Abstract
The Research Domain Criteria, launched by the National Institute of Mental Health, is a new dimensional and interdisciplinary research framework for mental disorders. The Research Domain Criteria matrix is its core part. Since an ontology has the strengths of supporting semantic inferencing and automatic data processing, we would like to transform the Research Domain Criteria matrix into an ontological structure. In terms of data normalization, which is the essential part of an ontology representation, the Research Domain Criteria elements (mainly in the Units of Analysis) have some limitations. In this article, we propose a series of solutions to improve data normalization of the Research Domain Criteria elements in the Units of Analysis, including leveraging standard terminologies (i.e. the Unified Medical Language System Metathesaurus), context-combining queries, and domain expertise. The evaluation results show the positive (Yes) percentage is more than 80 percent, indicating our work is favorably received by the mental health professionals, and we have formed a good data foundation for the Research Domain Criteria ontological representation in the future work.
Keywords
Introduction
In the realm of mental health, there are two major diagnostic criteria:
The RDoC framework focuses on dimensional psychological constructs (concepts), which may reflect mechanisms implicated in psychopathology. 6 At the core of the framework is the RDoC matrix. The rows of the matrix represent various constructs, grouped hierarchically into broad domains of function. The columns denote different Units of Analysis, which are the methods used to investigate and understand constructs. Currently, there are five domains: Negative Valence Systems, Positive Valence Systems, Cognitive Systems, Social Processes, and Arousal and Regulatory Systems, as well as eight types of Units of Analysis: Genes, Molecules, Cells, Circuits, Physiology, Behavior, Self-report, and Paradigms. The elements of the matrix were generated based on extant research and vetted by over 200 researchers from relevant fields. 7 Table 1 shows examples of elements in the current version of the RDoC matrix.
Element examples of the RDoC matrix. a
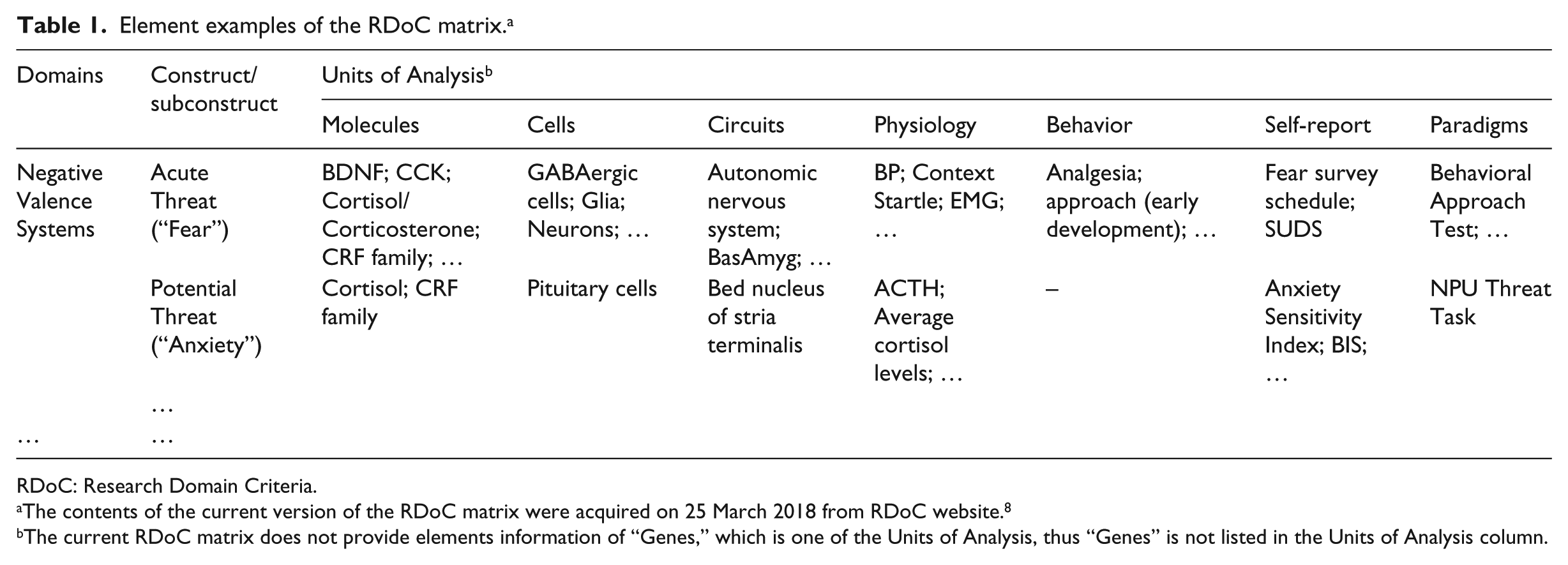
RDoC: Research Domain Criteria.
The contents of the current version of the RDoC matrix were acquired on 25 March 2018 from RDoC website. 8
The current RDoC matrix does not provide elements information of “Genes,” which is one of the Units of Analysis, thus “Genes” is not listed in the Units of Analysis column.
Since its inception, RDoC has attracted extensive attention from the psychiatric communities, who actively discuss its values and applications. Lebowitz et al. 9 found that applying the RDoC model had the potential to build upon the advances of etiology and treatment through integrating previously siloed research areas and to facilitate translational research. Smucny et al. 10 demonstrated that an RDoC-based framework could understand the important aspect of mental disorders, that is, cognitive control dysfunction might be described as a continuum of deficits across schizophrenia and bipolar disorder. Wildes and Marcus 11 concluded that the RDoC framework provided an exciting alternative for treatment development research in eating disorders.
From the perspective of knowledge representation, it is worth to transform the RDoC matrix into an ontological structure. An ontology is an explicit specification of a conceptualization, with the purpose of knowledge sharing and interoperation among programs based on that shared conceptualization. 12 It can realize machine-readability, consistency checking, and semantic inferencing and may further support automatic data processing and knowledge discovery. 13 One of the most representative ontologies in the biomedical field is Gene Ontology, 14 which has played fundamental roles in dynamic maintenance, sharing, and interoperation of genome-related information. 15 Alike, the ontological representation of the RDoC may significantly support automatic data processing (annotating, extracting, and integrating) and facilitate intelligent diagnosis of mental disorders, especially in the big data era. 16
The RDoC is a dynamic initiative 17 and open to the community for ideas and efforts. The RDoC dimensions have been instantiated as experimental constructs that are expected to change rapidly over time on the basis of new data. 18 A lot of constructive suggestions are constantly made by mental health peers. Mittal and Wakschlag 19 discussed a new way to conceptualize development within the RDoC framework. Watson et al. 20 proposed some intriguing revisions to the domain of Negative Valence System. Based on detailed analyses of methodological and conceptual challenges of RDoC, Lilienfeld 21 suggested some modifications to the design of constructs. Using terminological/ontological assessment criteria, Ceusters et al. 22 pointed out some shortcomings of the current RDoC, including element terms lacking face value and absence of exactly represented relationships. To summarize, the suggested improvements could be classified into two aspects: content (constructs design) and form (data normalization). For ontology construction, data normalization is an essential necessity, and also the core part our article is currently focusing on.
In this article, we propose informatics-based approaches to facilitating data normalization and standardization of the RDoC elements. The rest of article is organized as follows. In the “Methods” section, we analyze the limitations existing in the RDoC elements and propose our normalization principles, rules, and methods. In the “Results” and “Discussion” sections, we present domain experts’ evaluations and the normalized results, discuss the issues awaiting better solutions and plans for future efforts. We conclude in the “Conclusion” section.
Methods
In this section, we first review and summarize the problems of RDoC elements in terms of data normalization, then introduce four normalization principles and combinational methods to solve different kinds of problems. Figure 1 shows the overall flowchart of data normalization and standardization of the RDoC elements in the Units of Analysis.
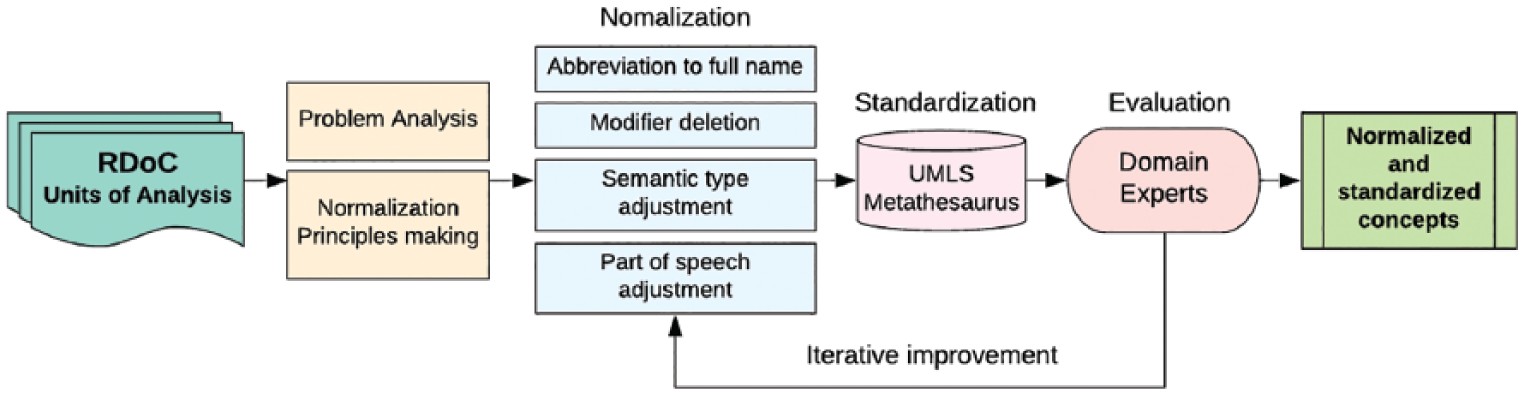
Overall flowchart of data normalization and standardization of the RDoC elements in the Units of Analysis.
Problem analysis
Based on the assessment criteria of terminology/ontology, 23 we summarized the main problems in terms of data normalization, as shown in Table 2.
Problems of data normalization.
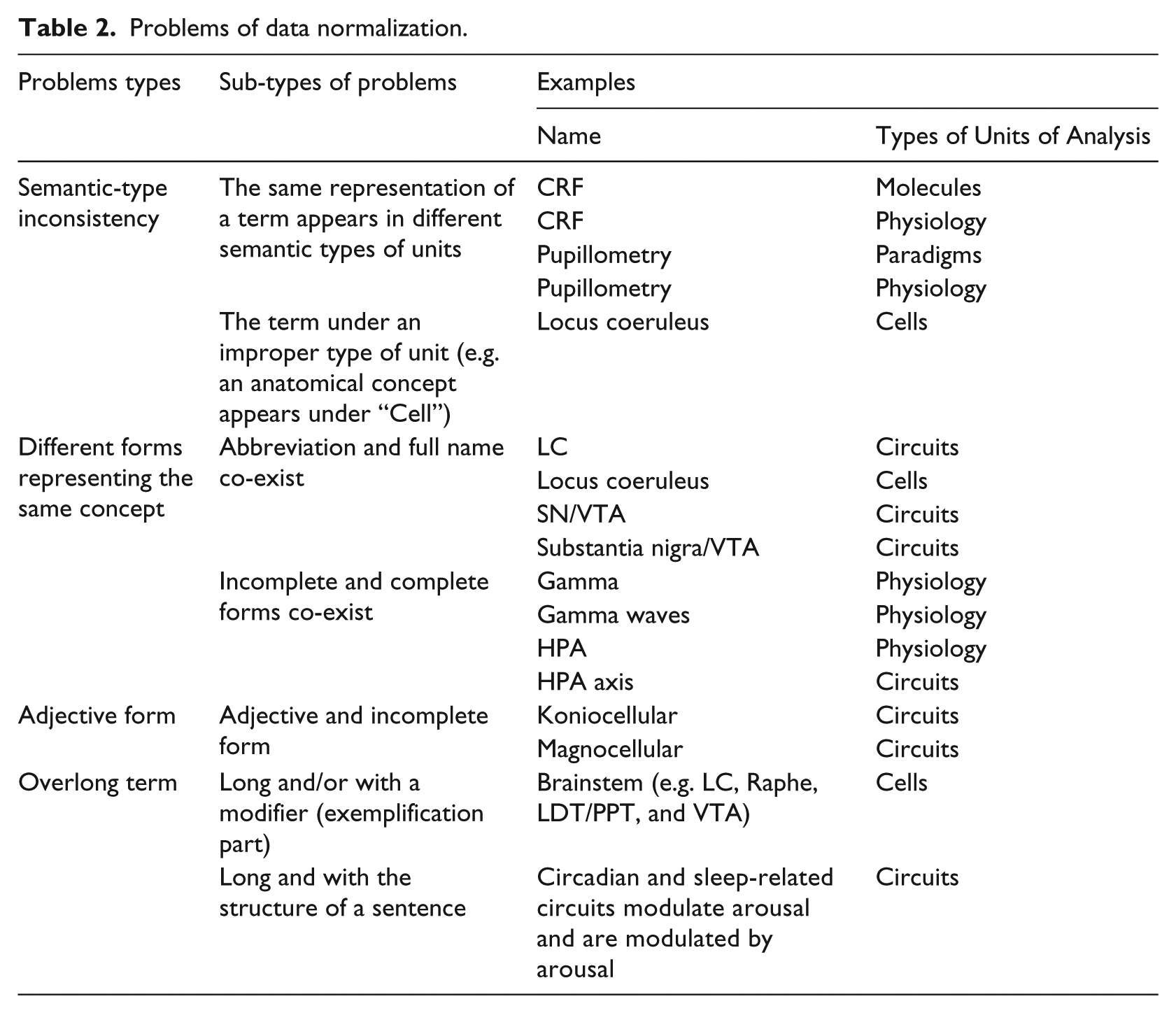
Normalization principles
In general, the criteria of ontology design include clarity, coherence, extendibility, and minimal encoding bias. 24 To guide our normalization work, we established four principles as follows.
Preserve the original intention of the RDoC elements
Understanding and preserving the original intention of the RDoC elements is our first and fundamental principle, since all these elements were carefully selected by the RDoC workgroup. 25 What we would like to contribute is to improve their normalization and standardization, rather than to change their content.
Keep semantic type consistency
Many elements appearing under specific Units of Analysis are not consistent with their head semantic types (e.g. Molecules, Cells, and Physiology). We try to adjust their forms to keep consistency of the asserted semantic type.
Reduce ambiguity
An abbreviation could map to different full names, which may cause ambiguity. Thus, we try to extend the abbreviations to their full names. To ensure a right extension, we need to consider the specific context under which the element appears.
Improve standardization
The Unified Medical Language System (UMLS) Metathesaurus is the most comprehensive and recognized biomedical thesaurus. It is organized in the unit of concepts and links similar names for the same concept from nearly 200 different vocabularies. 26 One term from among the various names within a concept is identified as the preferred term. Preferred terms are computed from a list of ranked source vocabularies and can represent this concept. 27 We treat the Metathesaurus as our normalization gold standard and select preferred names as normalization forms. For the terms that cannot map to the Metathesaurus, the reliable information source (medical literature, official websites, and monographs) is the alternative reference for standardization.
Data pre-processing
Data acquisition by HTML parser
Since the elements in Units of Analysis are located in different web pages of domains/constructs, 8 we used an HTML parser 28 to acquire all the elements and stored them in an Excel sheet, while preserving their corresponding domains/constructs/subconstructs information for the relation construction in the next step.
Data merging
A single term often appeared several times under the same or different types of units. For example, “Acetylcholine” appeared seven times under “Molecules.” For the same terms under the same unit, we assumed that they represent the same meaning and retained one of them as an independent concept. By contrast, for the same terms appearing under different units, we assumed that they represented different meanings, thus kept all of them.
Normalization rules and methods
Regarded as the normalization standard, we map all terms to the Metathesaurus. Different situations (no matching, partial matching, or exact matching) would occur, which also constitute our normalization rules and steps. Figure 2 shows the flowchart of data normalization:
Step 1: map terms to the Metathesaurus: if no match is found, we combine a proper contextual term to search online together, to find the most possible form. The most possible term means a good match that is not only in the right form but also has the right semantic meaning. More specifically, it should be well congruent with the environment where the term appears (the domain/construct/subconstruct and the Unit of Analysis). Furthermore, it needs to be supported by more than three reliable information sources (medical literature, official websites, or monographs), for example, the term appears in three medical papers in PubMed. If a match is found, go to Step 2.
Step 2: if a match is found: if there are one or more exact matchings (since one abbreviation may correspond to several full names), choose the most appropriate one which would best fit the semantic type (e.g. Molecules, Cells, or Circuits) of the term and use Metathesaurus preferred name as the normalized form. If there is no exact matching, select the possible segments in the Metathesaurus results which might match the semantic type of term alike. Then, search the semantic segments online, to iteratively look for the right form which can both align with the context and be supported by more than three reliable information sources, the same with Step 1.
Step 3: domain expert evaluation: two domain experts review the modified results (obtained from Steps 1 and 2), to evaluate and give suggestions. Based on their evaluations, we reference Metathesaurus again and finalize the normalization.
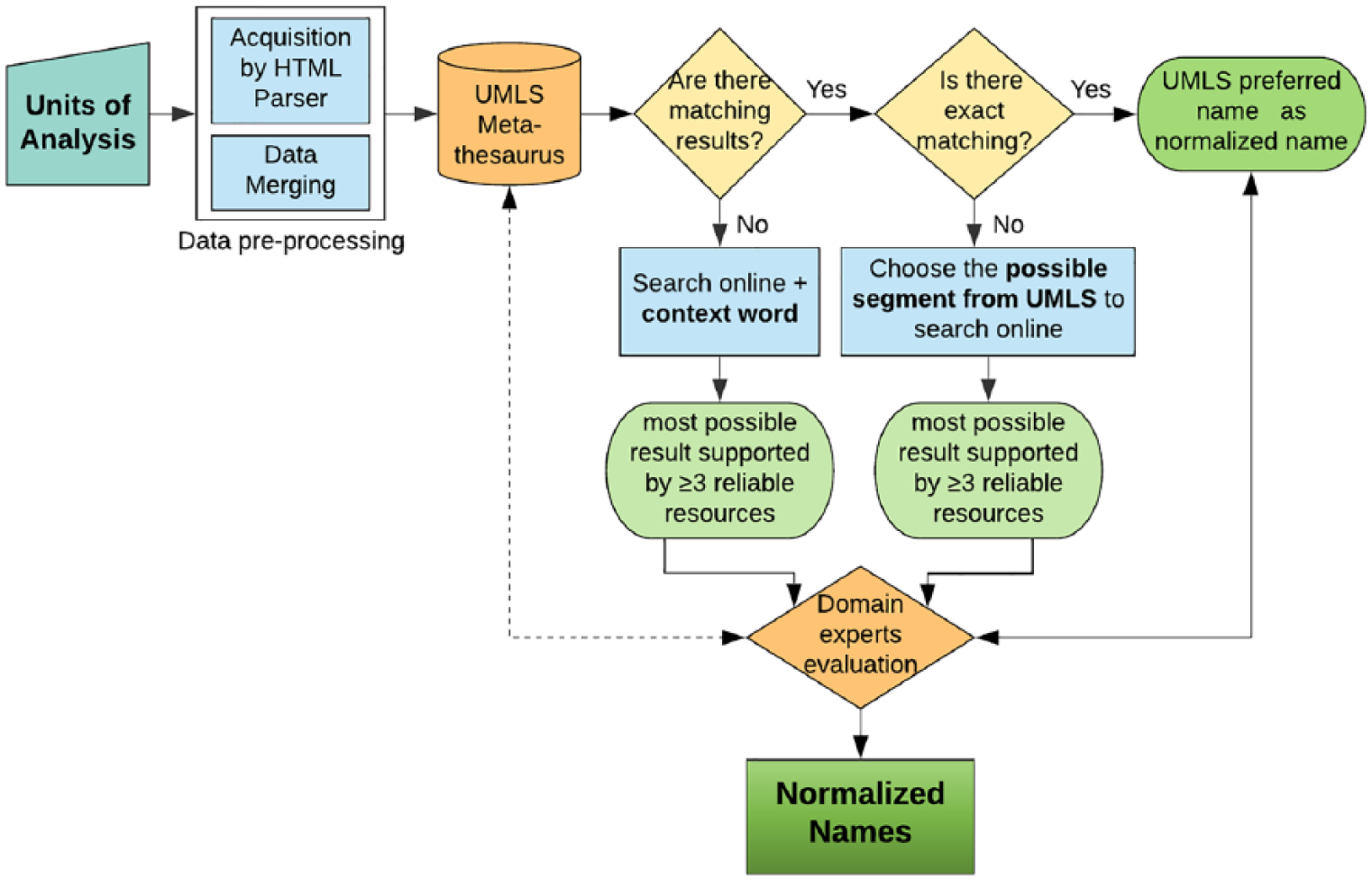
Flowchart of data normalization.
Following these rules, we implemented four types of data normalization.
Keep semantic type consistency
A lot of terms were not congruent with their head semantic type. For example, in the “Cells” units, there were several anatomical concepts:
Extend abbreviations to full names
There were a lot of abbreviations. In particular, some terms were very short, such as A1, D2, and N1, which made the normalization work more difficult. To understand their intended meaning and acquire the right full names, we designed a query strategy that combined proper contextual words. As the RDoC framework conceptualized mental illnesses as brain disorders, 25 we chose the contextual words related with the brain/central nervous system (“cortex,” “nucleus,” “pathway,” etc.) to jointly search online. Furthermore, we shifted among these contextual words to align with their specific environment.
For example, “A1” appeared under “Circuits.” When querying it in the Metathesaurus (2017AB version), we got 1391 results, making it hard to determine a reasonable full name. Then, we selected “brain” as the possible contextual term and searched together with “A1,” we found the “A1 cortex” that appeared in several medical papers and had a formal definition. Based on domain knowledge, “A1 cortex” could be congruent with the semantic type of “Circuits.” So, “A1 cortex” was chosen as the candidate full name. Furthermore, we matched “A1 cortex” with the Metathesaurus again and got the preferred form “Primary auditory cortex” as its final normalized name.
Adjust parts of speech
There were several adjective and incomplete terms in RDoC, such as
Delete modifiers to make terms clean
Several RDoC terms were long, and some of them had modifier parts, for example, “Brainstem (e.g. LC, Raphe, LDT/PPT, VTA),” “EEG features e.g. evoked gamma,” and “Photoplethysmography (skin color measure of capillary dilation; temperature).” In general, concepts in the ontology/terminology seldom contain the exemplification parts in their preferred forms. Therefore, we deleted the modifiers to make terms clean and adjusted their semantic type accordingly. Meanwhile, we preserved the modifier information in the annotation property of the concept in the ontology. For example, we modified “Brainstem (e.g. LC, Raphe, LDT/PPT, VTA)” to “Brainstem neurons” since it appeared under “Cells.”
Results
We checked through all terms of Units of Analysis. For the terms with improper forms as described above, we modified them and provided the modification descriptions in detail.
Evaluation by domain experts
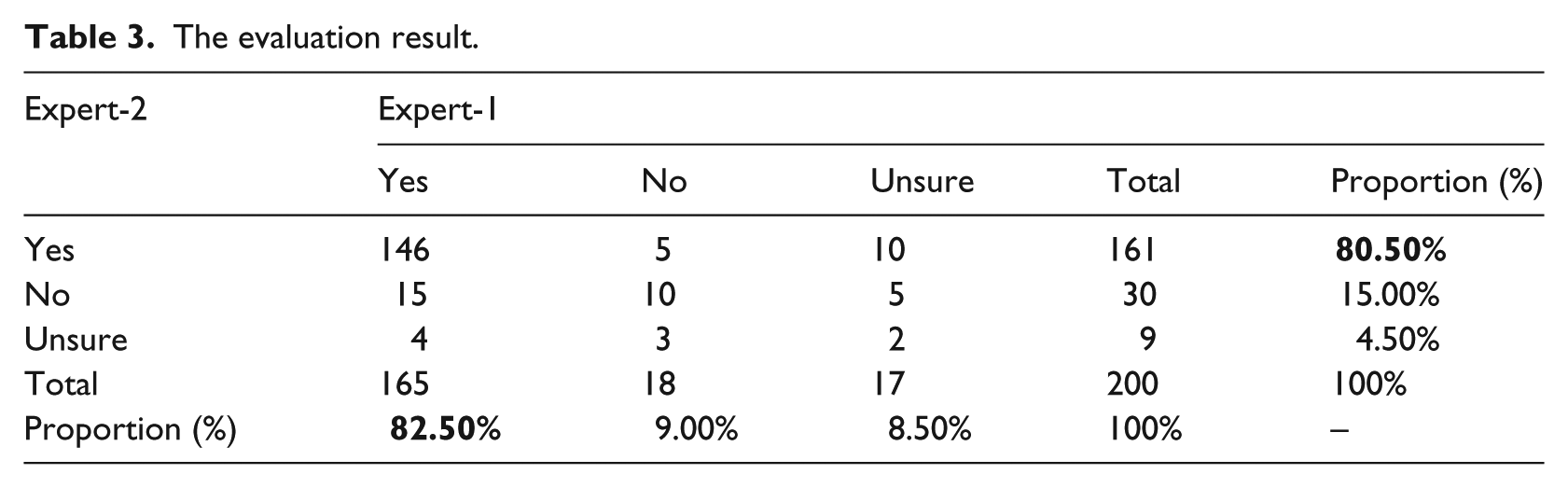
To guide normalization direction and improve accuracy, we invited two mental health professionals (our co-authors Dr S.S. and Dr J.E.H) as domain experts to evaluate the normalization results. Among 795 terms in total, we randomly selected 200 terms. The two experts provided their confidence levels (“Yes,” “No,” and “Unsure”) on the suggested modifications. In the cases of “No” and “Unsure,” they would provide “Suggested Modification” as better solutions. Their combined evaluation results are shown in Table 3: the columns represent the evaluation results of Expert-1 and the rows represent that of Expert-2. In the “Yes,” “No,” and “Unsure” part, the numbers indicate their combined results. For example, “146,” corresponding to two “Yes,” indicates that there are 146 normalization terms that both experts judged “Yes.” Next to it, “5” indicates that there are five terms that Expert-1 judged “Yes” while Expert-2 judged “No.” In the “Total” and “Proportion” sections, the numbers are statistical results of a single expert. For example, “161” means that there are 161 terms in total that Expert-2 judges “Yes” (sum of 146 + 5 + 10); next to it, “80.5%” means that the “Yes” by Expert-2 occupies the proportion of 80.5 percent (161/200). It can be seen that both experts gave more than 80 percent “Yes” to the results, that is, 82.5 percent (Expert-1) and 80.5 percent (Expert-2), respectively.
The evaluation result.
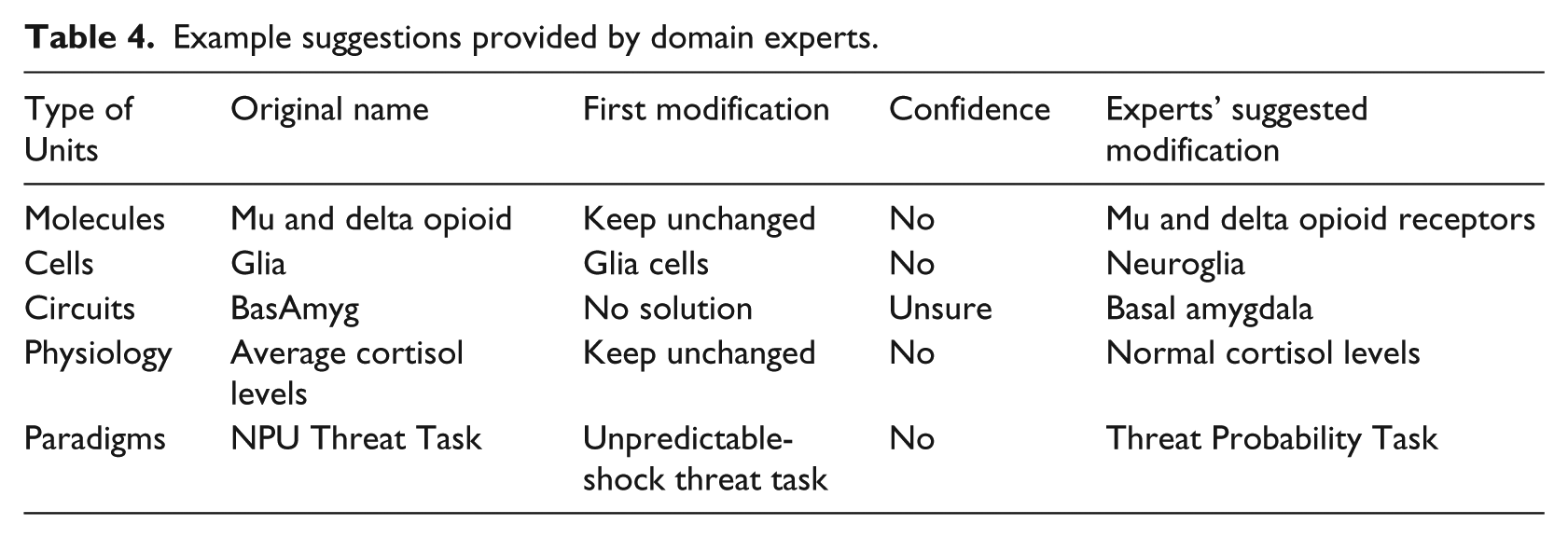
For the terms rated by “No” or “Unsure,” our domain experts provided their suggestions of more appropriate normalization, for example, adding “receptors” to “Mu and delta opioid” and extending “BasAmyg” to its full name “Basal amygdala.” Table 4 shows some examples.
Example suggestions provided by domain experts.
Normalization results
Based on domain experts’ evaluation and suggestions, we re-checked all terms and matched them with the UMLS Metathesaurus again. If they could be matched exactly, we chose the preferred name as the normalization forms; if there were no exact matches, we adopted the names suggested by domain experts. The normalization results (representative examples) are shown in Table 5.
Normalization results (partial).
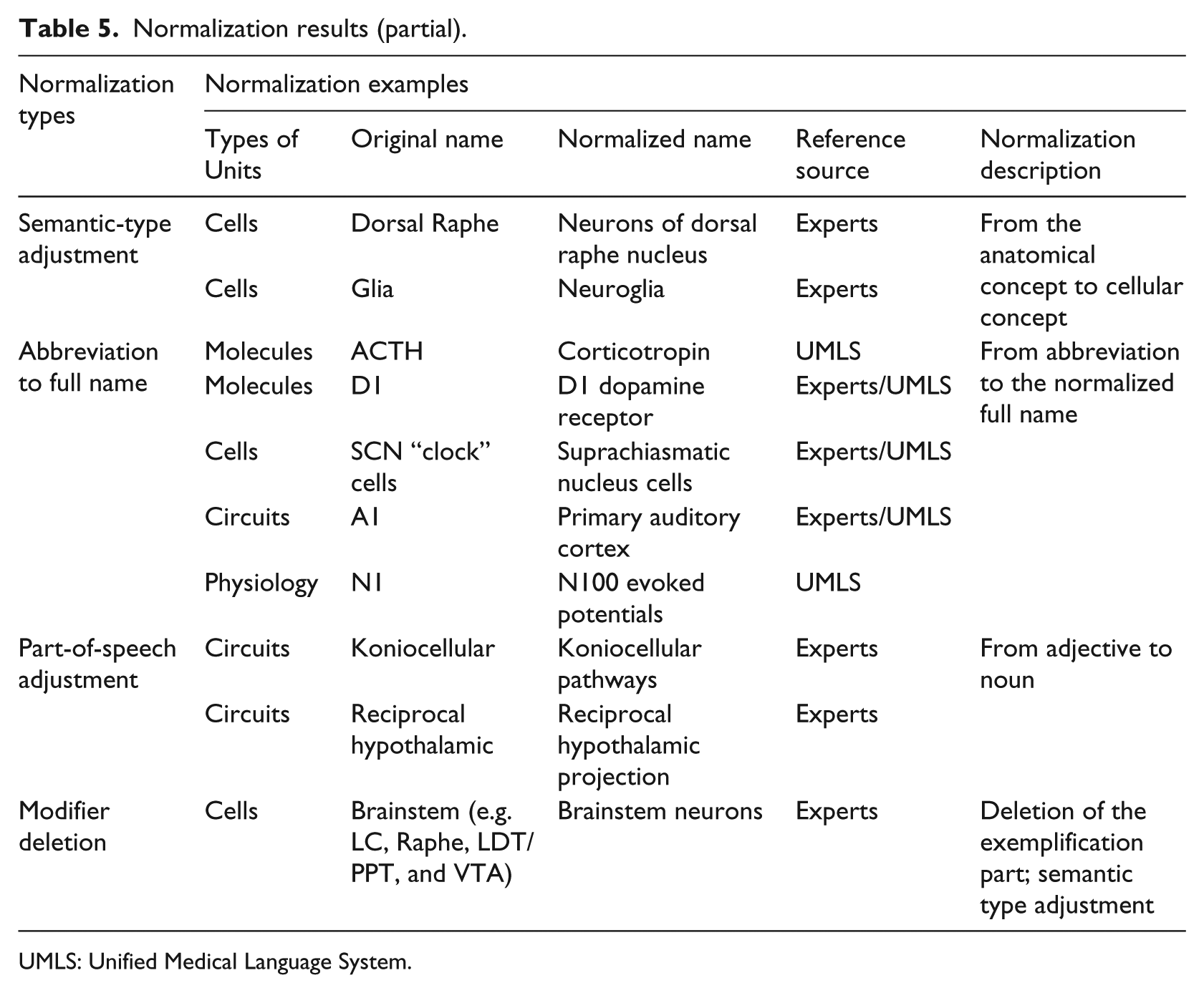
UMLS: Unified Medical Language System.
Discussion
Issues awaiting better solutions
There are still some issues awaiting better solutions.
The same terms appearing under different units
Whether the same term can belong to different semantic types of Units is an issue worthy of deep consideration. For example, “CRF” appears under both “Molecules” and “Physiology.” Currently, we retained both of them as individual concepts. Whether transforming the “CRF” under “Physiology” into “corticotropin-releasing hormone changes” can represent a proper physiological concept needs to be further discussed.
Overlong terms
For the overlong terms without modifier parts, we kept them unchanged, for example, “time-series of response times to extract variability and frequency domain analyses visual search” and “Circadian and Sleep-related circuits modulate arousal and are modulated by arousal.” How to modify these overlong terms also awaits a good solution.
Combinational terms
There are several terms that use “and” to combine two or more concepts, which would lead to no exact matching in the Metathesaurus or other terminologies/ontologies. For example, “dopamine and dopamine-related molecules” and “Lateral, perifornical, and dorsomedial hypothalamuses” consist of more than two terms. To follow Common Data Models (CDMs) rationale 29 and split them into a couple of independent concepts would be an alternative solution.
Future efforts
Constructing a useful ontology is never an easy task. In the future, we would like to make our efforts in the following ways:
Further data normalization and organization
Since there are several terms that cannot be mapped to the Metathesaurus, we will use other recognized vocabularies/ontologies that are not included in the Metathesaurus (e.g. ontologies from the OBO Foundry and the NCBO BioPortal) to normalize them. In addition, a controlled vocabulary often has a well-designed structure. Different from the domain/constructs which have explicit hierarchies, the elements in the Units of Analysis are currently without a systematic design. We would build hierarchies for them, referencing MeSH 30 and other related vocabularies/ontologies, to allow them to better serve as classes in ontology. What’s more, logical hierarchies may support granularity control of the terms.
Ontological representation
Semantic relationships are critical parts for an ontology. Currently, the RDoC matrix provides no information about relations among domain/constructs and Units of Analysis. So, we would like to construct their possible relationships, leveraging the UMLS Semantic networks. Based on the normalized terms and explicit semantic relationships, we will further build the ontological representation of the RDoC.
Conclusion
In this article, we proposed a combinational approach to normalize and standardize the RDoC elements. We mainly realized four types of modifications: ensuring semantic type consistency, extending abbreviations to full names, adjusting part-of-speech, and deleting modifiers to make terms clean. The results were favorably recognized by our domain experts. The normalized data would be further represented in an ontology, which can support automatic data processing and intelligent application of the RDoC. Our research may promote the realization of the ultimate goal of the RDoC, which is to foster translational science and improve the outcomes of mental disorders.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was partially supported by the National Library of Medicine of the National Institutes of Health under Award Number R01 LM011829 and the Cancer Prevention Research Institute of Texas (CPRIT) Training Grant #RP160015. The content is solely the responsibility of the authors and does not represent the official view of the funding agencies.
