Abstract
A natural language processing (NLP) application requires sophisticated lexical resources to support its processing goals. Different solutions, such as dictionary lookup and MetaMap, have been proposed in the healthcare informatics literature to identify disease terms with more than one word (multi-gram disease named entities). Although a lot of work has been done in the identification of protein- and gene-named entities in the biomedical field, not much research has been done on the recognition and resolution of terminologies in the clinical trial subject eligibility analysis. In this study, we develop a specialized lexicon for improving NLP and text mining analysis in the breast cancer domain, and evaluate it by comparing it with the Systematized Nomenclature of Medicine Clinical Terms (SNOMED CT). We use a hybrid methodology, which combines the knowledge of domain experts, terms from multiple online dictionaries, and the mining of text from sample clinical trials. Use of our methodology introduces 4243 unique lexicon items, which increase bigram entity match by 38.6% and trigram entity match by 41%. Our lexicon, which adds a significant number of new terms, is very useful for matching patients to clinical trials automatically based on eligibility matching. Beyond clinical trial matching, the specialized lexicon developed in this study could serve as a foundation for future healthcare text mining applications.
Introduction
There is a growing need to efficiently transform these free text clinical research eligibility criteria into computable formats to support the subject recruitment process. Various approaches have been proposed to achieve high-performance text analysis of clinical trial eligibility criteria. Prior work has typically used the Bag of Words (BOW) model for text analysis.
A BOW is a way of extracting features from text data for use in natural language processing (NLP). The approach is simple and flexible and can be applied in a myriad of ways for extracting features from documents. It is called “bag” of words, because any information about the order or structure of words in the document is discarded. The model is only concerned with whether known words occur in the document, not with where they appear in the document.
However, the BOW approach does not recognize multi-word terms, which are common in the medical and healthcare domains. The use of the general BOW model results in a loss of word ordering information and the semantics of multi-word terms. The word n-gram model does take into consideration the context information of a word, which depends on the previous or the next word. 4 The n-gram is a contiguous sequence of n items from a given sample of text.
Although the word n-gram model improves text analysis performance, it decreases the performance if the number of words (n) is greater than 3. 5
Lexical resources play an important role in text mining, information retrieval, and NLP tasks. In the healthcare domain, specialized and modularized lexicon resources must be developed to support high quality text analysis. Additionally, expert knowledge can contribute to improving Named Entity Recognition (NER) performance, either by customizing existing lexicons or by supplementing them with new terms. 6 The specialized dictionary that we have developed would be useful for NER across different cancer domains. 7
Text mining and NLP applications require knowledge about words. One of the most time-consuming and labor-intensive tasks in healthcare text mining research is the creation, compilation, and customization of the necessary lexicons. 8 If automated procedures could be developed for large-scale extraction and formalization of lexical data from existing resources, healthcare information systems would be able to exploit the power of lexicons in their applications.
At present, there is no lexicon resource available for identifying the word n-gram terms in breast cancer clinical trial eligibility. The main objective of the study is to build a specialized lexicon for breast cancer clinical trials, which would be valuable for future text mining and NLP applications. To that end, we build a specialized lexicon that facilitates analysis of eligibility criteria for breast cancer clinical trials. We believe that this lexicon will eventually help improve patient recruitment and enrollment by automatically matching patients with clinical trials. To the best of our knowledge, such a study has not been carried out before. The hybrid methodology that we use to develop the specialized lexicon is innovative. It combines the knowledge of domain experts, terms from multiple online dictionaries, and text mined from sample clinical trials. It could be effectively leveraged in future healthcare studies employing NLP and text mining. We use the specialized lexicon for the selection and reduction of word n-gram features and generating clinical trial clusters, to reduce search time, and, in the future, help automatically match patients with clinical trials.
Background and related work
Development of specialized lexicon
Overview of related text mining tasks
The concept of n-grams was first discussed in 1951 by Shannon. 9 Since then, character n-grams have been used in many areas, such as spelling-related applications, string searching, prediction, and speech recognition. 10
Word n-gram is a sequence of consecutive tokens, with a length of n. 11 Word n-gram feature induction, sometimes also referred to as feature extraction, induces features on textual data based on a set of word n-grams. With feature induction, the textual data is represented in a feature space, usually encoding the existence of these word n-grams or their frequency. The word n-grams to be used as features may be chosen by either using a data-driven approach or a dictionary-based approach.
In data-driven feature induction, every word n-gram combination from the textual data is created. Thus, the feature size equals the word n-gram vocabulary size. Such a data-driven feature induction does not require prior domain knowledge to recognize meaningful word n-grams.
In the dictionary approach, word n-gram tokens are selected based on a specialized lexicon database that focuses on a specific domain. This approach proposes that a word n-gram feature selection that maps all bigram and trigram tokens to the specialized lexicon database be used.
Named Entity Recognition (NER) is the task of identifying and classifying entities such as person names, place names, organization names, etc., in each document. Named entities play a major role in information extraction. A well-performing NER is an important NLP technique. Many techniques have been applied in English for NER. Some of them are rule-based systems, 12 which make use of dictionary and patterns of named entities. Other techniques applied for NER are Decision trees, 13 Hidden Markov Model (HMM), 14 Maximum Entropy Markov Model (MEMM), 15 and Conditional Random Fields (CRF). 16
NER has been done generically but can also be specialized where a finer tagset is needed to describe the named entities in a domain. Specialized NER has been in existence for a long time in the bio-domain 17 for identification of protein names, gene names, DNA names, etc. The NER task is also viewed as the first step of information extraction of free text clinical studies describing shock, trauma, inflammation, and other related states. 18 The proposed specialized lexicon also supports the NER process in the healthcare domain.
Specialized lexicon
In linguistics, a lexicon is a language’s inventory of lexemes and it is the vocabulary of a person, language, or branch of knowledge. Lexicon plays a critical role in NLP and text mining, providing foundational reference resources. In many NLP and text mining tasks, such as NER, syntax alone is not enough to build a high-performance system; an external source of information such as a dictionary is also required. 19 Most state-of-the-art approaches for NER use information from lexicons and semi-supervised learning in the form of word clusters. A specialized lexicon, which is a list of word types related to the desired named entity types, is considered an essential component in state-of-the-art NER systems. 19 The word n-gram model is used for most state-of-the-art language models, but it requires very large corpora and lexicon resources.
Accrual to clinical trials
Clinical trials are essential for testing new treatments, advancing medical knowledge, and improving outcomes for patients. However, less than 5% of the adult cancer patients are enrolled into clinical trials. 20 Several studies have described what prevents patients from participating in clinical trials.21,22 Lovato et al. 23 investigated intervening factors in patient recruitment and identified major obstacle categories, such as diverse populations, recruitment strategies, overall planning and management, patient and physician attitudes, generalizability and adherence, and cost. Ross et al. 24 reported other barriers to recruitment, some related to clinicians and others to patients: time constraints, lack of staff and training, concern about the impact on doctor-patient relationship, concern for patients, loss of professional autonomy, and difficulty with consent procedure, lack of rewards and recognition, and insufficiently interesting questions. Comis et al. 25 demonstrated that one of the primary problems with recruitment of patients in clinical trials is patient disqualification due to eligibility criteria.
With rapid advances in information technology, more and more studies have focused on IT capabilities for improving clinical trial accrual. 26 Andronikou et al. 27 analyzed the clinical trial design process and presented an approach for the automatic selection of patients eligible to participate in clinical trials. Weng et al. 28 surveyed the literature and identified five aspects of eligibility criteria knowledge representations that could be used in research and clinical decision support systems: the intended use of computable eligibility criteria, the classification of eligibility criteria, the expression language for representing eligibility rules, the encoding of eligibility concepts, and the modeling of patient data. Milian et al. 29 proposed an approach for automatic extraction of clinical trial eligibility criteria meaning and evaluating that for a patient. They employed the semantic entity (e.g. diseases, treatment, measurements, etc.) detecting technique for text analysis, using ontology annotators and semantic taggers. Their study proposed a pattern detection algorithm for contextual information and its assessment.
There are several studies that applied lexicon resources to clinical trials with the purpose of enhancing analysis of textual data and selection of patients for subject matching. Patel et al. 30 described a case study that explored an ontology reasoning technique in the medical domain and investigated whether it was possible to use semantic technologies to aid the selection of patients for clinical trials based on their health records and the SNOMED CT ontology. Cuggia et al. 31 reviewed the contributions and limitations of decision support systems for automatic recruitment of patients to clinical trials and characterized important features of clinical trial decision support systems. They also evaluated the effectiveness and potential of such systems for improving trial recruitment rates.
Xu et al. 32 developed a semi-automatic framework combining text analysis techniques with manual review to extract genetic alteration information from cancer trial text. Their framework was used to classify cancer treatment trials from ClinicalTrials.gov and extract gene and alteration pairs from the Title and Eligibility Criteria sections of clinical trials.
Park et al. 33 developed and utilized a clinical trial management system (CTMS) that complies with standardized processes from multiple departments/units, controlled vocabularies, security, and privacy regulations. It is the first CTMS developed in-house at an academic medical center side that can enhance the efficiency of clinical trial management in compliance with privacy and security laws.
NLP and text mining in clinical trial eligibility
During the last decade, several studies on clinical trials have used the text mining approach. One of the salient research streams is the formal representation of eligibility criteria. 28 Tu et al. 34 examined formalizing eligibility criteria in a computer-interpretable language to facilitate eligibility determination for study subjects and the identification of studies on similar patient populations. ERGO (Eligibility Rule Grammar and Ontology) annotation is used for capturing the semantics of eligibility criteria. Luo et al. 35 examined a semi-automatic process for extracting Common Data Elements (CDEs) from eligibility criteria of clinical trials. Their study is the first study using text mining in CDE discovery from free-text clinical trial eligibility criteria.
There have been foundational studies on enhancing eligibility criteria representation. Luo et al. 36 presented a corpus-based approach to create a semantic lexicon for clinical research eligibility criteria using the Unified Medical Language System (UMLS). The main purpose of their research was to reduce the ambiguity in UMLS semantic-type assignment while building a semantic lexicon for clinical trial eligibility criteria. A total of 20 UMLS semantic types, representing about 17% of all the distinct semantic types assigned to corpus lexemes were identified, covering about 80% of the vocabulary of the test corpus.
Feature selection and summarization of clinical trials is an emerging research topic. Boland et al. 37 investigated the feasibility of feature-based indexing, clustering, and search of clinical trials. They argued that concept-oriented eligibility features could enhance user search effectiveness, facilitating meaningful and efficient indexing for clinical trials. In their study, concept-oriented eligibility features are a clinically meaningful atomic patient state—such as diagnosis, symptom, or demographic characteristics—which is derived from eligibility criteria. The goal of eligibility criteria analysis is to facilitate the automatic matching of clinical trials and patients. Wilcox et al. 38 presented a model, electronic Participant Identification and Recruitment Model (ePaIRing), which uses patient information to enhance patient recruitment in clinical trials.
Over the past decade, as noted above, several studies have been conducted on clinical trial and subject eligibility criteria. However, none of those studies tried to generate a specialized lexicon resource, even though a specialized lexicon is fundamental to medical text analysis and serves as a foundation for NLP and text mining.
Han et al. 39 developed electronic data capture systems and electronic case report forms (eCRFs), for clinical research. They developed a natural language processing–driven medical information extraction system (NLP-MIES) based on the i2b2 reference standard and evaluated the application with 24 eligible participants. According to the reported results, the system can improve both the accuracy and efficiency of the data entry process.
Zeng et al. 40 developed OCTANE (Oncology Clinical Trial Annotation Engine), an informatics framework, and implemented an instance of OCTANE at a large cancer center. OCTANE consists of three modules. The data aggregation module automates retrieval, aggregation, and update of trial information. The authors argue that OCTANE can be helpful for establishing and maintaining a comprehensive database necessary for automating patient-trial matching.
Lexicon building in biomedical domain
Domain-specific lexicons are extensively used in biomedical research for tasks such as natural language processing, information retrieval, and text mining. Most lexicons in the biomedical domain were created manually by expert curators; there is a growing demand for an automated lexicon building and compiling method in the biomedical domain.
Parai et al. 41 implemented the Lexicon Builder web service using all English ontologies in UMLS and a subset of the NCBO BioPortal ontologies. These ontologies offer a dictionary of 4,222,921 concepts and 7,943,757 terms. Thompson 42 built BioLexicon and provided an overview of method for the design, construction, and evaluation of a large-scale lexical and conceptual resource for the biomedical domain. BioLexicon integrates different types of terms from several existing data resources into a single, unified repository and contains over 2.2 million lexical entries and over 1.8 million terminological variants, as well as over 3.3 million semantic relations, including over 2 million synonymy relations.
Nguyen et al. 43 proposed the use of text mining methods for the automatic construction of a large-scale biodiversity term inventory using distributional semantic models (DSMs) to identify names that are semantically related to any given species name. They conducted a comparative evaluation of various types of DSMs with four different metrics and found that prediction-based DSMs performed significantly better than count-based DSMs. Their term inventory contains more than 288,000 scientific and vernacular names of species.
Sasaki et al. 44 demonstrated that a large-scale lexicon, tailored for the biology domain, is effective in improving question analysis for genomics Question Answering (QA). Rinaldi et al. 45 combined multiple biomedical knowledge bases and extracted entity names from the collected resources. For evaluating their work, they determined the accuracy of automatic protein name detection using the IntAct corpus and found that the performance of their approach was competitive.
Percha et al. 46 proposed a radiology lexicon curation approach using distributional semantics algorithms. They applied Word2vec, a distributional semantics software package, to the radiology notes to identify synonyms for RadLex, a structured lexicon of radiology terms. They found that distributional semantics algorithms can assist lexicon curation, saving researchers time and money.
For online-based biomedical lexicon research, Hsieh 47 proposed a method for estimating semantic similarity by exploiting the page counts of two biomedical concepts returned by the Google AJAX web search engine, because there is a limitation in the traditional ontology-based methodologies when we measure semantic similarity between two terms. The proposed method for estimating semantic similarity was evaluated by applying support vector machines on two datasets.
In this study, we develop a breast cancer-specific multi-gram lexicon by inducing high-impact multi-gram terms from clinical trial descriptions and integrating heterogeneous online resources.
Methods
We divided the specialized lexicon development process into two phases: corpus-based lexicon generation and collection of existing online dictionaries. The first step of corpus-based lexicon generation is the collection and organization of the corpus, which is a large body of natural language text used for natural language analysis. Data for 378 breast cancer related clinical trials, including four subsections—purpose, eligibility, contacts/locations—were collected from ClinicalTrials.gov; only the eligibility section was used for the corpus.
No patients were involved as subjects in the study and no actual clinical data were used for this study. The extracted corpora were processed for tokenization using the Stanford Tokenizer, which divides the text inside the corpora into a sequence of tokens that roughly correspond to “words.” Next, we performed lemmatization using the Stanford CoreNLP suite, which converts different forms of a word to its base or dictionary form, known as the lemma. Stop word removal is the last step in the preprocessing of corpus-based lexicon generation. We created a stop word list of 429 items based on the snowball stop word list and applied the stop word removal process.
A bag of words was derived from the preprocessed corpus. The bag of words was converted to a matrix form to create a vector space model representing text documents as vectors of identifiers. Next, to identify high impact n-gram words, we calculated term frequency–inverse document frequency (TF-IDF) scores for the words. TF-IDF score is a numerical statistic that reflects how important a word is to a document in a collection or corpus. It is a popular approach for weighting factors in information retrieval, text mining, and user modeling.
In the second phase, we collected n-gram words from multiple online resources using a web crawler developed for this study and merged them into the corpus-based lexicon. Considering lexicon accessibility, comprehensiveness, and recentness, we selected four major online lexicon resources. The online lexicon resources are in dictionary format; thus, all the dictionary items were included in the breast cancer specialized lexicon, except for duplicate items.
The first online resource used was the National Cancer Institute (NCI) dictionary. NCI is the federal government's principal agency for cancer research and training. It created and maintains a dictionary of cancer and biomedical terms, which are defined in a non-technical language. The terms and definitions are reviewed by a multidisciplinary panel of reviewers, and approximately 50 new and 50 revised terms are included each month (https://www.cancer.gov/publications/dictionaries/cancer-terms). A total of 4704 terms was collected from NCI’s Dictionary of Cancer Terms.
The second resource from which we collected terms is Breastcancer.org, a nonprofit organization dedicated to providing the most reliable, complete, and up-to-date information about breast cancer (http://www.breastcancer.org/). A total of 910 terms were collected from the Breastcancer.org glossary section.
The third resource we used was the web site of American Cancer Society (ACS), which is a nationwide, community-based voluntary health organization dedicated to eliminating cancer as a major health problem. We collected 155 terms from the glossary section of ACS’s site (https://www.cancer.org/cancer/glossary.html).
The last online resource we used is emedicinehealth.com, which is an online medical news network and contains over 900 health and medical articles with a focus on emergency medicine, written by physicians for patients and consumers. We collected 28 terms from the emedicinehealth.com site. Table 1 shows the number of dictionary items collected by source.
Number of items by source.

Implementation of the breast cancer lexicon
Figure 1 depicts the steps we followed for building the specialized lexicon for breast cancer. We collected the data in two phases. In phase I, we identified 378 clinical trials using the search term “Breast Cancer” from ClinicalTrials.gov between January 1, 2010 and January 1, 2011 and downloaded all the related information as a collection of individual XML files. In phase II, we collected additional 664 clinical trials from the same repository, using the same search term, between January 1, 2018 and January 1, 2019.
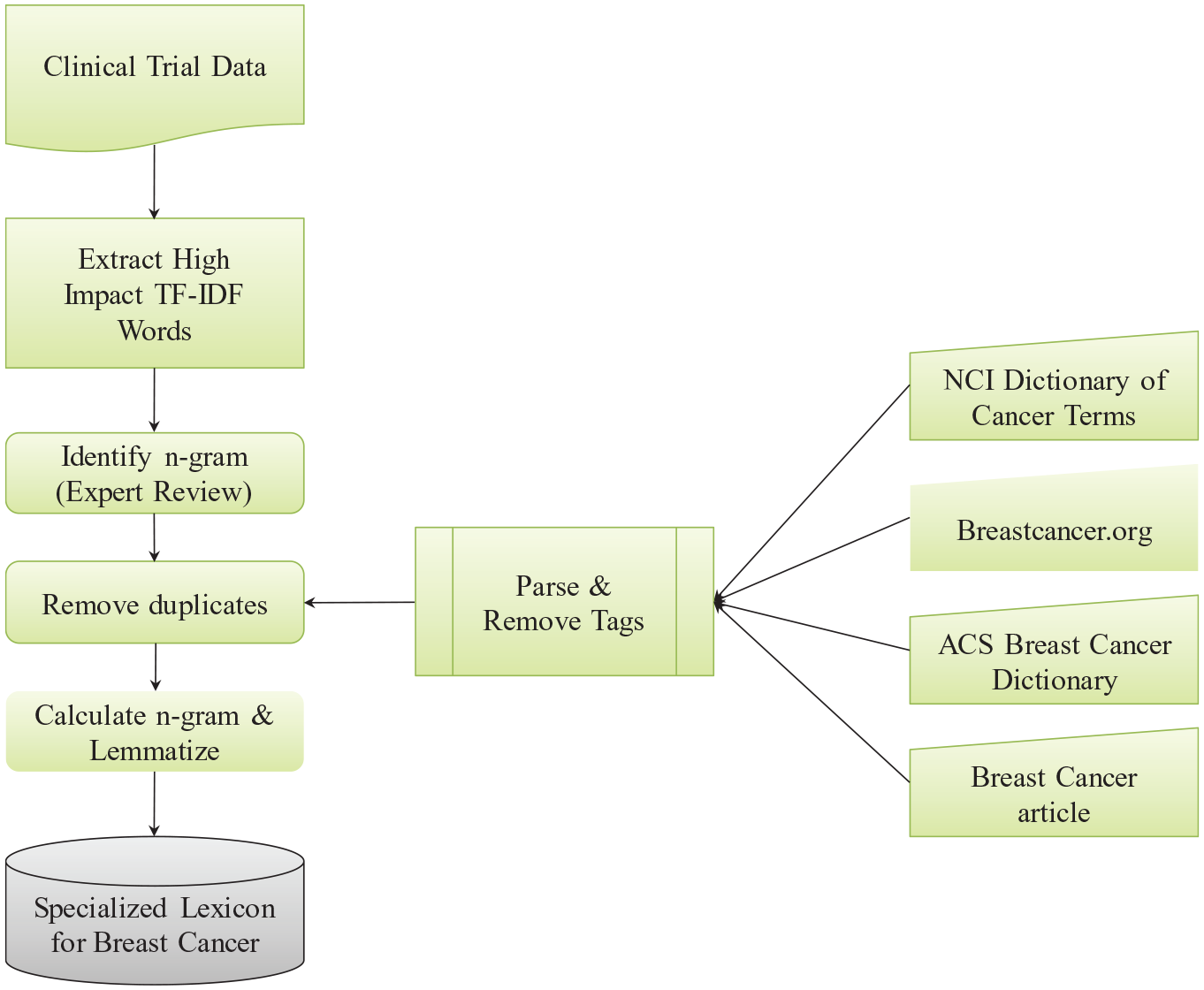
Steps for building specialized lexicon.
All XML tags and metadata were removed and only the <eligibility>—<criteria>—<textblock> section was extracted. Since subject eligibility criteria text is in free text format and contains two opposite criteria, “inclusion” and “exclusion,” we separated subject eligibility criteria text blocks based on the key words “Inclusion criteria” and “Exclusion criteria.”
We constructed a specialized word n-gram term lexicon for the breast cancer domain. The lexicon was based on high TF-IDF score words from the clinical trial eligibility data set and other online resources (i.e. NCI Dictionary of Cancer Terms, Breastcancer.org, and ACS Breast Cancer Dictionary).
First, during pre-processing, we performed tokenization, lemmatization, and stop word removal over the selected data set. Second, we calculated the TF-IDF scores for all the unigram features that were drawn from the breast cancer clinical trial eligibility text data set. Three experts reviewed the unigram list, organized in descending order by the TF-IDF score, and manually identified bigram and trigram terms from the unigram list; the review was conducted sequentially. Two of the authors involved in the study, one with an MD and a PhD in Biomedical Informatics and the other with a PhD in Information Systems, were used as domain experts for this study. Both of them have more than 5 years of experience in multidisciplinary breast cancer care and information system design and testing for clinical decision support systems for breast cancer tumor boards.
When the identified bigram and trigram terms are relevant to breast cancer clinical trials, they were selected as candidates for the lexicon. The output of the first reviewer was reviewed by the second reviewer and the output of the second reviewer by the third reviewer. The results of the sequential review were then discussed among the three reviewers until all the reviewers agreed on the selected bigrams and trigrams. The final review was conducted by the expert who is a medical doctor. After the expert review, a total of 1506 multi-gram terms were identified. Only adjacent words were considered. For example, the adjacent terms “pregnant” and “woman” were used to generate the bigram “pregnant woman.”
Only bigram and trigram terms were considered in the expert review process. Since n-gram language models have a practical limitation with respect to using terms of higher order than trigrams, bigram and trigram models are common in NLP. 10
Third, we developed an online medical term crawler in the Ruby language to gather breast cancer terms from web sites of breast cancer domains such as NCI (https://www.cancer.gov/publications/dictionaries/cancer-terms). The crawler automatically collected web documents from the targeted sites listed in Medical Library Association (https://www.mlanet.org/) 48 as online cancer information resources; those documents were then reviewed by the first author. The URL for each targeted site was provided to the crawler. The HTML format cancer term glossary documents were collected from the four online resources (NCI, Breastcancer.org, ACS, and emedicinehealth.com) and the crawler parsed the HTML documents to remove unnecessary HTML and CSS tags. Only medical terms were extracted from the targeted website.
Evaluation of the lexicon
Two experiments were conducted to evaluate the efficiency of the specialized lexicon.
First, all items in the lexicon were directly matched with SNOMED CT in the UMLS Metathesaurus to examine the uniqueness of the specialized lexicon items.
SNOMED CT is considered one of the most comprehensive medical terminology products in the world (http://www.snomed.org/snomed-ct) 49 and is designated as one of the standards worldwide. SNOMED CT is also a required standard in interoperability specifications of the U.S. Healthcare Information Technology Standards Panel (https://www.nlm.nih.gov/healthit/snomedct/) 50 . Finding new lexicon items and supplementing SNOMED CT would therefore increase the interoperability and performance of the electronic exchange of clinical health information. 51
Only English terms in SNOMED CT were used in the evaluation. We created a database query in the Structured Query Language (SQL) and ran the query to evaluate uniqueness. The SQL query selected all the items in the specialized lexicon and matched each item with terms in SNOMED CT.
Second, the items in the specialized lexicon and in SNOMED CT were matched with a test data set to validate the usefulness of the lexicon for processing clinical trial data. A total of 1,058 clinical trial studies from January 1, 2011, to January 1, 2013 were collected from ClinicalTrial.gov and the eligibility criteria section was divided into two parts: inclusion criteria and exclusion criteria. These two data sets were pre-processed with tokenization, lemmatization, and stop word removal. All possible trigram and bigram combinations were generated to match with the proposed specialized lexicon and SNOMED CT.
Results
The crawler collected 4704 terms from the NCI dictionary of Cancer Terms, 910 terms from Breastcancer.org, 155 terms from the American Cancer Society Breast Cancer Dictionary, and 28 terms from breast cancer glossary of terms in emedicinehealth.com. All the collected items were stored in a MySql database and duplicates were removed by an SQL query. The specialized lexicon includes a total of 7303 items: 707 trigrams, 2098 bigrams, 4162 unigrams, and 336 n-gram terms consisting of more than three words. Table 2 shows the number of dictionary items by the type of n-gram.
Number of items and number of unique items by n-gram type.
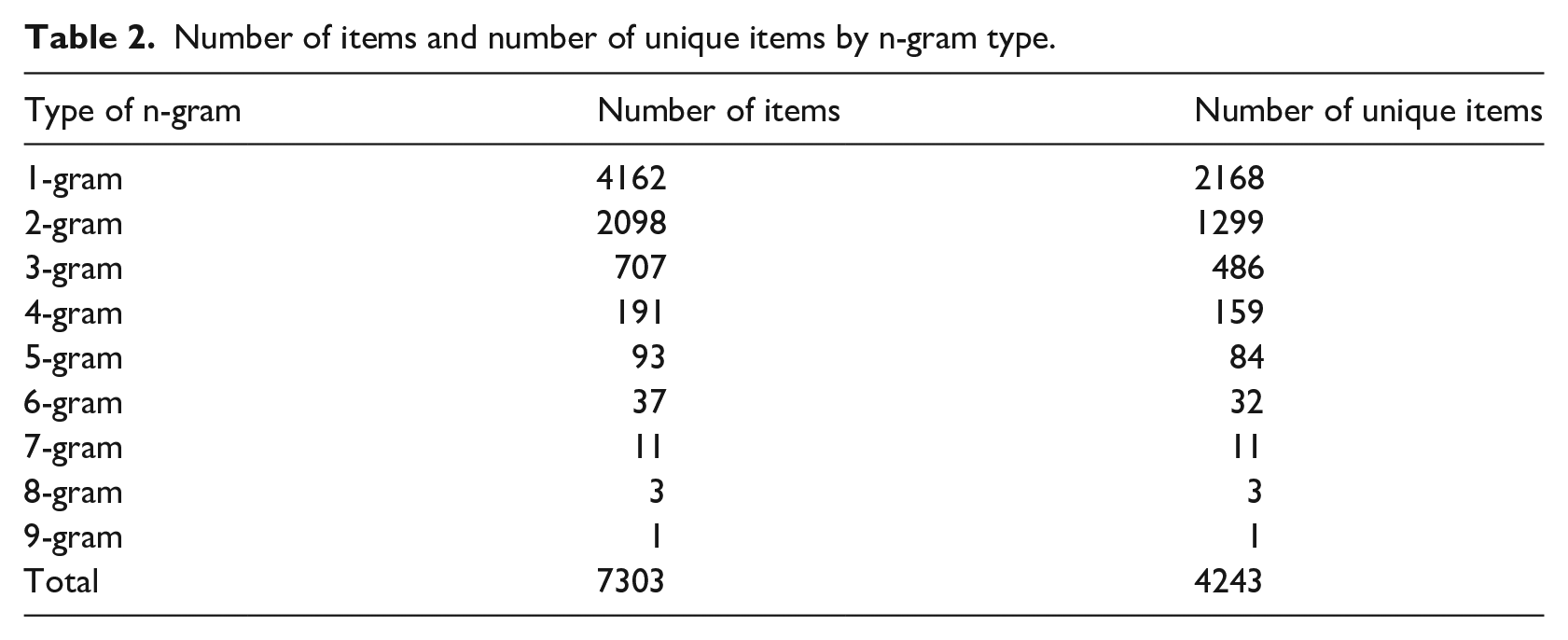
Based on the query results, 4243 items in the specialized lexicon were found to be unique, with 3060 items overlapping with SNOMED CT. Table 2 shows the number of unique items for different types of n-gram in the specialized lexicon. The evaluation results show that around 58% of the specialized lexicon items were newly introduced as part of the lexicon resource by using our methodology.
The matching results for trigram and bigram are presented in Table 3. SNOMED CT matched most items for both trigrams and bigrams, as reasonably expected, since the size of SNOMED CT is much larger than the specialized lexicon. However, the number of matched items by using both the specialized lexicon and SNOMED CT is greater than the number of matched items using SNOMED CT only. Table 3 shows that an additional 777 trigram matches were accomplished by adding the specialized lexicon to the SNOMED CT only match. This represents a 41% increase over the SNOMED CT only match. For example, the trigram “3 Tesla MRI,” which was included in our specialized lexicon, does not appear in SNOMED CT.
Trigram and bigram matching.
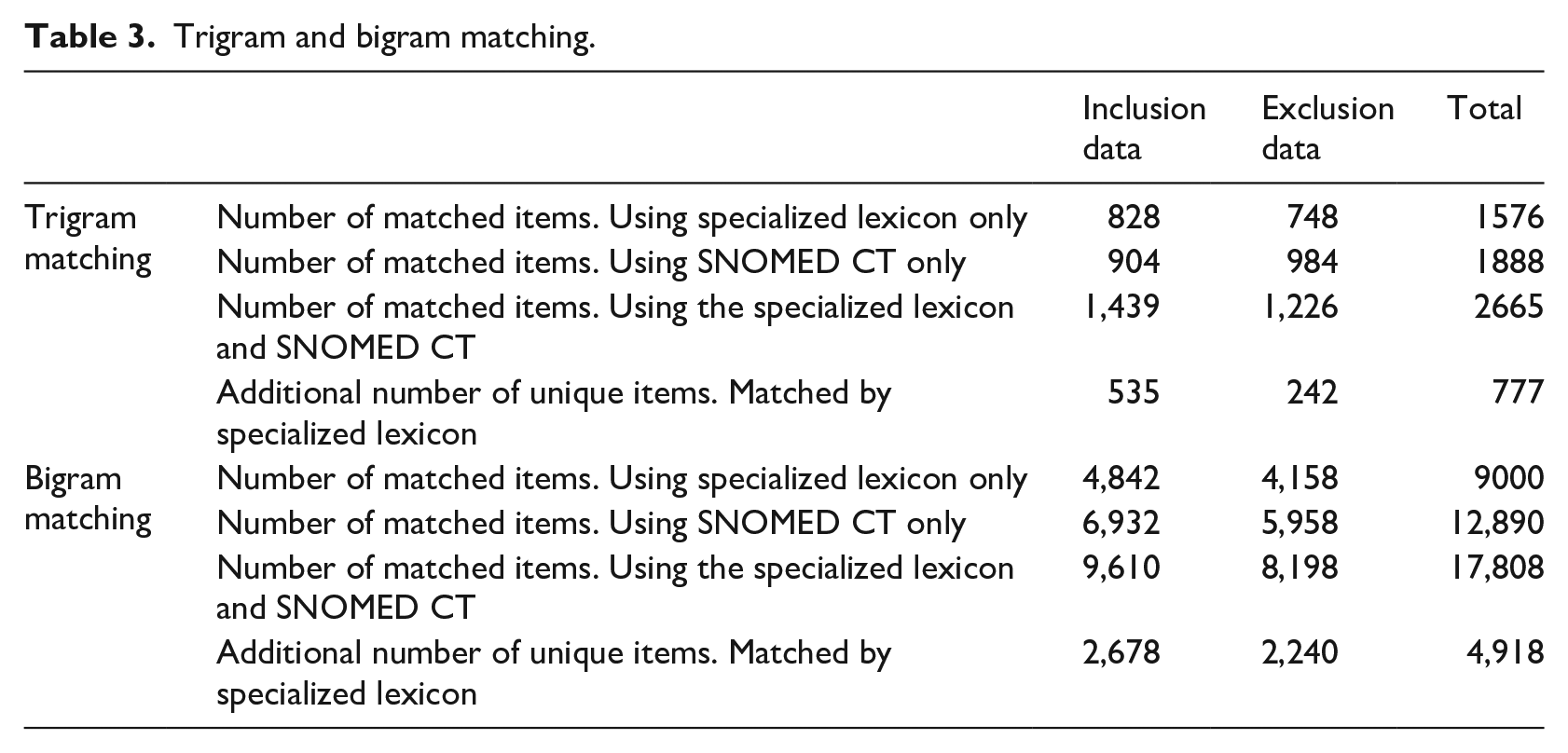
Similarly, as shown in Table 3, an additional 4,918 bigram matches were made by adding our lexicon to the SNOMED CT only match. This represents a 38.6% increase over the SNOMED CT only match. We can therefore conclude that our specialized lexicon helps to improve the clinical trial subject eligibility matching process substantially. Figure 2 shows a graphical comparison of the matching results for the different lexicon resources.
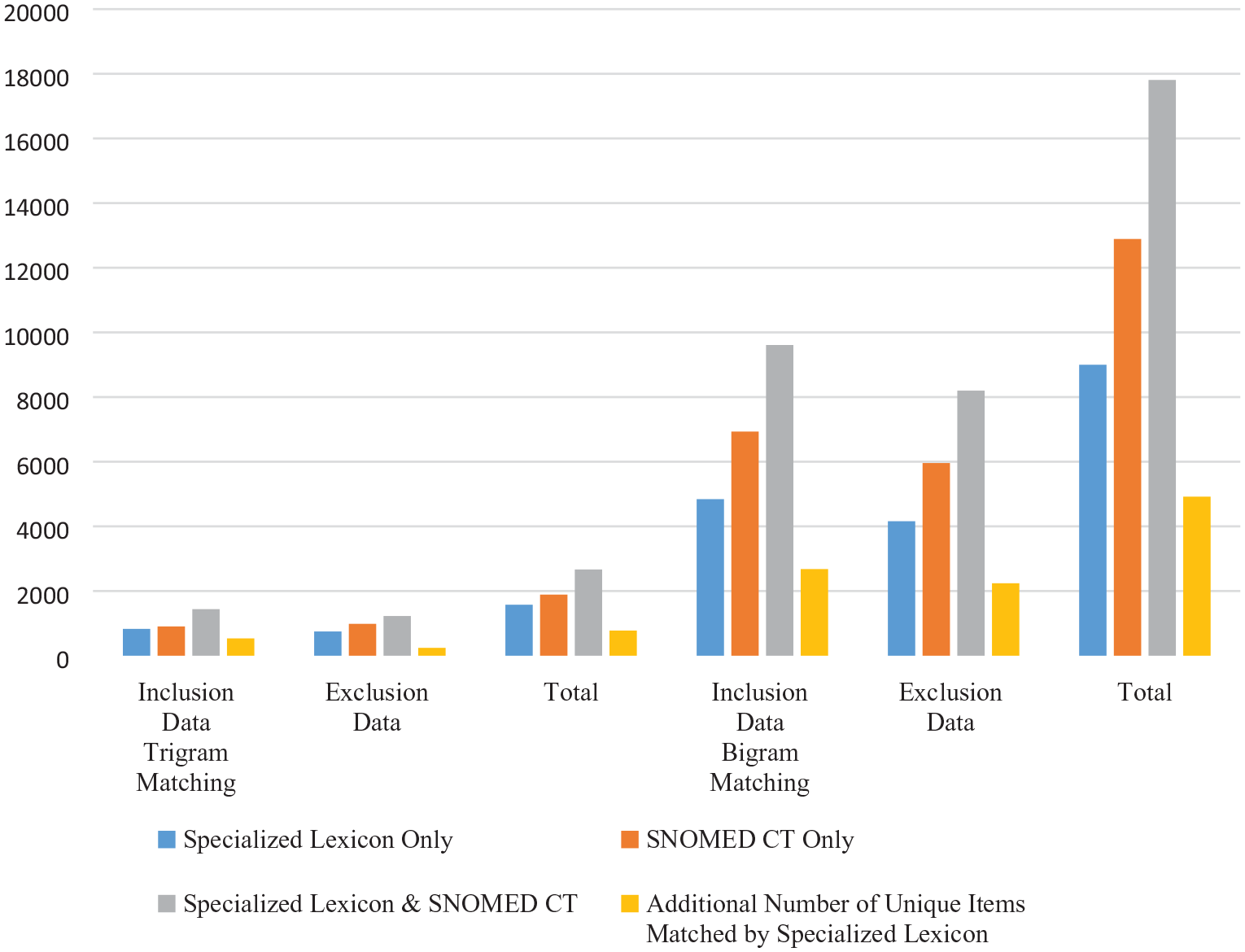
Trigram and bigram matching.
To check the robustness of the results obtained using our specialized lexicon, we used all the unigrams from the phase II data set, covering breast cancer trials from January 1, 2018 to January 1, 2019. First, we pre-processed phase II data, performing tokenization, lemmatization, and stop word removal. Then, we calculated the TF-IDF scores for all the unigram features and ordered them by the TF-IDF scores. All unigrams from the phase I and phase II data set were then compared. We found that 84.2% of phase II terms were covered by the phase I data set. Next, we randomly selected a sample of the additional terms from the phase II data set; this sample was then reviewed by the medical expert. Many of the additional terms were not considered to be meaningful for breast cancer clinical trial analysis. After removing those terms, we found that only 7.9% of phase II data can be genuinely considered to be valid and thus eligible for updating the specialized lexicon.
The difference between phase I and phase II mainly results from new treatments and new drugs introduced after phase I period. For example, unigram term ‘Atezolizumab’, ‘pembrolizumab, and ‘Nivolumab’ were found in the phase II dataset, but it was not included in phase I since they were introduced after 2014; ‘immune checkpoint-blockade,’ a trigram about regulators of the immune system, was found in phase II, but not in phase I since Immune checkpoint blockade therapy has been actively treated only after 2015.
The large overlap between the terms for the two phases demonstrates the robustness of our approach. To embrace incremental new treatment and drug terms, we plan to update the Specialized Lexicon for Breast Cancer Clinical Trial once every 2 years with the latest clinical trial text data.
Discussion
Some of the most time-consuming and labor-intensive tasks in text mining research are the creation, compilation, and customization of necessary lexicons. 52 Lexical resources are required for improving the performance of text mining, especially in NER. For healthcare informatics researchers, it is important to implement modular systems because the healthcare domain is highly specialized; thus, it is impossible to cover the entire healthcare domain with a single system. Therefore, the building of customized lexical resources is necessary for these highly domain-specific systems. 53
In this study, we built a specialized lexicon for breast cancer and demonstrated the semi-automated lexicon building process. We found a total of 4243 unique lexicon items in the breast cancer domain using a semi-automated lexicon building approach. These lexicon items are not present in SNOMED CT, one of the most comprehensive, clinical healthcare terminology products.
The evaluation of the breast cancer specialized lexicon using clinical trial subject eligibility documents revealed that even though the total number of matched items using the specialized lexicon is less than the number of matched items using SNOMED CT, about 30% of the matched items using our custom lexicon and SNOMED CT were derived from the lexicon. Our lexicon contributes to existing work in the area by introducing several new multigram medical terms, which can be used for the automatic patient matching process. The results demonstrate the critical role played by the specialized lexicon and expert knowledge in the development of lexicon resources.
Text datasets for breast cancer clinical trial subject eligibility were collected in two phases. The robustness of the results using phase I data was tested using the more recent phase II data. The phase II data confirmed that the analysis with phase I data has a significant contribution as foundational work for future breast cancer clinical trial research; the proposed lexicon needs to be updated periodically with terms for newly developed treatments and drugs.
Our research has some limitations. First, the coverage rate of our specialized lexicon is relatively low. The lexicon includes limited online resources. Use of more extensive resources should result in higher performance. Second, for validating the usefulness of our lexicon, we only determined the number of matched terms with the test data set. If an annotated data set as the gold standard is available, more sophisticated evaluation metrics could be included.
Conclusion
In this study, we built a specialized lexicon by using a hybrid approach: corpus-based lexicon generation and collection of online resources. The evaluation results show that the breast cancer specific lexicon could be exploited for NLP and text mining applications in the healthcare domain to improve the efficiency of clinical trial subject eligibility matching by automating the patient matching process. The lexicon resources used to capture multi-gram complex medical terms emerged as key to enhancing automatic patient matching effectively. The breast cancer specialized lexicon developed in this study could be used for future clinical trial NLP and text mining studies.
A future research direction is to use our breast cancer clinical trial specialized lexicon for automatic patient matching with clinical trials capturing NER in clinical trial subject eligibility text and patient electronic health records to enhance the performance and quality of text mining and NLP tasks.
