyword to allow the NLP algorithm to proceed with further semantic analysis for extracting the lesion type. The lesion type is extracted by matching the text to the predefined R list. Equation (1) illustrates the structure of the R list, which contains a set of the mammography lexicons used for pattern matching.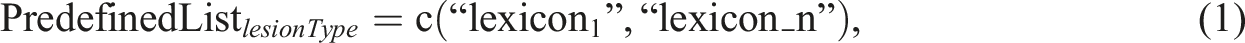
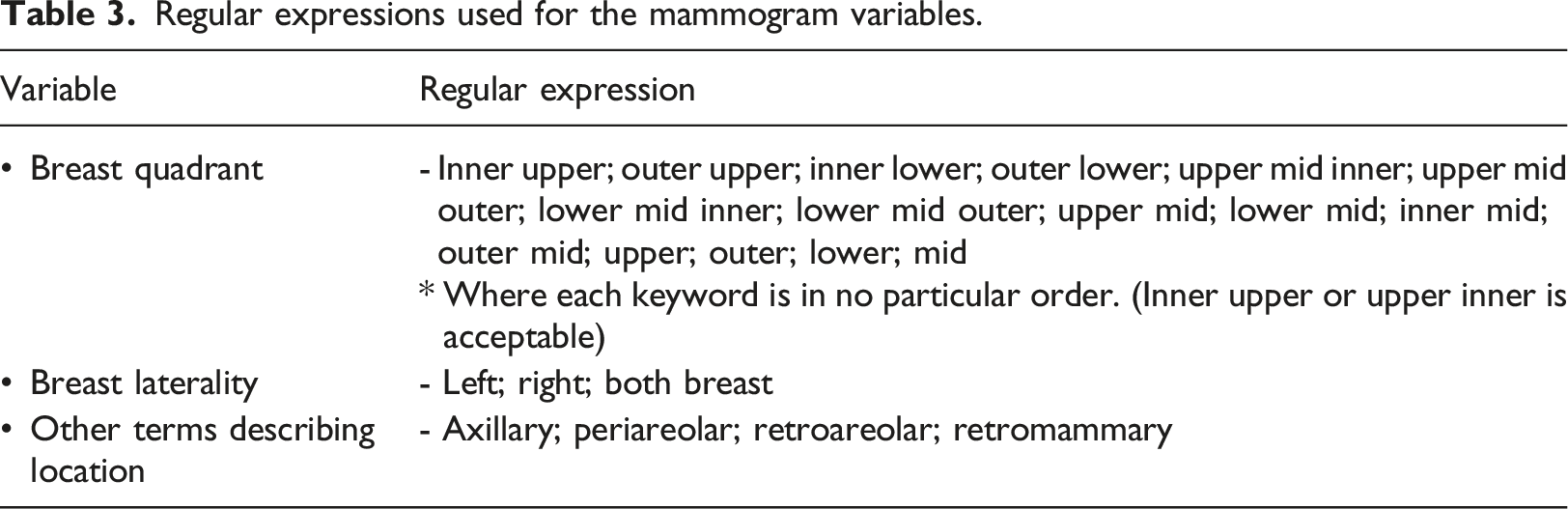
The lexicon used for lesion type is illustrated as a hierarchy tree in Figure 3. During algorithm refinement, terms that were not present in the ACR BIRADS classification system were added to reduce false negatives in the training data. Table 3 illustrates the regular expressions used for identifying the lesion type. Mammogram lexicons. Adapted from ACR BIRADS classification system 5th edition. Regular expressions used for the mammogram variables.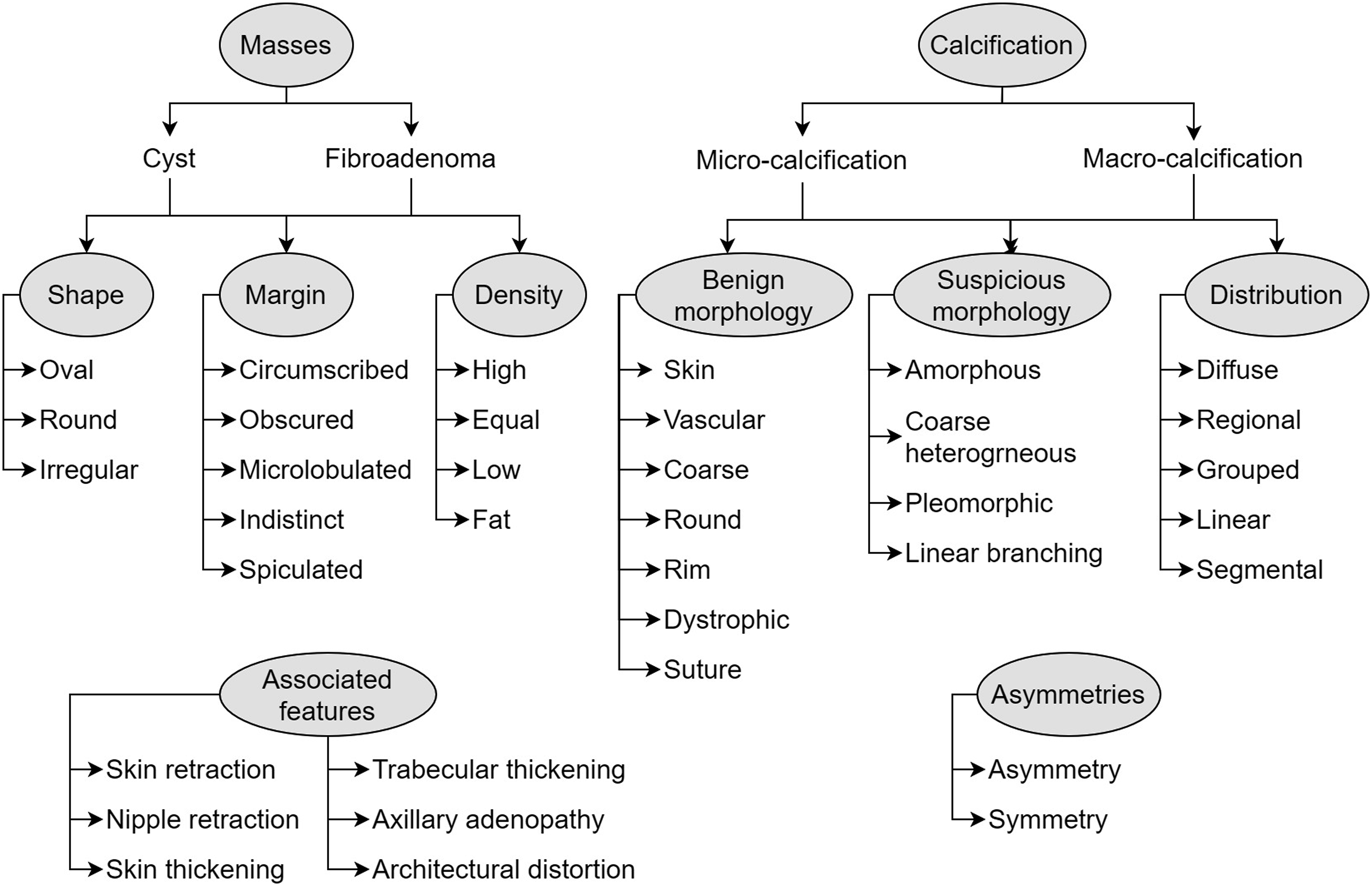

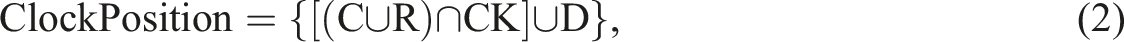
Regular expressions used for the ultrasound variables.
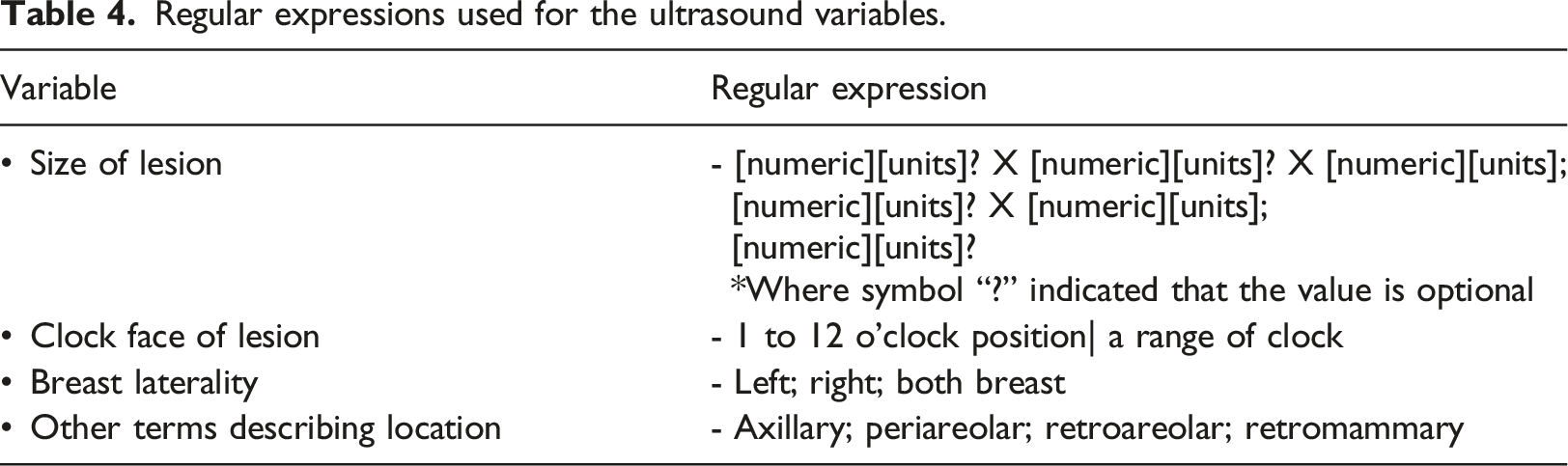

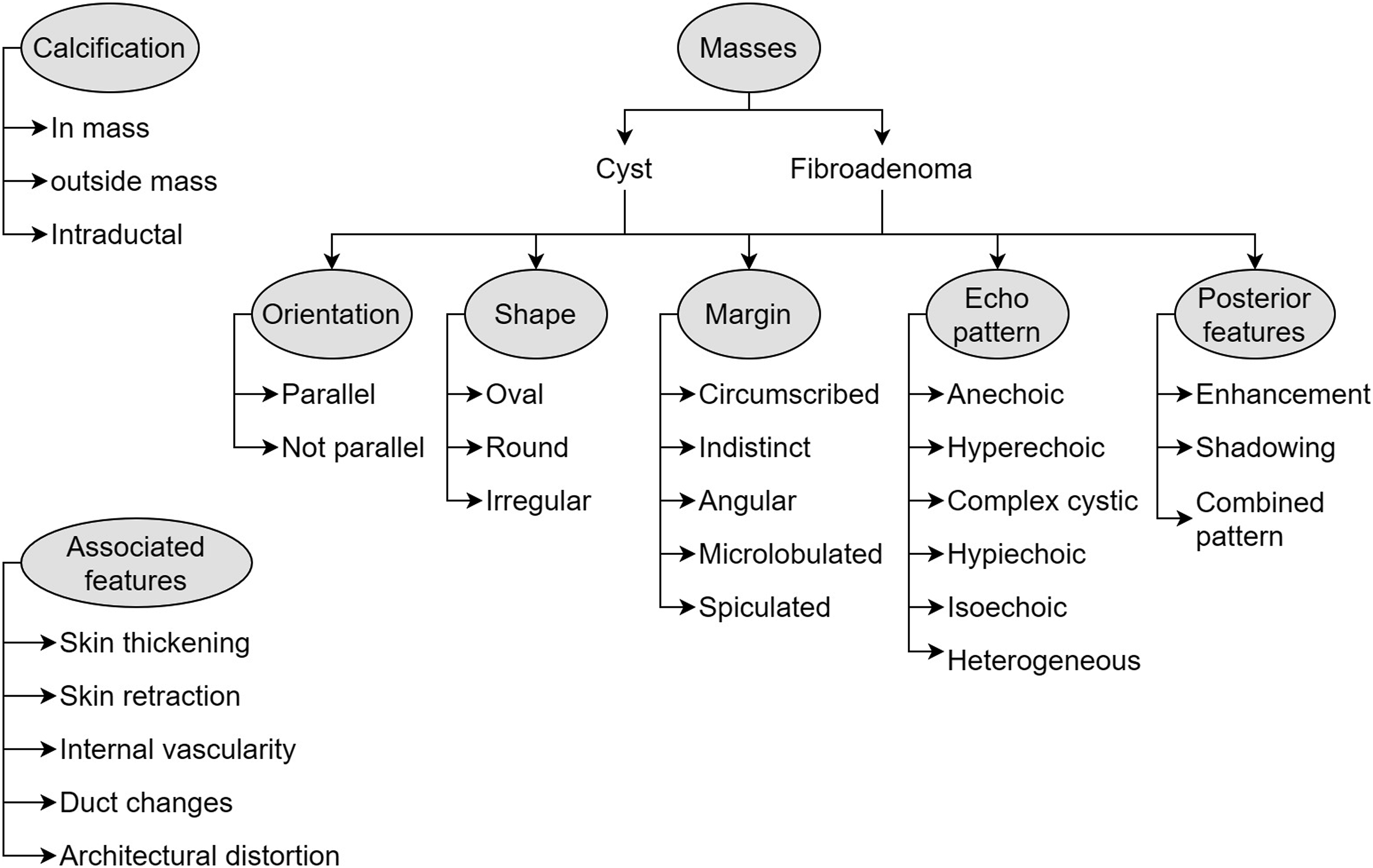
Ultrasound lexicons. Adapted from ACR BIRADS classification system 5th edition.
Dates can be reported in various forms, with the year, month, and day written in different sequences and expressed in numeric or alphabetical format. To simplify the extraction process, the R package “lubridate” was utilized.
24
The algorithm extracts dates and converts them into the Year/Month/Day format to ensure consistency. The keyword “birads” (B) is used to identify the value for BIRADS category and BIRADS density. The algorithm extracts the alphabetical value (A) or numerical value (N) that follows the keyword “birads” as shown in Equations (3) and (4).

Narrative keywords for BIRADS density score.

The outcomes of the information extraction step consist of a set of responses, along with their corresponding data elements, which are stored in a data frame for further utilization in predictive analytics. Using the extracted information, a grammatically correct sentence was generated to produce a synoptic breast radiology report.
Graphical user interface for rule-based NLP algorithm
A simple graphic interface for the text mining program was developed using Python 2.7.18
25
along with the Tkinter module.
26
This interface aims to assist clinicians in auditing reports by presenting the information in a summarised format. Figure 5 shows the design of the interface. Text mining program interface design.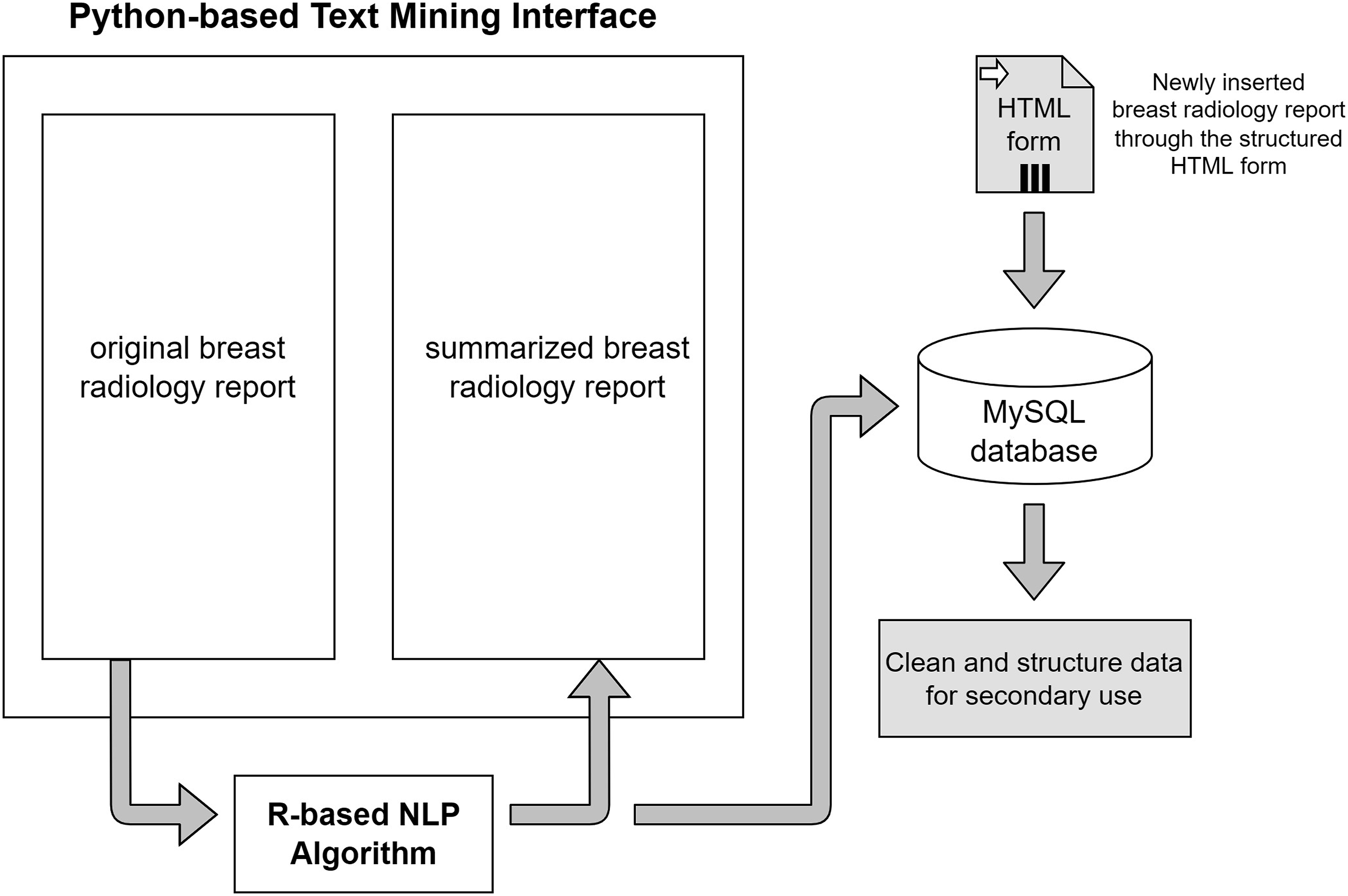
Evaluation of the NLP algorithm
The information extraction ability of the NLP algorithm was assessed using a human validation method. A radiologist was provided with both free-text reports and structured radiology reports generated by the proposed algorithm. During the evaluation, the radiologist determined the correctness of the algorithm-extracted findings and also identified any information missed by the algorithm. In information extraction tasks, the emphasis is on accurately identifying and extracting specific information of interest. The evaluation metrics used, such as precision, recall, and F1 score, measure the accuracy and completeness of the extracted information compared to the ground truth. True negatives, which represent correctly identified absence of information, are not typically included in these metrics. The focus is on maximizing the extraction of relevant information rather than evaluating the absence of it. To measure the algorithm's performance in information extraction, five metrics known as precision, recall, false discovery rate (FDR), false negative rate (FNR), and F1 scores were used.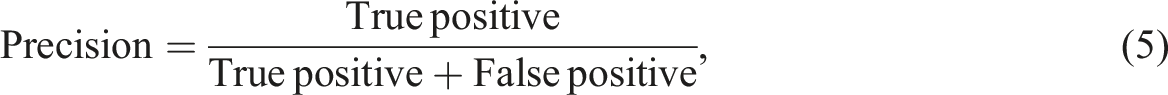
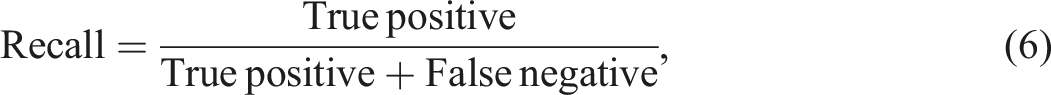
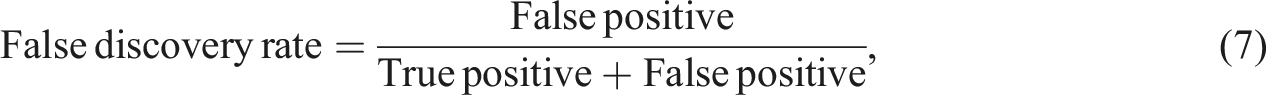
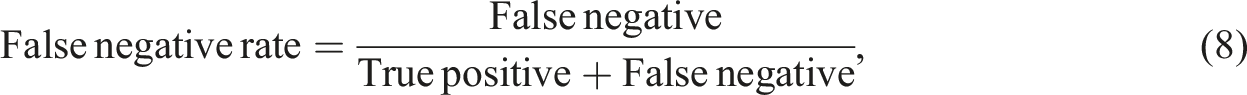
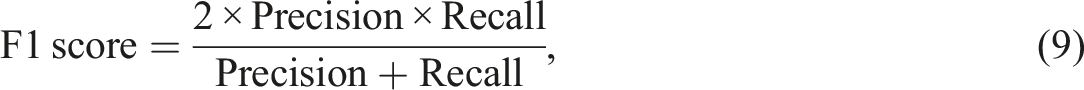
Borderline cases can occur when there are amendments made to the free-text report. These amendments are typically added at the bottom of the report. In such cases, the extraction process can be less accurate and potentially result in increased false-negative or false-positive values. To address this issue, a reminder message will be displayed when the keyword “amendment” is detected in the report. This serves as a prompt for clinicians to pay closer attention to any potential issues or changes that have been made.
Predicting BIRADS category using variables extracted by NLP algorithm
Datasets and variables
Variables involved in predictive analytics.
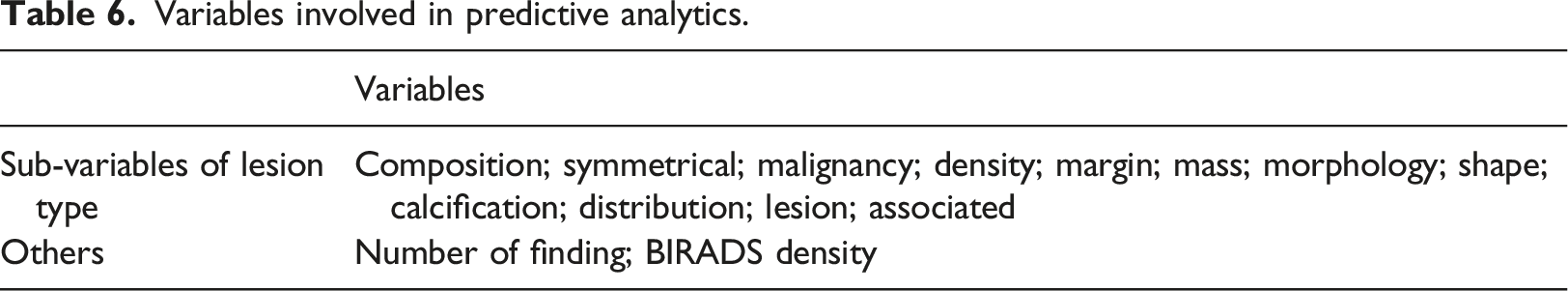

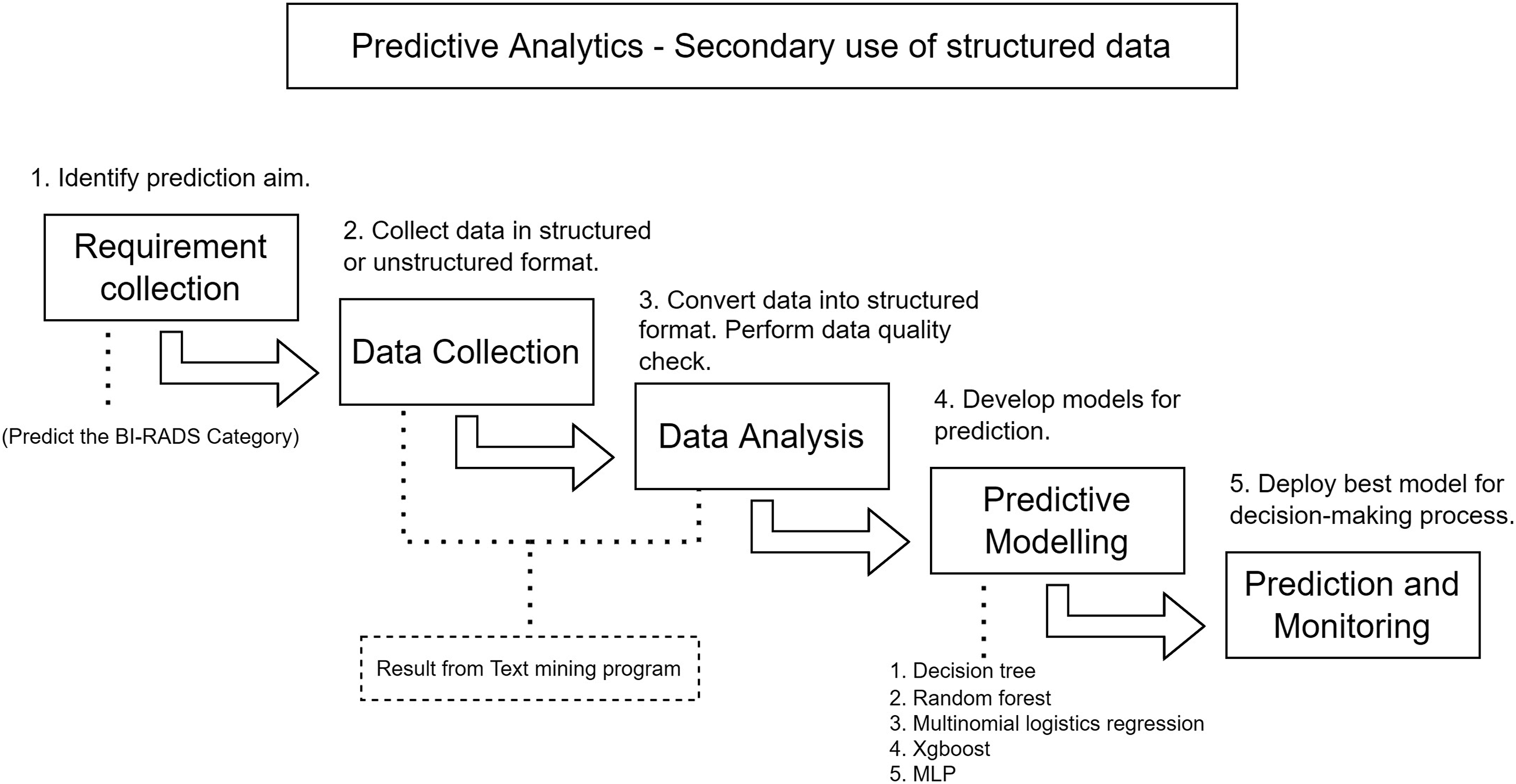
Overview of predictive analytics process.
Feature selection
Feature selection is a critical step in developing a predictive model. Utilizing only the features with higher importance scores can enhance the accuracy of the model. The R package “caret” provides the recursive feature elimination (RFE) method, which facilitates the removal of unimportant features. 27 By eliminating non-significant attributes, the chance of overfitting during model training is reduced. RFE ranks feature importance using the mean decrease of accuracy (MDA) metric, which calculates the accuracy loss after randomly permuting the values of each input in each iteration. 28 Consequently, variables with higher importance result in a greater decline in the prediction accuracy. As a result, the variables with the lowest importance are eliminated, resulting in a minimal set of variables with the best predictive ability.
Predictive model
Five machine learning-based predictive models were selected for this study: decision tree, random forest, multinomial logistics regression, extreme gradient boosting, and multiple layer perceptron. The dataset was randomly split into a training set and a testing set using the data splitting function provided by the R package “caret”. The training set consisted of 70% of the data (160 mammogram findings), while the testing set comprised the remaining 30% (69 mammogram findings).
Decision tree
The decision tree model for BIRADS category prediction was generated using the classification and regression tree (CART) function provided by the R package “tree”. 29 CART builts the decision tree by recursively partitioning the dataset into subsets based on different predictor variables. At each split, the algorithm selects the predictor variable that best separates the data into homogeneous subsets. This process continues until a stopping criterion is met, resulting in terminal nodes. Each terminal node in the decision tree represents a specific combination of predictor variables and provides a prediction for the target variable, which, in this case, is the BIRADS category. 30 By using the CART function from the “tree” package, the decision tree model captures the relationships between predictor variables and the BIRADS category, enabling accurate predictions for new, unseen data.
Random forest
The random forest model, developed using the R package “RandomForest,” is based on Breiman and Cutler's original Fortran code. 31 The algorithm generates multiple independent decision trees using various random subsets of the training set. By generating multiple trees with varying subsets, the random forest model captures a range of possible relationships between the input variables and the target variable. During prediction, the model combines the outputs of these trees and determines the final prediction by selecting the most frequently occurring output among the individual decision trees. 32 This ensemble approach enhances the robustness and accuracy of the predictions in a wide range of applications.
Multinomial logistics regression
The R package “nnet” was utilized to develop the multinomial logistics regress model. 33 Multinomial logistics regression is a classification method that generalizes logistics regression to handle multiple discrete outcomes. During operation, one of the categories was chosen as the base category, and the remaining categories are individually regressed against the base category to determine their probabilities. The coefficients of the regression model represent the influence of each predictor variable on the log-odds of each outcome. By fitting the multinomial logistic regression model, predictions and classifications of new instances into one of the discrete outcome categories can be made. This approach is particularly useful when dealing with categorical response variables that have more than two levels.
Extreme gradient boosting
The gradient boosting machine is an ensemble supervised learning technique that accepts matrices for model evaluation. It is a powerful supervised learning method that is capable of handling both regression and classification problems. It is particularly effective when dealing with complex and diverse types of variables. The technique works by sequentially building a series of weak prediction models, with each subsequent model focusing on minimizing the errors made by the previous models. This iterative process targets areas where the previous models performed poorly, gradually improving the overall accuracy. The final prediction is obtained by combining the predictions of all the weak models.34,35 The R package “xgboost” was used to perform the gradient boosting machines. 36
Multiple layer perceptron
This study utilized multiple layer perceptron, which is a feed-forward and supervised learning technique for artificial neural networks, implemented using the R package “h2o”. 37 The multiple layer perceptron consists of at least three layers, namely input, hidden, and output layers. It is known as universal approximator, as demonstrated by Cybenko's theorem, which states that a neural network with a single hidden layer and a sufficient number of neurons can approximate any continuous function. The multiple layer perceptron works by propagating input data forward through the network, applying weights to the connections between neurons, and activating each neuron using an activation function. During training, the network adjusts the weights based on the difference between the predicted and actual output, optimizing its ability to make accurate predictions. 38 This technique is effective in capturing complex relationships and patterns in the data, making it suitable for various regression and classification tasks.
Model evaluation
Similar to information extraction tasks, the concept of true negatives is not commonly utilized in multi-class classification scenarios. True negatives are primarily applicable to binary classification problems, where the goal is to correctly classify instances into two categories. In multi-class classification, the emphasis is on assessing the model's ability to accurately identify and differentiate between multiple classes or categories. Therefore, evaluation metrics in multi-class classification typically focus on measures such as accuracy, precision, recall, and F1 score using R package “caTools”,
39
which provide insights into the model's overall performance across the different classes. In the model’s performance assessment, the accuracy of each model was evaluated by: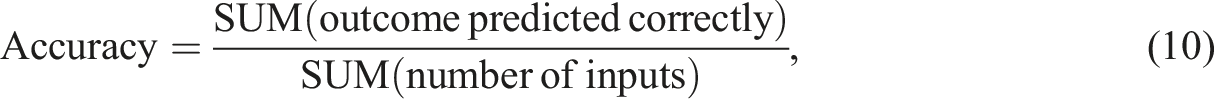
In addition, the agreement between the predicted outcomes and the actual outcomes was further evaluated using the kappa coefficient. The kappa coefficient measures the level of agreement between two raters or evaluators, taking into account both the observed agreement and the agreement expected by chance. It is calculated as: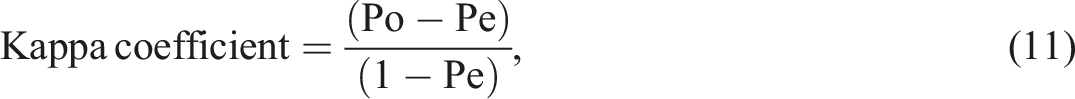
Result
NLP algorithm to extract important variables from breast radiology report
Evaluation result of training data.
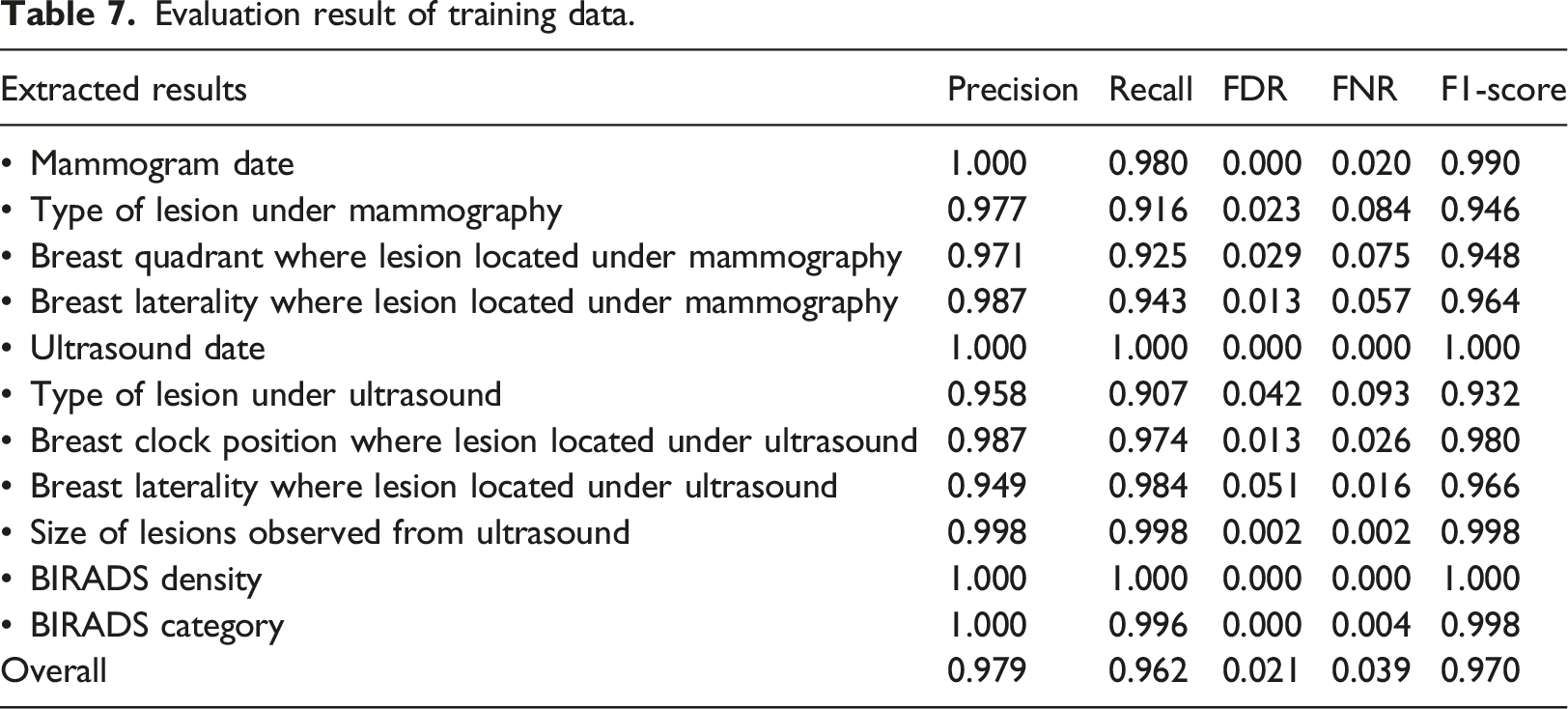
Evaluation result of testing data.
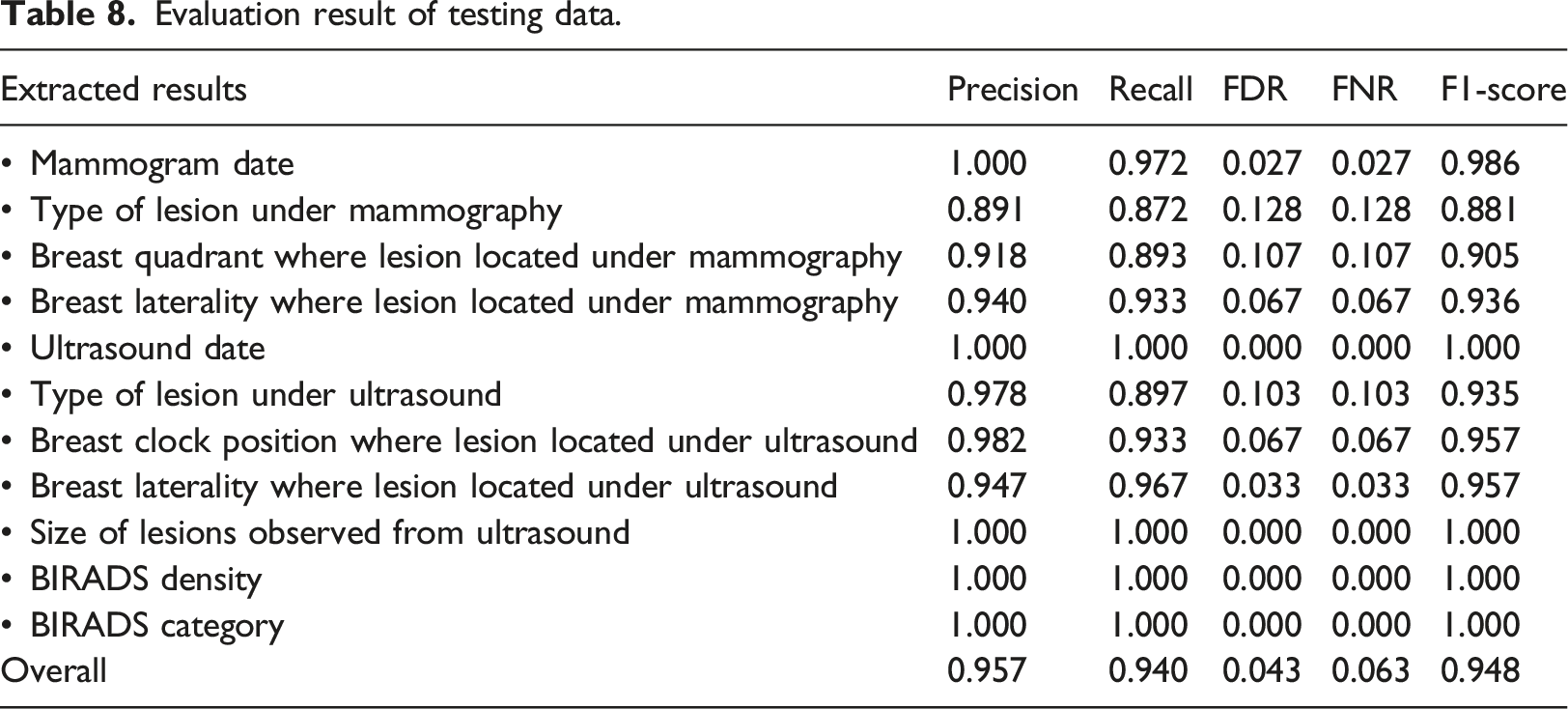
Rule-based NLP algorithm
The integration of the R package “lubridate” eased the process of examination date extraction. However, in some cases, false extraction of the date can occur. For instance, in the case of a date reported as “1.2.18”, the month value can be interpreted as 1 or 2, depending on whether the month or the day is checked first. Consequently, this leads to an increase in the false positive, resulting in reduced precision.
During the lesion type extraction process, the NLP algorithm performs pattern matching within the input text using predefined lists. Here are some examples of false positive and false negative cases that occurred throughout development of the NLP algorithm (Figure 7). In Example 1, the algorithm incorrectly extracted the lesion type as “with calcifications” for a lesion with a size of 1.3 x 0.8 x 0.8 cm in Example of sentences in free-text report.
Multiple lesions can be present on the same breast, and the radiologist may only mention the breast laterality once before reporting the multiple lesions observed. In Example 1, there are two same-type lesions with different sizes located on the same breast as reported in
In some cases, radiologists report the BIRADS density in numerical or narrative format instead of using the category system from A to D. Henceforth, the algorithm is designed to automatically convert the numeric value to an alphabetic value in the sentence where the keywords “density” and “BIRADS” are present. For the BIRADS density described in narrative format, the algorithm converts it into an alphabetic value by searching for keywords that describe each BIRADS density, as shown in Table 5.
The proposed NLP algorithm is capable of analyzing sentences with more complex structure, including those that involve multiple lesions with different sizes, position, and type within the same sentence. Figure 8 demonstrates an example where the algorithm successfully identifies the boundary between two different lesions when there are two measurements and the keyword “and” is present between two clock positions in a single sentence. Multiple size, type and position extraction in same sentence.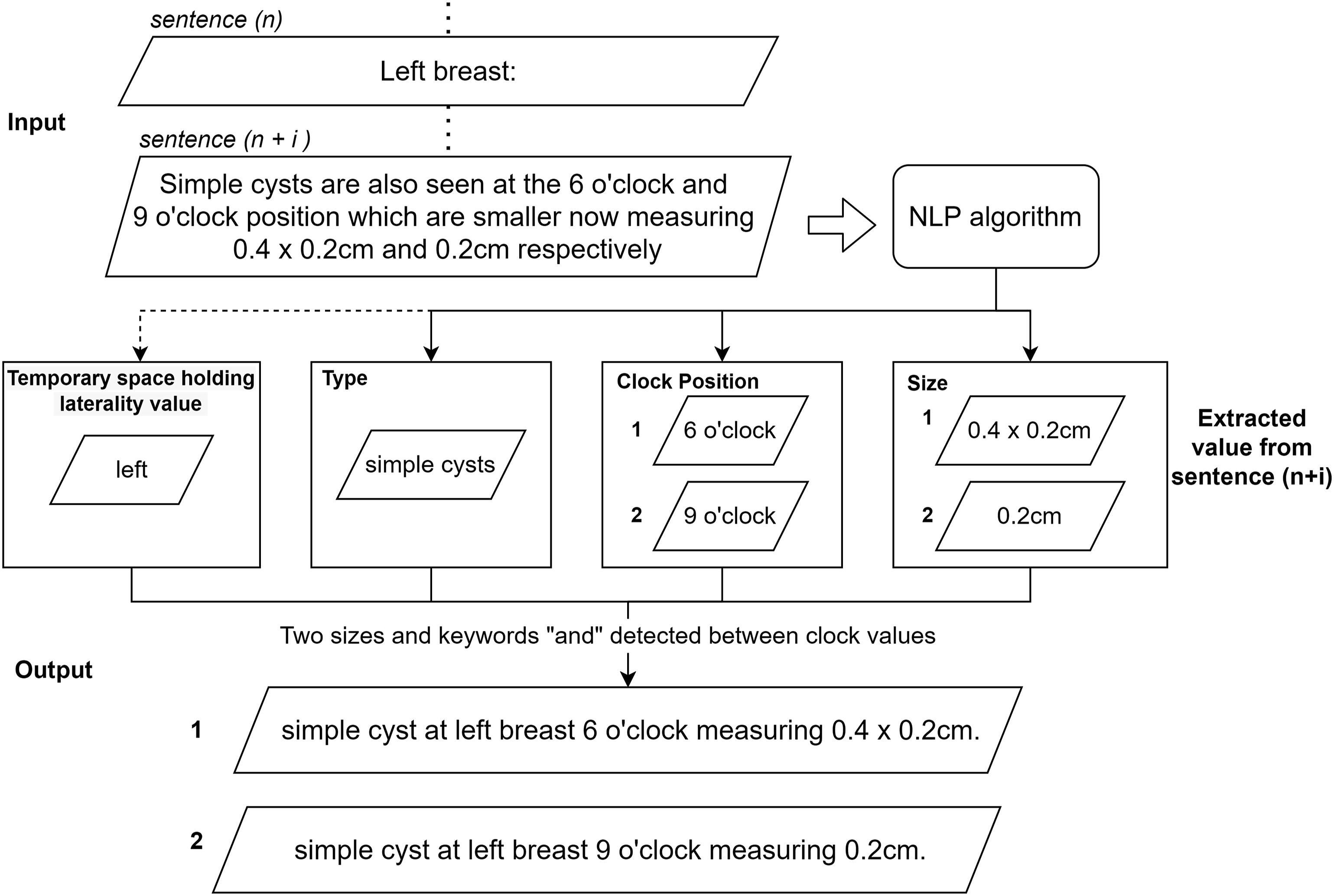
Graphical user interface for rule-based NLP algorithm
The interface of the text mining program consists of two main panels (Figure 9). The left panel is responsible for displaying the original inputted breast radiology report, while the right panel shows the summarized synoptic breast radiology report generated by the NLP algorithm. Although the displayed input report may contain some meaningless characters due to the exporting process across different platforms, this does not affect the extraction process. These unwanted characters are removed in the preprocessing step. Graphical user interface of rule-based NLP algorithm.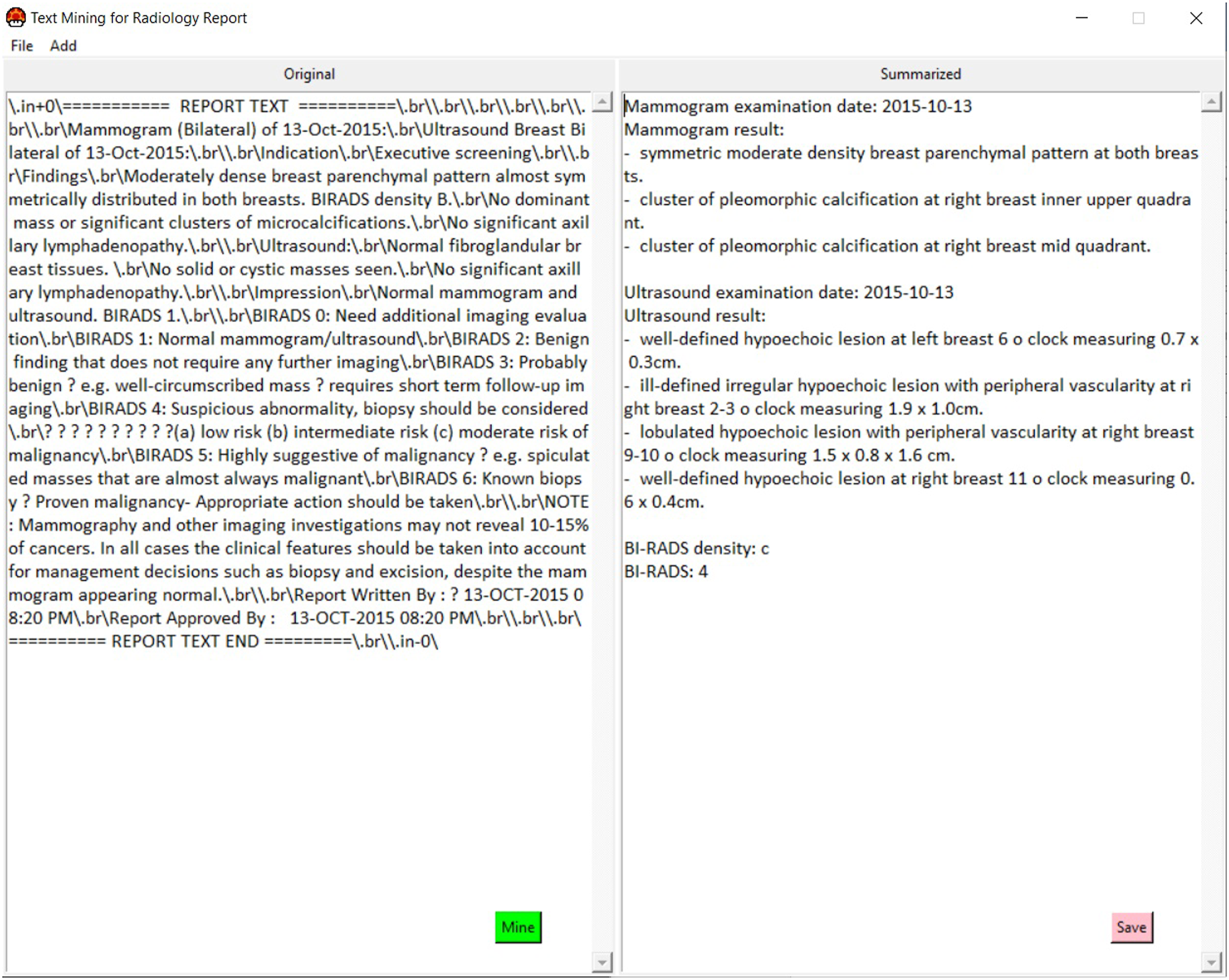
Predicting BIRADS category using variables extracted by NLP algorithm
Feature selection with RFE with 5-fold cross validation
RFE has identified that all 14 features are significant in predicting the target variable, BIRADS category value. These features collectively contribute to the best predictive model with the highest accuracy of 0.86 and a kappa value of 0.81. Table 9 shows the details of the accuracy and kappa value of each set of variables after RFE. The 14 significant features identified are: 1. Calcification; 2. Composition; 3. BIRADS density; 4. Number of finding; 5. Density; 6. Calcification distribution; 7. Associated; 8. Margin; 9. Symmetrical; 10. Lesion; 11. Morphology; 12. Malignancy; 13. Shape; 14. Mass. Resampling performance over subset size.
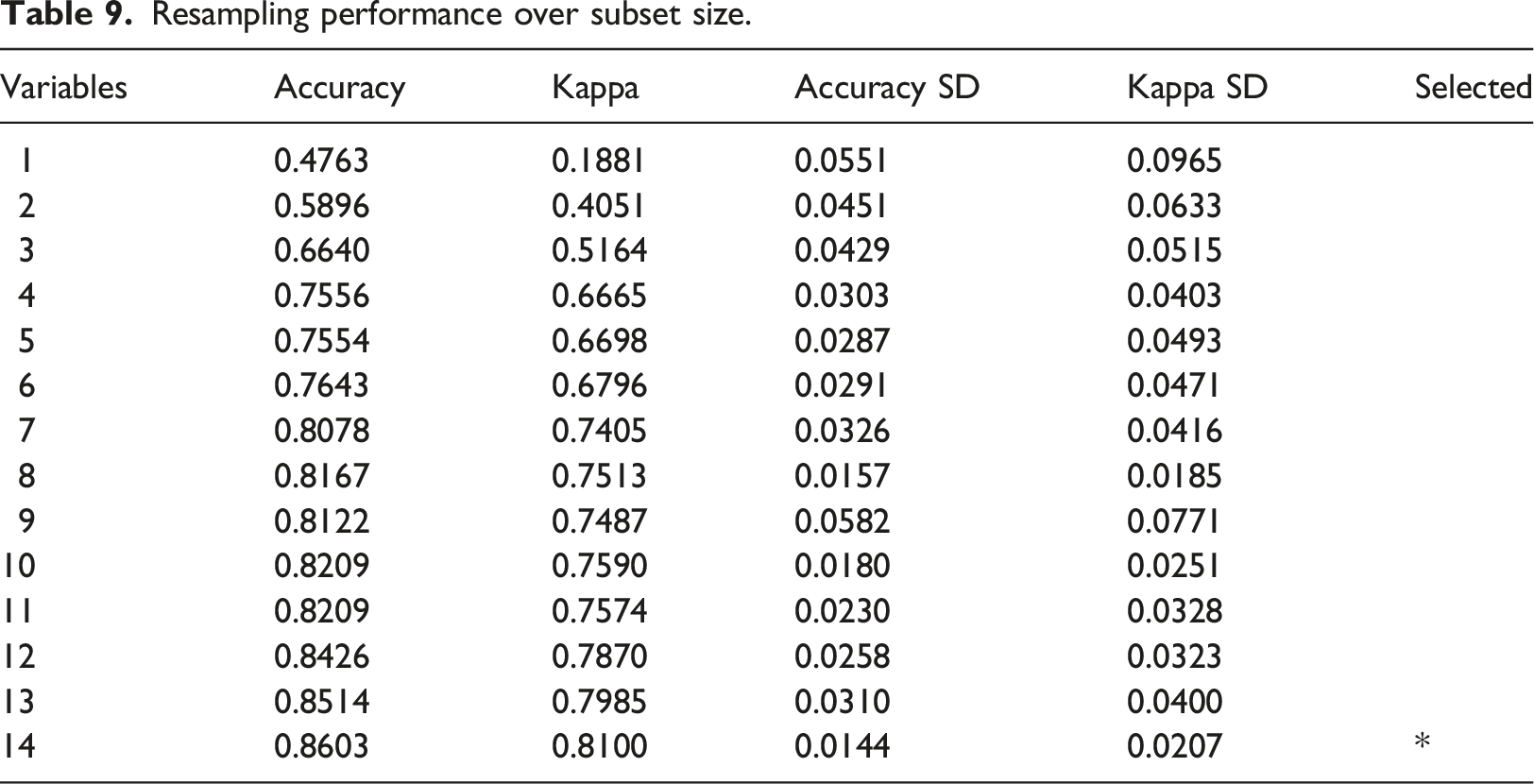
These variables have been determined to have a strong association with the target variable and play an important role in accurately predicting the BIRADS category value.
Model evaluation
Model accuracies.
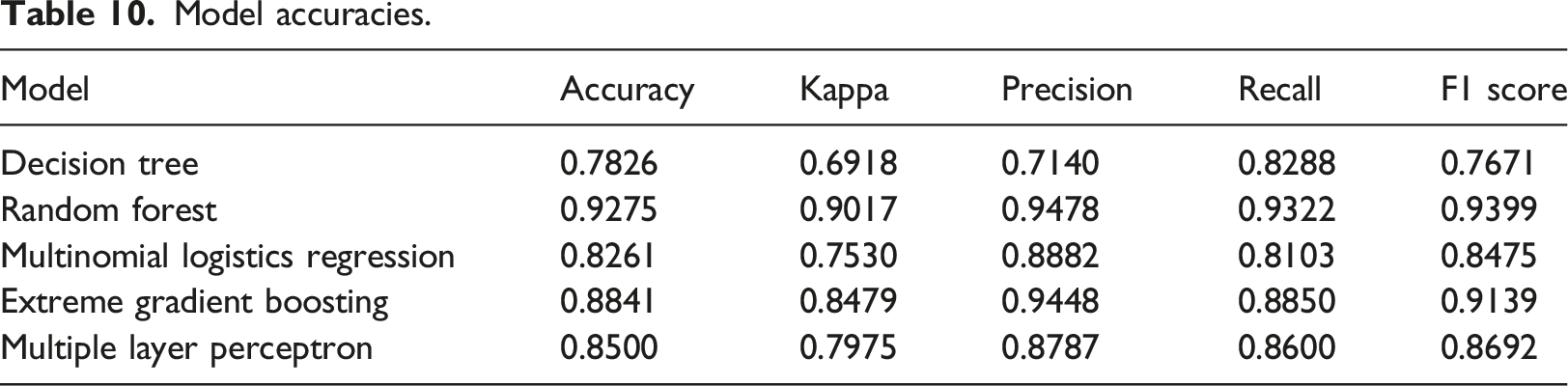
Among the five predictive models, the random forest model achieved the highest accuracy of 0.9275, followed by the extreme gradient boosting model (0.8841), multiple layer perceptron models (0.85), and multinomial logistic regression model (0.8261), and the decision tree model with the lowest accuracy of 0.7826. These predictive models were trained to predict the BIRADS category value ranging from 1 to 5. The BIRADS category value 0 and BIRADS category value six were excluded from the analysis for two main reasons. Firstly, datasets with BIRADS categories 0 and six comprised a small portion of the overall dataset. Secondly, BIRADS category 0 indicated an incomplete mammogram, while category six required confirmation through a biopsy. Overall, these findings demonstrate the performance of the predictive models in accurately predicting the BIRADS category value, while considering the specific exclusions and limitations of the dataset.
Discussion
The rise of EMRs has generated enormous data that is rich in information. However, most medical reports are written in narrative format and separated into different sections, rather than reporting each data element in distinct categories. Consequently, clinicians require more time to interpret and determine the subsequent treatment process. Clinical data stored in free-text have different database structures from structured data, which increases the difficulty of performing data mining. Therefore, NLP, whether rule-based or machine learning-based techniques, provide an opportunity in the clinical domain to reduce the costly manual training data annotation processes.8,9
The highly unstructured radiology reports have limited opportunities for data mining due to the requirement of structured data for most data mining methods. Henceforth, we propose a rule-based algorithm that aimed to reduce the workloads of the clinicians in the keypoint highlighting process and provides an opportunity for further analysis with minimal human interference. With the aid of the graphical user interface, clinicians can utilize the rule-based NLP algorithm to generate a synoptic breast radiology report in the auditing process without requiring any programming skills.
In order to facilitate the auditing and research purpose, achieving a high F1 score (with a value of 1 being the best and 0 being the worst) was crucial. 40 Our NLP algorithm yielded promising results in information extraction tasks, with a test set F1 score of 0.947. Data elements with straightforward and minimal ambiguity, such as a dates, BIRADS category, and BIRADS density, showed higher F1 score. The formats used to write these variables are limited, which eliminates the need for complex rules to extract the required information. Conversely, the lesion type, which can be described in a variety of terms, shows the lowest F1 score in both training and test data. The complexity and ambiguous reporting style and lexicon when describing a lesion increased the false positive and false negative values, consequently impacting the overall accuracy.
Lately, the increasing use of machine learning techniques has shown great potential in achieving higher accuracy for automated outcome classification, especially when large volume of data is available. Spandorfer et al. conducted a similar study where they structured unstructured computed tomography pulmonary angiography (CTPA) report. In the study, 91.6% of statement were correctly labelled by the algorithm. 17 Although the algorithm showed promise in achieving a high accuracy in the automated labelling process, it primarily focused on providing section labels to sentences, which is considered the most fundamental structure. However, this approach necessitated the manual annotation of section labels on a large amounts of training data by a clinical expert.
The rule-based NLP algorithm continues to be prominent in clinical NLP, despite the growing popularity of machine learning approaches in the broader NLP community. Clinical NLP has distinct requirements compared to other domains, as machine learning-based NLP primarily focused on prediction, estimation, and association mining. In contrast, the rule-based system offers simplicity and facilitates interactive refinement through clinician feedback, making it easier to debug. Meanwhile, machine learning-based systems often operate as black boxes, making interpretation and correction more challenging. Consequently, rule-based NLP remained favoured in the clinical field. 41
In a study carried on by Hammami et al., rule-based NLP was employed to extract cancer morphology codes from pathology reports, achieving an impressive accuracy of 98.14%. 19 Another example of rule-based NLP is the extraction of family history information, including associations with diseases and the living status of the family members, from the EMRs. 42 Bozkurt et al. compared different NLP algorithms, including deep learning, rule-based, and hybrid models for the classification of prostate cancer severity. Their findings indicated that rule-based model (accuracy=0.86) is outperformed by the deep model (accuracy=0.73) and a hybrid model (accuracy=0.75). 18 Rule-based NLP is not limited to classification tasks alone; it can also be utilized in medical diagnosis systems to suggest diagnosed diseases from user’s self-input. 43
Although rule-based NLP relies on handcrafted rules, it offers the advantage of incorporating domain knowledge from knowledge bases or experts. In contrast, machine learning-based NLP requires a large set of manually annotated and well-structured training data to ensure the accurateness in a given task. Rule-based NLP, on the other hand, reduces the need for costly manual annotation process required in machine learning-based techniques. For example, the labels for training data in a model that performs smoking status and hip fracture classification are generated by the rule-based NLP. 11
In this study, we demonstrated the real-world application of our NLP algorithm in assisting the predictive analytics process. We developed five predictive models for BIRADS category prediction based on mammogram features extracted using the NLP algorithm. All variables were found to be significant, resulting in an accuracy of 0.8603 after performing features selection using RFE methods. In addition, the kappa value of 0.81 indicated strong agreement with the data reliability. 44 Among the models, the random forest model has successfully predicted the BIRADS category value with the highest accuracy of 92.75%. The other models also demonstrated accuracies ranging from 78% to 88%. These successful predictions of the BIRADS category provide further evidence of the effectiveness and feasibility of our NLP algorithm in data mining research.
The main limitation of this study is the use of small dataset from a single institution. The use of a closed-architectural third-party RIS makes obtaining the breast radiology report troublesome and laborious. The rules generated from the dataset, which is limited to a single hospital may not cover all possible scenarios. Besides, different terminology is applied in different medical centers, which can increase the false negative value. Therefore, further research is needed to minimize human workload by automating the process of retrieving and annotating the breast radiography report from the RIS. Furthermore, the exploration of increasing the dataset from multiple sources would improve generalizability and minimize algorithm bias.
Conclusion
The study presented demonstrates the practical application of NLP in routine clinical practice. The ideas presented here not only digitize free-text reports into a minable form, but also prospectively digitize data collection. NLP enables the conversion of narrative reports into a structured format automatically. However, human review and validation are still necessary on utilizing the NLP application in these clinical reports, which is significant for clinical documentation for care and medico-legal purposes. Take Europe as an example, the European Commission published a white paper emphasizing human dignity and privacy protection in medical AI. It aligns with their AI approach based on fundamental rights and values, recognizing the importance of these aspects in using medical AI technologies. 45
The utilization of AI in clinical domain including radiology field will soon be getting more popular in future. However, human expertise is still indispensable. Henceforth, a balance and cooperation between AI and human intelligence is important in solving problem with legal requirements. Sorantin et al. have conducted a comprehensive discussion on the distinctions between AI and human intelligence in problem-solving, highlighting the significance of achieving a balance and fostering cooperation. They introduce the concept of “explainable AI” as a means to exemplify this approach. 46
In conclusion, this work assists clinicians by extracting significant points from free-text reports and serves as a low-cost preliminary step in machine learning-based analysis or data mining. It enables the automated annotation process of training data, facilitating efficient and accurate data labelling. By leveraging NLP, this study contributes to the advancement of clinical practice, improving the extraction of meaningful insights from free-text reports. Furthermore, the integration of NLP techniques in machine learning-based studies opens up new possibilities for automated annotation and analysis of clinical data, providing valuable support to healthcare professionals and researchers.
Supplemental Material
Supplemental Material - Natural language processing in narrative breast radiology reporting in University Malaya Medical Centre
Supplemental Material for Natural language processing in narrative breast radiology reporting in University Malaya Medical Centre by Wee Ming Tan, Wei Lin Ng, Mogana Darshini Ganggayah, Victor Chee Wai Hoe, Kartini Rahmat, Hana Salwani Zaini, Nur Aishah Mohd Taib, and Sarinder Kaur Dhillon in Health Informatics Journal
Footnotes
Acknowledgements
The authors thank the Breast Cancer Research group (patients, physicians, nurses, technical staff, research assistants and hospital administration staff at UMMC) for providing relevant data to conduct this study.
Author contributions
Conceptualization, W.-M.T., W.-L.N., M.D.G., N.A.T., V.C.-W.H., K.R., H.S.Z. and S.K.D.; methodology, W.-M.T. and S.K.D. validation, W.-M.T., W.-L.N., M.D.G.; formal analysis, W.-M.T.; resources, H.S.Z. and W.-L.N.; writing—original draft preparation, W.-M.T.; writing—review and editing, W.-M.T. and S.K.D.; supervision, S.K.D.; project administration, S.K.D., K.R. and N.A.T. All authors have read and agreed to the published version of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical statement
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
