Abstract
Background:
Lung adenocarcinoma (LUAD) is the most common lung cancer, associated with high metastasis and low survival rates. Identifying reliable biomarkers is essential for better prognosis and treatment.
Methods:
In this study, we analyzed RNA sequencing data, mutation information, and clinical data from the TCGA-LUAD cohort and other multicenter datasets to investigate the role of Holliday junction recognition protein (HJURP) in LUAD. We employed immunohistochemistry in tissue microarray cohort to validate the prognostic significance of HJURP. The DepMap project was used to validate the effect of HJURP knockout in vitro.
Results:
Holliday junction recognition protein was identified as an adverse prognostic factor in the TCGA-LUAD cohort and diverse ethnic groups. Its expression correlated with poor immunotherapy outcomes, and HJURP knockout suppressed cancer cell proliferation. High HJURP expression was linked to increased mutation frequency, particularly in TP53 and TTN. Pan-cancer analysis also indicated HJURP as a poor prognostic factor in various solid tumors.
Conclusions:
Holliday junction recognition protein emerges as a significant biomarker in LUAD, consistently associated with poor prognosis across multiple cohorts. Its role in various oncogenic pathways and correlation with advanced disease stages underscore the potential of HJURP as a target for therapeutic intervention and as a marker for prognosis in LUAD.
Keywords
Introduction
Lung cancer remains a predominant concern in oncology, consistently leading in both prevalence and fatality rates, and represents a significant challenge to worldwide public health systems. 1 Specifically, lung adenocarcinoma (LUAD), accounting for more than 40% of lung cancer cases, presents a pronounced systemic risk due to its proclivity for distant metastases and an average overall survival (OS) duration of less than 5 years. 2 Furthermore, conventional treatment modalities such as surgery, radiotherapy, and chemotherapy, while standard, are often encumbered by limitations or associated with a range of adverse effects. 3 Given the substantial mortality associated with lung cancer, the development of effective prognostic markers is imperative to facilitate tailored therapeutic approaches for patients with LUAD.
The current era of medical science is distinguished by a significant reshaping of our comprehension of cancer biology. The emergence of specific biological markers for cancer has enabled a more individualized treatment paradigm, enhancing the traceability of cancer’s malignant progression and paving the way for novel defensive strategies against this disease.4-6 In this context, the enhancement of multi-omics has been a crucial factor, offering robust support in the search for key genes implicated in LUAD. This development provides fresh perspectives in the quest for innovative biomarkers and the identification of new therapeutic targets.
In the sphere of oncological research, Holliday junction recognition protein (HJURP) has been identified as playing a tumorigenic role in various cancers, including glioma, oral cancer, melanoma, and breast cancer.7-10 Studies have elucidated the involvement of HJURP in critical aspects of tumorigenesis, such as cellular proliferation, chromosomal instability, and oncogene activation. Mechanistically, HJURP functions as a chaperone for centromere protein A (CENPA) histone, facilitating its placement during the late mitosis to early G1 phase of the cell cycle. 11 Kato et al 12 used microarray profiling to identify HJURP and reported its overexpression in both lung cancer cell lines and tissue samples. Their findings revealed that elevated HJURP expression in non–small cell lung cancer (NSCLC) was associated with poorer OS.
Within the context of LUAD, HJURP has been the subject of limited research. The work of Wei and colleagues indicates HJURP’s role as a tumor promoter in NSCLC through the activation of the Wnt/β-catenin pathway. 13 Furthermore, research by Chen et al 14 suggests HJURP’s potential as a prognostic molecular biomarker indicative of poor prognosis in LUAD. However, these studies are predominantly single-omics in nature and lack validation through clinical samples from their respective centers. To further investigate the role of HJURP in LUAD, our study uniquely combines multicenter, multi-omics sequencing data with a tissue microarray (TMA) cohort, aiming to explore the feasibility of HJURP as a biomarker in LUAD.
Materials and Methods
Data collection and processing
Our study commenced with the retrieval of RNA sequencing (RNA-seq) data, accompanying mutation information, and clinical data for Lung Adenocarcinoma (TCGA-LUAD) and pan-cancer cohorts from the UCSC Xena database (https://xenabrowser.net/datapages/). To facilitate this analysis, we employed the R packages “clusterProfiler” 15 and “org.Hs.eg.db” for converting the RNA-seq Ensembl IDs to SYMBOL IDs. For validation purposes, we gathered datasets for LUAD from the GEO database (http://www.ncbi.nlm.nih.gov/geo/), specifically the GSE31210, GSE50081, GSE68465, and GSE37745 datasets. The datasets used in this research are detailed in Table S1.
Prognostic evaluation
The assessment of prognosis in this study was based on the estimation of hazard ratios (HRs) through Kaplan-Meier (KM) analysis and the application of time-dependent receiver operating characteristic (ROC) curves. For these analyses, the R package “survival” was used for conducting KM analyses and Cox regression modeling. In addition, the R package “survivalROC” was employed to calculate the area under the ROC curves (AUCs). 16 The construction of the nomogram was facilitated using the “rms” package. In this study, a P-value less than .05 (2-sided) was considered indicative of statistical significance.
Functional enrichment
In our research, the association between HJURP expression and the expression of all genes was analyzed using the Spearman correlation method. To ascertain the functional enrichment and delineate the associations with KEGG pathways, we employed the “clusterProfiler” R package. 15 This approach facilitated the identification of biologically relevant pathways potentially linked to HJURP.
Mutation landscape
We aimed to investigate whether there are genomic mutation differences between high and low HJURP expression groups. Utilizing the “maftools” and “ComplexHeatmap” R packages, we visually represented the mutation landscape of significantly mutated genes (SMGs) in LUAD using mutation waterfall plots.17,18 In addition, we conducted a visual analysis of the amplifications (Amp) and deletions (Del) in chromosomal segments, as well as the copy number variations (CNVs) of genes, through CNV waterfall plots in LUAD. The differences in CNV among HJURP expression subgroups were calculated and confirmed using the χ2 test. This approach allowed for a detailed and clear visualization of the mutational variances associated with different HJURP expression levels in LUAD.
Immunohistochemistry
In this study, LUAD tissue samples were from a TMA cohort which is a commercially cohort (ZL-Luc601) sourced from Shanghai Zhuoli Biotech Company. This TMA cohort comprised 31 valid LUAD tissue samples along with corresponding clinical data. The procurement of these tissue samples was conducted with ethical approval from the Ethics Committee of Shanghai Zhuoli Biotech Company (approval number SHLLS-BA-22101102). Written informed consent was obtained from each participant for the collection and use of their samples in this research.
Prior to embedding in paraffin, samples were fixed in 4% formaldehyde. Tissue sections, sliced at a thickness of 4 μm, were treated with a 1% H2O2 solution and subsequently blocked with nonimmunogenic goat serum. The sections were incubated overnight with primary antibodies at 4°C, followed by a 30-minute room temperature incubation with biotinylated secondary antibodies to bind with the primary antibodies. The specific IHC staining procedure was consistent with our previous research. The final scoring was determined by adding together the percentage of positively stained cells and the intensity score of the staining. The scoring for the percentage of immunoreactive cells was defined as follows: 0 (0%), 1 (1%-10%), 2 (11%-50%), and 3 (>50%). The visual scoring and grading of staining intensity were as follows: 0 (negative), 1 (weak), 2 (moderate), and 3 (strong). The antibody used in this study was the anti-HJURP antibody (Proteintech, 15283-1-AP).
Evaluating HJURP dependency in vitro
To assess the reliance of various tumor cell lines on HJURP, we used CRISPR gene editing data sourced from the Broad Institute’s Dependency Map (DepMap) portal (https://depmap.org/portal/). Our analysis involved 2 primary modules from the DepMap resource—“Perturbation Effects” and “Predictability”. These modules facilitated a detailed examination of gene dependency and co-dependency. For this purpose, we downloaded the “Achilles_gene_effect.csv” and “sample_info.csv” files from the DepMap Public dataset, which were instrumental in conducting our analysis.
Statistical analysis
To evaluate associations between variables, Spearman correlation analysis was used for computing correlation coefficients. For comparing 2 continuous variables, either the Wilcoxon rank-sum test or Student t-test was implemented, depending on data distribution. In addition, for analyzing differences between categorical variables, the χ2 test or Fisher exact test was applied.
Reporting guidelines
The reporting of this study follows the REMARK guidelines, see supplementary materials. 19
Results
HJURP is identified as an unfavorable prognostic factor in LUAD
The work flow of this study, as depicted in Figure 1, began with the collection of multi-omics data from The Cancer Genome Atlas (TCGA) and 3 large LUAD datasets from the Gene Expression Omnibus (GEO): GSE31210, GSE50081, GSE68465, and GSE37745. Analysis of these datasets suggested the potential of HJURP as a biomarker for LUAD. Subsequent stages involved prognostic analysis, TMA cohort immunohistochemistry (IHC) validation, multi-omics analysis, and pan-cancer assessment, confirming HJURP as an unfavorable prognostic factor in LUAD.

The workflow of this study.
Initially, to evaluate the prognostic capability of HJURP in LUAD, KM survival analysis was performed across multiple center cohorts. This analysis, based on different prognostic endpoints such as OS, progression-free interval (PFI), disease-free survival (DFS), and progression-free survival, revealed that higher expression of HJURP was associated with poorer prognosis in LUAD (Figure 2A and B, P < .05). Subsequently, both univariate and multivariate cox regression assessments were conducted. The results indicated that HJURP in the multivariate Cox regression analysis was not statistically significant in GSE31210-OS (HR = 1.266, P = .094) and GSE50081-DFS (HR = 1.212, P = .315). However, in other datasets, higher expression levels of HJURP were demonstrated to be an independent prognostic risk factor in LUAD (Figure 2C and D, P < .05).

HJURP is an unfavorable biomarker in LUAD. (A) Kaplan-Meier survival analyses for HJURP in multicenter datasets. (B) The scatter diagram depicts the prognostic distribution among different patients. (C) Univariate Cox regression assessing HJURP in multicentre datasets. (D) Multivariate Cox regression assessing HJURP in multicenter datasets. P values < .05 were considered statistically significant. Abbreviations: HR, hazard ratio; CI, confidence interval. Data points represent individual patients; red = high-expression group, blue = low-expression group.
Constructing a nomogram with HJURP expression
To enhance clinical applicability, we developed a nomogram incorporating HJURP expression alongside clinical information such as age, tumor stage, and pathological T stage, to prognosticate OS in patients with LUAD (Figure 3A). The accuracy of this model was demonstrated through calibration curves, which revealed a strong concordance between the nomogram-predicted and actual 1-, 5-, and 8-year OS (Figure 3B). The AUC values for the 1-, 5-, and 8-year OS predictions were 0.746, 0.7, and 0.713, respectively (Figure 3C), underscoring the reliability and precision of our nomogram. Furthermore, the decision curve analysis illustrated that our nomogram exhibited superior predictive capabilities for 1-, 5-, and 8-year outcomes when compared with other benchmarks (Figure 3D). In summary, the developed nomogram, integrating HJURP expression, demonstrates robust and consistent predictive efficacy in assessing the prognosis of patients with LUAD.

Nomogram. (A) Development of a nomogram. (B, C) Performance evaluation by calibration and ROC curve. (D) DCA curve.
IHC validation of HJURP in TMA cohort
To further validate our observations derived from public datasets, an IHC assay was conducted on TMA cohort. This assay was crucial in confirming the prognostic relevance of HJURP in LUAD through TMA analysis. In LUAD samples, HJURP was predominantly found in the nucleus, with a secondary distribution in the cytoplasm (Figure 4A). Notably, increased HJURP expression was associated with more advanced disease stages (Figure 4B). Kaplan-Meier survival analysis was applied, revealing that high expression of HJURP is linked to a poor prognosis in terms of OS (Figure 4C). Time-dependent ROC analysis demonstrated that HJURP had robust predictive power for the third and fifth years of survival (3-year AUC = 0.83, 5-year AUC = 0.90, Figure 4D). Univariate and multivariate Cox regression analysis revealed that HJURP was an independent risk factor for OS (Figure 4E). These comprehensive approaches underscored the effectiveness of HJURP as a prognostic indicator for survival outcomes in patients with LUAD.

HJURP protein is elevated indicates worse survival in LUAD TMA cohort. (A) This section features illustrative images from IHC staining of HJURP in LUAD. (B) Evaluation of HJURP IHC scores across different clinical stages of LUAD, using the Wilcoxon rank-sum test. (C) The KM curves for survival analysis, illustrating OS in LUAD, with a distinction between groups having low (n = 17) and high (n = 14) HJURP expression. (D) Time-dependent ROC is conducted to assess the efficacy of HJURP in predicting 3-year and 5-year OS. (E) Cox regression analysis of OS. The box represents the interquartile range (IQR, Q1-Q3), the line inside the box indicates the median, and whiskers extend to 1.5×IQR. P < .05 was considered statistically significant.
Knocking out HJURP inhibits tumor proliferation in vitro
To elucidate the in vitro role of HJURP, we investigated its susceptibility to CRISPR/Cas9-mediated disruption across a broad array of human cancer cell lines, drawing upon data from the DepMap project. The gene effect scores, or dependency scores, serve as indicators of cell growth and survival diminution following gene depletion. A score of −1 denotes the median effect observed in all essential genes commonly affected. Crucially, HJURP knockout exhibited a suppressive impact on the proliferation of various cancer cell lines, including those of lung cancer, with the median dependency score approaching −1 (Figure 5A). The top 10 cell lines most sensitive to HJURP knockout, namely NB7, TMK1, OCUBM, PCI38, HT29, SNU16, KPNYS, NB17, NH12, and NCIH1437, all demonstrated dependency scores below −1.5. The species of these cell lines are displayed in Figure 5B, with NCIH1437 being a LUAD cell line. In addition, an analysis of the Top Co-dependency Pearson correlations of HJURP revealed CENPA, GTF3C5, OIP5, THG1L, GTF2H3, CPSF3, RFC5, WDR18, UTP23, and MIS18A as the top 10 co-dependencies (Figure 5C). These findings suggest that HJURP may play a potential oncogene effect in multiple cancer types, which warrants further experimental validation.

Exploring HJURP dependency in cancer cells using the DepMap project. (A) The impact of CRISPR/Cas9-mediated HJURP knockout was assessed across a spectrum of human cancer cell lines, with each line depicted by a distinct circle symbol in the analysis. (B) Identification of the top 10 cancer cell lines exhibiting heightened sensitivity to HJURP knockout. (C) Analysis of the top 5 co-dependencies observed in tumor cell lines following HJURP deletion via CRISPR/Cas9.
Functional enrichment analysis of HJURP
Following the identification of the prognostic significance of HJURP in multicenter LUAD cohorts and its oncogenic effects in vitro, we sought to investigate its potential biological functions. Using GSEA based on KEGG pathways, we discerned the activity profile of HJURP (Table S2). Our findings, as illustrated in Figure 6A and B, revealed that HJURP predominantly activated pathways associated with cell cycle, glycolysis/gluconeogenesis, spliceosome, ribosome, focal adhesion, and DNA replication. Conversely, pathways such as olfactory transduction, vesicular transport, allograft rejection, glycosaminoglycan degradation, asthma, and lysosome were primarily inhibited by HJURP, as shown in Figure 6C and D.

Functional enrichment profiling of HJURP. (A, B) Top 6 KEGG pathways depicting activation, ranked by normalized enrichment score (NES). (C, D) Top 6 KEGG pathways indicating suppression, ranked by NES. P < .05 was considered statistically significant.
HJURP in TCGA pan-cancer
In our investigation of HJURP, we observed its marked prognostic relevance in LUAD. To determine if this predictive potential extends to other cancer types, we conducted a pan-cancer analysis using TCGA data encompassing 32 distinct cancer types. This analysis revealed elevated HJURP expression in cancers such as testicular germ cell tumors, uterine serous carcinoma, diffuse large B-cell lymphoma, cervical squamous cell carcinoma, and thymoma (THYM). In contrast, lower expression levels of HJURP were observed in kidney chromophobe (KICH), thyroid carcinoma, pheochromocytoma and paraganglioma, kidney renal papillary cell carcinoma (KIRP), and low-grade glioma (Figure 7A). Univariate Cox regression analysis revealed HJURP as a risk factor in various tumors including adrenocortical carcinoma, KICH, kidney renal clear cell carcinoma, KIRP, among others, with detailed results presented in Figure 7B and C. Intriguingly, HJURP served as a protective factor only in the PFI of THYM. These findings indicate that HJURP is not solely a unique marker for LUAD prognosis but also possesses predictive relevance in certain solid tumors.

Prognostic value of HJURP in TCGA pan-cancer. (A) A boxplot representation is employed to depict the expression levels of HJURP across a pan-cancer spectrum. (B) Univariate Cox regression assessing HJURP expression and PFI across different tumor types. (C) Univariate Cox regression assessing HJURP expression and OS across different tumor types. The error bars indicate the 95% confidence interval (CI).
Multi-omics analysis regard to HJURP
In our study, primarily focused on transcriptomics, we endeavored to analyze the role of HJURP in LUAD at the DNA mutation level. We investigated high-frequency mutated SMGs and loss of CNV using mutational genomics (Figure 8). In addition, we conducted differential testing based on HJURP expression groups (Figure 8). In LUAD, we identified the top 5 high-frequency mutated genes as TP53, TTN, MUC16, APOB, and ASXL3, with TP53 and TTN mutations occurring at frequencies of 49% and 45%, respectively. Furthermore, in the HJURP high-expression group, we observed a higher mutation frequency in TP53, TTN, MUC16, HMCN1, and FRAS1 compared with the low-expression group (Figure 8, P < .05). Regarding CNVs, higher gains were noted at 8q24.21.Amp in the HJURP high-expression group (P < .05), whereas greater losses were observed at 13q12.11.Del and 4q34.3.Del (Figure 8, P < .05). Overall, higher HJURP expression was associated with increased heterogeneity at the mutation level.
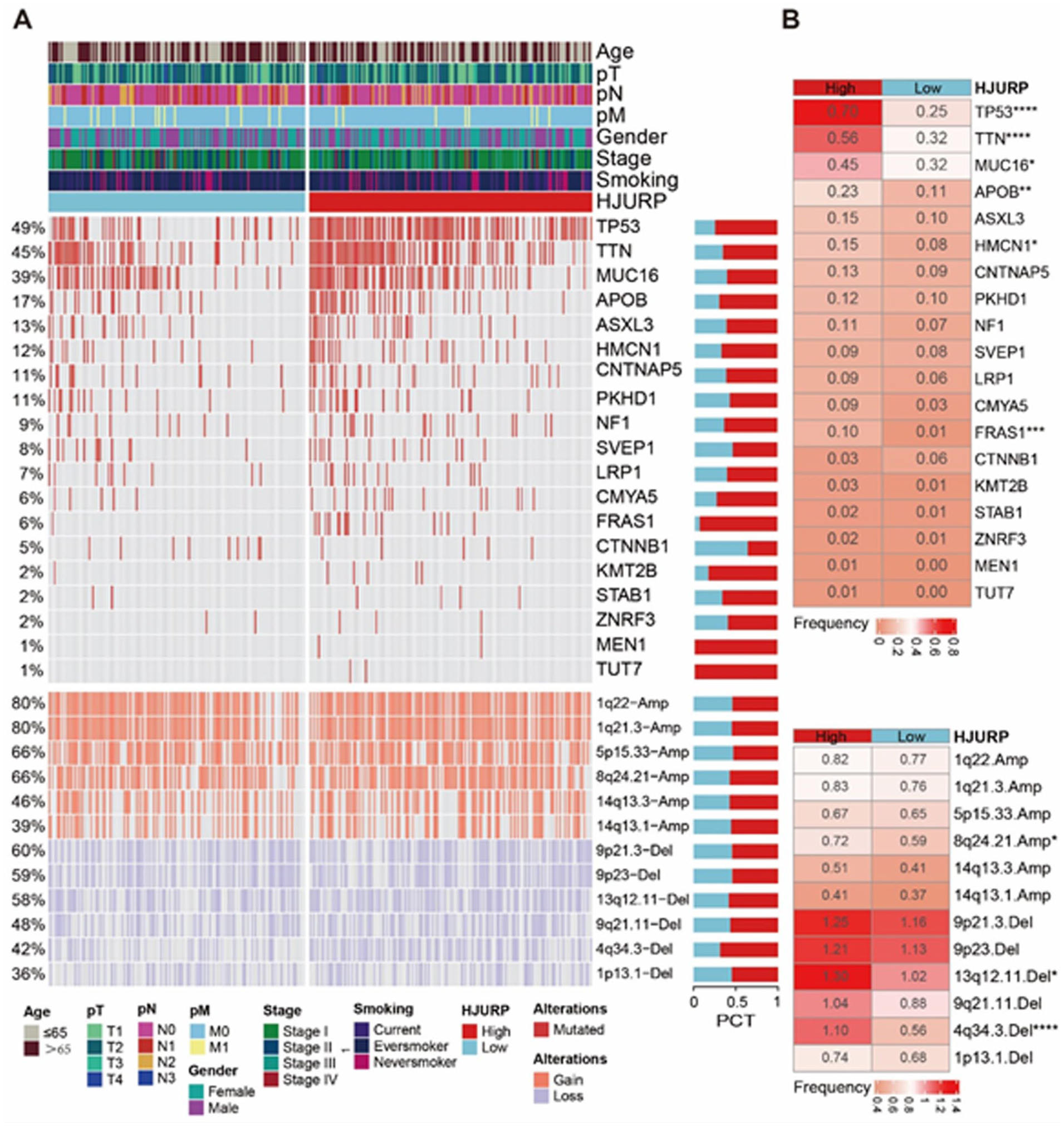
Genomic insights into HJURP. (A) This figure presents an integrated genomic landscape, showcasing the relationship between HJURP expression, SMGs, and CNV. (B) This section provides a comparative mutational analysis, emphasizing the differences in SMGs and CNV patterns among various subgroups classified by HJURP expression levels. *P < .05; **P < .01; ****P < .001.
HJURP is related to the immunotherapy response
Immune checkpoint inhibitors (ICIs) have emerged as a cornerstone in the treatment of various malignancies, with patients with lung cancer being among the primary beneficiaries. 20 Given the identification of HJURP as a prognostic marker across multiple cancers, we sought to investigate its association with immunotherapy outcomes. Using the online tool BEST (https://rookieutopia.com/app_direct/BEST/), we conducted comprehensive analyses. The Cho cohort, consisting of NSCLC patients undergoing anti-PD-1 therapy, revealed that nonresponders exhibited significantly higher HJURP expression levels. The AUC for HJURP’s predictive value in treatment response reached 0.764, and patients with elevated HJURP expression demonstrated shorter survival durations (Figure 9A). Similar trends were observed in other cancer cohorts treated with ICIs. For instance, elevated HJURP expression was associated with poorer outcomes in patients receiving CAR-T cell therapy (Figure 9B). Moreover, comparable patterns were evident in the anti-CTLA-4 cohort. Although KM survival analyses did not achieve statistical significance (P > .05), there were notable differences in survival rates between patients with high and low HJURP expression levels (Figure 9C). Collectively, these findings highlight the potential of HJURP as a predictive biomarker for immunotherapy efficacy.

HJURP is a potential biomarker for immunotherapy response. (A) Therapy response analysis of HJURP in Cho cohort. (B) Therapy response analysis of HJURP in CAR-T cohorts. (C) Therapy response analysis of HJURP in anti-CTLA-4 cohorts. The box represents the interquartile range (IQR, Q1-Q3), the line inside the box indicates the median, and whiskers extend to 1.5×IQR. P < .05 was considered statistically significant.
Discussion
Lung adenocarcinoma, recognized as the most prevalent subtype of lung cancer, is notably susceptible to metastasis and recurrence, factors that contribute to its relatively low 5-year survival rate.2,21 The collaborative efforts in establishing TCGA and GEO databases have resulted in a continuous influx of diverse cancer data. The expansive and rich nature of this big data has significantly enhanced our understanding and characterization of human cancers.22,23 Leveraging these invaluable data resources, our research identified HJURP as a potential biomarker in LUAD.
Despite previous scattered reports suggesting HJURP as a factor indicative of poor prognosis in LUAD, there was a lack of validation through comprehensive multi-omics, multicenter, and TMA cohort. Our current study addresses this research gap. Holliday junction recognition protein, CENP-A chaperone, plays a crucial role in the localization of CENP-A on centromeric chromatin, thereby facilitating centromeric assembly. 24 Some research has indicated the pro-oncogenic role of HJURP in various cancer types. Notably, HJURP exhibits overexpression in a diverse array of cancers, including those of the pancreas, kidney, breast, glioma, prostate, stomach, bladder, colorectal, myeloma, oral cavity, and liver.7,8,25-33 Our findings reveal that HJURP acts as an unfavorable prognostic factor not only in the TCGA-LUAD cohort, predominantly consisting of American patients, but also in the cohorts from Japan, Sweden, Canada, and TMA cohort Chinese population. This consistent role of HJURP across diverse ethnicities underscores its potential as a universal biomarker in LUAD prognosis.
Subsequent to our epidemiological findings, we delved into investigating the impact of HJURP on tumor cells. Our observations revealed that HJURP knockout not only influences lung cancer but also exerts a suppressive effect on the proliferation of diverse cancer cell lines within the DepMap project. Notably, the dependency scores for the NCIH1437 LUAD cell line were among the top 10. These analyses are in alignment with several published in vitro studies, suggesting an oncogenic effect of HJURP across multiple cancer types. 34 Coincidentally, in our pan-cancer analysis, HJURP emerged as a factor indicative of poor prognosis in various solid tumors. What is more, HJURP expression was found to be significantly associated with poor immunotherapy outcomes across multiple cancer cohorts, including anti-PD-1 therapy in NSCLC and CAR-T cell therapy. Patients with higher HJURP expression consistently demonstrated shorter survival durations and reduced treatment efficacy. These findings underscore the potential of HJURP as a predictive biomarker for response to ICIs. Compared with the preliminary findings by Kato et al, our study extends beyond a single population by using large-scale expression and survival data from global multicenter public datasets. In addition, we integrated nomogram modeling, CRISPR-Cas9 in vitro validation, and multi-omics analyses to comprehensively investigate the role of HJURP in NSCLC. 12 This finding lays a significant foundation for future research extending the role of HJURP to other cancer forms.
On identifying the oncogenic effect of HJURP in various cancers, our focus shifted to deciphering the biological mechanisms underlying its action. Functional enrichment analyses revealed that HJURP predominantly activates pathways related to the cell cycle, glucose metabolism, and DNA repair. Previous studies showed that overexpression of CENPA often correlates with HJURP overexpression, leading to aberrant CENPA deposition.35,36 This anomaly contributes to mitotic defects, centromeric dysfunction, and chromosomal instability, ultimately promoting cancer development and progression. Heo et al 37 identified that HJURP down-regulation regulated cellular senescence through the p53-dependent pathway. Besides, HJURP’s role extends beyond genetic mechanisms in cancer progression to encompass epigenetic influences. Lai et al 27 discovered that HJURP modulates cyclin-dependent kinase inhibitor 1 (CDKN1A) via the GSK3/JNK signaling pathway in prostate cancer, impacting tumor cell growth. Holliday junction recognition protein has also been implicated in modulating tumor cell sensitivity to radiotherapy through its involvement in DNA repair processes.38,39 Filipescu et al 40 reported that HJURP is highly expressed in p53-deficient tumors and promotes tumor growth. Building on this finding, we extended the investigation of HJURP to NSCLC and systematically evaluated its association with somatic mutations in multiple key driver genes. Unlike prior studies that mainly focused on TP53 mutations, our analysis included a larger sample size and a broader mutation spectrum. Building on this, our mutational genomics analysis revealed distinct patterns in the HJURP high-expression group. This group exhibited an increased frequency of mutations in genes such as TP53, TTN, MUC16, HMCN1, and FRAS1 when compared with the group with lower HJURP expression. Moreover, a significant upsurge in chromosomal gains was observed at the locus 8q24.21.Amp in the high-expression group. Conversely, more pronounced losses were detected at chromosomal regions 13q12.11.Del and 4q34.3.Del. Based on these discoveries, it is posited that HJURP plays a critical role in epigenetic modifications, which manifest at the initial stages of tumor development. The functional impact of HJURP across various tumor types is mediated by diverse mechanisms of action.
Moreover, this study acknowledges certain limitations. Due to constraints in time and funding, an in-depth mechanistic exploration of HJURP’s role in LUAD was not feasible. In addition, TMA cohort comprised only 31 cases, a relatively small sample size compared with larger public datasets. Future research endeavors are planned to methodically address and rectify these shortcomings.
Conclusions
This study found HJURP as a robust prognostic biomarker in LUAD, consistently associated with poor OS across diverse multicenter cohorts, including TCGA-LUAD and populations from Japan, Sweden, Canada, and China. Multi-omics analyses reveal HJURP’s role in promoting tumor progression through pathways related to cell cycle, DNA repair, and glucose metabolism, with higher mutation frequencies in genes such as TP53 and TTN observed in high HJURP expression groups. Furthermore, elevated HJURP expression correlates with unfavorable immunotherapy outcomes, highlighting its potential as a predictive marker for ICI response. These findings underscore HJURP’s potential as a therapeutic target and a reliable biomarker for risk stratification and personalized treatment in LUAD.
Supplemental Material
sj-xlsx-1-onc-10.1177_11795549251388872 – Supplemental material for Unveiling HJURP as a Biomarker of Poor Prognosis and Immunotherapy Resistance in Lung Adenocarcinoma: A Multicenter Study
Supplemental material, sj-xlsx-1-onc-10.1177_11795549251388872 for Unveiling HJURP as a Biomarker of Poor Prognosis and Immunotherapy Resistance in Lung Adenocarcinoma: A Multicenter Study by Qinglin Tan, Peiliang Kong, Guobiao Chen, Chen Chen, Huiting Mo, Yuancheng Huang, Manman Zhang, Yanmin Cai, Hanbin Zhang, Jianming Lu and Yifen Wu in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
The authors express their sincere appreciation to the TCGA (The Cancer Genome Atlas) and GEO (Gene Expression Omnibus) databases for making lung cancer data publicly available, which has substantially aided the research conducted in this study.
Ethical considerations
Lung tissue microarrays used in this study were commercially purchased from Shanghai Zhuoli Biotech Company (Product ID: ZL-Luc601). All samples were collected by the supplier with prior informed consent from the patients and ethical approval from the supplier’s institutional review board. As the authors were not directly involved in the collection of clinical samples, no additional patient recruitment or interventions were conducted in this study.
Author contributions
Yifen Wu played a pivotal role in securing funding and designing the study. The public datasets collection and analysis were diligently conducted by Qinglin Tan, Peiliang Kong, Guobiao Chen, Yanmin Cai, and Manman Zhang. The initial draft of the manuscript was prepared by Qinglin Tan, Peiliang Kong, Chen Chen, Huiting Mo, and Yuancheng Huang. The immunohistochemistry was completed by Qinglin Tan and Guobiao Chen and Hanbin Zhang. The manuscript underwent thorough refinement and revision by Yifen Wu and Jianming Lu. Each author made significant contributions to the development of the article and has given their approval for the final version.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Guangdong Basic and Applied Basic Research Foundation (Project’s number: 2020B1515120063).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
The datasets utilized in this study has been delineated within the Materials and Methods section. For any further inquiries, please reach out to the corresponding author.
Informed consent statement
All clinical samples were obtained through routine diagnostic procedures with written informed consent from all patients prior to use.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
