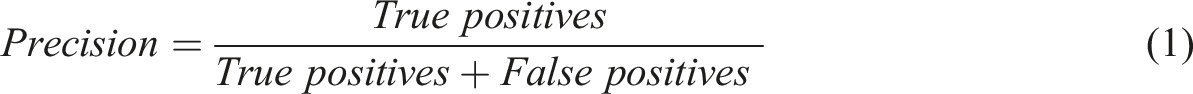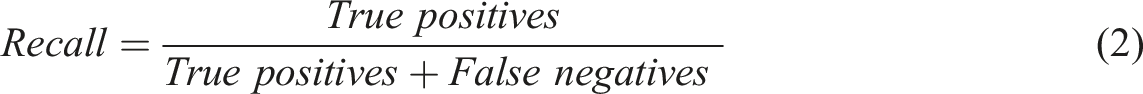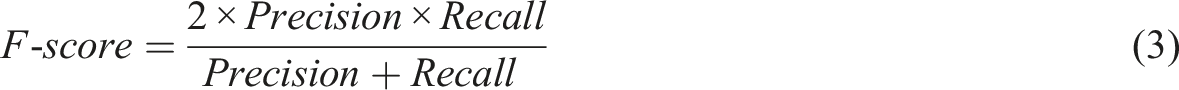Abstract
Defining profiles of patients that could benefit from relevant anti-cancer treatments is essential. An increasing number of specific criteria are necessary to be eligible to specific anti-cancer therapies. This study aimed to develop an automated algorithm able to detect patient and tumor characteristics to reduce the time-consuming prescreening for trial inclusions without delay. Hence, 640 anonymized multidisciplinary team meetings (MTM) reports concerning lung cancers from one French teaching hospital data warehouse between 2018 and 2020 were annotated. To automate the extraction of eight major eligibility criteria, corresponding to 52 classes, regular expressions were implemented. The RegEx’s evaluation gave a F1-score of 93% in average, a positive predictive value (precision) of 98% and sensitivity (recall) of 92%. However, in MTM, fill rates variabilities among patient and tumor information remained important (from 31% to 100%). Genetic mutations and rearrangement test results were the least reported characteristics and also the hardest to automatically extract. To ease prescreening in clinical trials, the PreScIOUs study demonstrated the additional value of rule based and machine learning based methods applied on lung cancer MTM reports.
Keywords
Introduction
Lung cancer represents the leading cause of cancer mortality, 1 with more than 30,000 cases and 33,000 deaths in France in 2018. 2 Manual review of medical records is essential in clinical trials, representing a high consuming task in terms of human and financial resources.3,4 For instance, the anatomic extent of disease described by the TNM classification reflects the prognosis of patients with lung cancer and helps to determine cancer staging and treatment.5,6 Difficulties in subjects inclusion in clinical trials are increasing, potentially causing delay and opportunity loss for patients 7 due to the numerous and highly specific criteria to check before inclusion.
The computerization of the multidisciplinary team meeting (MTM) reports is a prerequisite to automate the classification of lung cancers. 8 Lung cancer patients are reviewed by a multidisciplinary team meeting (MTM), including a quorum of experts in pulmonology, oncology and thoracic surgery, in a reference centre. 9 Multidisciplinary approach is required to validate diagnosis and to propose a treatment plan according to patient’s profile, described by several clinical or paraclinical criteria. These criteria are also verified to assess clinical trials eligibility. Automatic qualification of eligibility criteria could reduce the workload and lead to higher efficiency compared to the manual process. 10
Since early 1990s, natural language processing (NLP) has been used as computerized data acquisition tool to recognize grammatical meaning, analyze (i.e., divide sentences and words into smaller parts) and synthesize (i.e., classifying it into groups) human speech. 11 Applied in oncology, algorithms based on NLP could be implemented to automatically extract clinical information for patient prescreening with high accuracy.12–14 Methods based on regular expressions (RegEx) with patterns’ recognition can extract a specific set of characters or a pattern in a piece of text or a text file with high sensitivity.15,16 RegEx is a NLP technique that recognizes a specific word (or sentence) in the human written text using a defined character string. This standard technique is created by software developers working with domain experts and supported by most programming languages. The RegEx can detect keywords in electronic health records to identify treatment or health condition.17,18 The free-text electronic medical reports represent one major information source for NLP. 15 Machine learning (ML) automatically classifying documents depending on lung cancer characteristics does already exist. 13
However, only few studies have attempted to develop lung cancer information extraction based on multiple NLP methods (RegEx and NBC).14,16 The novelty of this work concerns the combined method applied on a wide spectrum of eligibility criteria. To the best of our knowledge, no system based on a text classification model and rule based methods has been applied on MTM reports for various clinical and histologic characteristics. Using combined NLP approaches on MTM reports for 8 characteristics, this study aimed to develop and assess a prescreening tool in pulmonary oncology trials as a use case. Lung cancers were studied because of three practical concerns: large amount of MTM reports available as one of the most frequent cancer, standardized treatment strategies (TNM) that concerns first line chemotherapy because of the severity of the disease.
Material and methods
We performed a monocentric, observational study to implement and evaluate natural language processing models based on the reuse of clinical reports. Study population concerned all adult patients who underwent a MTM in thoracic oncology between 2018 and 2020 in one French university hospital. The MTM reports reused were stored in the Tours hospital clinical data warehouse (CDW) based on the eHOP® model.
19
For each class of the 8 characteristics (Appendices, Table 3), the method consisted of 2 steps: (i) RegEx implementation to extract a pattern for each class and (ii) multiclass classification by ML and (iii) evaluation of both methods, with estimation of performances metrics. A preliminary preprocessing step was necessary to normalize free text from MTM reports. The steps (i) and (ii) were performed on the same MTM reports between the train data and test data splits to compare the results. The step (iii) estimated 3 metrics: recall (also known as sensitivity), the fraction of relevant instances that were retrieved (equation (1)); precision (also called positive predictive value), fraction of relevant instances among the retrieved instances (equation (2)) and F-score, their harmonic mean (equation (3)).
Finally, for each multiclass factor, macro-average metrics were calculated (based on metrics computed independently for each class, treating all classes equally). Rare classes were as important as frequent classes to automate patient’s characteristics extraction easing subjects’ inclusion in trials (especially for rare patient and tumor profiles).
Free text preprocessing
Were extracted from the hospital data warehouse all the “thoracic MTM reports” filled between 2018 and 2020 (n = 1224). Only complete reports concerning “Tracheo Bronchial Tumor” (n = 1033) were selected. Manual annotation was performed on a sample of 640 records (62%) based on 2 distinct free text fields “history of the disease” and “history” in the MTM reports (Figure 1). Flow chart of Thoracic MTM reports from Hospital Data Warehouse Extraction to Annotated and Normalized Train and Test Corpora.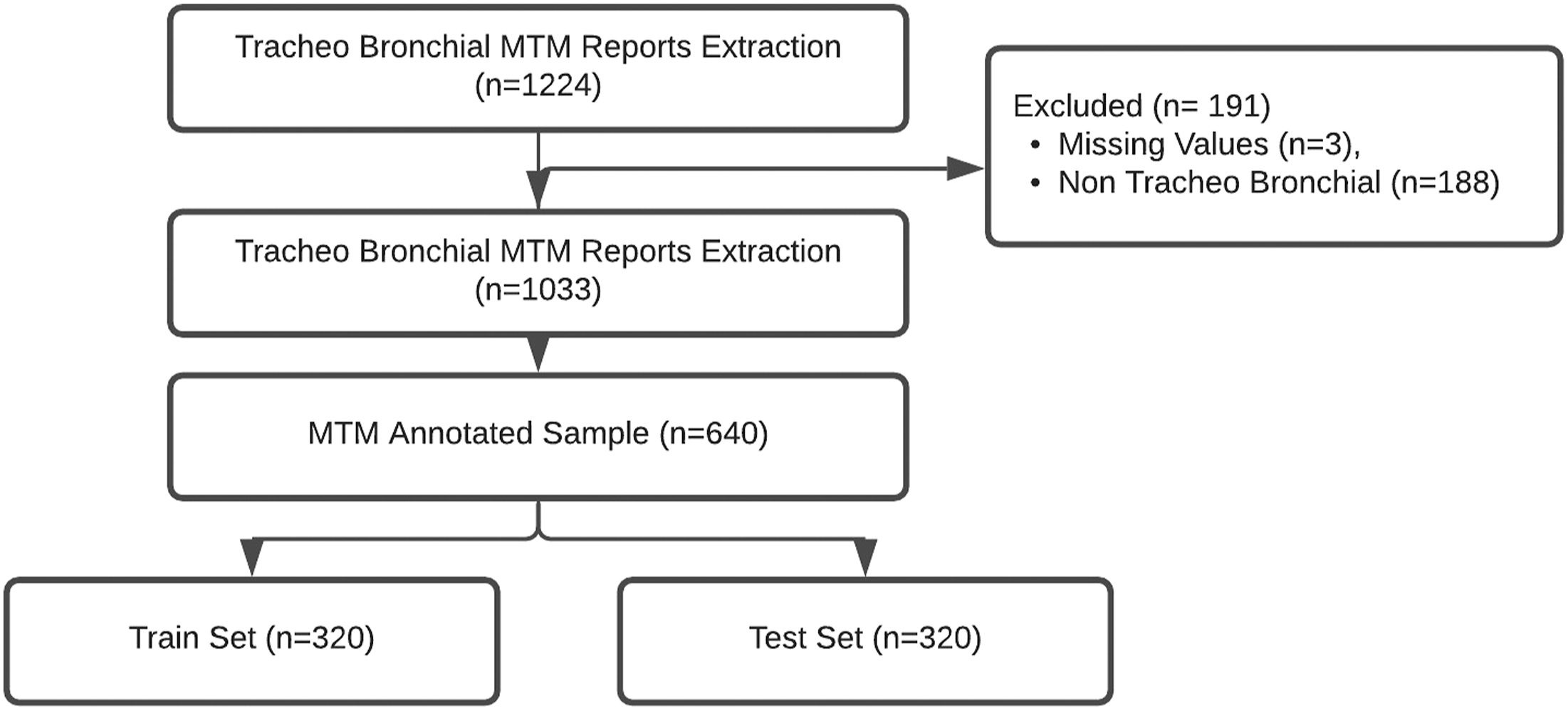
Eight characteristics concerning patient and tumor profile were extracted from the MTM: a) World Health Organization Performance status (WHO PS), b) tumor extension with the c) T, d) N and e) M classifications, f) TNM stage, g) histological type, criteria indicating the prescription of specific targeted therapies in lung cancers such as h) ALK gene rearrangement and i) EGFR receptor mutations.20,21
Each of the 8 factors had multiple classes, corresponding to international nomenclatures (appendix, Table 3). The “Histology” classes corresponded to one or more codes from the International Classification of Diseases, 10th edition for oncology “ICD-O 10”. The rarest histology types were gathered in the same “other” class. The factors “T”, “N” and “M” corresponded to the clinical tumor extension assessment (TNM 2017 classification) based on clinical and paraclinical examinations before any tissue sampling. The factor “TNM stage”, was imputed from the combination of the 3 factors “T”, “N” and “M” (according to the TNM 8th edition).
The molecular biology testing of ALK gene rearrangement and EGFR gene mutation testing had the same classes: positive test, negative test, absence of testing (i.e., “Not tested” or “nt”) and absence of information in report (i.e., “Not Retrieved” or “nr”). For each factor (but the WHO PS and the TNM stage), “nr” class was assigned by default, when none of the other classes were retrieved. Absence of information was considered as a class itself to train the system to recognize missing values also. Natural Language Processing (NLP) methods were performed on MTM free text to normalize grammar, spelling and abbreviation.
Regular expressions
To be recognized in the MTM free text, each class (n = 52) corresponded to a distinct pattern (n = 45) except for the deducted classes (n = 7, e.g. “no reference”, “not available”). Abbreviations, synonyms, spelling variations, confounding RegEx patterns (e.g., “t2” vertebra instead of “t2” stage) for the same class were taken into account. Concordance rates (false/true positives and false/true negatives) between the RegEx extractions and the gold standard (manual annotations) were calculated. At the implementation stage, multiple RegEx patterns (between 2 to 10 versions) were implemented step by step (addition/removal of specific or sensitive patterns) and tested on the training set (n = 320). Concordance rates were evaluated for the different versions of the patterns and the one with the best performance on the train set was finally chosen. At the evaluation stage, the RegEx and the ML models were once applied on the test dataset (n = 320).
RegEx were implemented using the R package “stringr”.
Supervised machine learning
Unlike the data split use in the ML literature, there was as much data in the train set as in the test set. As the data of the test set was never used to train the model, a category that was absent in the train test could not be assigned by the model. To be operated by machine learning programs turning textual data into tabular format was necessary. A document term matrix (DTM) was built from the observations of the corpus (320 MTM-lines) and terms from the train descriptor dictionary (5087 terms-columns). The dictionary presented all of the free texts terms extracted from the train set only, with the “bag of words” technique. 22
The DTM values (for each MTM report and each word of the dictionary, n = 320 × 5087) were the term frequency-inverse document frequency (TF-IDF) weights. They were calculated from the frequency of the term in the document weighted by the rarity of the documents containing this term (i.e. the logarithm of the inverse of the proportion of documents in the corpus that contains this term).23,24 The specificity of text categorization consisted of a very sparse DTM with many descriptors for a low number of observations leading to a vast majority of null values in DTM (97%).
To predict the class of each MTM reports, Naive Bayes classifier (NBC) models were performed. NBC is a family of ML linear classification model based on Bayes theorem. With strong independence assumptions between the parameters, Bayes theorem can be used for prediction of the characteristics as a classifiers decision rule. In our study, for each of the 8 characteristics, MTM reports have been classified based on their conditional probabilities to belong to the class. 25 For each of the 8 factors of interest, NBC models were implemented from the R package “naivebayes” (with the “naive_bayes” function) with specific parameters. 26 The TF-IDF numerical statistic was used to reflect how important a word was to a MTM in our corpus.27,28 Laplace smoothing function was used to avoid multiplying probability by “zero” in case of a new term in the test corpus.
Each NBC model was tested multiple times for different sets of Laplacian estimators to tune this hyper-parameter. At the first step, five Laplacian estimators’ values (from 0.001 to 1000) were compared according to the F1-score obtained by each of the five models. At the second step, a set of five values was tested framing the one with the highest F1-score. Eventually, the second step was once repeated. The optimal Laplacian estimator value was considered as the one giving the highest F1-score after these 3 steps.
Results
Distribution of the factors and classes in the MTM reports
Co-occurrence in multidisciplinary team meeting reports test sample for t, n and m factors.

For the majority of the corpus, a “TNM Stage” was imputed based on the presence of all three factors “T”, “N” and “M” for almost half of them (48,7%). Incomplete stage imputation was possible for 6% of the reports if at least the “M” factor was present in the text (e. g. each M1a are stage IV-A whatever their “T” or “N”). For the rest of the MTM reports (n = 45%), “TNM” stage was not imputed because of insufficient information about T, N and M in text (Table 1).
Performances metrics of RegEx and NBC
Macro-average performance metrics between the two methods.
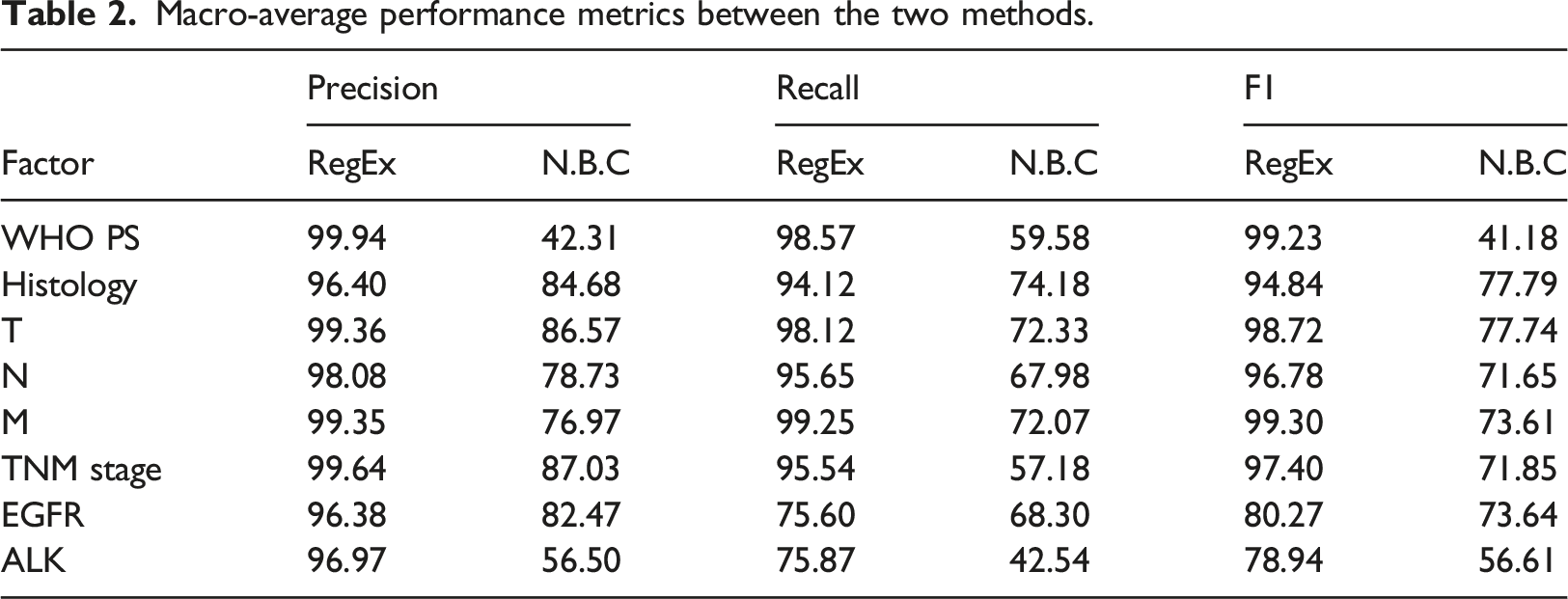
With the NBC method, two factors (“EGFR” and “T”) presented a macro-average precision rate higher than 85% and (“TNM stage”) higher than 90% for one factor. The factors associated to the lowest macro-average precision rates with NBC (“WHO PS” and “ALK”) obtained the main gain in performance rates with the RegEx (+57.6 and +40.5 respectively). The precision rates with RegEx method were all above 96%. The best precision rate over the two methods was for “WHO PS” (>99.9%).
Concerning the macro-average recall rates of the NBC method, they reached more than 74% for two factors (“Histology” and “TNM stage”). The lowest recall rates were obtained for “WHO PS” and “ALK” but they also presented the main gain in performance rate with the RegEx. RegEx method enabled higher recall rates (>76%) than NBC and 5 factors presented rates higher than 95% (“WHO PS”, “T”, “N”, “M” and “TNM stage”). The best macro-average recall rates were obtained for the “M” classification (99.2%), the “TNM stage” (99%) and the “WHO PS” score (98.6%). Over all the 8 factors, the best recall rate was for “M” classification with RegEx method.
The best F1-scores (>99%) were calculated for “M”, “WHO PS” and “TNM stage” with the RegEx method. With the NBC method, most of the factors presented macro-average F1-scores higher than 73%. The lowest F1-scores were obtained for “EGFR” mutation and “ALK” rearrangement tests results. Few factors (n = 4) presented precision or recall rates lower than 60% (Table 2). Overall, RegEx showed higher results than NBC supervised machine learning method (Figure 2). Macro-average F-score Per Factor compared between the Naive Bayes Classifier (NBC) model and the RegEx Method. For each factor, a mean of each of its classes F1-score rate is calculated to obtain (not weighted) macro-average F1-score Rate. When a F1-score Rate was not available for a class of a Factor, it was excluded from the calculation of the macro-average for the factor.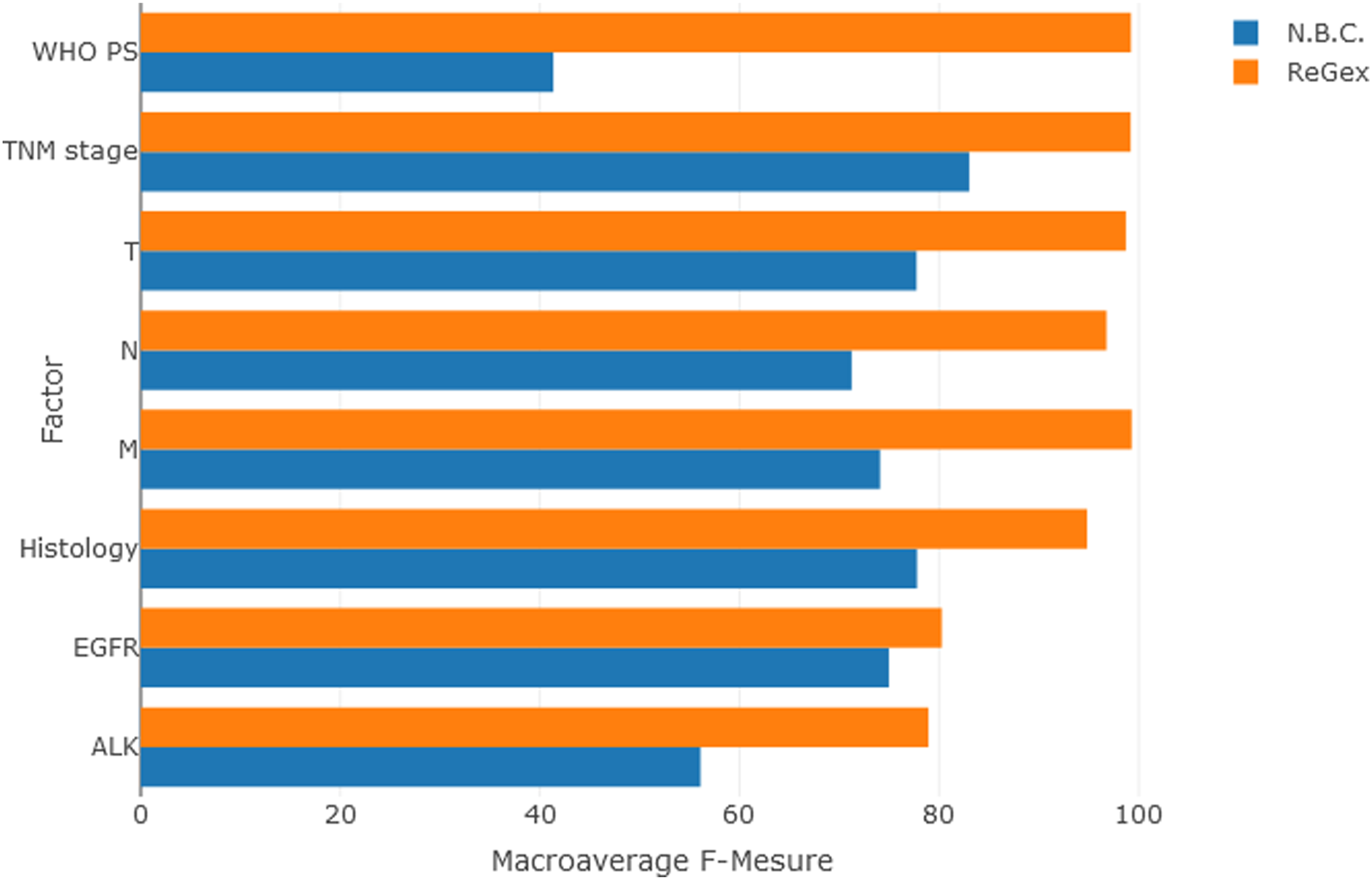
The macro average F1-score deltas between the two methods were smaller than 26 points except for the “WHO PS” and “ALK” factors. The NBC method failed to correctly classify the rarest classes especially (Appendix, Table 5, “WHO PS: 3”, “WHO PS: 4”,“ALK: 1” and “ALK: nt”). The class “EGFR: nt”, presented recall rate twice higher with the NBC than with the RegEx method.
Discussion
This study allowed the implementation and assessment of an automatization model based on NLP integrated in a Hospital Data Warehouse to ease information extraction about patients with lung cancer. The RegEx and the ML models could automatically and robustly screen electronic medical records to increase and accelerate recruitment for clinical trials, target therapy prescription especially in lung cancer and potentially improve patient outcomes. The NLP model could ease clinical trials prescreening task with high performances by classifying MTM reports on patient and cancer profiles. As a use case, eight different characteristics were extracted by RegEx with precision rates up to 99.9% and recall rates up to 98.6%. To classify cancer TNM stage, the NLP model showed higher performances in terms of macro-average recall (or sensitivity) and precision than published expert rules based and machine learning systems.12,13,15
Multiple tumor characteristics within multiple classes (n = 52 different information overall) individually were extracted with performance at least as high as that of validated rule-based classification systems for single task. 13 Information extracted referred to international standard terminologies used in healthcare systems, allowing the adaptation of this NLP model in different facilities. So far, the studies about NLP on clinical reports has been conducted more frequently on English or Chinese language as shown in the systematic review of Ford et al. 29
Heterogeneous RegEx performance was obtained across the different classes of each factor studied, depending on their frequency in reports or the structuration of the text in MTM report. For some factors, supervised machine learning models can represent supports for the rule-based methods as demonstrated with the “EGFR: nt” retrieval, doubling the recall rate.
However, NBC showed limitations to classify the rarest classes caused by inconsistencies in medical reporting. Because of their scarcity, two classes were missing in the training set (i.e., “Stage: Ia2” and “ALK: nt”) and were recoded “no retrieval” (“nr”). Moreover, the NBC model was based on the probability of being classified in a class for an observation also called “posterior probability”. This probability was proportional of the class occurrence in the text corpus “prior probability”, nearly null for the rarest classes.
The 8 factors annotated presented unbalanced distributions in text reports among their different classes (Appendix, Table 4). The most frequent classes were “WHO PS: 1” (48%) and “Histology: adenocarcinoma” (44%), “Tumor: t4” (21%), “Node: n3” (20%) and “Metastasis: m0” (21%), “Stage: Iva” (24%) in thoracic MTM reports. Concerning the genetic tests, the most frequent class was an absence of EGFR mutation (27%) as for ALK (37%). Among the 52 classes extracted in the MTM report sample, five of them were present in only 1% of the reports (T “tis”, TNM stage “Ia1” and “Ia2” and ALK “1” and “nt”). Information about EGFR status results were more frequently reported in texts than for ALK, as described in literature. 30
The study had some limitations. Because of the paper-based nature of the data source, all of the information exchanged in MTM might not be recorded in electronic forms. Since 2007 MTM reports are manually completed by the oncology coordination center, into the oncology communicating file with semi-structured fields. Data concerning facultative fields such as: stage TNM/pTNM, histology is less reported than mandatory fields such as: sex, age, tumor localization, OMS-PF, treatments, surveillance, G8 is most of the time reported.
The manual annotation method is questionable because it was realized by a unique annotator. The RegEx implementation was realized by the professional who did the annotations possibly leading to RegEx overfitting. This limitation implies RegEx adaptations processes to be applied on MTM reports from other centers. For the majority of the eight factors extracted, RegEx patterns were simple and short (two words at the most) because the texts were semi-structured. This represents an advantage to adapt RegEx on heterogeneous reports writing styles and reports structures.
Methodological choices about evaluation might also present several limitations. First, the NBC model was evaluated with holdout method whereas a stratified cross validation could have reduced the risk of variance in performance metrics’ estimations. The holdout method was necessary to respect the train and test split also used for RegEx implementation. Secondly, instead of weighted macro-average metrics often used for unbalanced classes, the evaluation concerned macro-average metrics. When a factor presented better results for its rarest classes (ALK and EGFR) than its frequent classes, the (not weighted) macro-average performances metrics were higher.
In the free text medical reports, abbreviations are common and different information can be written in the same way (e.g., “t1” for tumor stage classification or “t1” for first thoracic vertebrae). RegEx patterns implementation had to avoid confusion between patient medical history and actual cancer parts in text. Misleading patterns caused false positive results as shown in previous study using only a keyword search. 31 To avoid misclassification, we used a rule-based NLP algorithm that excluded confounding patterns (e.g., each “t4” adjacent to “vertebra”).
In conclusion, training the algorithm on a larger MTM report sample could increase recall and precision to recognize the rarest classes’ patterns, thanks to higher proportions of each class to extract. However, the MTM reports may vary between healthcare settings in text redaction structure, reducing the reproducibility of the performances. Further analyses must be performed to estimate its performances in subject selection for a clinical trial in real life, comparing its results with actual lists of selected subjects for trials conducted in centers.
To be applied on different hospital data warehouses, external validity of the algorithm has to be assessed. Adaptation of the algorithm on other cancer localization must be studied. The PreScIOUS study demonstrated the interest of combined artificial intelligence methods to accelerate the prescreening and selection of eligible patients with lung cancer in clinical trials. The major expectation of this tool will be to help researchers to include faster new eligible patients, reducing delay and improving prognostic, from routinely reported bio-clinical information of a hospital clinical data warehouse.
Supplemental Material
Supplemental Material - Prescreening in oncology trials using medical records. Natural language processing applied on lung cancer multidisciplinary team meeting reports
Supplemental Material for Prescreening in oncology trials using medical records. Natural language processing applied on lung cancer multidisciplinary team meeting reports by Marie Ansoborlo, Christophe Gaborit, Leslie Grammatico-Guillon, Marc Cuggia and Guillaume Bouzille in Health Informatics Journal
Footnotes
Acknowledgements
The content is solely the responsibility of the authors. Research reported in this publication was supported by the University François Rabelais Tours. We thank the Region Centre-Val de Loire, Region Bretagne and the Cancéropôle Grand Ouest (Oncology bid data sharing for Research, ONCOSHARe project).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
According to French regulations, all patients were informed of the potential reuse of their data for research purposes, could refuse to participate and no signed consent was required. Data from eHOP CDW are de-identified and linked to a unique anonymous identifier. The non-interventional researches from the Tours Hospital eHOP CDW were approved by the French Data Protection Agency (CNIL; N 2,212,853).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.