Abstract
Introduction
Interest in artificial intelligence (AI) and machine learning (ML) in healthcare is rapidly growing, and there is hope for incorporation of ML into everyday medical practice. 1 Natural language processing (NLP) is a particularly enticing tool for clinicians in search of important patterns buried in the massive amount of unstructured text stored in the electronic health record (EHR).2,3
In psychiatry, NLP is especially promising, as diagnostic criteria rely heavily on patient report and are often contained in the narrative text of a clinical encounter.4–6 Several excellent recent reviews have summarized the history of AI and ML, including NLP, and its use in psychiatry, which has accelerated in the past 10 years.3,4,7–12 A recent review showed a sizeable percentage of ML studies were from psychiatry (11%), with only imaging and cardiology specialties contributing a larger single proportion. 13 Experts have developed several powerful tools to make ML and NLP more accessible.14–20 There have been many successful demonstrations of NLP use for classification and prediction within a psychiatric clinical context, the breadth of which we cannot fully describe here.21–26 Studies of NLP in psychiatry typically use unstructured data from either the EHR or social media sites, with the goal of extracting or establishing a diagnosis, monitoring changes over time, or predicting onset of disease. 4
While NLP research in mental health has rapidly grown, there are challenges to incorporating NLP into routine clinical practice. Reasons include limited familiarity of ML to clinicians, limited availability of validated tools and methods at clinical sites, and prohibitive costs for widespread adoption of NLP into patient care. 7 Effective implementation of NLP requires knowledge and skills that are not regularly taught during medical training.27,28 The Informatics for Integrating Biology and the Bedside (i2b2) Center issued an NLP challenge in 2016 that demonstrated how complicated and time-consuming it can be to develop well-performing algorithms for psychiatric symptom classification. 29 The time constraints of clinical work make it challenging to overcome the learning curve that most clinicians face if they want to use NLP in research studies.
The Clinical Annotation Research Kit (CLARK) is an ML software kit that classifies patient charts based on unstructured data. It was developed for clinicians with varying degrees of expertise in NLP modeling and informatics. 30 CLARK has been successfully used to answer clinical questions within various medical specialties, but it has not been used in a psychiatry context.30,31 The primary aim of CLARK is to serve as an open-source, easy to use, graphical interface for machine learning-based classification of structured and unstructured patient health records. Broadly, CLARK takes inputs of unstructured clinical notes from patients of known health outcomes, paired with expert-defined lexicons of terms associated with the outcome, to train classifiers based on the occurrence of the terms within the notes. To do so, CLARK transforms the clinical notes into multi-dimensional feature vectors based on the given lexicons, and then applies a user-selected machine learning model on these vectors. After validating the model on a prespecified subset of the patients, the classifier is then applied to a held-out set of additional patients to assess performance metrics such as accuracy. Full technical details on CLARK are previously described. 30 CLARK can allow an end-to-end clinician-led process that relies on little outside technical expertise and can be rapidly implemented, thus circumventing several of the common challenges of NLP implementation mentioned above.
In this study, we tested CLARK’s ability to classify unstructured clinical notes of solid organ transplant recipients based on the presence or absence of a depression diagnosis and/or substance use diagnosis, and compared the results generated by CLARK to a gold standard classification by mental health experts. Additionally, to compare between classical statistical learning NLP methods and more recent transformer models, we compared CLARK’s performance to that of a bidirectional encoder representations from transformers (BERT) model from the open-source Hugging Face library. 32 The impact of pre-transplant psychiatric disorders on solid organ transplant recipient outcomes have been widely studied and debated.33–43 Psychiatric diagnoses are common pre-transplant; for example, 20%–40% of candidates will experience depression pre-transplant.33,34,39,40 However, accurate retrospective extraction of pre-transplant psychiatric diagnoses is a time-intensive task. While all organ transplant recipients undergo psychosocial evaluation, diagnoses are rarely included in the problem list. Therefore, the population provides ample representation of cases and controls; methods to accelerate accurate retrospective identification of cases from the EHR could benefit research efforts within the transplant community; and the results could inform future development of algorithms to identify psychiatric disorders in the wider population of medically ill patients. This work demonstrated the first use of CLARK in a psychiatry context and showed that an NLP software kit can aid research by clinicians with a wide range of knowledge in ML techniques.
Methods
All adults (age 18 or older) who received a solid organ transplant at a tertiary care center over a four-and-a-half-year timeframe were included in the analysis. The final dataset included 652 unique individuals.
Demographics, transplant wait list duration, and transplant-specific medical history were extracted from a central data repository containing clinical, research, and administrative data sourced from our health care system, with the ability to query most data elements as far back as mid-2004.
Psychiatric diagnosis was manually abstracted from the electronic health record (EHR) to establish a “gold standard” labeled dataset. The EHR was reviewed by a content expert and licensed psychologist and two graduate-level research assistants. Research assistants completed thorough training in standardized chart review methods to discern whether individuals met criteria for psychiatric disorders based on the Diagnostic and Statistical Manual of Mental Disorders, Fifth Edition (DSM-5). 44 To ensure inter-rater reliability, research assistants were required to demonstrate 100% agreement in data extraction of diagnoses in a parallel record review with the psychologist prior to completing independent reviews. After research assistants demonstrated consistent expertise, the psychologist completed continuous randomized reliability checks on 10% of all records. Depression diagnoses included Major Depressive Disorder (MDD), and Other Depressive Disorder (not meeting MDD criteria). SUD diagnoses included all SUDs other than Tobacco Use Disorder but did not include isolated episodes of substance use not meeting disorder criteria.
We trained models to detect the occurrence of substance use disorder or depression diagnoses in the EHR using CLARK, an open-source, interface-guidable NLP toolkit developed for computable phenotyping in unstructured data such as clinical notes. CLARK has been described in detail previously. 30 Broadly, CLARK takes input consisting of unstructured clinical notes paired with user-defined lexicons of regular expression (regex) terms. CLARK builds a classification model to link these terms with the potential outcomes and give a probabilistic confidence of the outcomes for each record. All modeling was performed using CLARK v1.0. We compared CLARK model performance to that of a BERT model based off of an open-source model, Hugging Face. 32
To create the corpus, we extracted clinical notes from our EHR and central data repository. Clinical notes were extracted from 2 years prior to the transplant date until the date of transplant and filtered by author credentials, author department, or encounter department to include those from psychiatry, psychology, and case management (as available). The corpus was duplicated to create separate classification models for depression and substance use diagnoses. Within each corpus, each record was labeled as case or control based on the “gold standard” generated by content experts. The corpus and gold standard labels for each diagnostic category were combined into a JavaScript Object Notation (JSON) formatted corpus for use with CLARK, and into tokenized inputs for use with Hugging Face. 32 We did not perform any language pre-processing techniques for CLARK analyses. For tokenization and use in Hugging Face, text was processed using the DistilBERT tokenizer 45 to convert to inputs usable with BERT models, and each note is reduced to the first 512 tokens in a sequence for compatibility and speed of model training with Hugging Face.32,46 Each fully labelled patient record was randomly assigned to one of two datasets: a dataset used to train the classification model (“training data set”) and a held-out evaluation (or “test”) dataset used to assess model performance. The distribution of categorical variables across training and evaluation datasets was assessed with Pearson’s chi-square; continuous variables were assessed with t-tests (if parametric) or two sample Kruskal-Wallis (Mann Whitney) tests (if nonparametric). Statistical significance was set at alpha = 0.05.
Regex terms developed for depression and substance use disorder models.

Within CLARK, we trained our classification models using 5- and 10-fold random versus stratified cross-validation and then applied a random forest, gaussian naïve bayes, or linear support vector machine classifier to predict each outcome. 47 CLARK uses the default setting for the Python scikit-learn Random Forest Classifier. 48 After achieving a satisfactory result with the training dataset, we input the evaluation dataset using the same classifier and cross-validation settings to evaluate the performance of the models.
Within Hugging Face, 32 we loaded the pretrained DistilBERT model for sequence classification, a faster and smaller model runnable on devices locally. 45 We set a learning rate of 0.00,002, batch sizes of 16 for both training and evaluation, weight decay of 0.01, and 50 epochs of retraining.
Performance metrics including prediction accuracy, sensitivity, specificity, balanced accuracy (average of sensitivity and specificity), positive predictive value (PPV), negative predictive value (NPV), Cohen’s kappa, and F1 were computed to assess performance. We also computed Receiver Operating Characteristic (ROC) curves and their corresponding area under the curve (AUC). To determine how well the models within CLARK would perform when a depression or SUD diagnosis was not available in the EHR problem list, the models’ performance were assessed when restricted to patients without the corresponding diagnosis in the problem list. Similarly, the models’ performance within CLARK were compared when restricted to recipients of kidney transplant versus other organ recipients.
Statistical analysis
Statistical analyses were performed using Microsoft Excel, 49 and R v. 4.2.1 50 ; the following R packages were used: pROC v.1.18.0, 51 ROCR v. 1.0-11, 52 caret v. 6.0-92, 53 and cvms v. 1.3.4. 54 Hugging Face hub 0.23.4 and CUDA 12.6 were used for the BERT model. Study data were collected and managed using Research Electronic Data Capture (REDCap).55,56
Results
Characteristics of solid organ recipients, stratified by training and evaluation sets a .
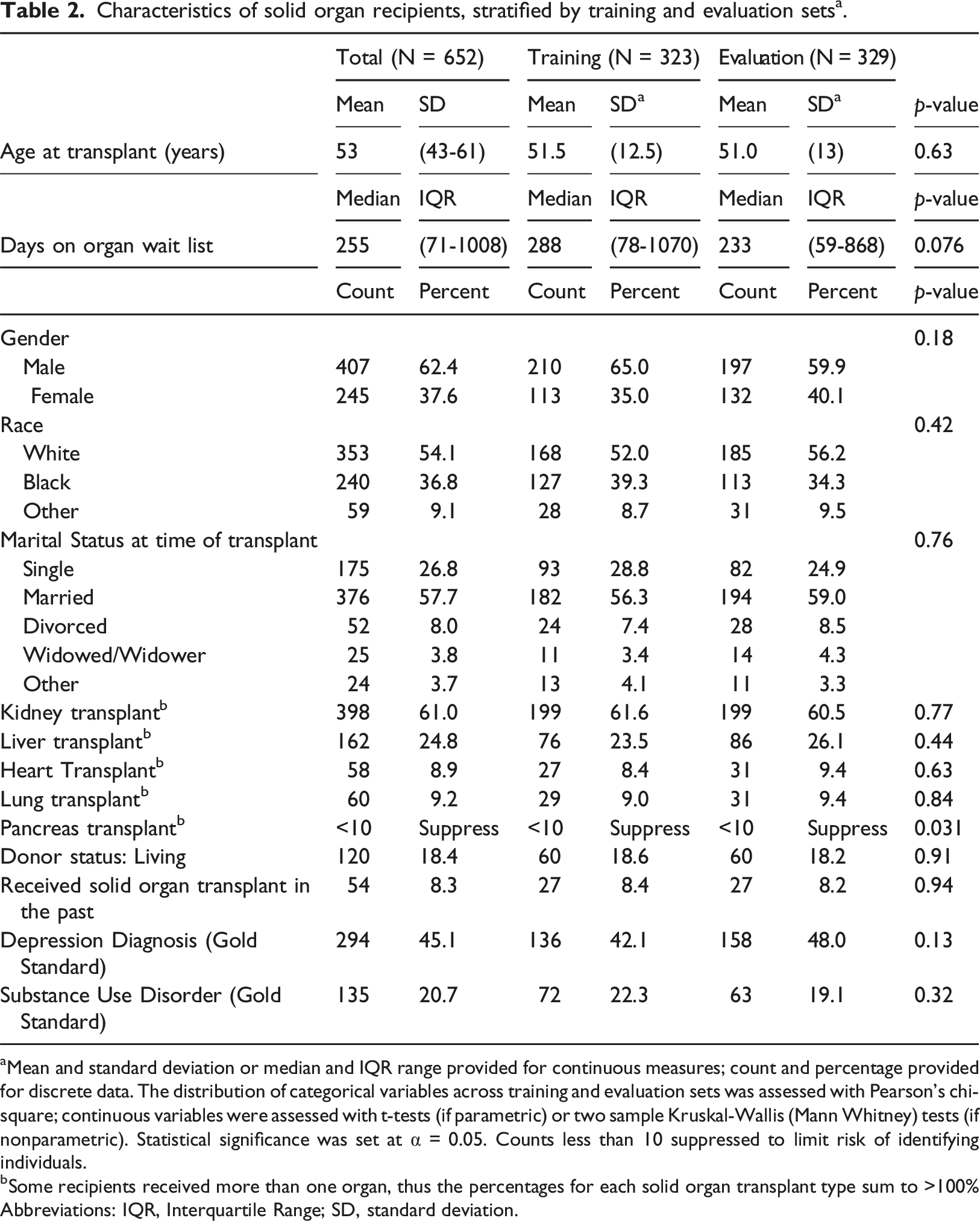
aMean and standard deviation or median and IQR range provided for continuous measures; count and percentage provided for discrete data. The distribution of categorical variables across training and evaluation sets was assessed with Pearson’s chi-square; continuous variables were assessed with t-tests (if parametric) or two sample Kruskal-Wallis (Mann Whitney) tests (if nonparametric). Statistical significance was set at α = 0.05. Counts less than 10 suppressed to limit risk of identifying individuals.
bSome recipients received more than one organ, thus the percentages for each solid organ transplant type sum to >100% Abbreviations: IQR, Interquartile Range; SD, standard deviation.
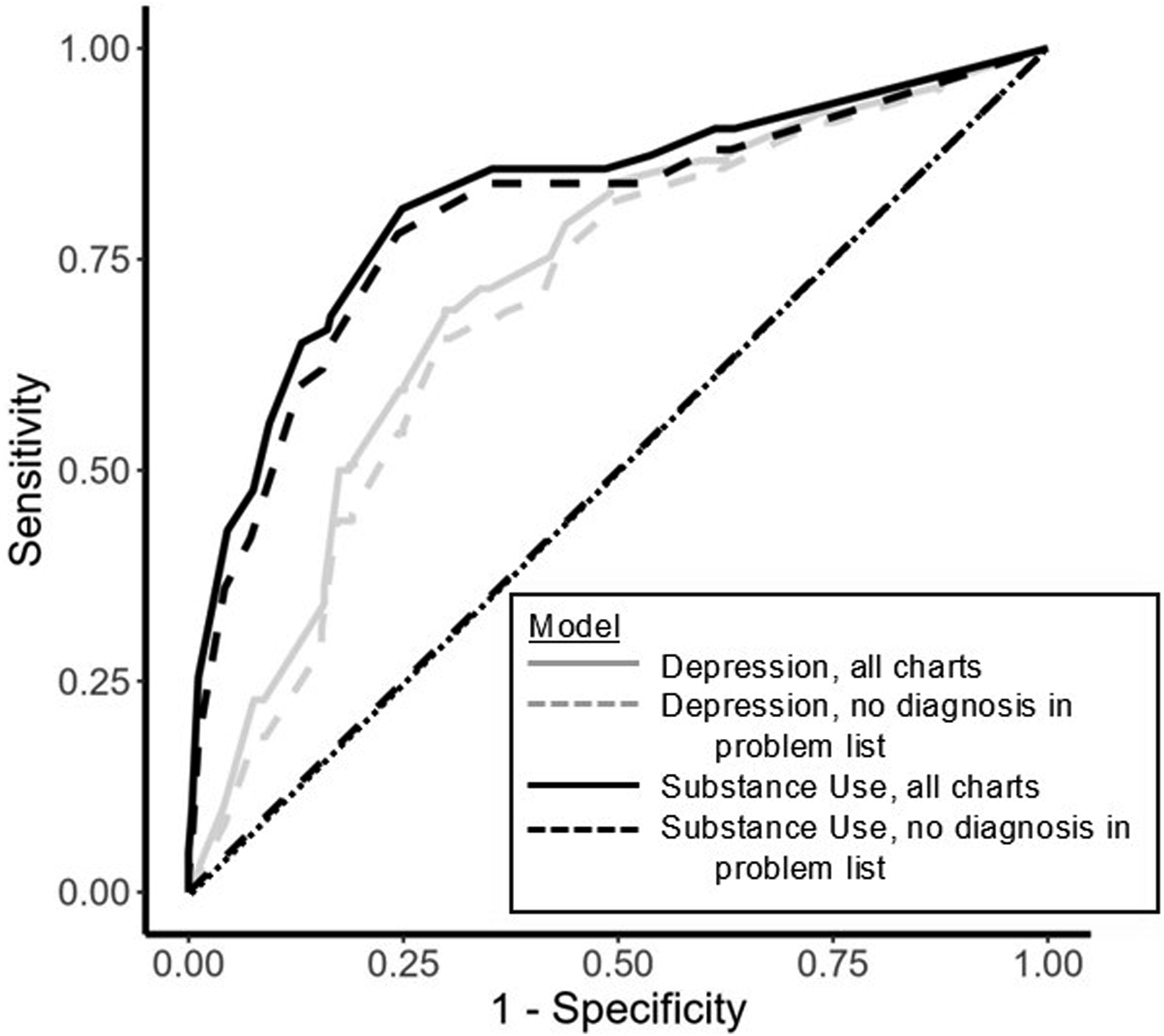
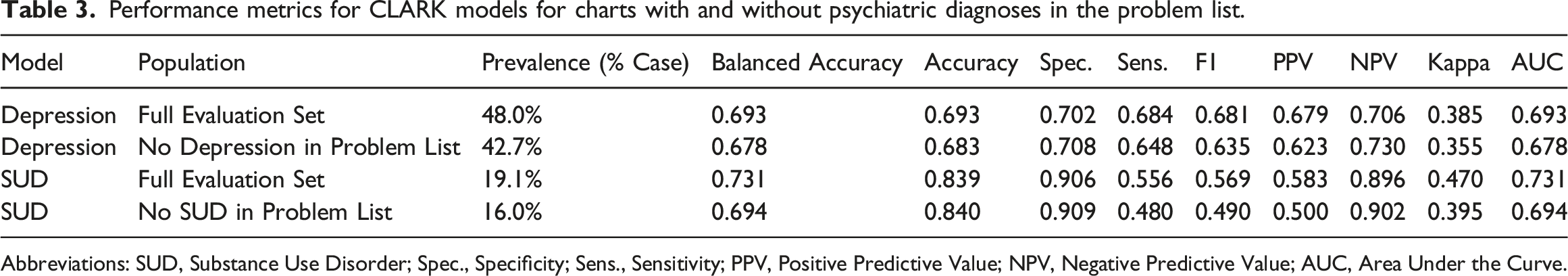
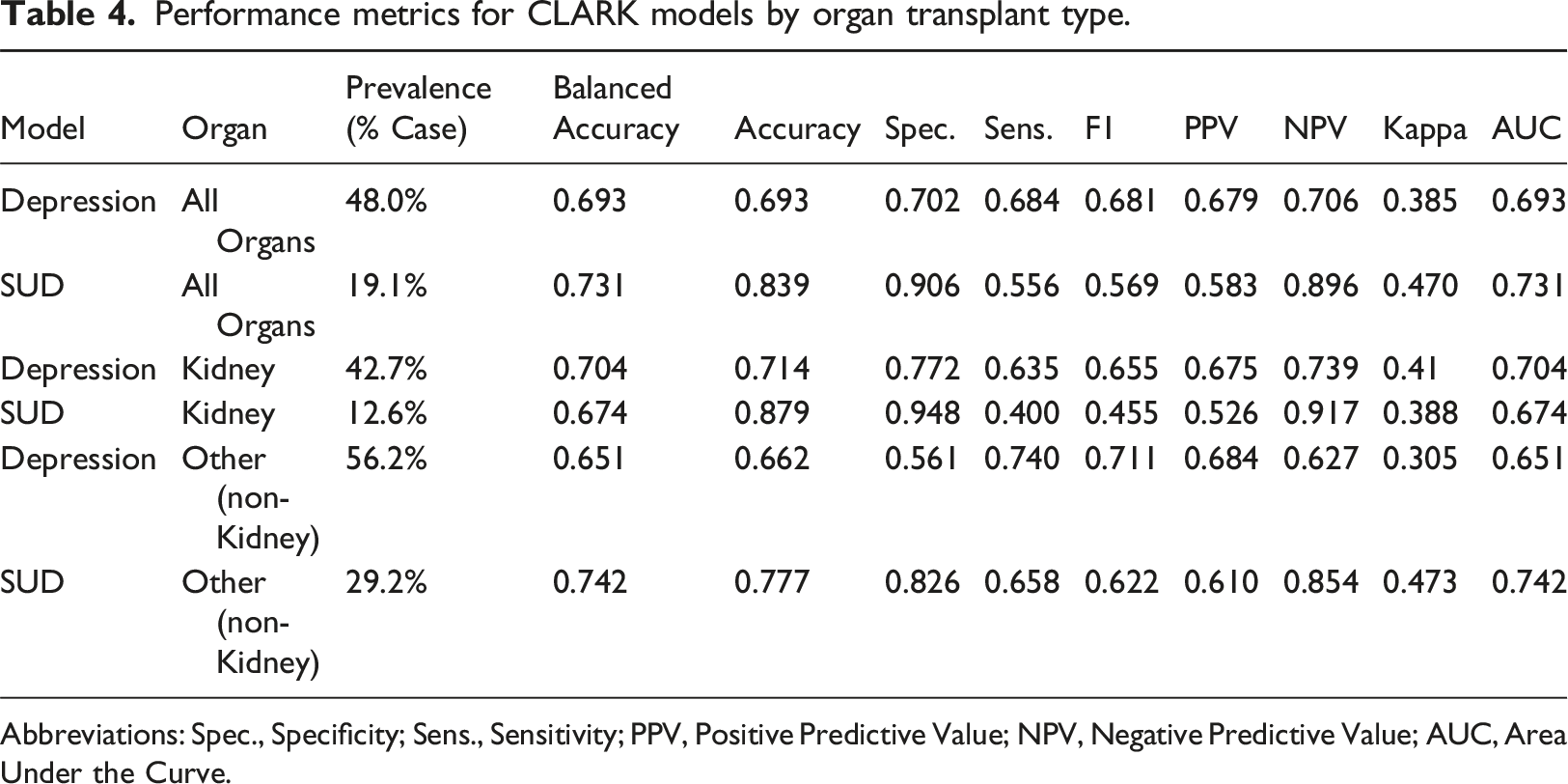
Within CLARK, using the random forest machine learning classifier with 5-fold random cross-validation, the SUD model had a specificity of 91% (and negative predictive value of 90%), but a sensitivity of 56% (and positive predictive value of 58%). Overall, the SUD model had an accuracy of 84%, balanced accuracy of 73%, and an F1 of 0.57, with AUC of 0.73 (Table 2, Figures 1 and 2). In contrast, the depression model had an accuracy of 69%, balanced accuracy of 69%, F1 score (0.68) and AUC (0.69) (Table 3, Figures 1 and 2). Sensitivity and specificity were comparable at 70% and 68%, respectively. In a subgroup analysis of patients without a mental health diagnosis in the EHR problem list, the models’ performance was similar to that of the main analysis (Table 3, Figures 1 and 2). In a subgroup analysis of kidney transplant recipients versus other organ transplant recipients, the depression model performance was similar across both transplant groups. The substance use model had a notably lower F1 for the kidney transplant subgroup, mirroring the very low substance use prevalence in this group (Table 4). Results using 5- (vs. 10-fold) random (vs. stratified) cross validation were comparable, emphasizing the robustness of our model across different choices of cross-validation hyperparameters; results from the random forest machine learning classifier were superior for both models compared to those using gaussian naïve bayes and linear support vector machine classifiers (data available upon request). Confusion Matrices for each CLARK Model. This figure provides a visual and numeric summary of each model’s performance for the full cohort (plots a and b) and for those charts without a depression or substance use diagnosis in the problem list (plots c and d, respectively). Depression models are in plots a and c. Substance Use Disorder models are in plots b and d. Receiver under the operator curves for the depression and substance use disorder models, by presence vs. absence of the corresponding diagnosis in the problem list. This figure provides the receiver under the operator curves for the following models: Depression, all charts (light grey, solid); Depression, no diagnosis in problem list (light grey, dashed); Substance Use Disorder, all charts (black, solid); Substance Use Disorder, no diagnosis in problem list (black, dashed). Performance metrics for CLARK models for charts with and without psychiatric diagnoses in the problem list. Abbreviations: SUD, Substance Use Disorder; Spec., Specificity; Sens., Sensitivity; PPV, Positive Predictive Value; NPV, Negative Predictive Value; AUC, Area Under the Curve. Performance metrics for CLARK models by organ transplant type. Abbreviations: Spec., Specificity; Sens., Sensitivity; PPV, Positive Predictive Value; NPV, Negative Predictive Value; AUC, Area Under the Curve.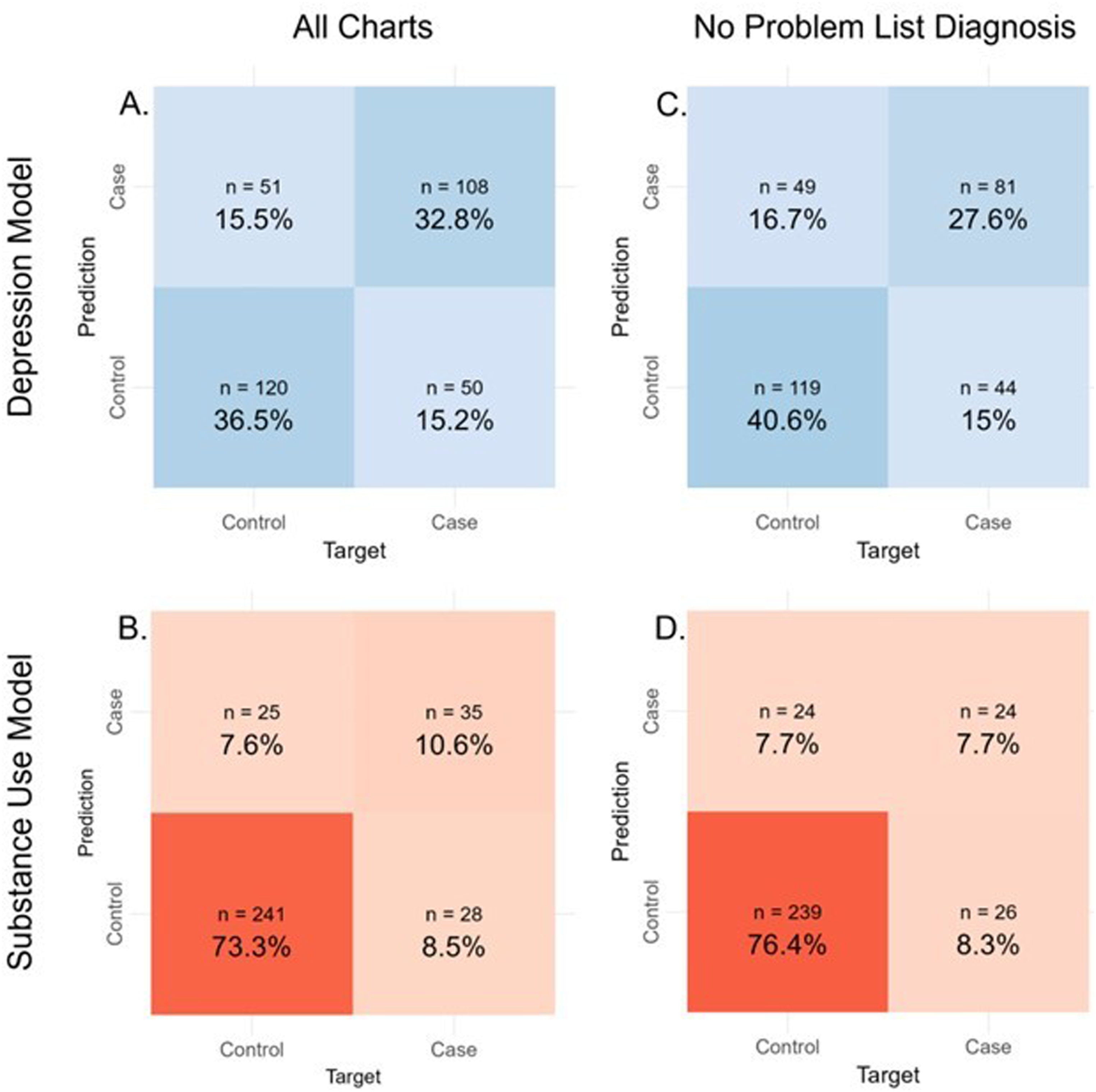
When comparing CLARK model performance to that of Hugging Face, 32 the BERT model did not perform as well as CLARK, though the patterns of performance across the two models were similar with a notably high specificity and low sensitivity for the SUD model, while the Depression model provided a more balanced performance (Supplemental Table 1). The BERT Depression model had notably low NPV, as compared to the SUD model which had high NPV but low PPV.
Discussion
This study describes the first application of the machine learning toolkit, CLARK, to classification of unstructured EHR data by psychiatric diagnosis. When comparing model performance, the depression model performed a more balanced job predicting the test set, with metrics for cases and controls being very similar. The low prevalence of SUD as compared to depression diagnoses led to a naturally higher SUD model specificity. The high specificity and NPV of the SUD model are in line with other studies diagnosing SUDs,57–59 but the model’s sensitivity of 56% (and positive predictive value = 58%), indicated the model struggled to identify those with substance use. Our models’ performance was not affected by the presence of a diagnosis in the problem list or medical history, as sensitivity analyses only including records without a depression or SUD diagnosis in the problem list revealed similar performance metrics for both models.
We aimed to test the performance of CLARK without the addition of other text processing or ML tools. Therefore, when compared to the existing literature, our study is also notable for its lack of pre-processing of text and reliance on unstructured notes. 4 Pre-processing text can improve model accuracy, but also represents a significant practical barrier for clinicians with a limited NLP skillset. 4 Overall, our models performed well given our approach, though it is difficult to directly compare results across the NLP literature given the numerous combinations of data, methods, classifier types, and platforms. 4
When considering barriers to building an accurate model performance, there are several issues to consider. There is a high prevalence of neurovegetative symptoms in the medically ill. Clinical notes for the medically ill often contain references to fatigue, low appetite, weight change, and poor concentration; all of which are also DSM-5 criteria for major depressive disorder. 44 This overlap in symptomatology could make it harder to find defining features in notes of individuals with or without a depressive disorder diagnosis. Aside from symptom overlap, words like “depressed” may show up within an illness description such as “depressed cardiac functioning.”
Bias and limitations
Given that our population was limited to solid organ transplant recipients, sampling bias was inherently present, and there may be characteristics unique to individuals who receive a solid organ transplant that impact our results. Our sample size was restricted to the population of transplant recipients over the specified timeframe; no sample size calculation was performed a priori. Race and ethnicity are known to influence psychotic disorder diagnosis. 60 When creating the gold standard, reviewers were not blinded to any aspect of the medical record, and unconscious bias related to patient demographics could have influenced their diagnoses. Because NLP of unstructured text inherently relies on the accuracy of the text, bias may be introduced by the author of the note 6 ; a recent study showed that clinicians may use different language in notes depending on patient’s race. 61 Although we do not have the issue of “black box” algorithm creation that happens in unsupervised ML, a supervised learning approach can still create or perpetuate bias depending on the quality of the input. 62
The notes in our corpus reflect the documentation style of transplant clinicians at a large academic medical center in the southeastern United States which uses one EHR, and generalizability of our lexicon may be limited by these factors. 3 The type of note templates and proportion of unstructured text relative to discrete data fields may also differ relative to other hospitals. Substance use can vary dramatically based on location, 63 making our lexicon subject to overfitting. The reviewers creating the gold standard could see each record’s problem list, medication list, and other structured data, along with more unstructured data than was extracted for analysis by CLARK. As a result, a record could have been labeled as positive for a diagnosis with data not included within the corpus CLARK was analyzing. While this was likely an uncommon event, it is important to consider when designing future studies involving NLP.
While CLARK is easy to navigate, the limited range of customizability and reliance on regular expressions are limitations. The user can choose between four classifier methods and quickly edit regular expressions to get the best combination of classifier and terms. However, using more complex, but widely used, preprocessing methods such as clinical Text Analysis and Knowledge Extraction System (cTAKES) allows investigators to take advantage of medical vocabulary dictionaries and phenotypes that have already been developed. 64 Using regular expressions without preprocessing can also lead to unintended inclusions, for example adding “thc” as a term without properly denoting word borders can pick up “forthcoming,” potentially leading to inclusion of a chart in the substance use category as a mistake. That being said, further use of CLARK for psychiatry diagnosis categorization could also lead to the development of more generalizable and accessible regular expression libraries.
Robustness is a key factor in model trustworthiness, 65 and while CLARK does offer four options for cross-validation of the algorithm, additional cross-validation options and validation using an external dataset would allow further demonstration of robustness when comparing results. We had just one dataset available, which prevented us from comparing performance on data from other institutions.
At a more fundamental level, most NLP tools require an English corpus, 3 making it difficult to include unstructured data containing direct quotations or wording from non-English speaking patients in the US. Additionally, our classification approach was based upon the United States (US) based classification system of psychiatric disorders, the DSM-5, as opposed to an international system. Even within the US, there has long been disagreement regarding the classification of psychiatric disorders, with other systems such as the Research Domain Criteria (RDoC) proposed as alternatives that rely on neurobiological mechanisms. 66
Newer learning models are being increasingly used in NLP research involving mental health, including transformer-based methods. 67 There are potential healthcare uses for transformer-based pretrained language models, such as BERT, and large language models (LLM), such as ChatGPT, but there is a large cost in terms of time, expertise, and data storage needed for training these models. 67 Indeed, when comparing CLARK performance to a BERT model based upon the open-source Hugging Face model, 32 CLARK demonstrated a more robust overall performance. Notably, many open-source BERT models, including Hugging Face, 32 require equal character length across all samples, either truncating or “padding” input in order to optimize model processing speed. 46 In general, the more accessible options (such as Hugging Face) have shorter character limits, and this likely affected our BERT model’s performance when compared to CLARK, which can manage input with variable character length. Running models that can account for longer sequences, such as our EHR free-text notes, are significantly more computationally expensive. In addition to these barriers, there remain substantial concerns about bias, privacy, and interpretability when using deep learning in healthcare. 67 While there is ongoing effort to make LLMs more usable and more transparent, in our experience, this type of learning model is a less accessible way for a clinician to perform text classification.
Clinical application and future Directions
CLARK’s approach to ready-made NLP software could be a key step to make ML and NLP techniques accessible at early levels of training. Medical students and other trainees could develop targeted projects using CLARK and become better versed in NLP as they advance in their career. CLARK is an excellent toolkit that provides an introduction to coding through regular expressions. As CLARK is used more broadly, more regular expressions libraries can be created and shared. Free toolkits like CLARK could help overcome several known barriers to widespread NLP use. 7
Explainability of identified features is important for being able to interpret results from machine learning algorithms, and therefore enhances how trustworthy the model is. 65 While CLARK does allow review of each patient record to see the highlighted regular expression features, it does not include a built-in method to determine the importance of each feature on the prediction outcome. The addition of more feature importance metrics to CLARK would both help the clinician refine feature selection and improve explainability.
CLARK has several applications for mental health clinicians. We used NLP models to classify medical records by diagnosis. A similar approach could identify specific diagnoses within a population. If combined with expert review, using CLARK or similar software with a higher confidence threshold could be especially helpful for sorting study participants based on psychiatric diagnoses or for screening large patient databases for population health or research purposes. Indeed, CLARK could be used as part of a two-step screening process: charts not meeting an established confidence threshold could be then flagged for further analysis by a clinician. Of note, the confidence of assignment to a certain outcome, such as depression or SUD, depends on the predictive model being used. In the random forest-based model within CLARK, confidence is defined as the proportion of decision trees that predict a patient to have a certain outcome.68,69 For the Linear Support Vector Machine and Gaussian Naive Bayes models within CLARK, the confidence thresholds were generated using the Python sklearn function “predict_proba” to generate the confidences (or probabilities) for each of the classifier types. 70 Confidence for BERT is calculated by taking a softmax operator (a transformation to turn a score to a probability) on the logits (the raw prediction scores from a Transformer) output by the model. 71 CLARK allows users to set the confidence threshold cut-off; by raising the confidence threshold, only records meeting this threshold are included in the output and the model accuracy is drastically improved. A confidence threshold can also be set in order to maximize performance over specific model metrics, such as accuracy or specificity. However, as this leaves a portion of records uncategorized, we opted to keep the default confidence threshold setting for our analysis.
CLARK does not replace the expertise of a clinician. In our population under study, the transplant population, the decision to place a patient on the waiting list for a solid organ transplant is a life and death decision. Any data incorporated into this decision must be of the highest accuracy, thus NLP methods do not appear appropriate for this clinical application at this time. However, in a research scenario, it could lessen the work burden on manual reviewers and focus the efforts of statistical or informatics experts when there are limited time and funds for these supports.
Conclusions
The role for NLP in psychiatry will likely only grow. Experience with NLP will enable psychiatrists to understand the strengths and limitations of this tool. Software for NLP will have to find the best balance between accessibility and accuracy, as well as accessibility and generalizability. Our work has demonstrated that highly accessible software toolkits can produce helpful and meaningful results. Unstructured data can be cumbersome to manipulate, but incorporation of NLP techniques with unstructured data has been shown to improve algorithm results. 64 As NLP is used more broadly in psychiatry, the classification schemes may even inform a better overarching classification model for our field.
Supplemental Material
Supplemental Material - Using a natural language processing toolkit to classify electronic health records by psychiatric diagnosis
Supplemental Material for Using a natural language processing toolkit to classify electronic health records by psychiatric diagnosis by Alissa Hutto, Tarek M Zikry, Buck Bohac, Terra Rose, Jasmine Staebler, Janet Slay, C Ray Cheever, Michael R Kosorok and Rebekah P Nash in Health Informatics Journal
Footnotes
Author contributions
A.H. drafted and edited the manuscript and created and tested the regular expressions in CLARK. T.Z. performed statistical analyses, drafted sections of the manuscript, and edited the manuscript. B.B. provided education and guidance around use of CLARK as applied to this particular project, including formatting EHR data for use. T.R., J.S., and J.S. created the gold standard. T.R. edited the manuscript. C.R.C. collected key aspects of the dataset and edited the manuscript. M.K. provided guidance around analyses performed. R.P.N. generated initial project concept and provided project leadership, collected data, generated the gold standard, edited the manuscript, and contributed to statistical analyses. All authors gave approval for the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by the National Center for Advancing Translational Sciences (NCATS), National Institutes of Health (UL1TR002489), and the Foundation of Hope for Research and Treatment of Mental Illness (Seed Grant). Michael Kosorok and Tarek M. Zikry are supported by UL1TR002489. Tarek M. Zikry is additionally supported by NIH (F31HL156464-03). Rebekah Nash is supported by the National Institute of Mental Health (K23MH128613), the Foundation of Hope for Research and Treatment of Mental Illness (Seed Grant), and the Doris Duke Charitable Foundation grant # (2020143). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH or other funding sources.
Ethical statement
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
