Abstract
Background
Prompt diagnosis of bacteremia in the emergency department (ED) is of utmost importance. Nevertheless, the average time to first clinical laboratory finding range from 1 to 3 days. Alongside a myriad of scoring systems for occult bacteremia prediction, efforts for applying artificial intelligence (AI) in this realm are still preliminary. In the current study we combined an AI algorithm with a Natural Language Processing (NLP) algorithm that would potentially increase the yield extracted from clinical ED data.
Methods
This study involved adult patients who visited our emergency department and at least one blood culture was taken to rule out bacteremia. Using both tabular and free text data, we built an ensemble model that leverages XGBoost for structured data, and logistic regression (LR) on a word-analysis technique called bag-of-words (BOW) Term Frequency-Inverse Document Frequency (TF-IDF), for textual data. All algorithms were designed in order to predict the risk for bacteremia with ED patients whose blood cultures were sent to the laboratory.
Results
The study cohort comprised 94,482 individuals, of whom 52% were males. The prevalence of bacteremia in the entire cohort was 9.7%. The model trained on the tabular data yielded an area under the curve (AUC) of 73.7% for XGBoost, while the LR that was trained on the free text achieved an AUC of 71.3%. After checking a range of weights, the best combination was for 55% weight on the XGBoost prediction and 45% weight on the LR prediction. The final model prediction yielded an AUC of 75.6%.
Conclusion
Harnessing artificial intelligence to the task of bacteremia surveillance in the ED settings by a combination of both free text and tabular data analysis improved predictive performance compared to using tabular data alone. We recommend that future AI applications based on our findings should be assimilated into the clinical routines of ED physicians.
Introduction
Community acquired bacteremia is a considerable cause of in-hospital mortality
Bacteremia is a life-threatening condition with very high rates of mortality if left untreated. 1 The 28-day mortality rate for bacteremia can be as high as 13.2% 2 ; therefore a prompt diagnosis and treatment of bacteremia are of the utmost importance for patients. A bacteremia diagnosis relies upon taking blood cultures from the patients early on in their workup, prior to antibiotic administration. The median time for a blood culture to arrive at the microbiology laboratory from the time the blood was drawn is approximately 3.5 h. 3 After arriving at the lab, the culture needs to be incubated and prepared for analysis. The average time from starting incubation to first clinical finding range from 1 to 3 days. 4 A study performed in Germany with a cohort of over 300,000 emergency department (ED) visits showed that 88% of patients were discharged within 6 h from ED arrival; 5 therefore, it is clear that blood culture results could not serve the physicians in their decision whether to hospitalize or discharge patients.
Artificial intelligence appliance in the task of bacteremia diagnosis
Alongside a myriad of scoring systems for occult bacteremia prediction, 6 efforts for applying artificial intelligence (AI) in this realm are still preliminary: Tsai and company demonstrated the ability to attain an adequate rate of bacteremia prediction among patients in the setting of the emergency department. Nevertheless, they included only febrile patients in their analysis. The AI methodology they used included two AI models applied on the same dataset: random forest followed by logistic regression (LR), and these resulted in area under the curve (AUC) values higher than 70%. 7 Febrile patients were also the target population of another study by Tsai and company, although this time they targeted pediatric patients. Here also, they established the ability to use AI for the purpose of better predicting those children coming to the ED with bacteremia. 8 Lee and company applied another AI methodology (MLP, multi-layer perception) to detect bacteremia in a retrospective manner on the whole population of two tertiary hospitals. They also achieved a considerably high AUC and even succeeded in predicting specific pathogens, namely, Acinetobacter in cases of pneumonia. 9 Choi and company applied AI algorithms on a pre-determined data set (42 items) and succeeded in achieving a high AUC for bacteremia prediction. They used an extreme gradient boosting model and compared it with the random forest and multivariable logistic regression models. 1 All of the above studies concentrated on AI-based analysis of multiple patients’ numeric parameters, but none of the above studies analyzed the potentially abundant and rich yield of the free-text data within patients’ electronic medical records (EMR).
Aim of the current study
The aim of the current study was to a apply state-of-the-art AI tools in the aforementioned realm; prompt, ED diagnosis of occult bacteremia. We used a large cohort collected in Israel's largest tertiary medical center for the purpose of development and validation of a combined AI methodology. We included an extreme gradient boosting model (XGB) with Natural Language Processing (NLP) algorithm that would potentially increase the yield extracted from clinical ED data, for example, when teeth chattering, noted as shivering, is the only textually documented symptom in patients’ EMR. We aimed to detect those patients that eventually had positive blood cultures (blood stream infection, BSI), either hospitalized or discharged.
Patients and methods
Study population
Our investigation focused on a retrospective cohort of adult patients, aged 18 years and above, who addressed or were referred to the emergency department care (ED) of the Chaim Sheba medical center between January 2017 and September 2023. Patients’ medical records, which serve for clinical purposes and are therefore considered highly reliable, were addressed, and patients’ data were retrieved only after an institutional review board (IRB) permission was granted (# 0348-23-SMC), and patients’ consent was waived due to the retrospective nature of this study.
Blood cultures were taken from these patients to assess for bacteremia, identifying 7414 hospitalizations (12.6%) and 1769 discharged cases (4.9%) with positive blood cultures. The study cohort comprised 94,482 individuals, of whom 52% were males. Of the whole cohort 38% (35,815) were discharged from the ED after initial workup. Of the discharged patients, 1769 were later confirmed positive for BSI, accounting for 19.2% of this subgroup. When considering the entire cohort, the prevalence of bacteremia was 9.7%, reflecting the significant impact of this condition on the emergency healthcare landscape. Figure 1 shows the CONSORT flow of patients in our cohort.
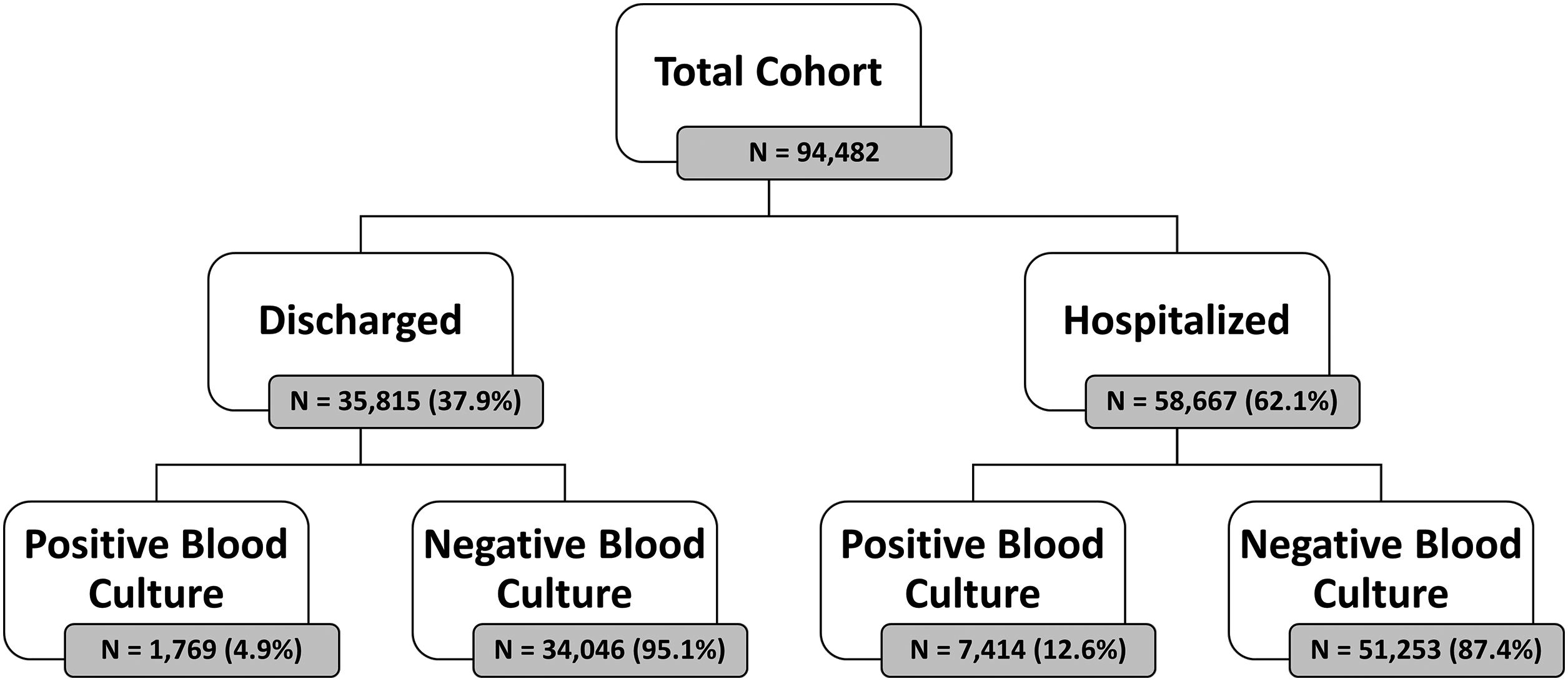
CONSORT flow of patients.
Patients’ data management
In a retrospective analysis of patients’ data, we defined a blood culture as positive if it came back positive for at least one of the following tests: gram stain (5.7% turned positive), anaerobic blood culture bottle (6.8% turned positive) and aerobic blood culture bottle (7.3% turned positive). We extracted the patients’ tabular data including demographic data, triage measurements, ED ESI (emergency severity index) score, time of admission, laboratory results, alongside free text data from the nurse's triage notes and the ED physicians’ clinical free texts, all originally written in Hebrew. Segmenting Hebrew words into parts is more challenging compared to English. To address this, we found that using TF-IDF on Hebrew text was an effective solution. We leveraged the characteristics of TF-IDF by defining the maximum and minimum thresholds, which allowed us to exclude high frequency terms such as stop words as well as very low frequency words.
All tabular data was converted into a numeric format. This transformation was crucial in order to insert data into the machine learning (ML) algorithms. There was no need of filling null fields as the selected algorithm could deal with empty values. Some patients were called back to the ED from their homes, after discharge, when their blood cultures’ results came back positive. These events were not taken into account in order to ensure data integrity and relate only to the first ED encounter. Also, we excluded erroneously created or duplicated patient records.
Statistical analysis
Machine learning models
Using both tabular and free text data, we built an ensemble model that leverages XGBoost (eXtreme Gradient Boosting) for structured data, and logistic regression (LR) on a word-analysis technique called bag-of-words (BOW) TF-IDF, for textual data. All algorithms were designed in order to predict the risk for bacteremia in ED patients whose blood cultures were sent to the laboratory. The algorithms were programmed using Python (Version 3.8.3 64 bits) and the XGBoost open-source library (version 1.6.1) with the scikit-learn wrapper (version 1.3.1). The XGBoost model, a form of gradient boosting, is a powerful machine learning algorithm that leverages the concept of training multiple weak learners, specifically tree-based classifiers, to augment each other and yield superior results. In this process, at each stage, a new decision tree is learned with the specific aim of correcting errors made by the existing ensemble of trees. The strength of the XGBoost algorithm lies in its robust prediction capabilities, achieved through an iterative process of prediction summation. Each decision tree in the ensemble is designed to fit the residual error of the prior ensemble, thereby continuously improving the model's accuracy. With the free text data, we employed a multi-step NLP (natural language process) to analyze the text. Initially, all of the relevant text notes associated with each ED visit combined into a single comprehensive text string for each ED encounter. Then, the text is processed using the BOW approach, which converts the text into a vector representation, where each row corresponds to a document. By that, we can convert every text into a vector representation. Following this, we applied the Term Frequency-Inverse Document Frequency (TF-IDF) technique into this matrix. TF-IDF is a statistical measure that evaluates how relevant a word is to a document within a large collection of documents (the corpus). The resulting TF-IDF matrix, which now held numerical representations of the text data, was then fed into a logistic regression model, which is known as a very efficient tool in classification assignments. The last step included running a range of weights between the model predictions to find the best combination that brings the best performances.
Models training and testing
The complete dataset generated, as described earlier, was split randomly to a test set of 18,897 (20%) samples and the remaining 75,585 (80%) samples were split randomly to a train set of 60,468 (64%) and a validation set of 15,117 (16%). The XGBoost hyper-parameters that we used in the training were: n_ests = 100 (ultimately delivering the best performance without overfitting), max depth = 4, n_ min_child_w = 50. The XGBoost model handled imputations of missing values. The TF-IDF hyper-parameter sets were min_gram = 2, max_gram = 4, min_df = 50, max_df = 0.5. The maximum features were limited to 10,000. The logistic regression was based on the defaults’ hyper-parameter except max_iter that was defined to 1000. By trying to optimize the model we trained the initial logistic regression with 10,000 features. Then we excluded the words that included only numbers and selected the top 8000 words that had the absolute highest coefficients. With that vocabulary, we trained the final model once again.
The model's performance was assessed using the AUC (area under the curve) metric. We have calculated the AUC both for training, validation, and test sets. Youden's index was employed to find the optimal cutoff point on the ROC curve in order to calculate sensitivity, specificity, false positive rate (FPR), negative predictive value (NPV), and positive predictive value (PPV) of the final models. We also used the cross-validation technique in order to ensure that there was no overfitting and no leakage of information in the models.
Results
Descriptive differences between groups
Detailed demographic and clinical data of the study population are presented in Table 1. As anticipated, patients whose blood cultures turned positive, were older, had lower ESI scores, and had higher incidence of hospitalization from ED rather than discharge. They also had higher body temperature, heart rate and respiratory rate and lower systolic and diastolic blood pressures although with only minuscule differences in absolute values. Regarding the laboratory parameters they had higher CRP blood concentrations, higher absolute neutrophil counts, worse kidney function tests and lower blood albumin concentrations. Although differences between our study groups achieved statistical significance, these should be attributed to the size of our study cohort rather than to the absolute values’ differences.
Patients’ characteristics according to blood cultures’ results.

ESI: emergency severity index; N: number; SD: standard deviation.
Model explainability and main features
The impact of each one of the features on the final model output is presented in Figure 2, using a Shapley additive explanations (SHAP) plot. SHAP is a method to explain the feature importance of machine learning models. It is based on the concept of shapely value in game theory (named after Lloyd Shapley who first described it in 1951 10 ). The shapely value is the solution to a problem where a group of players play a game and get a certain result. The values are the relative contribution of each player to the result. The meaning of “additive explanations” is that the explanation is made by running the model while adding features one at a time until the result is reached. This contrasts with other explanation models, for example, models which run all the possible feature combinations and compare them to the result. The features are ordered according to their ranking importance, from top to bottom, regarding their contribution to the raw probability of blood cultures coming back positive. The horizontal spread pattern represents the impact degree on the final model probability (SHAP value). The horizontal location represents the direction each value affects the final model probability—dots located on the right of the plot increase the probability of the model's prediction for positive BSI, and dots located on the left decrease the probability of its prediction. Red dots represent high parameter value, while blue dots represent low values.
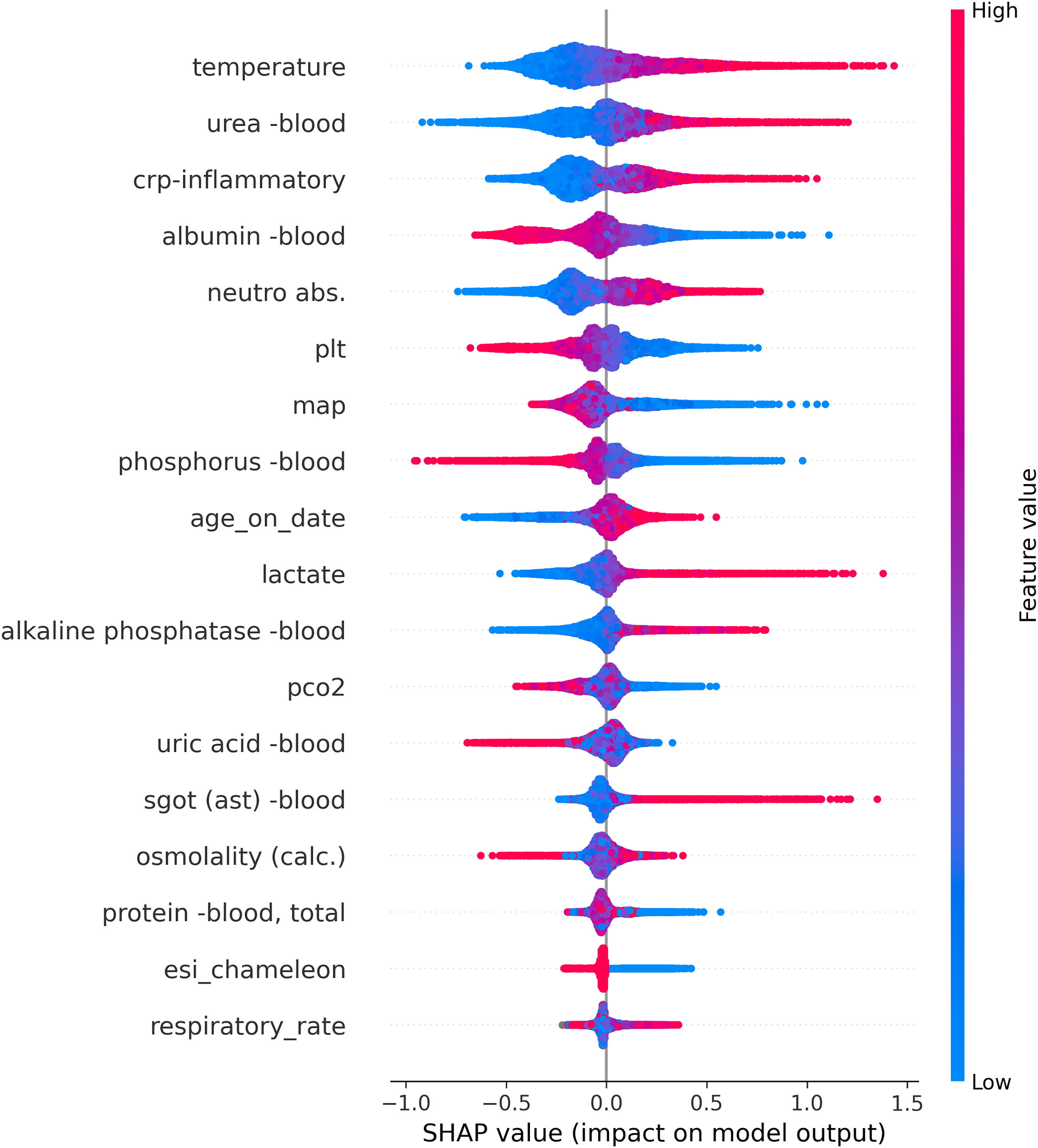
SHAP summary plot ranks each numeric feature by their impact on predictions.
As shown in Figure 2, the model analysis of the structured data delineated factors such as age over 80, fever, hypotension, elevated blood urea nitrogen, and lactate levels as significant predictors of positive blood cultures.
Similarly, Figure 3 shows the SHAP values for the NLP-algorithm’s chosen wordings according to their predictive values for positive or negative blood cultures. Note that the algorithm was trained on Hebrew texts, while the wordings in this figure represent an English translation of the original Hebrew wordings.

SHAP summary plot ranks each textual feature by their impact on predictions.
AI-driven performance: evaluating XGBoost, LR, and the ensemble model
The model trained on the tabular data yielded an AUC of 73.7% for XGBoost, while the LR that was trained on the free text achieved an AUC of 71.3%. After evaluating various weight combinations ranging from 45% to 70% for the models, all ensemble models performed similarly, with an improvement over the baseline models (based on tabular or text data). The selection of the weights was carried out according to the research question, taking into account the specific performance indicators of sensitivity versus specificity. The optimal combination was found to be 55% weight on the XGBoost prediction and 45% weight on the LR prediction. The final model prediction yielded an AUC of 75.6% as shown in Figure 4.

AUC curves for the three models’ performances.
The addition of free text into the tabular data improved the predictive performance compared to using the tabular data alone. The sensitivity, specificity, positive-predictive-value (PPV), and negative-predictive- values are presented in Table 2.
Evaluating classification metrics across XGBoost, LR, and ensemble model.

Abbreviations: AUC, area under the curve; LR, logistic regression; NLP, natural language processing; NPV, negative predictive value; PPV, positive predictive value; TNR, true negative rate; TPR, true positive rate.
Discussion
Rule-based bacteremia predictions
Prompt identification of patients at high risk of bacteremia during their stay in the emergency department, as it is with other life-threatening medical conditions, is very important and has a potentially profound impact on the ED physicians’ diagnostic and treatment measures taken. Previous publications addressed the potential of harnessing AI in the realm of bacterial infections, not necessarily bacteremia, in the setting of ICU, 11 for the purpose of diminishing unnecessary antibiotic usage in critically ill patients 12 and by integrating clinical texts’ analyses for enhancement of infection diagnosis in critically ill patients. 13 In their 2022 editorial, Retamar-Gentil, and Lopez-Cortez, 14 on the prediction of bacteremia in the emergency room, describe different rule-based systems developed in order to identify patients at risk. Such rule-based systems include the model published by Julian-Jimenez et al. 15 based on the combination of temperature over 38.3°C, Charlson index score, high respiratory rate, increased leukocytes count, and increased pro-calcitonin blood values. This model achieved high accuracy grades but was restricted only for patients who presented with signs and symptoms suggestive of a urinary tract infection. A similar model was suggested by Gonzalez et al., once again achieving high levels of accuracy according to their AUC–ROC analysis but was also restricted to a specific patients’ population—this time, those suffering from solid neoplasia. 16 Both systems did not and could not take into account parameters that were not pre-conceived by the researchers and did not consider out-lying parameters. Previously described statistical models referred to the drawbacks of conventional computing, including poor reproducibility in prediction accuracy and inconsistency in predictor selection and offered the application of Bayesian prediction: Jin et al. 17 presented a system of 20 predictors that were analyzed according to the Bayes’ theorem achieving a ROC–AUC value of 70%. All the above methodologies have two main drawbacks: all rely on rule-based tabular data analysis, and none related to the potential benefits of natural language processing.
From rule based to AI analysis
Clinical decision support systems based on artificial intelligence are rapidly developing in many clinical disciplines, including in the field of emergency medicine. 18 AI applications are available in a wide variety of architectures, representing variable combinations of knowledge-based rules, restricting the span of clinical recommendations generated by machine learning algorithms. The extent to which machine learning (ML) is restricted by rules is associated with the extent that physicians and regulators will allow flexibility. In their scoping review of such hybrid architectures, Kierner, Kucharski, and Kierner 19 name five optional structures: rules that are embedded in ML architecture (REML), ML pre-processes input data for rule-based inference (MLRB), rule-based method pre-processes input data for ML prediction (RBML), rules influence ML training (RMLT), and parallel ensemble of rules and ML (PERML). All these are certainly appropriate in cases AI is expected to generate clinical treatment recommendations. Nevertheless, AI systems that would predict bacteremia do not necessarily have to rely on any rule-engaging architecture. Applying AI analysis without restriction of pre-defined rules would probably do better. This is the main reason that in the current study we used AI algorithms without any rule-based reliance.
Using the XGBoost methodology in medicine is not new. Key features of XGBoost algorithm include its ability to handle complex relationships in data, regularization techniques to prevent overfitting, and incorporation of parallel processing for efficient computation. It is widely used in various domains due to its high predictive performance and versatility across different datasets. 20 In the realm of predictive medicine, the XGBoost algorithm knows how to deal well with missing data 21 and shows superiority over other AI models in the prediction of chronic kidney disease patients’ survival, 22 acts as an explainable AI model for the prediction of myocardial infarct in large and diverse populations, 23 and more.
The potential benefits of harnessing NLP to the mission
In light of the fact that most of the medical data related to patients’ interactions with their physicians is verbal, the prospects from automatic text analysis in the medical AI are huge. 24 Applications that use NLP exist in many clinical realms: de-identification of clinical files’ data, 25 analyzing patient notes, assisting patients in navigating the healthcare system, and supporting clinical decision-making when combined with human oversight. 26
In the current research, we applied the NLP in the task of screening patients’ ED notes for verbal indicators of a positive blood culture. We expected several specific wordings to emerge, for example, “low blood pressure,” “looking sick” and mainly “shivering.” The NLP model, which combines TF-IDF with logistic regression, was integrated into our predictive model and identified several hundreds of words and text fragments as predictive of either positive or negative blood cultures, as specified above. Choi et al. implemented AI algorithms for the same task, 27 but they did not focus on NLP tools but rather on neural networks. They concluded that AI should be best implemented in cases of physicians’ uncertainty, a feature that has also the potential to benefit by NLP implementation.
Advantages and innovation of the current model
Only few previous studies have addressed the challenges we faced in the current research. Mahmoud et al. did a retrospective analysis of 36,405 blood cultures of already hospitalized patients. In trying to predict positive blood cultures they found that indeed, rule-based scoring systems had low efficiency. They applied AI algorithms that had high specificity but rather low sensitivity. Nevertheless, they did not integrate the NLP technique into their AI model. 28 Boerman A. W. et al. published their study results in this quest. In their 598 out of 4885 (12.2%) ED visits, at least one blood culture returned positive. They used both a gradient boosted tree model and a logistic regression model that showed good performance in predicting blood culture results with an AUC–ROC of 77% (95% CI 0.73 to 0.82) in the test set. In the gradient boosted tree model, they claim to have 69% accuracy regarding negative cultures with a negative predictive value of over 94%. They also did not include an NLP component in their AI modeling.
Limitations of our model
In contrast to the options available in English, 29 currently, the new language models do not provide a satisfactory response to the text in Hebrew, and especially in medical Hebrew. Another complexity of the text in the medical files stems from the fact that the texts are not written in a standard language in terms of syntax, grammar, and spelling errors, alongside the combination of medical terms and abbreviations in two languages (English and Hebrew). We are in the process of working on a dedicated model for the medical files, but at this stage, the choice of a model based on TF-IDF and BOW allowed us to provide a fast and computationally efficient answer with good performance to the research question.
Another future challenge would be finding the appropriate scheme for implementing our findings into the clinical workflow. We foresee our results as serving this purpose, maintaining the need for physicians’ vigilance, as hailed by Finlayson et al., 30 regarding this sacred domain of bacteremia identification.
Conclusions
Our findings highlight the value of combining structured data with the nuanced information contained in free text to better predict BSI. The integration of free text data notably improved our model's performance compared to relying solely on structured data. Overall, our approach aims to provide a more reliable tool for an early identification of bloodstream infections, potentially leading to improved patient outcomes. This study was done in a single center but on a large and diverse population; therefore, we conclude that our findings would be reproducible in other clinical fields and disciplinary models. Decision support systems based on artificial intelligence will likely be deployed in emergency medicine practices worldwide. We foresee a place for combined XGBoost algorithms with NLP modeling to be applied in this realm.
Footnotes
Contributorship
All authors contributed to this manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
This study was approved by an IRB: (# 0348-23-SMC). Patients’ consent was waived due to the retrospective nature of this study.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
