Abstract
The ability to automatically categorize submitted questions based on topics and suggest similar question and answer to the users reduces the number of redundant questions. Our objective was to compare intra-topic and inter-topic similarity between question and answers by using concept-based similarity computing analysis. We gathered existing question and answers from several popular online health communities. Then, Unified Medical Language System concepts related to selected questions and experts in different topics were extracted and weighted by term frequency -inverse document frequency values. Finally, the similarity between weighted vectors of Unified Medical Language System concepts was computed. Our result showed a considerable gap between intra-topic and inter-topic similarities in such a way that the average of intra-topic similarity (0.095, 0.192, and 0.110, respectively) was higher than the average of inter-topic similarity (0.012, 0.025, and 0.018, respectively) for questions of the top 3 popular online communities including NetWellness, WebMD, and Yahoo Answers. Similarity scores between the content of questions answered by experts in the same and different topics were calculated as 0.51 and 0.11, respectively. Concept-based similarity computing methods can be used in developing intelligent question and answering retrieval systems that contain auto recommendation functionality for similar questions and experts.
Keywords
Introduction
A great number of people seek their health-related questions from online resources when they face a specific health condition and do not have enough knowledge to answer it. 1 These types of healthcare consumers play a more effective role in managing their disease(s) and treatment plans. Therefore, they need to gather and process health-related information from various resources and make a final decision.2,3 Also, it has been shown that 71 percent of health consumers between the ages of 18 and 20 years look for health information in online resources. 4 However, despite the availability of healthcare information on various websites, consumers are often confused and cannot find what exactly they are looking for.5,6 Several studies showed7–9 that most consumers tried to find health information by searching two or three keywords that hardly describe their actual medical condition. The keywords used in these simple searches typically do not match to the structured content of online databases; hence, the query returns unrelated results. Therefore, the performance of searching health information by consumers in online search engines is typically low.
Online question and answering (Q&A) communities are rapidly growing. Users can ask their healthcare questions in various topics in Q&A websites, sometimes anonymously. They usually can find answer for their questions quickly without any cost. History of questions which has been sent to online communities is a valuable resource to capture consumers’ vocabularies in a given domain. However, the lack of standard concepts in such questions makes it difficult for consumers to find question relevant to a particular health problem. Therefore, one of the most challenging hurdles in online Q&A communities is duplicated questions. Such duplicates cause a large number of similar questions piling up on websites which makes the search and browse for other consumers quite frustrating. Similarly, medical experts need to find comparable questions and respond with relevant information. In these instances, online community’s support team need to find a way to classify similar Q&As through a well-defined content management framework. Therefore, identification of similar questions in Q&A retrieval systems is essentially important from both consumers’ perspective and medical experts.
A retrieval system can automatically suggest similar Q&As to users by considering inter-topic similarity. Also, instead of having a backup team manually sorting questions into different topics, the system can automatically categorize questions in various topics based on inter-topic similarity. The goal of this study was to compare inter- and intra-topic similarity between Q&As submitted to the online health communities. We used a concept-based similarity computing method to facilitate finding similar healthcare-related questions and medical experts.
Method
Figure 1 shows the step by step process of Q&A gathering, concept extraction, and similarity calculation.
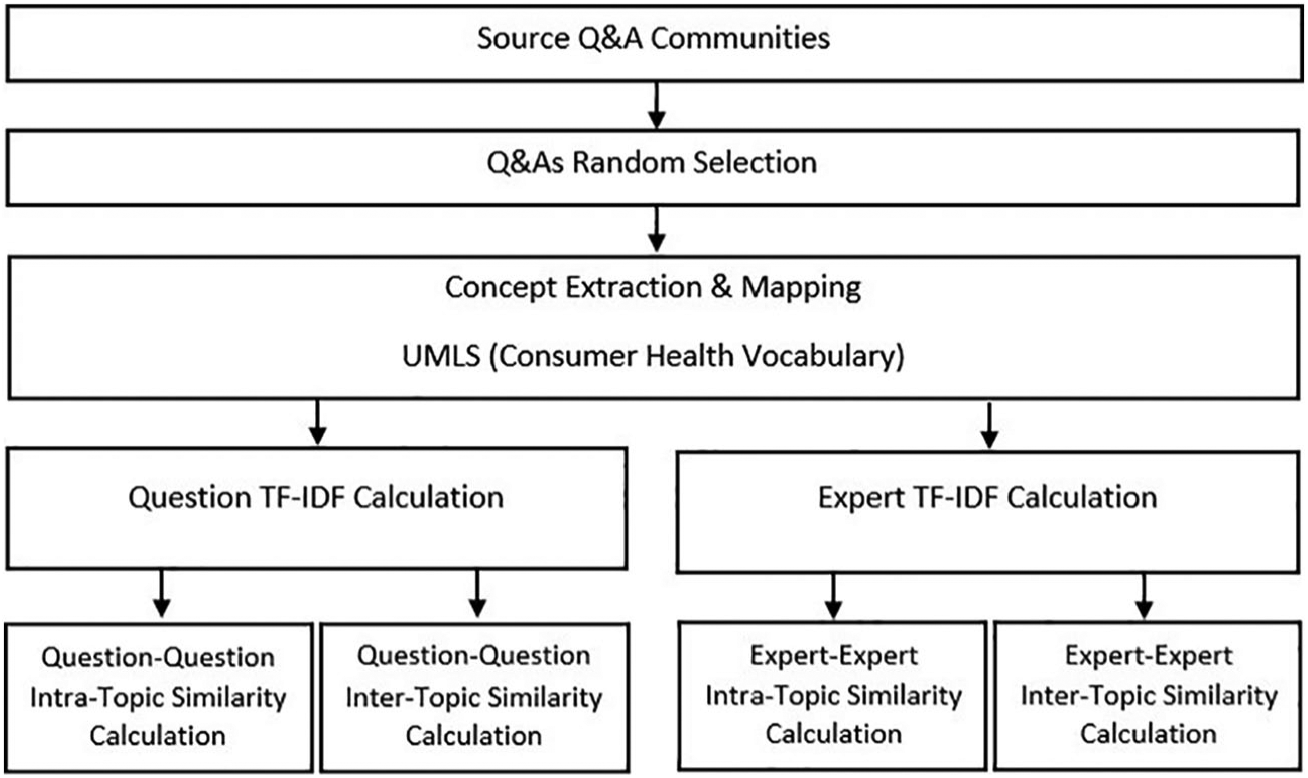
Steps of Q&As gathering, concept extraction, and similarity calculation.
Step 1: source Q&A communities
Yahoo Answers was launched in 2005 as a major social Q&A website that covers a wide range of domains and has the most number of visits among Q&A websites in United States. 10 Health category in Yahoo Answers is a major topic with more than 20 subcategories such as diabetes, women’s health, and cancer. 11 WebMD is another major health information website that was created more than 20 years ago. 12 This online health community has more than 100 million visitors per month that seek diverse health-related services such as Q&A on various health topics, medication information, and clinical symptom.13,14 NetWellness is a non-profit community health service with around 500 healthcare subject matter experts. The website was funded by three universities, including Case Western Reserve University, Ohio State University, and the University of Cincinnati. More than 70,000 questions in distinctive health domains have been answered by medical experts since 1995 on the website. 15 Content evaluation of NetWellness, based on feedbacks received from end users, shows Q&As posted on this website were greatly helpful for healthcare consumers. 16
Step 2: Q&A random selection
A PHP script was written to collect Q&As from source communities and store them in a MySQL database (version 5.6). Subsequently, 3000 Q&As were randomly selected for our study. The health topics of the three sources (Yahoo Answers, WebMD, and NetWellness) were not exactly the same. Therefore, we randomly selected 75 Q&As (five Q&As for each topic) to calculate the inter-topic and intra-topic similarities. Also, 400 Q&As were randomly selected to calculate expert–expert similarities. For example, questions 1 and 2 (provided in the Online Supplementary Material) are two sample questions extracted from NetWellness community. Our objective was to extract Unified Medical Language System (UMLS) concepts from these questions and calculate similarity scores among them based on the steps depicted in Figure 1.
Step 3: concept extraction and mapping
In this step, selected Q&As were inserted into a concept extraction and mapping pipeline where MetaMap was used to extract clinical entities and map them to UMLS Concept Unique Identifiers (CUIs).17,18 MetaMap data source was restricted to Consumer Health Vocabulary (CHV) terminology as the most relevant resource for mapping consumer healthcare terms to the equivalent controlled vocabularies. MetaMap processed the two sample questions (provided in the Online Supplementary Material). Each health consumer term was mapped to a UMLS CUI and equivalent preferred term from controlled vocabularies. MetaMap assigns a threshold of up to 1000 which indicates centrality and coverage of concept in the processed text. Supplementary Table 1 contains the output of concept extraction and mapping phase for the content of sample questions.
Step 4: term frequency–inverse document frequency calculation
Term frequency–inverse document frequency (TF-IDF) score indicates the importance of a term based on the number of word occurrences in a document. 19 Considering this frequency indicator, a concept that appears several times in a question gets a higher weight. On the other hand, a general concept that appears frequently in most of the questions is considered as a common concept with low weight.19–22 In this phase, TF-IDF weighted vectors were calculated based on extracted concepts for questions (question TF-IDF calculation) and experts (expert TF-IDF calculation). Equation (1) shows how to compute TF-IDF weight of a concept

where the value of
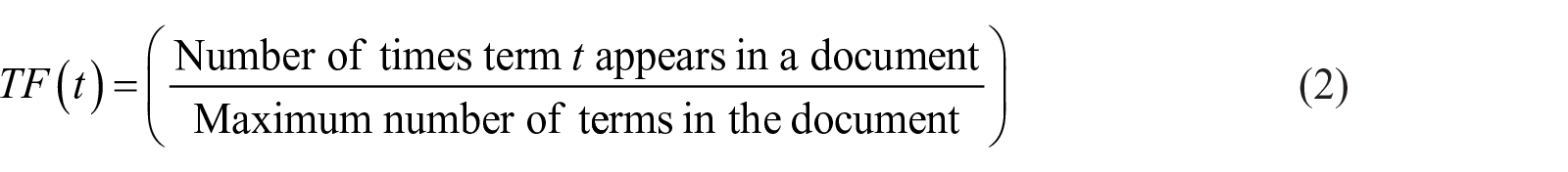

For example, TF-IDF for concept “C0023884” in question 1 which maps to “Liver” preferred term is calculated as follows

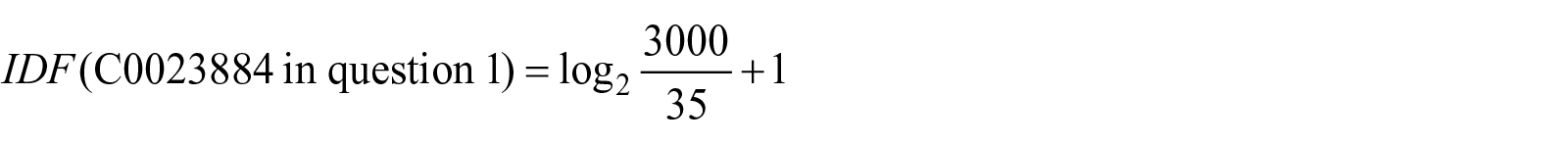
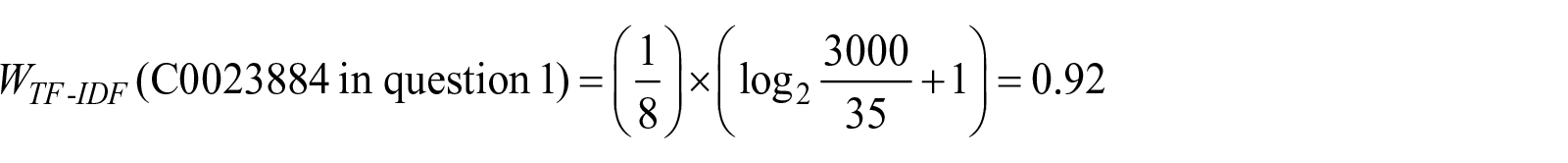
All these steps were performed on concepts extracted from selected Q&As. Supplementary Table 2 shows TF-IDF numbers for extracted CUIs of sample questions.
MetaMap weights (threshold numbers) were normalized in the range of [0–1]. Then, the total weight of a concept is obtained by multiplying the MetaMap normalized weight and the TF-IDF weight.
Step 5: similarity calculation
The similarity measure is a function that computes the degree of similarity between two vectors of UMLS concepts. UMLS concepts relevant to a question and expert are considered as weighted vectors. So, similarity measure was used to represent the similarity between a pair of questions and experts.
Question–question similarity calculation
In this phase, similarity between two questions is calculated. This similarity is divided into question–question intra-topic similarity and question–question inter-topic similarity. In question–question intra-topic similarity, Q&As of the same topics were randomly selected from our corpus and the similarity between them were computed. Also, to compute question–question inter-topic similarity, Q&As from different topics of source community were selected and cosine similarity between them was considered. Supplementary Figure 1 shows the calculation method of question–question similarity between vectors of questions concepts.
Expert–expert similarity calculation
We generated profile for some experts based on answered questions. Then we computed the similarity between profile of experts in same (expert–expert intra-topic similarity) and different topics (expert–expert inter-topic similarity). Expert–expert similarity displays the similarity of the concepts in the questions answered by the experts. This similarity can help find experts with similar profiles and activities. Intra-topic similarity shows the similarity between content of questions answered by experts in the same topic. On the other hand, inter-topic similarity calculates the similarity between concepts of questions that are answered by experts in different health topics. Supplementary Figure 2 shows the calculation method of expert–expert similarity between vectors of extracted concepts related to experts.
For example, cosine similarity between the TF-IDF vectors of the two sample questions was calculated based on equation (4)

where

Table 1 shows the final weighted vectors for questions 1 and 2. MetaMap assigns a threshold number to each extracted concept from 600 to 1000. This value indicates coverage and centrality of the concept in the processed text of the question. In this table, MetaMap average is the average of threshold numbers assigned by MetaMap for extracted concepts. Also, total weight is an aggregated weight that for each concept was calculated by multiplying MetaMap weight (semantic weight) by TF-IDF weight. Finally, TF-IDF weight is a numerical value that indicates the importance of a concept based on TF-IDF method.
Final weighted vectors for questions 1 and 2.
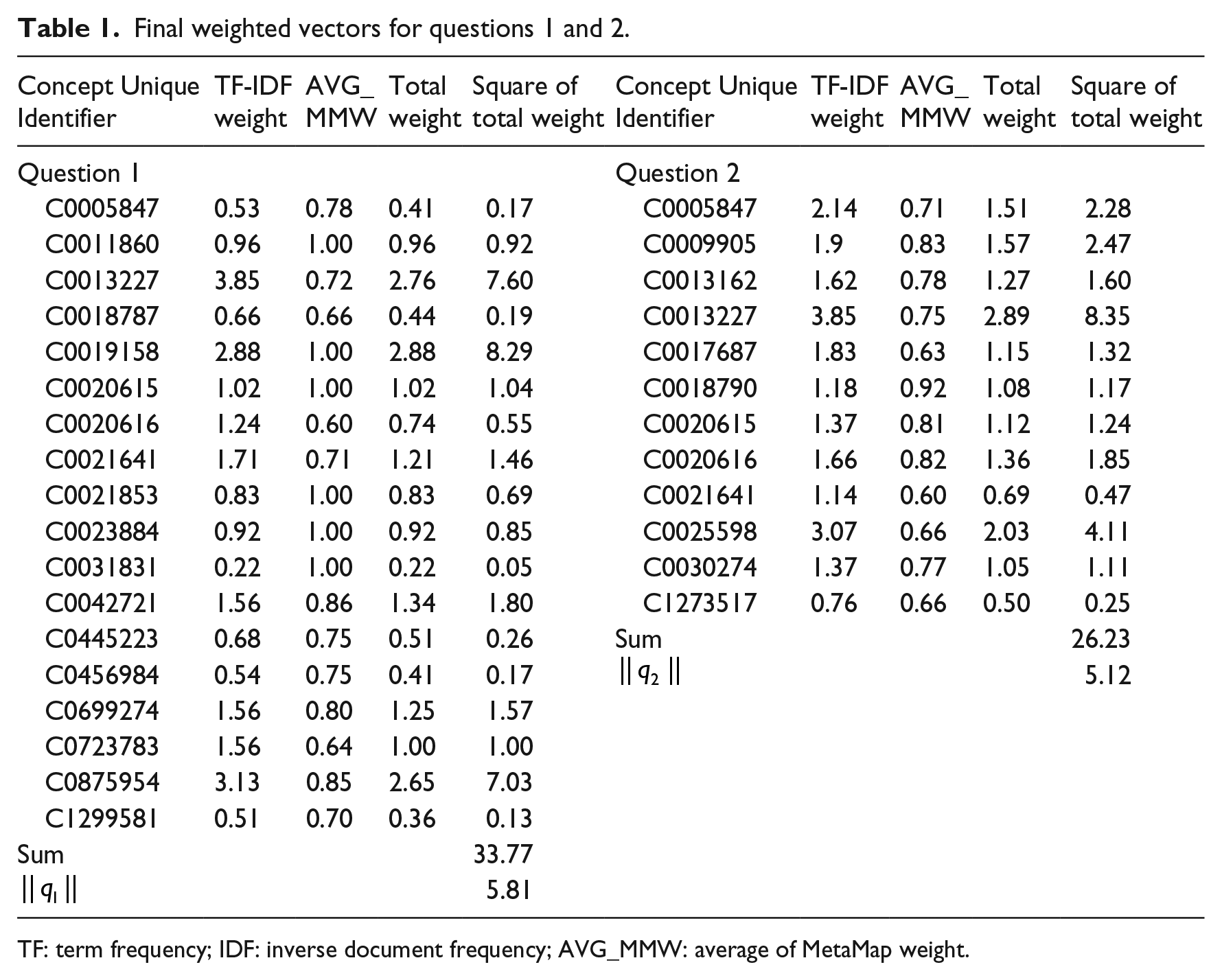
TF: term frequency; IDF: inverse document frequency; AVG_MMW: average of MetaMap weight.
Finally, the similarity between these two sample questions based on extracted concepts is 0.38

The last step is calculating cosine similarity between two questions or experts. Supplementary Figure 1 illustrates all of these steps.
Result
In our study, Yahoo Answers, WebMD, and NetWellness were considered as the source of Q&A communities. One thousand questions, together with their answers, were randomly selected from each source. First, question–question similarity was calculated in two steps: intra-topic similarity and inter-topic similarity. In inter-topic similarity, 75 Q&As from 15 different topics (five Q&As in each topic) were randomly selected from our corpus and the similarity between questions of the same topic for each community was computed. For example, the similarity between selected Q&As of “Breast Cancer” topic in NetWellness community was 0.157. Also, the average of intra-topic similarity was calculated separately for each source community. Supplementary Table 3 shows computed values of intra-topic similarity for selected topics.
Subsequently, 75 questions were selected from 15 topics mentioned above to compute the similarity between questions of different topics. Supplementary Table 4 contains the result of inter-topic similarity for a few selected topics. For example, the average similarity between Q&As of “Diabetes” and “Breast Cancer” in NetWellness was 0.015. Supplementary Figure 3 shows the similarity distribution in details.
Our result showed a considerable gap between intra-topic and inter-topic similaritys. This difference in the similarity of the two groups was apparent in nearly all three studied sources.
In the second phase of our study, we computed expert–expert similarities. To calculate the similarity between questions that had been answered by an expert, 400 answered questions by 20 experts (20 Q&As for each expert) were randomly selected from NetWellness. Supplementary Table 5 indicates distribution of selected experts in different health topics.
After the concept extraction phase for selected Q&As, TF-IDF weighting vectors were computed based on equation (2) for each expert. Similarity between experts was also calculated according to the cosine similarity between their TF-IDF vectors. Table 2 shows similarity numbers as a 20 × 20 matrix considering expert-related topics. Since the similarity matrix is symmetric, the similarity results were represented by an upper triangular matrix which all the entries below the main diagonal are zero. The value of each cell in this matrix indicates the similarity between experts in related row and column. The similarity between a vector and itself is 1; therefore, the values on the main diagonal of similarity matrix are all 1. As an example, the similarity between expert 2 and expert 3, who are both represented “Breast Cancer” topic, is 0.64. On the other hand, the similarity between expert 3 and expert 9, who are assigned to other topics “Breast Cancer” and “Diet and Nutrition,” are calculated as 0.04.
Expert–expert similarity matrix.
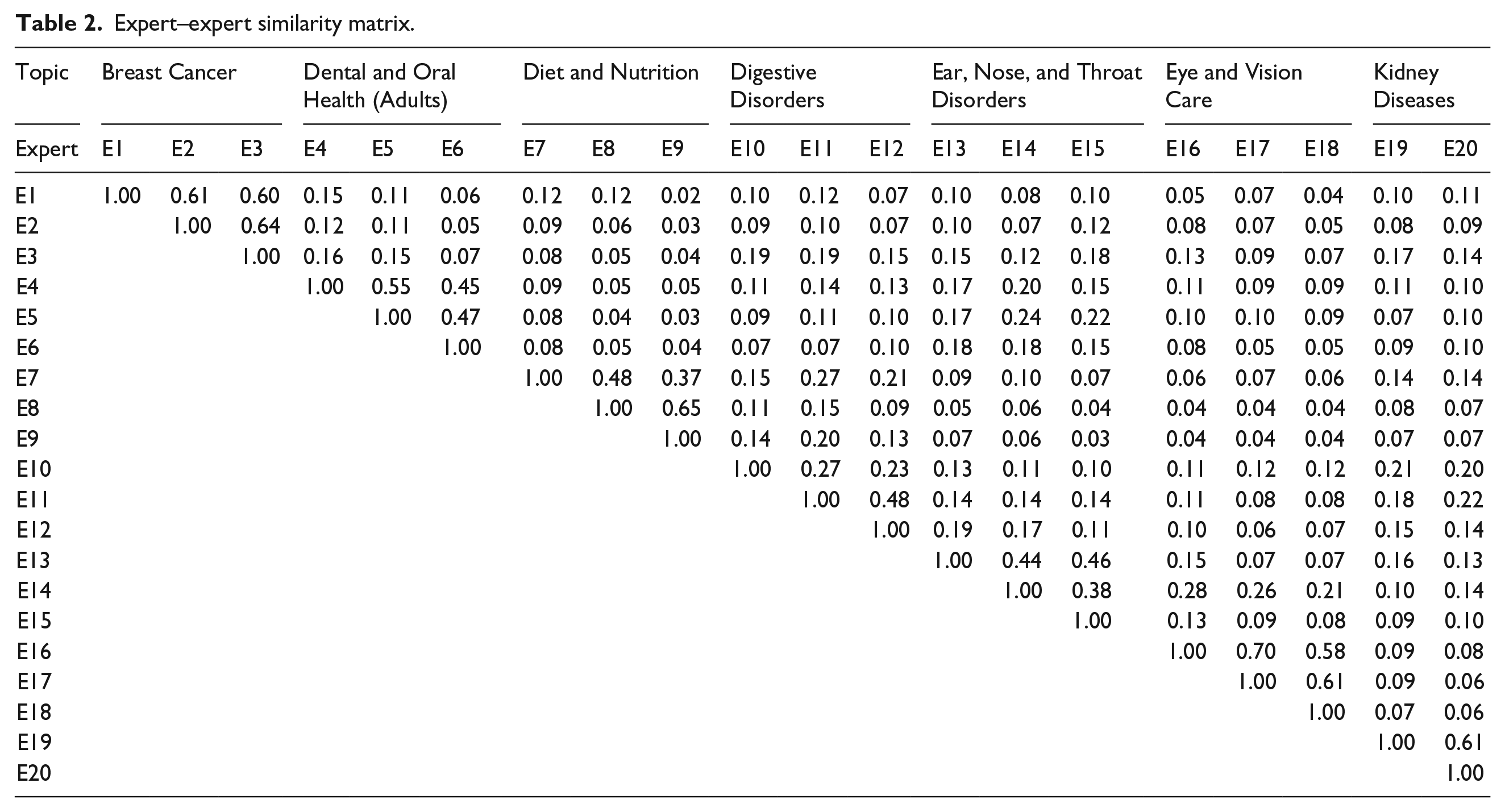
Figure 2 shows the integrated intra-topic and inter-topic similarity of the mentioned experts. The size of the circles represents the degree of similarity. Larger circles indicate greater similarity, where they can be seen in the main diagonal.
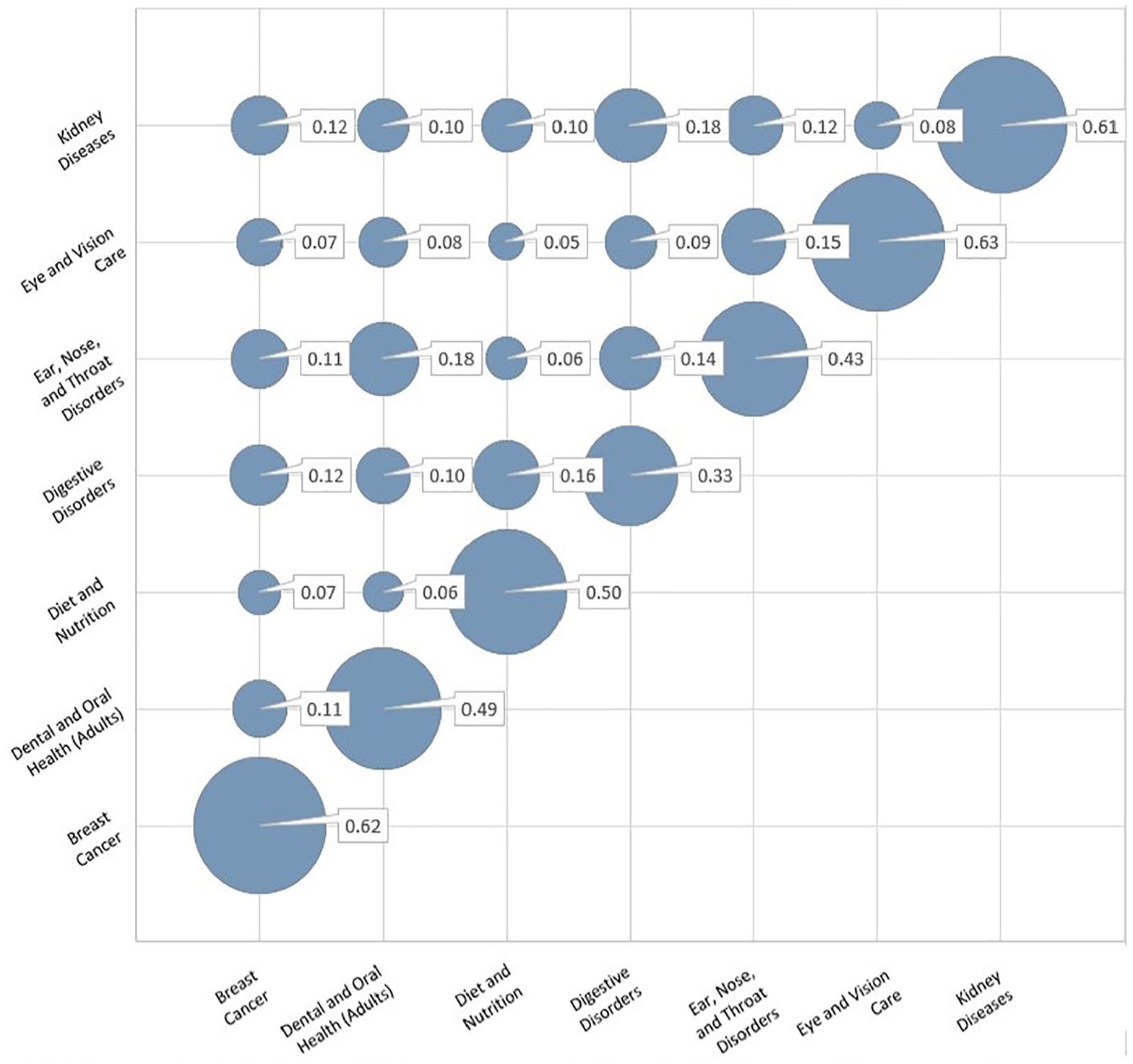
Intra-topic and inter-topic similarity between multiple domains experts based on the answered questions.
Discussion
In this study, the similarity between submitted questions by healthcare consumers in online communities was investigated. There was a significant difference between intra-topic similarity and inter-topic similarity which could be helpful for finding similar questions and experts. Therefore, it seems that concept-based similarity computing method has potentials in developing intelligent Q&A retrieval systems that contain auto recommendation of similar questions and experts.
There are three main steps in health information search process: (1) selecting information sources, (2) evaluating collected data, and (3) processing and preparing information for users. 23 People usually ask about their health related problems from several sources. The result of a study 24 which examined consumer health information searching practice showed that online users usually consult more than three sources during their request for information. The majority of consumers (90.5%) will usually look to a second source, where more than half of them (76.2%) consult a third source to find their intended information. There are several reasons for such behavior. First, they do not get a compelling answer from a given resource. Second, most of health consumers seek for a more comprehensive answer or read additional opinions. Third, some people hop from one website to another in order to find similar answer, for a given health problem, that confirms their earlier findings. In this study, comparing the results of intra-topic similarity and inter-topic similarity indicated the similarity between questions from same topics was significantly higher than the similarity between questions from different topics (provided in Supplementary Tables 3 and 4). It means the overlap between extracted concepts of questions from different topics is little. So using concept-based method can classify questions of different topics automatically. Therefore, concept-based similarity computing methods can help find similar questions and experts from different online resources and collect comprehensive answers.
Consumers usually evaluate the quality of online information (like Q&As seen on the websites) based on the credibility of author(s) and references. Also, appearance of similar results for a health condition search in several online sources of information may indicate the validity of the information. Thus, allocating an online environment where users can consolidate information from multiple resources with reliable references increases their confidence about such information. Concept-based similarity computing method can help developers to create a “Meta” Q&A health community from available online Q&A health societies. A lot of questions from several sources can be automatically integrated in various topics by similarity between their concepts. This method will simplify information search process from multiple sources and facilitates information retrieval among a huge number of historical Q&A that are stored as valuable knowledge bases.
Comparing the results of intra- and inter-topic similarity could indicate that the similarity among questions of the same topics is significantly higher than the similarity between questions from different topics. Such results can be achieved by gathering questions from several Q&As communities as the source of information from various health topics. Also, expert–expert similarity has been calculated for different health topics. Our results indicate the similarity between the content of questions that were answered by experts for the same topic (intra-topic similarity) is higher than the similarity between concepts of questions that were answered by experts in a different health topic (inter-topic similarity) (Figure 2). Therefore, identification of similar experts can be done in an automated fashion. Online community’s web services, by using our method, can recommend similar experts to consumers who could not get compelling answer for their health problem from other experts. Finally, from the viewpoint of the online community organizer, calculating cosine similarity based on extracted concepts can help community support team programmatically define a process for categorizing questions and experts by topics.
Limitation
The sample questions in this study were only gathered from three popular online health communities. Considering submitted Q&As from more online communities can lead to a more accurate and comprehensive results.
Conclusion
Concept-based similarity computing methods can be used in developing intelligent Q&A retrieval systems that contain auto recommendation functionality for similar questions and experts. Also, applying concept-based similarity methods have potentials to integrate multiple communities where users can find additional information from similar questions that were asked in other communities.
Supplemental Material
Supplementary_Figure_1 – Supplemental material for Similarity of medical concepts in question and answering of health communities
Supplemental material, Supplementary_Figure_1 for Similarity of medical concepts in question and answering of health communities by Hamid Naderi, Sina Madani, Behzad Kiani and Kobra Etminani in Health Informatics Journal
Supplemental Material
Supplementary_Figure_2 – Supplemental material for Similarity of medical concepts in question and answering of health communities
Supplemental material, Supplementary_Figure_2 for Similarity of medical concepts in question and answering of health communities by Hamid Naderi, Sina Madani, Behzad Kiani and Kobra Etminani in Health Informatics Journal
Supplemental Material
Supplementary_Figure_3 – Supplemental material for Similarity of medical concepts in question and answering of health communities
Supplemental material, Supplementary_Figure_3 for Similarity of medical concepts in question and answering of health communities by Hamid Naderi, Sina Madani, Behzad Kiani and Kobra Etminani in Health Informatics Journal
Supplemental Material
Supplementary_Table_1 – Supplemental material for Similarity of medical concepts in question and answering of health communities
Supplemental material, Supplementary_Table_1 for Similarity of medical concepts in question and answering of health communities by Hamid Naderi, Sina Madani, Behzad Kiani and Kobra Etminani in Health Informatics Journal
Supplemental Material
Supplementary_Table_2 – Supplemental material for Similarity of medical concepts in question and answering of health communities
Supplemental material, Supplementary_Table_2 for Similarity of medical concepts in question and answering of health communities by Hamid Naderi, Sina Madani, Behzad Kiani and Kobra Etminani in Health Informatics Journal
Supplemental Material
Supplementary_Table_3 – Supplemental material for Similarity of medical concepts in question and answering of health communities
Supplemental material, Supplementary_Table_3 for Similarity of medical concepts in question and answering of health communities by Hamid Naderi, Sina Madani, Behzad Kiani and Kobra Etminani in Health Informatics Journal
Supplemental Material
Supplementary_Table_4 – Supplemental material for Similarity of medical concepts in question and answering of health communities
Supplemental material, Supplementary_Table_4 for Similarity of medical concepts in question and answering of health communities by Hamid Naderi, Sina Madani, Behzad Kiani and Kobra Etminani in Health Informatics Journal
Supplemental Material
Supplementary_Table_5 – Supplemental material for Similarity of medical concepts in question and answering of health communities
Supplemental material, Supplementary_Table_5 for Similarity of medical concepts in question and answering of health communities by Hamid Naderi, Sina Madani, Behzad Kiani and Kobra Etminani in Health Informatics Journal
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by Medical Research Council of Mashhad University of Medical Sciences (grant number 951317).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
