Abstract
Structured Product Labels follow an XML-based document markup standard approved by the Health Level Seven organization and adopted by the US Food and Drug Administration as a mechanism for exchanging medical products information. Their current organization makes their secondary use rather challenging. We used the Side Effect Resource database and DailyMed to generate a comparison dataset of 1159 Structured Product Labels. We processed the Adverse Reaction section of these Structured Product Labels with the Event-based Text-mining of Health Electronic Records system and evaluated its ability to extract and encode Adverse Event terms to Medical Dictionary for Regulatory Activities Preferred Terms. A small sample of 100 labels was then selected for further analysis. Of the 100 labels, Event-based Text-mining of Health Electronic Records achieved a precision and recall of 81 percent and 92 percent, respectively. This study demonstrated Event-based Text-mining of Health Electronic Record’s ability to extract and encode Adverse Event terms from Structured Product Labels which may potentially support multiple pharmacoepidemiological tasks.
Keywords
Introduction
Structured Product Labels (SPLs) support the distribution of up-to-date and comprehensive information regarding marketed medical products in a computerized format for use in health care information systems. 1 SPLs contain boxed warnings, indications for use, contraindications, warnings and precautions, Adverse Reactions, and other information for medical products. 2 Even though SPLs may adequately support tasks that were unfeasible with previous labeling formats, such as the automated comparison of texts by section, their current organization with the complex list of Adverse Event (AE) terms in various sections cannot easily convey the information to the medical experts. 3 This is a critical disadvantage for many post-market safety surveillance tasks at the US Food and Drug Administration (FDA), such as the review of safety reports submitted to spontaneous reporting systems. For example, the medical experts at the FDA evaluate all types of AEs, labeled and unlabeled, but always try to identify new or unexpected unlabeled AEs that may appear in the post-market reports. It is, therefore, critical to present the key information of the SPLs to the medical experts in the most efficient and automated manner.
A number of systems have been used to extract information from the FDA SPLs, such as the Structured Product Label Information Coder and Extractor (SPLICER) 4 and the Structured Product Labels eXtractor (SPL-X). 5 Most of these systems focus on the extraction of AE terms from particular sections using various approaches, such as named-entity recognition and rule-based techniques, in combination with existing platforms, such as the NegEx algorithm 6 and the National Library of Medicine (NLM) MetaMap tool. 7 Several other groups have focused on creating repositories of drug-related information obtained, at least, in part, from the FDA SPLs. For example, Smith et al. 8 processed the “Indications for Use” and the “Adverse Reaction” section from the FDA SPLs as part of their effort to create the Drug Evidence Base.
The Side Effect Resource (SIDER) is the largest resource of product labeling data and contains information on 1430 marketed medicines and associated AEs from various public sources including the FDA SPLs. 9 SIDER includes a list of Medical Dictionary for Regulatory Activities (MedDRA) terms for a large set of medical product labels for the 1430 products. 9 MedDRA is the terminology that is widely used at the FDA to encode the information in safety report narratives and may support the comparison between the unlabeled and labeled information in post-market reports. SIDER has been used in several studies for various purposes, such as the prediction of drug side effect profiles, the identification of multi adverse drug events, the generation of process-drug-side effect networks, and the prediction of drug label changes.10–14 We used SIDER as the basis for the generation of a comparison dataset that included a list of MedDRA terms for a certain number of SPLs.
The main purpose of this work was to evaluate the ability of an existing natural language processing (NLP) system to extract AEs from product labels and encode them to MedDRA Preferred Terms (PTs).We previously developed the Event-based Text-mining of Health Electronic Records (ETHER) system to process the narratives of safety reports submitted to the FDA and translate the extracted AEs into MedDRA terms. 15 We further expanded ETHER’s functionalities by including a capability to process certain sections from SPL files and generate MedDRA PTs in an automated fashion. We particularly compared the PTs extracted by ETHER with those listed in SIDER for the same set of SPLs and evaluated the differences through a qualitative error analysis (QEA).
Methodology
Comparison dataset
We downloaded the SIDER database, version 4.1, on 21 October 2015. This version contained 1430 unique medical products. Each product was linked to one or more labels, not necessarily in SPL format. As SPL is the standard adopted by the FDA, we identified the labels following the SPL standard only. This was accomplished by matching for “Set IDs,” which are 32-character hexadecimal strings unique to an SPL with an associated date and selected the most recent based on the date in the string. In the case of a tie, the hexadecimal strings were sorted and the alphabetically last “Set ID” was selected. Labels in non-SPL format were excluded, and the initial set of SPLs was created.
The full XML files for these SPLs were then retrieved from the DailyMed website using the DailyMed REST API. 16 For any product “Set ID” that could not be found in DailyMed, the next most recent “Set ID” from the SIDER data file for the same product was searched instead. If none of the product “Set IDs” could be located in DailyMed, we excluded that product from our analysis and created the final set of SPLs. We subsequently retrieved the full list of MedDRA terms for the selected SPLs from SIDER, but used only the PTs to build the comparison dataset. It should be noted that the SIDER database contained both PTs and Lower Level Terms (LLTs), for each “Concept ID” found in each label. The “Concept ID” is a numerical representation of the Systematized Nomenclature of Medicine Clinical Terms terminology, which is often used in clinical documentation and reporting. 17
SPL processing
We processed the XML versions of the labels retrieved from DailyMed using ETHER and generated the corresponding list of AE terms and PTs. ETHER translates the extracted AE terms to MedDRA PTs in two subprocesses (Figure 1). First, the AE text is mapped directly to the PTs and LLTs based on the corresponding relationships in the MedDRA hierarchy. Second, the NLM MetaMap tool is used to map the AE text to MedDRA PTs. 7 If any LLTs are returned in either of the sub-processes, ETHER maps them to PTs based on the corresponding relationships in the MedDRA hierarchy. In the final step of the process, the ETHER and MetaMap lists are merged and duplicates are eliminated.
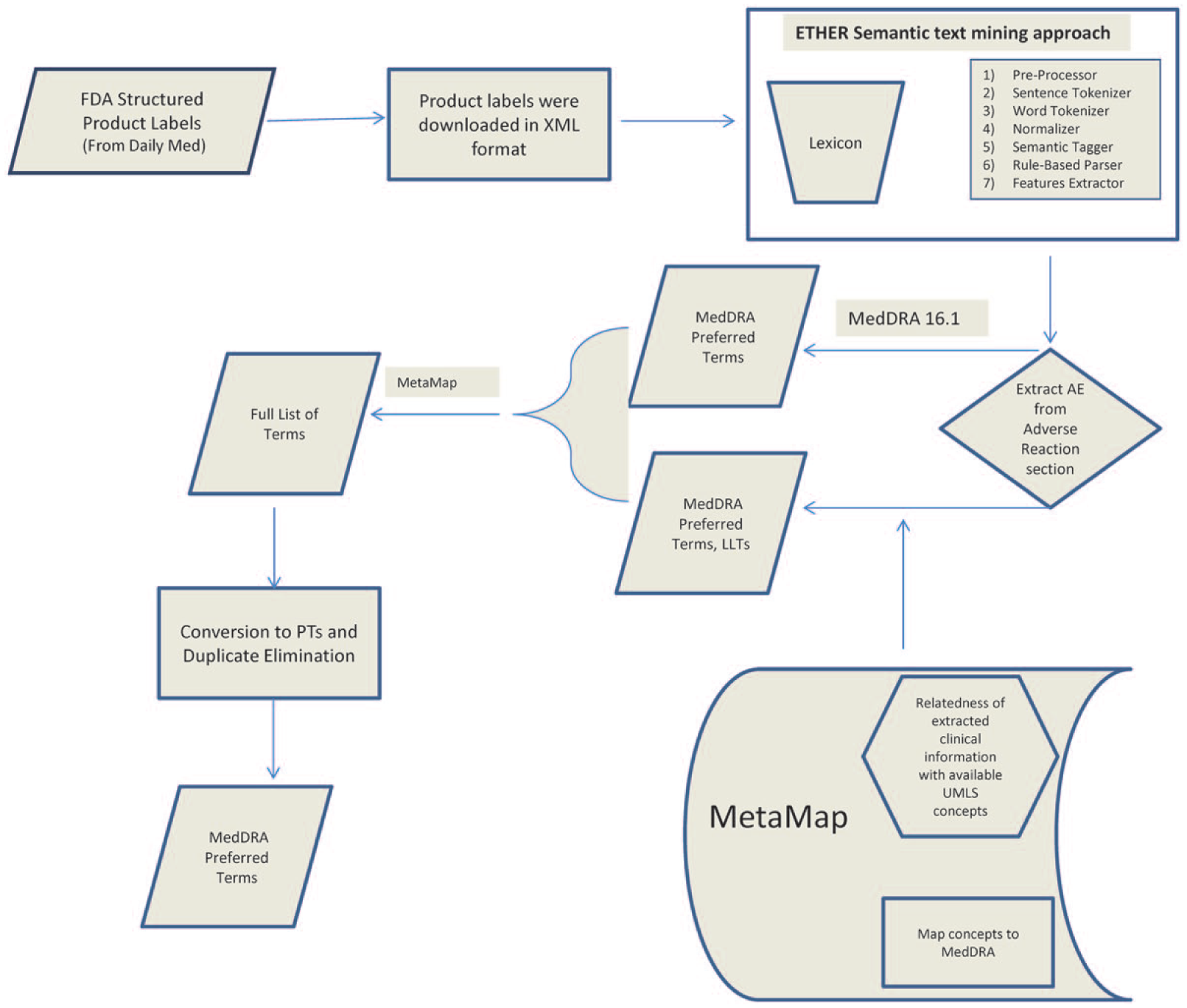
Processing of the Structured Product Labels (SPLs) with the Event-based Text-mining of Health Electronic Records (ETHER) system, and conversion of the extracted Adverse Event (AE) terms into Medical Dictionary for Regulatory Activities (MedDRA) Preferred Terms (PTs) with the assistance of MedDRA 16.1 and MetaMap.
The current local version of ETHER captures features from the “Boxed warning, Warnings and Precautions, Adverse Reactions and Use in Specific Populations” sections of SPLs. In this study, we used ETHER to extract the AE terms and their PT representation from the “Adverse Reactions” section only; this section may sometimes appear as “Adverse Events” or “Adverse Effects” section in the SPLs and their PTs.
Data analysis
We compared the ETHER PTs with the SIDER PTs for the final set of SPLs. The PTs that were missed and the extra PTs generated by ETHER synthesized the pool of false negative (FN) and false positive (FP) PTs, respectively. For a more detailed analysis, a subset of 100 labels was created by randomly selecting 25 labels from each quartile of the final set of SPLs ordered by the number of FNs. This sample was intended to be representative of the tool’s capability on the full dataset, but small enough to be manually reviewed in a reasonable amount of time. To evaluate the performance of our approach, we calculated the standard metrics of recall, precision, and F-measure. We subsequently performed a QEA to assess the differences. The FN and FP PTs for the 100 labels were analyzed and categorized in certain error categories and subcategories (Figure 2).
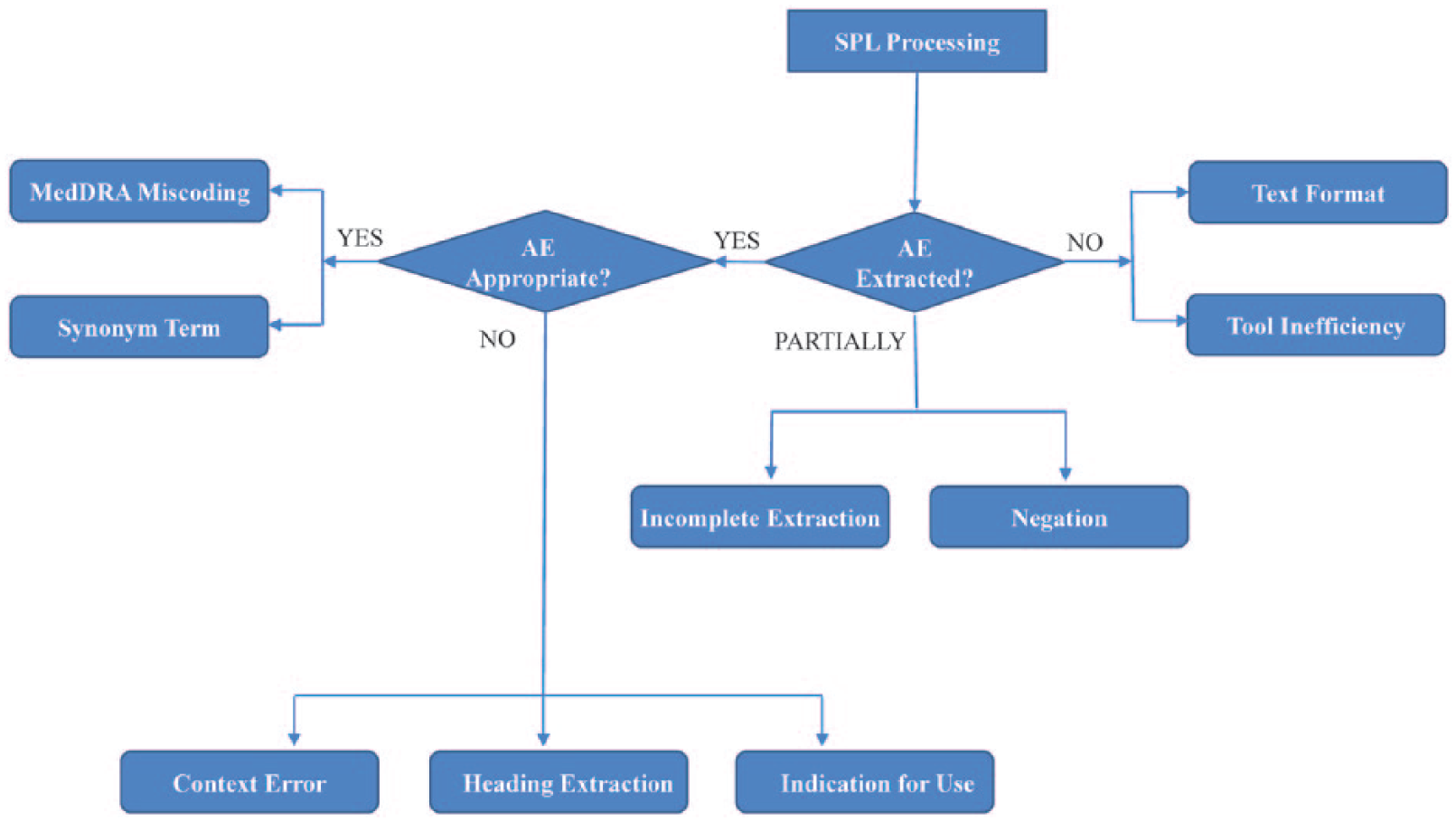
Error categories and subcategories for the qualitative error analysis.
These error categories were created to assess any system’s ability to extract and encode the expected AE terms from the label texts. As shown in Figure 2, three main categories were defined: (A) an AE term is extracted; (B) no AE term is extracted; (C) an AE term is partially extracted by the system. The subcategory hierarchy may span up to three levels. The first two levels of the subcategories for each of the main categories are described below:
An AE term is extracted. Two specific subcategories further classify the FN or FP terms found in this category: The appropriate AE is extracted.( An inappropriate AE is extracted.(
No AE term is extracted. This error type results in FN terms further categorized to the following subtypes:
An AE term is partially extracted. This error type results in FN or FP terms which are further categorized to the following error subtypes:
Terms that do not fall into any of the above error types are assigned to an “Unknown Error” type.
An expert pharmacologist used the above scheme and performed the initial QEA by reviewing the FN and FP PTs with the corresponding AE terms in the SIDER database. Then, a second pharmacologist reviewed the results of the initial QEA findings. Disagreements between the two reviewers were discussed and, after a consensus was reached, the final list of error types for the FN and FP PTs was generated. For the “Synonym Term” error type, we took an additional step and processed them with the online version of MetaMap; MedDRA 16.1 was the source used for the translation process. The MetaMap output was mapped to the MedDRA 16.1 PTs and supported the review of the list of synonyms. If a MetaMap-based PT and an ETHER-based PT matched for the same label-based AE term, we put the ETHER PT in the “Synonym Term” category. When all the synonym terms were verified through this process, we removed them from the FN and FP list. Based on the refined lists, we recalculated the performance of our approach.
Results
Among the 1430 products downloaded from SIDER, only 1171 of them had at least one label in the SPL format. When using the DailyMed REST API to download the XML files for the most recent labels, 36 of the products’ “Set IDs” could not be found. Out of those 36 products: for 24 products, the next most recent “Set ID” from the SIDER data file was used; for 1 product, the third most recent “Set ID” was used; and for 11 products, none of their “Set IDs” could be found in DailyMed and were therefore excluded from our analysis. We then processed the XML versions of the remaining labels (N = 1160) with ETHER. One XML file could not be analyzed by ETHER and was removed resulting in 1159 SPLs for analysis. This selection process has also been summarized in Figure 3.
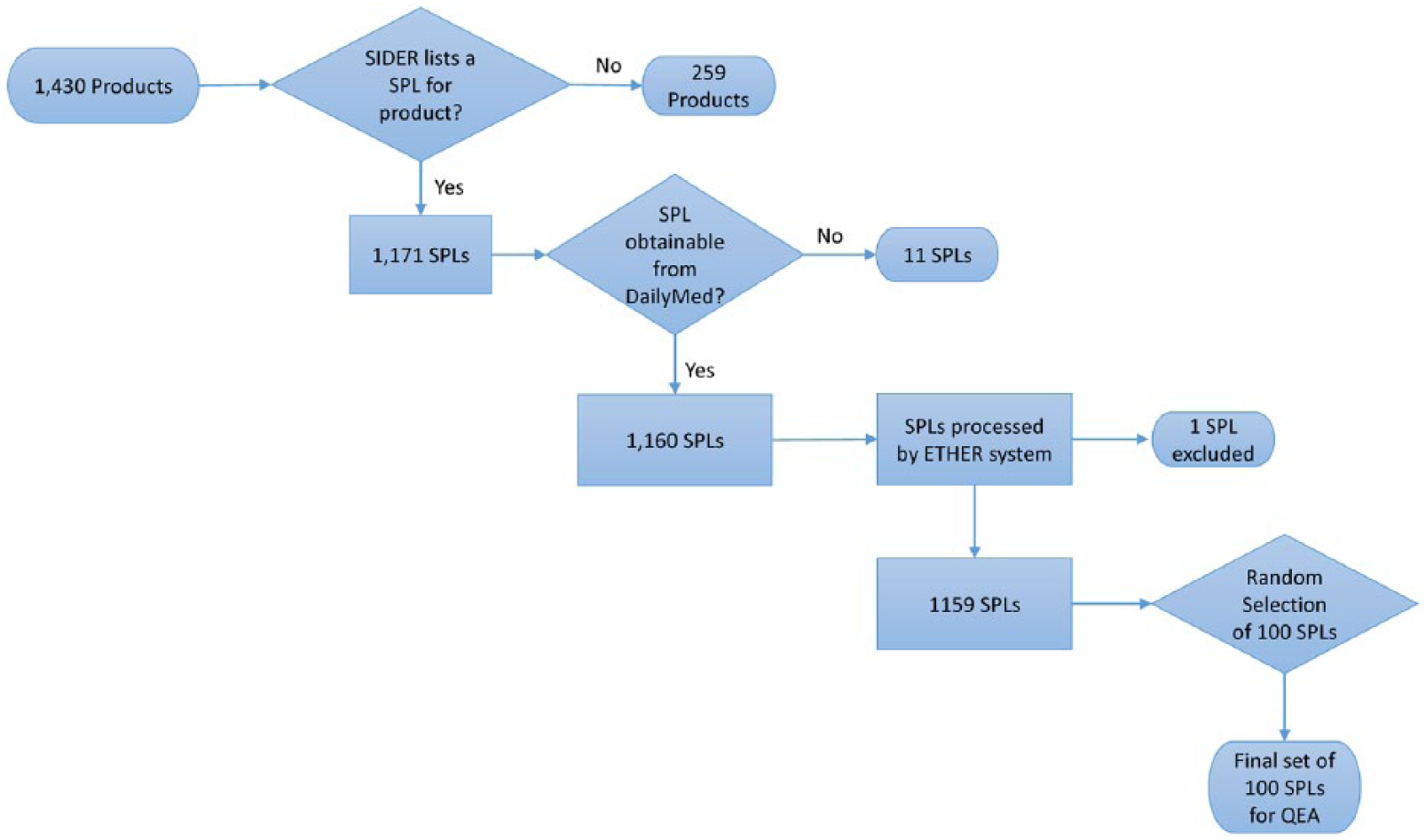
Flowchart of the selection process for Structured Product Labels (SPLs) for products found in the Side Effect Resource (SIDER).
The SIDER database contained both PTs and LLTs for each AE term found in the labels. We expected the LLT and the PT for a given concept to be linked in the MedDRA hierarchy with a child–parent relationship. However, this type of relationship does not exist for all the labels in SIDER, and it was not possible to directly retrieve the PTs for the included LLTs. We elected to use only the PTs from SIDER for the construction of the comparison dataset and not the MedDRA hierarchy for directly mapping any of the LLTs to different PTs. This seemed the most reliable strategy to capture the same information that SIDER makes available online. As a result, we found a total of 70,796 MedDRA PTs from SIDER for 1159 products.
The ETHER PTs generated for these 1159 product labels (N = 73,895) were compared with the SIDER PTs (N = 70,796) for the same labels. There were a total of 13,441 FN PTs and 16,504 FP PTs found for the 1159 product labels. The distribution of these FN and FP PTs per label is shown in Figure 4. ETHER generated MedDRA PTs with a precision, recall, and F-measure of 0.78 (standard error (SE) = 0.10), 0.81 (SE = 0.073), and 0.79, respectively. As it was not feasible to perform manual QEA over 1159 labels for this project, a set of 100 labels were randomly chosen for further analysis. The labels were ordered by the number of FN PTs, and 25 labels were randomly selected from each quartile. The range of FN PTs in each quartile was Q1: [0, 4]; Q2: [4, 9]; Q3: [9, 16]; and Q4: [16, 340] (see Figure 5). The ETHER performance over the 100 labels was similar to the full set of labels with precision, recall, and F-measure equal to 0.77 (SE = 0.088), 0.81 (SE = 0.081), and 0.79, respectively. There were 1082 FN and 1337 FP PTs identified for the 100 labels; these PTs were further reviewed and labeled with the appropriate error types as described below and in Supplement 1. The data are available in Spreadsheets A (FNs) and B (FPs) in Supplement 2.
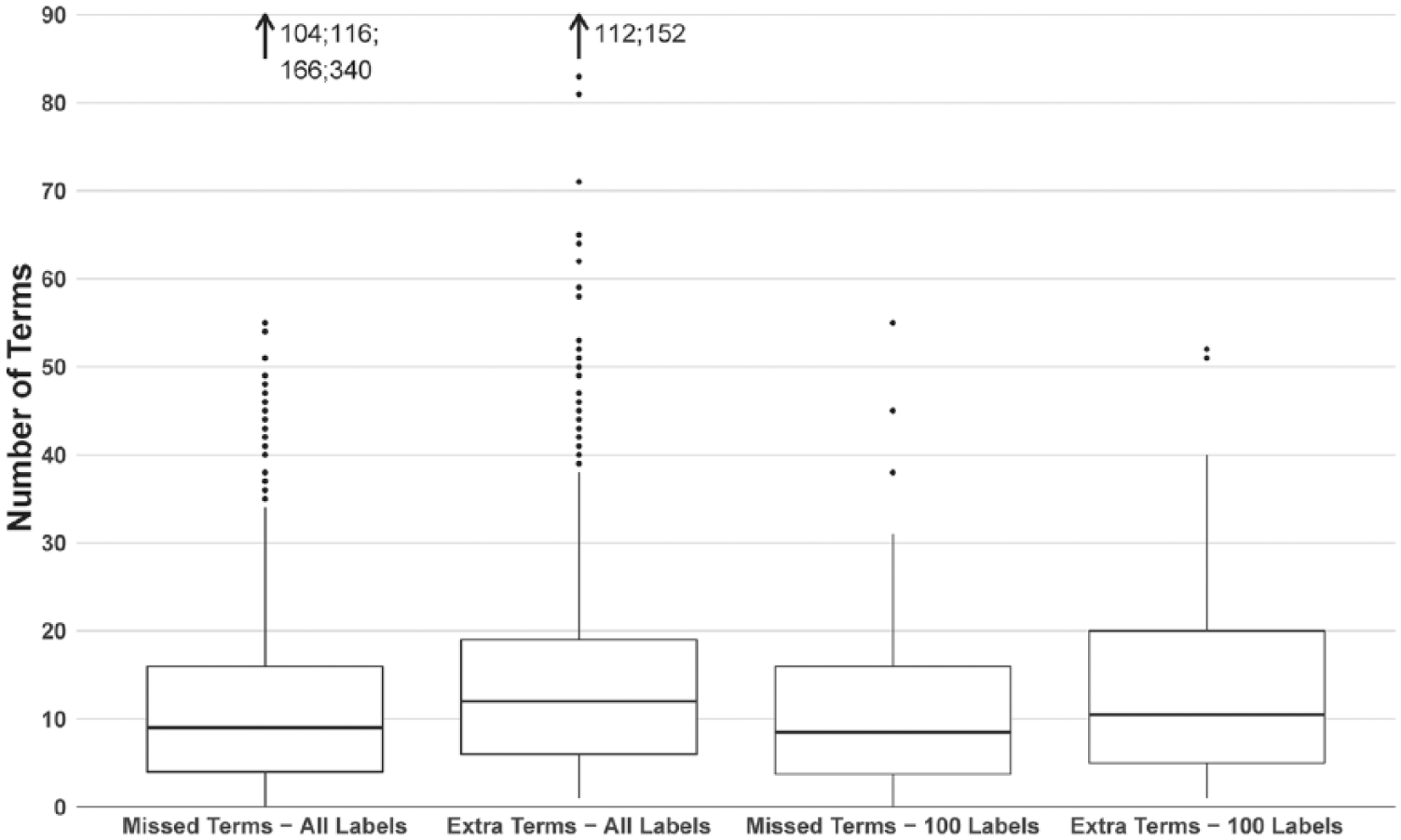
The box plot shows the distribution of the false negative (FN) Preferred Terms (PTs) in the full set of labels (N = 1159), alongside the distribution of false positive (FP) PTs generated by the Event-based Text-mining of Health Electronic Records. The rightmost boxes show the corresponding distributions of the FN and FP PTs from the set of 100 labels. The whiskers extend up to 1.5 times the interquartile range.
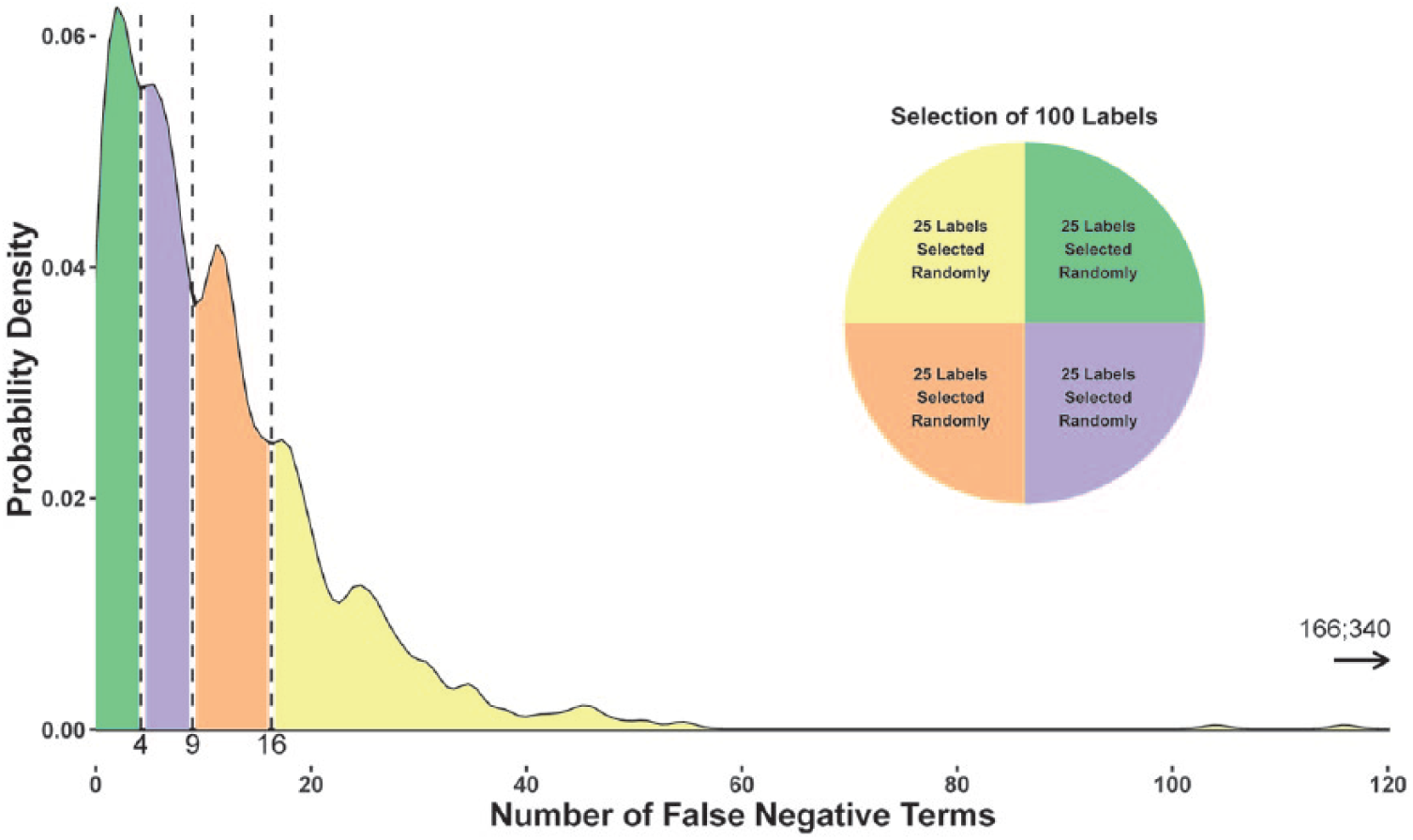
The probability density plot shows the distribution of the false negative Preferred Terms for all labels of the final set (N = 1159). Twenty-five labels were randomly selected from each bin and formed the subset of 100 Structured Product Labels that was further analyzed.
During the QEA for the FN and FP PTs of the 100 SPLs, we manually reviewed the Adverse Reaction section of these labels retrieved from DailyMed and the corresponding section of the SPLs that were available on the SIDER website. This revealed textual discrepancies for 18 of the 100 product labels, and we concluded that the product labels retrieved from DailyMed (and processed by ETHER) were published later than the product labels used by the current SIDER release (4.1). Since the dates included in the “Set ID” did not correspond to the date of publication in DailyMed, we were unable to conclusively retrieve the matching versions of the labels. We, therefore, decided to revise our results by introducing a new error category (“Text Discrepancy”) that was applied during the QEA for FN and FP PTs which were caused by these discrepancies in label texts.
In our QEA for the 100 SPLs, the most common error types for the FN terms (Figure 6) were the “Synonym Term” (43.81% of the total FN-related errors) and the “Tool Inefficiency” (15.34% of the total FN-related errors) error types; 16.54 percent of the total FN-related errors should be attributed to weaknesses of the comparison dataset, which are further described below. In the FP error analysis, the major error types were the “Context Error” (24.39% of the total FP-related errors), the “Unknown Error” (18.25% of the total FP-related errors), and the “Incomplete Extraction” (12.49% of the total FP-related errors) types (Figure 6).
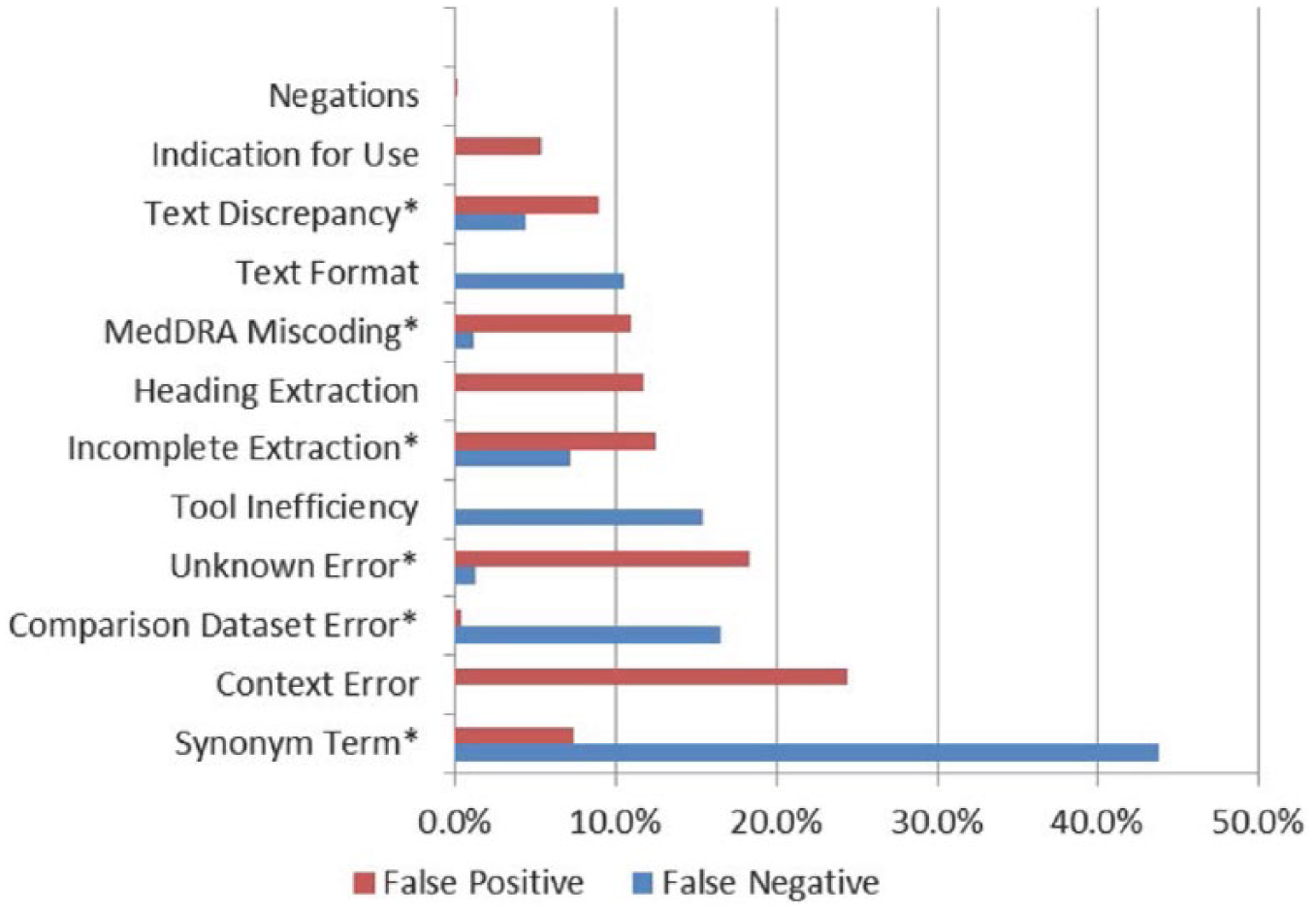
Result of the qualitative error analysis for the 100 product labels; this column chart shows the percentage of error types assigned to the Event-based Text-mining of Health Electronic Records–based false negative (FN) and false positive (FP) Preferred Terms (PTs) for the 100 product labels. The common error types found in both the FP and the FN PTs are marked with an asterisk.
In addition to the “Text Discrepancy” error type, the “Comparison dataset” error type was not included in the original QEA schema but was revealed as a potential weakness of the SIDER resource during the expert review process. We particularly identified the following weaknesses in the comparison dataset:
SIDER included two “Concept IDs” for a single PT. After a closer examination, only one of the two “Concept IDs” was found to be linked to the SIDER PT; the other “Concept IDs” could not be located in the label text and should be, therefore, considered as a comparison dataset error not as FN.
There were AE terms that were partially extracted (N = 4) by SIDER or represented indications or secondary complications—the assigned PTs (N = 46) were incorrectly included in the list of PTs for the Adverse Reaction section.
SIDER also provided inappropriate MedDRA PTs (N = 6) for four SPLs.
For a few SIDER PTs (N = 132), the corresponding label text could not be located in the 59 product labels.
Pairings of LLTs and PTs (N = 5) did not match the MedDRA 16.1 hierarchy for eight product labels.
Following the QEA over the 100 labels, we revised the list of FP and FN PTs in two stages based on our findings and recalculated ETHER’s performance. First, all FP and FN PTs categorized under the “Text Discrepancy” and “Comparison dataset” types were completely excluded from the analysis, as these types were unrelated to the system’s ability to extract AE information, and updated precision, recall, and F-measure were 0.79 (SE = 0.080), 0.84 (SE = 0.066), and 0.82, respectively. Then, we allowed FP and FN PTs under the “Synonym Term” category to count as matching PTs. All metrics were increased with precision, recall, and F-measure rising to 0.81 (SE = 0.082), 0.92 (SE = 0.055), and 0.86, respectively. Interestingly, recall was significantly increased by 8 percent points over the previous stage, indicating ETHER’s ability to accurately represent the labeled information. However, the number of FP PTs remained at almost the same level with a small improvement of 2 percent points.
Our QEA further supported the generation of a complete and validated (by two experts) collection of PTs for the Adverse Reaction section of the 100 labels. This dataset is available as a spreadsheet in Supplement 2 (Spreadsheet C) for research and other purposes.
Discussion
Our study examined the potential use of ETHER for the extraction of AE information from SPLs and the translation of this information to MedDRA PTs. We used the SIDER resource and DailyMed to retrieve the full text of SPLs and generate the comparison dataset with the AE and PT terms for this purpose. ETHER was able to generate the AE PTs for 100 product labels with an initial precision and recall of 0.77 and 0.81, respectively. After eliminating the PTs related to the “Text Discrepancy” and “Comparison dataset” error types for the reason above, precision and recall increased to 0.79 and 0.84, respectively. The further treatment of the “Synonym Term” category increased the precision and recall to 0.81 and 0.92, respectively. The initial ETHER performance over the 100 labels (precision: 0.77 and recall: 0.81) was similar to the full dataset of the 1159 labels (precision: 0.78 and recall 0.81). This indicates that our random sampling process efficiently captured the characteristics of the full dataset and was representative of ETHER’s performance over it. Findings on these 100 labels could be potentially extrapolated to the total SPL population.
Our study has some limitations. First, the “Adverse Reaction” section for some of the product labels that were retrieved from DailyMed was not identical to the same section found in the SIDER resource. Even though we accounted for all FP and FN PTs in the 100 labels that were affected by these text differences with the “Text Discrepancy” error type, it was not possible to repeat the same process for the final set with all the 1159 SPLs and revise our findings. Second, we evaluated the “Adverse Reaction” section only and did not consider the “Indication for Use” section that is also available in SIDER. The exclusion of PTs related to that section might have further improved the performance of our approach. Third, we performed the QEA over a subset only and not the full set of SPLs. However, it should be noted that the manual QEA for 1159 SPLs is a challenging task and requires the allocation of considerable resources. In any case, our sample appeared to be representative of the SPL population. Fourth, the comparison dataset errors revealed in the QEA may have introduced some bias to our study. We argue though that the removal of PTs related to these errors and the detailed manual review of the Adverse Reaction texts for the 100 labels was necessary to support a fair evaluation of our approach.
Apart from the weaknesses discussed as the “Comparison dataset” errors above, SIDER has a few more limitations that should be evaluated in future studies. First, the current SIDER release (the same used in our analysis) uses MedDRA 16.1 dictionary. It could be therefore argued that SIDER does not contain the most updated MedDRA terms given that MedDRA version 20.0 was recently released (at the time of writing this article). Whereas the changes from 16.1 to 20.0 are minor, this is an aspect that should be acknowledged. Second, the lack of PT mappings for some of the LLTs found in the SIDER database further reduces the amount of AE information that is coded to PTs. Third, the product labels are not continuously updated, and the latest published label may be different from the latest label in SIDER for the same product. Despite its limitations, SIDER remains the only publicly available database with MedDRA PTs for a large collection of SPLs and product labels in general. This explains its extensive use in previous studies and its selection for our analysis. The SIDER-related observations indicate the need for the development of a universal, validated, and continuously updated resource. Doubtless, this is an enormous task, however critical to support future efforts.
Culbertson et al. 18 extracted the AE information from black box warning by semantic processing with a precision of 94 percent and recall of 52 % SPLICER performed very well both in terms of precision and recall (95% and 93%, respectively) for the AE extraction from SPLs; 4 and SPL-X demonstrated precision and recall of 95 percent and 77 percent, respectively. 5 In this study, ETHER demonstrated a maximum precision and recall of 81 percent and 92 percent, respectively. Our results have shown that ETHER is on par with or superior to these systems in terms of extracting information from product labels. It should be emphasized though that our evaluation was based on an external comparison dataset, which is different from what was pursued in the other studies. Some of the relevant challenges and limitations were discussed above.
Our analysis revealed particular areas for improvement. Errors like “Incomplete Extraction,” “Text Format,” “Tool Inefficiency,” and “MedDRA Miscoding” could be treated with the appropriate enhancements in the ETHER algorithm, such as the addition of new rules or more lexical resources. For example, the “Adverse Reaction” section of product labels often contains laboratory test results, a type of information that is not necessarily an Adverse Reaction and is not processed by the current version of ETHER. It is therefore impossible to generate the corresponding PTs; however, this could be handled with the addition of the appropriate module to the ETHER algorithm. Similarly, ETHER is not programmed to process the “Indication for Use” section and exclude the corresponding terms from the “Adverse Reaction” section. There is also no process to exclude AE terms from headings or evaluate the overall context. Whereas the latter is often required to determine the appropriate PT for an AE term, it can be particularly challenging for an automated tool like ETHER. We argue that it is possible to efficiently handle certain context-related problems by performing additional development and further refining our system. The methodology followed in our QEA and the defined error types could be also reused in similar studies for systematically recognizing system weaknesses and identifying areas for improvement.
The improvement in ETHER’s ability to extract and translate AE terms to MedDRA PTs may assist the medical experts and epidemiologists at the FDA in using the SPL information in their pharmacoepidemiological research. We further believe that our work sheds some light into the processing and coding of other clinical texts, such as the clinical notes and discharge summaries found in electronic health records. The nature of clinical texts presents significant challenges to NLP applications and approaches of the kind may efficiently address them.
Conclusion
This study demonstrated ETHER’s potential contribution to the extraction and encoding of AE terms from SPLs toward the support of pharmacoepidemiological tasks. We have shown that the ETHER system can be applied to Adverse Reaction section of the SPLs and efficiently generate MedDRA PTs. The improvement in ETHER’s ability to extract and translate AE terms to MedDRA PTs may assist the medical experts and epidemiologists at the FDA in using the SPL information in their pharmacoepidemiological research.
Footnotes
Acknowledgements
The authors thank Michael Kuhn at the Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Germany, for his assistance with the Side Effect Resource (SIDER) and Carol Pamer at the Center for Drug Evaluation and Research, FDA, for her input and advice in this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This work was supported, in part, by the appointments of M.F., K.K., and A.P. to the Research Participation Program administered by ORISE through an interagency agreement between the US Department of Energy and the US Food and Drug Administration (FDA).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
