Abstract
Clinical practice guidelines are valuable sources of clinical knowledge for healthcare professionals. However, the passive dissemination of clinical practice guidelines like publishing in medical journals is ineffective in changing clinical practice behaviour. In this work, we proposed a framework to help adopting an active clinical practice guideline dissemination approach by automatically extracting clinical knowledge from clinical practice guidelines into a clinical decision support system–friendly format. The proposed framework is intended to help human modellers by automating some of the manual formalization activities in order to minimize their manual effort. We evaluated our framework using all recommendations from two clinical practice guidelines produced by the Scottish Intercollegiate Guidelines Network: the ‘Management of lung cancer’ clinical practice guideline and the ‘Management of chronic pain’ clinical practice guideline. We conclude that the proposed framework can be effectively used to formalize drug and procedure recommendation in clinical contexts.
Keywords
Introduction
Clinical practice guidelines (CPGs) as defined by the Institute of Medicine are ‘systematically developed statements to assist practitioner and patient decisions about appropriate health care for specific clinical circumstances’. 1 CPGs offer concise instruction on the optimal care for the patient based on the latest clinical findings. The main benefit of CPG is to improve the quality of care and the consistency of care. For a healthcare professional, a CPG can help offer explicit recommendations when a healthcare professional is uncertain about how to proceed. 2
It is been shown that passive dissemination of CPGs like publishing in a medical journal is ineffective in changing clinical practice behaviour. 3 Many healthcare practitioners are not aware of the existence of the CPG, and even when they are directed to the relevant CPG, they experience difficulties using it in their daily practice. 4 Nevertheless, integrating CPG knowledge into clinical systems, such as decision support systems, has shown to be more effective. 5 In order to best benefit from the CPGs’ knowledge by following an active CPG dissemination approach, an interest in formalizing medical knowledge contained in CPGs has grown. The heritage of the narrative text CPGs will remain a source of medical knowledge that awaits its formalized counterpart to be developed and integrated into clinical systems. This integration could be manifested in different technical facades like clinical decision support system (CDSS) or extension to the electronic health record (EHR) system used by healthcare facilities.
There are several formal languages developed to help modelling clinical guidelines into computer-interpretable guidelines (CIGs); the review study 6 presents some of the CIG modelling languages. Assuredly, the development of the guideline modelling languages is an important step towards facilitating the CPG formalization process, yet the formalization task remains laborious and complex, mainly because it requires human modellers with two different areas of expertise: a medical expertise to correctly interpret the medical knowledge of CPGs and a knowledge engineer expertise to correctly represent the medical knowledge using the syntax of the modelling language.
Nonetheless, CPG formalization approaches have been published.7–14 These approaches are either based on a set of manual steps to gradually convert narrative text CPGs into CIGs,7,8 or based on automated information extraction mechanisms frequently using linguistic patterns.9–14 While the accuracy of the manual approaches is straightforwardly controlled, as the resulted accuracy is as good as the input provided by the human modellers, these approaches are impractical to use in formalizing large numbers of CPGs. On the other hand, the automated and semi-automated information extraction–based approaches are more suited to formalize a relative large number of CPGs, but these approaches have not shown how expandable they are in handling a large number of heterogeneous CPGs, CPGs with different styles, granularity, and so on – a common challenge with all large-scale information extraction systems. 15 Therefore, the motivation of our work in building a CPG formalization framework is to enable the human modeller to control the expressiveness of the extraction rules in the formalization system without rebuilding the system.
The purpose of this work is to minimize the effort required by human modellers to bridge the gap between CPG and CIG by extending the automation of the formalization process of the drug recommendation and procedure recommendation clinical contexts. We followed a two-level clinical context extraction mechanism to assist the human modeller to better control the balance between accuracy and scalability with heterogeneous CPGs.
Methods
The proposed framework follows a multi-step approach, which has been shown to be a good strategy for CPG formalization. 16 We architected the framework to set boundaries around the aspects of the drug recommendation in the CPG formalization, where each aspect is implemented as a separate autonomous component in a CPG formalization pipeline. The framework is based on the Unstructured Information Management Architecture (UIMA). 17 In the following sections, we provide a description of each component in our CPG formalization pipeline as illustrated in Figure 1.
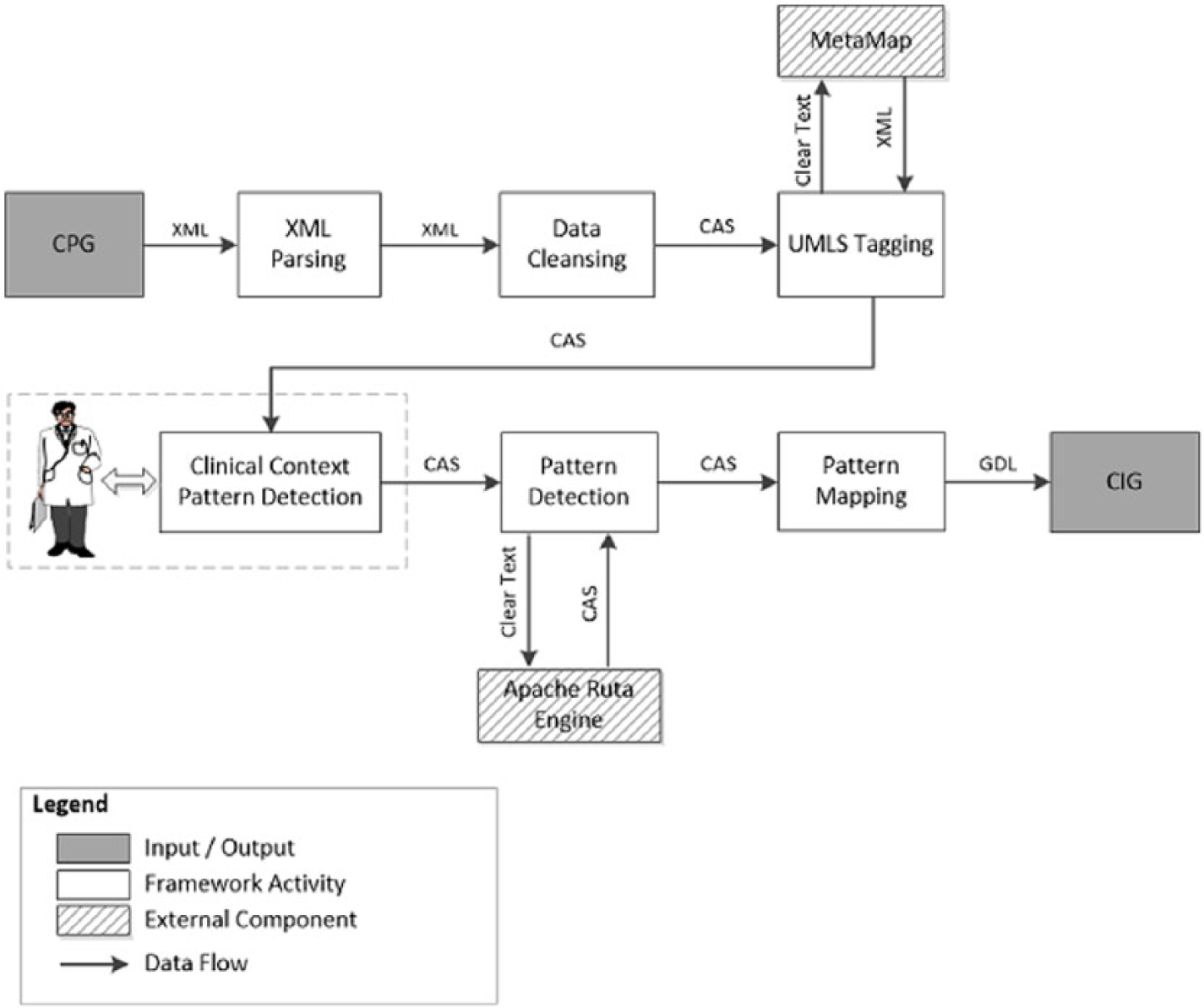
CPG formalization pipeline.
XML parsing
We used CPGs extracted from the National Guideline Clearinghouse (NGC) 18 in XML format. The XML parsing component extracts the content of the XML CPG documents into a structured object.
Text cleansing
Most of the sections extracted by the XML parsing component contain narrative text mixed with HTML tags. HTML tags are used by Web browsers to render text for visual display, but as we are not interested in composing the text for web browsers, we removed all HTML tags from the text.
Medical concept tagging
This is a component to map CPG text to a medical vocabulary; we used the Unified Medical Language System (UMLS) Metathesaurus as our biomedical vocabulary database. The UMLS Metathesaurus contains more than 2.6 million concepts each assigned to at least one semantic type from the set of 133 semantic types of the UMLS semantic network. We used MetaMap,19,20 to map CPG text to the UMLS Metathesaurus concepts. For integrating MetaMap with the UIMA framework, we leveraged the MetaMap UIMA Annotator 21 which is a wrapper that makes the MetaMap tool usable as an UIMA analysis engine.
Medical tags disambiguation
This is the process of finding the correct UMLS concepts, when multiple concepts are assigned by MetaMap with the same score. For example, the word
MetaMap UMLS concept mapping for the word
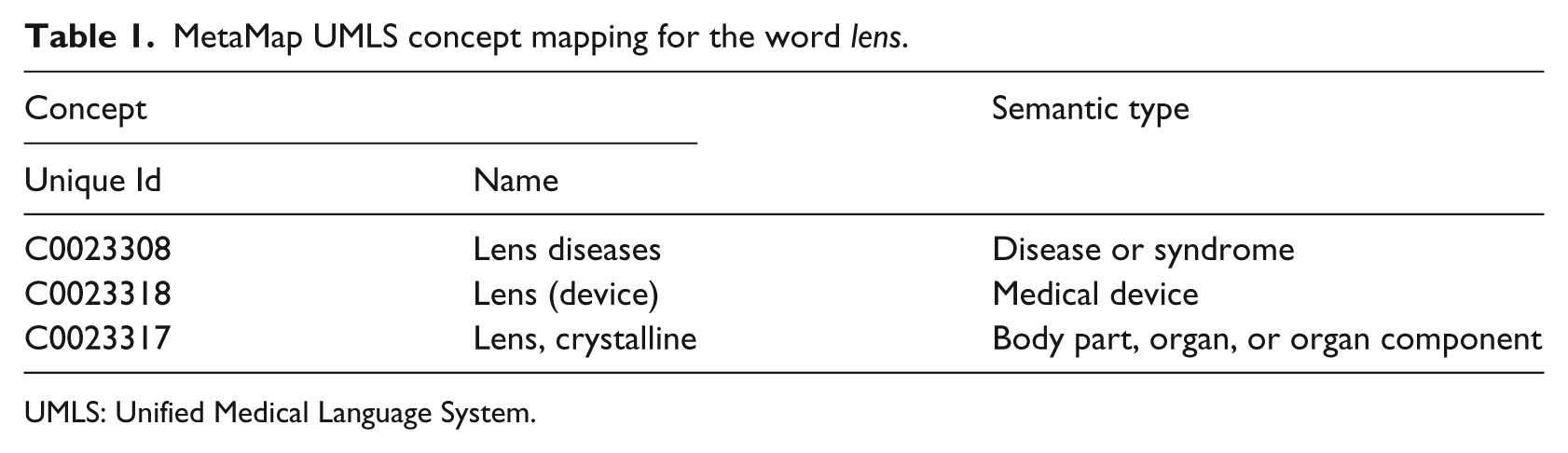

UMLS: Unified Medical Language System.
To solve this type of ambiguity, we used a graph-based disambiguation algorithm 22 to rank the generated MetaMap UMLS concepts based on their relatedness to the context of co-located text.
Clinical context pattern detection
This is a rule-based extraction component. This component is the first level of our clinical context extraction mechanism; its function is to extract text fragments that contain the minimum necessary features of the clinical context in question. Extracting clinical context based on the minimum necessary features follows the top-down approach 15 where only general rules that cover as many possible instances of clinical context need to be defined, which means rules that have high coverage and poor precision. Because general extraction rules tend to be in small numbers and are simple to define, the rule authoring task is a good fit for the medical experts who usually lack extensive knowledge in rule authoring. To further simplify the rule authoring task for the medical expert, we used UIMA Ruta 23 as it has a defined rule-based language with the ability to build rules against the text as well as against the semantic annotations of the text. We also defined a guideline of four steps to structure the effort required. In the following sections, we describe each of the four steps to author drug recommendation extraction rules with sample Ruta syntax highlighted in Figure 2:
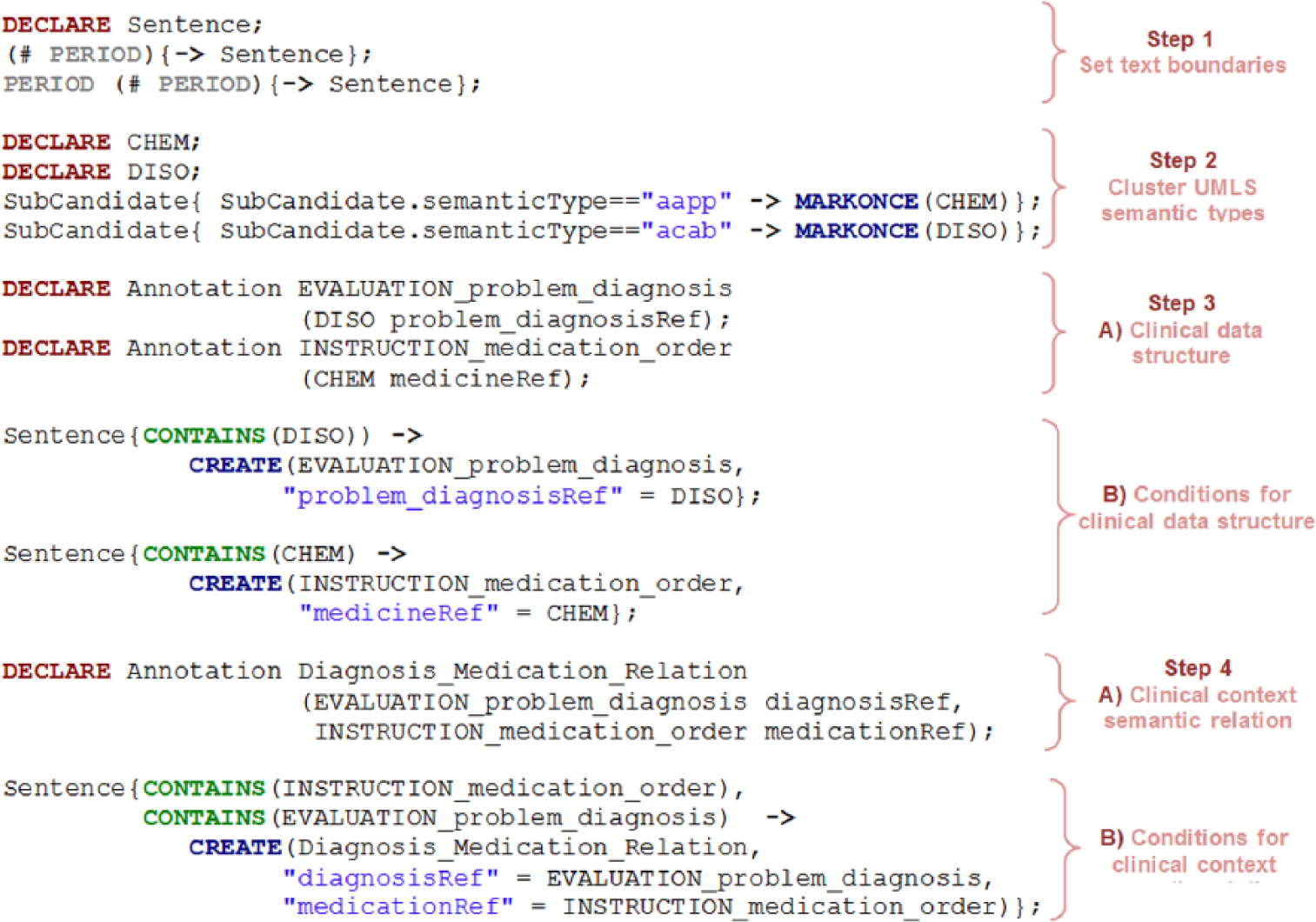
UIMA Ruta patterns for drug recommendation.
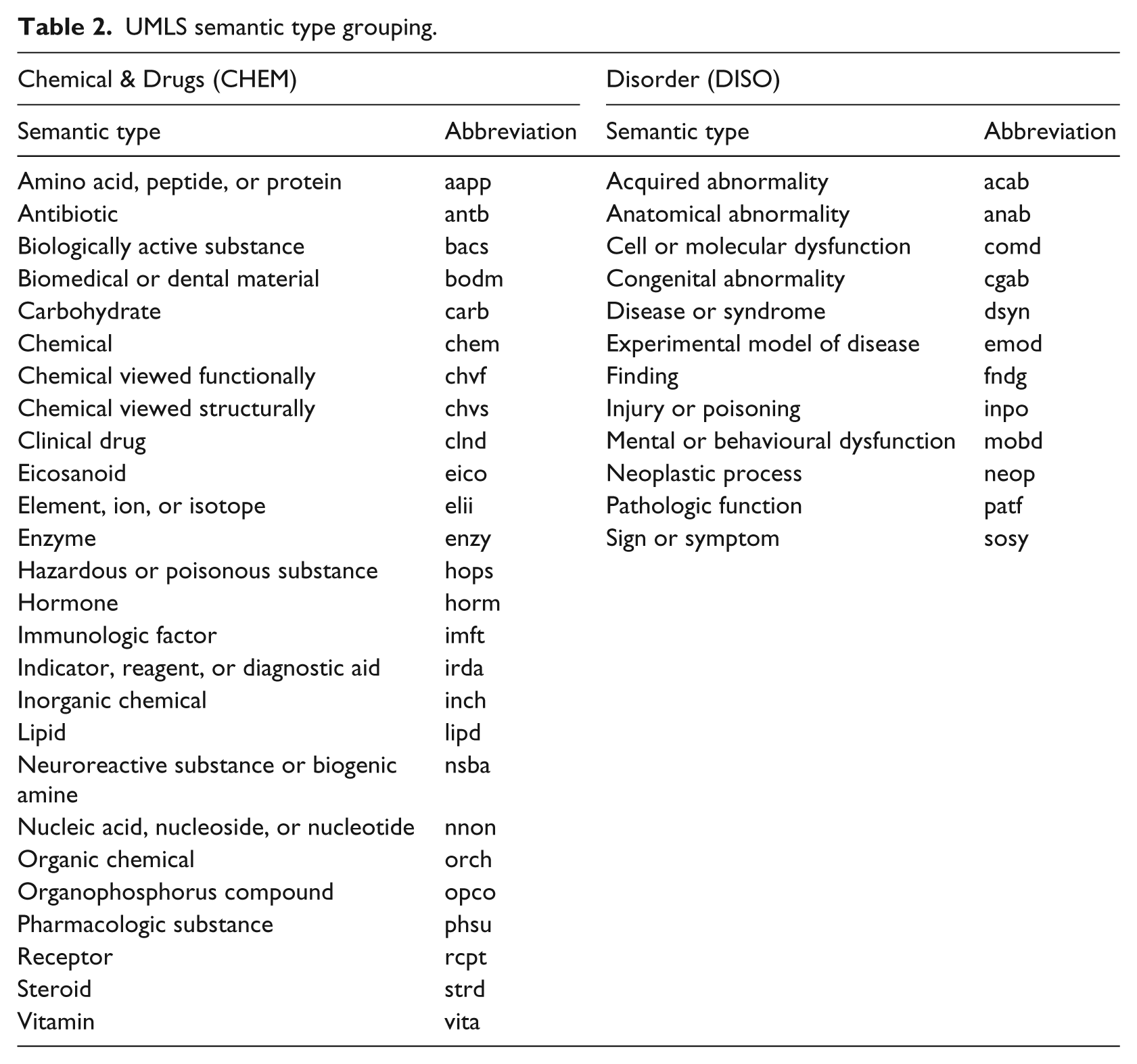
UMLS semantic type grouping.
Detecting an instance of the defined clinical data structure in the CPG text is achieved by annotating the CPG text with the clinical data structure types based on predefined conditions. The conditions could be based on specific lexicon, syntax, or previously annotated semantics; Step 3 in Figure 2 shows the Ruta code of our version of the ‘problem diagnosis’
In Figure 3, we show the ‘Fluoxetine (20–80 mg/day) should be considered for the treatment of patients with fibromyalgia’ sentence from the ‘Management of chronic pain: a national clinical guideline’. 26 CPG and the annotations are assigned based on the four clinical context pattern detection steps.

Annotations for the drug recommendation clinical context.
Clinical context filtering
This component is responsible for removing the clinical context instances wrongly labelled by the clinical context pattern detection component. We used a logistic regression 27 classification algorithm to decide on the correctness of the drug recommendation labels. Because this classification algorithm is supervised, which means it requires to be trained using a correctly annotated data set, we generated a training data set composed of 117 recommendation sentences extracted from the Yale Guideline Recommendation Corpus (YGRC). 28 The YGRC is composed of 1275 recommendations which cover a broad range of diseases and mental disorders extracted from the NGC. We annotated all YGRC sentences with MetaMap and then selected 117 sentences that have tokens in the DISO semantic group in addition to other tokens in the procedure or CHEM semantic group. We manually tagged each sentence as either drug/procedure recommendation or non-drug/procedure recommendation.
Clinical context mapping
This is a component to map instances of clinical context semantic relations to their target CIG constructs as a set of rules. We used the openEHR Guideline Definition Language (GDL) 29 as our target CIG; GDL leverages the designs of openEHR Archetype Model that we used in the pattern detection step.
Although the clinical context mapping is still a manual task to be done by the knowledge engineer, it can be fully automated if the medical expert modeller used a standard naming convention for the clinical data structures used in the clinical context pattern detection component.
Our evaluation was based on measuring the precision, sensitivity/recall, and specificity of the extracted drug recommendation and procedure recommendation rules form the input recommendation sentences. The precision, sensitivity/recall, and specificity are measured based on the correctness of our framework in finding instances for the UIMA Ruta patterns defined by the medical expert. More formally, assume that
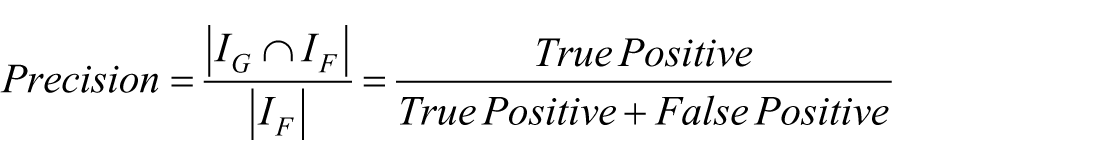
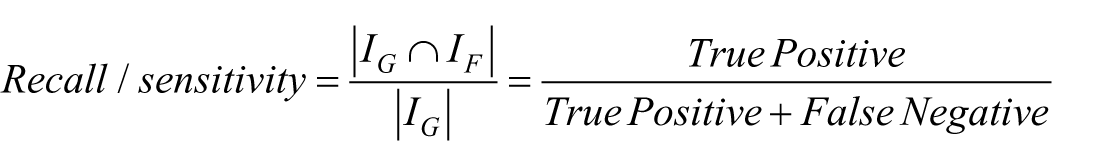
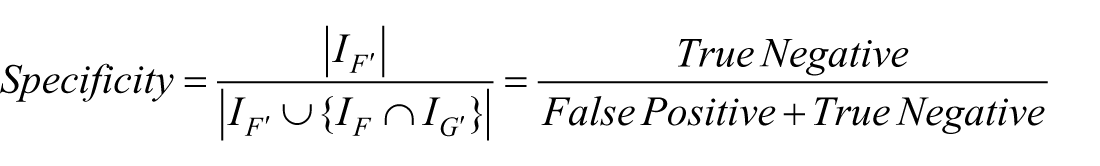
Results
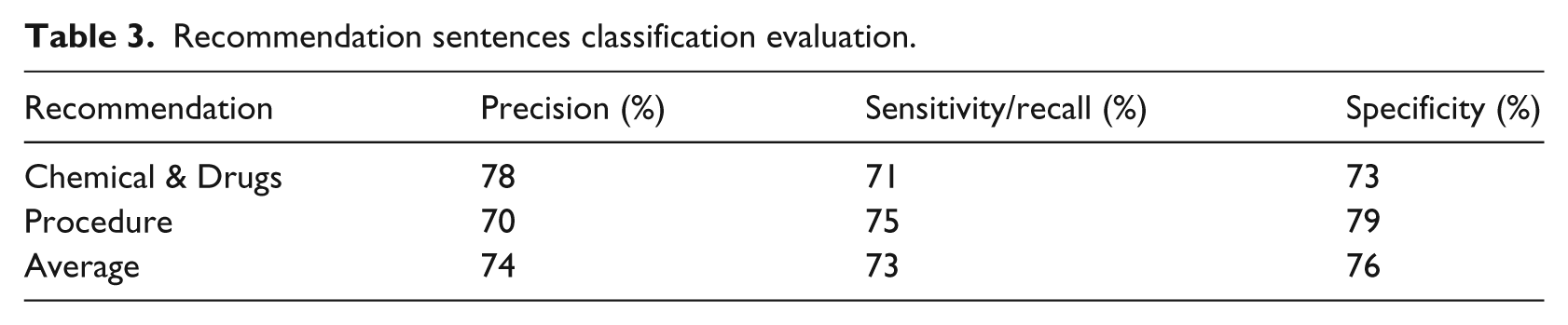
We implemented our formalization framework in JAVA (version 1.7) and integrated it with the GDL editor. 29 Due to the lack of access to independent human modellers, we could not measure the manual effort saving introduced by our framework; nevertheless, we evaluated the accuracy of the knowledge extracted by our framework for the drug recommendation and the procedure recommendation clinical contexts. To build our gold standard for the drug recommendation clinical context to measure against, we used all recommendations from the ‘Management of chronic pain’ CPG 26 and the ‘Management of lung cancer’ CPG, 30 and then we manually tagged each recommendation as either drug/procedure recommendation or non-drug/procedure recommendation. The resulted test data set is composed of 169 recommendation sentences. In Table 3, we show the accuracy of our framework on classifying the 169 recommendation sentences.
Recommendation sentences classification evaluation.
We evaluated the correctness of the formalized recommendations by manually checking the extracted rules and we assigned a coefficient of 1 for rules that are correctly coded and complete, 0.5 for rules that are correct but partial (e.g. not all elements of the rule conditions are captured), and 0 for rules that are wrong. In Table 4, we show the accuracy of the extracted rules based on the described metric. There are two main sources of errors for the wrong rules: either wrong MetaMap annotations or wrong classification from our clinical context filtering component using logistic regression.
Extracted rules’ accuracy.

Discussion
Using information extraction, techniques to extract biomedical knowledge have been applied in other non-CPG biomedical documents.31,32 The proposed framework is tailored to the specific needs of CPGs where the granularity of clinical knowledge differs between different types of CPGs; therefore, we designed the framework to enable the human modeller to control the granularity of the extracted clinical knowledge.
The proposed framework is based on a clinical context pattern detection component using UIMA Ruta, a rule-based information extraction mechanism to extract text fragments that contain the minimum necessary features of the clinical recommendations type in question. We believe that extracting clinical recommendation by following a top-down approach 15 where only general rules that cover as many possible instances of clinical recommendation need to be defined is suitable for CPG formalization. General extraction rules tend to be in small numbers and are simple to define making the rule authoring task a good fit for medical experts who usually lack extensive knowledge in rule authoring. Although general information extraction rules result in high coverage, they have a poor precision; therefore, we added the clinical context filtering component using logistic regression to filter out false-positives introduced by general information extraction rules defined in the clinical context pattern detection component. The framework was tested on extracting only two clinical context types due to the lack of pre-annotated test data sets. In the following discussion, we analyse the achieved results:
The precision is impacted by the size and the quality of our training data set; in the presented example, we used a training data set made of 117 recommendation sentences extracted from YGRC which is small to provide high precision. This issue could be lessened by feeding the outputted rules of the framework back to the training data set, a step that requires a minor manual tagging of which rules are correctly extracted and which ones are wrongly extracted.
The sensitivity/recall is impacted by how we split our CPG into smaller text chunks, for example, in the presented example we split CPG into sentences, but some drug recommendations within the CPG have the drug and the medication located in two separate sentences, and therefore, these ones are missed by our extraction rules. This issue could be lessened by changing the size of our unit of analysis from one sentence to two consecutive sentences or to the whole paragraph, but such a modification would hurt the precision unless we add more rules to handle cross-sentence extraction. Different cross-sentence extraction approaches can be applied. One approach would be to perform cross-sentence extraction when a sentence only contains one part of the clinical context such as a sentence with only a disease, followed by a sentence that only contains the other part of the clinical context such as a sentence with only a medication. This approach is very conservative and would not impact the precision of the in-sentence extraction rule. Incorporating other cross-sentence extraction approaches that have more coverage would likely interfere with other in-sentence extraction rules. Therefore, with every cross-sentence extraction approach, we need to evaluate the cross-sentence extraction precision gain to the in-sentence precision loss.
The specificity is impacted by how strict are our conditions for tagging a sentence with a specific semantic relation. In the presented example, we achieved high specificity because our conditions for tagging drug medication semantic relations are very strict.
The proposed framework can be effectively used to formalize the drug recommendation and procedure recommendation clinical contexts of CPGs into CDSS-friendly format. More significantly, it provides human modellers with a methodology to extend the framework to formalize other clinical context of CPGs. The framework is focused on automating the CPG common formalization steps while allowing the human modeller to stay in control of all the knowledge extraction steps. By configuring the
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
