Abstract
Objectives:
Lung adenocarcinoma (LUAD), a predominant form of lung cancer, is characterized by a high rate of metastasis and recurrence, leading to a poor prognosis for LUAD patients. This study aimed to identify and rigorously validate a highly precise biomarker, Cathepsin L (CTSL), for the prognostic prediction of lung adenocarcinoma.
Methods:
We employed a multicenter and omics-based approach, analyzing RNA sequencing data and mutation information from public databases such as The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GEO). The DepMap portal with Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR/Cas9) technology was used to assess the functional impact of CTSL. Immunohistochemistry (IHC) was conducted on a local cohort to validate the prognostic significance of CTSL at the protein expression level.
Results:
Our findings revealed a significant correlation between elevated CTSL expression and advanced disease stage in LUAD patients. Kaplan–Meier survival analysis and Cox regression modeling revealed that high CTSL expression is associated with poor overall survival. The in vitro studies corroborated these findings, revealing notable suppression of tumor proliferation following CTSL knockout in cell lines, particularly in LUAD. Functional enrichment revealed that CTSL activated pathways associated with tumor progression, such as angiogenesis and Transforming growth factor beta (TGF-beta) signaling, and inhibited pathways such as apoptosis and DNA repair. Mutation analysis revealed distinct variations in the CTSL expression groups.
Conclusion:
This study highlights the crucial role of CTSL as a prognostic biomarker in LUAD. This combined multicenter and omics-based analysis provides comprehensive insights into the biological role of CTSL, supporting its potential as a target for therapeutic intervention and a marker for prognosis in patients with LUAD.
Keywords
Introduction
Consistently occupying a leading position in cancer statistics, lung cancer poses an ongoing and severe public health challenge. 1 Within this context, lung adenocarcinoma (LUAD) has emerged as a significant subtype, representing a substantial proportion of lung cancer cases. Characterized by a high likelihood of distant metastasis and a mean overall survival (OS) of under 5 years, LUAD presents a formidable threat. 2 The limitations and side effects associated with traditional treatments such as surgery, radiotherapy, and chemotherapy further complicate the clinical management of this condition. 3 Therefore, there is an urgent need for the development and implementation of reliable biomarkers to guide specific treatment plans for LUAD, with the aim of improving patient outcomes.
A defining feature of contemporary medicine in this century is the transformative understanding of the biological landscape of cancer. 4 The advent of cancer biomarkers has revolutionized patient care, offering a tailored approach to treatment. 5 This evolution has rendered the process of cancer malignant transformation more discernible, leading to the development of innovative methods for combating the disease. 6 Advancements in high-throughput sequencing technologies have been instrumental in the identification of pivotal genes associated with LUAD. 7 This technological progress opens new avenues for the discovery of groundbreaking biomarkers and therapeutic targets.
Cathepsin L (CTSL), a cysteine protease family member, is pivotal for lysosomal degradation of intracellular and endocytosed proteins, intracellular protein breakdown, and SARS-CoV-2 infection processes.8,9 CTSL upregulation is frequently observed in a variety of human cancers and contributes to oncogenesis, tumor progression, invasion, and poor patient prognosis.10 -12 Moreover, the significant association of CTSL with cancer-related osteolysis detrimentally impacts both the quality and length of life of cancer patients. 13
Reports on CTSL in LUAD have been sporadic. Han and colleagues, in a preclinical study, demonstrated that EMT induced by CTSL upregulation was linked to chemotherapy resistance in A549 cells. 14 Additionally, higher expression levels of CTSL are correlated with shorter OS in patients with lung cancer and may indicate a poor response to immunotherapy. 15 To further our understanding of the role of CTSL in LUAD, our study is the first to utilize a comprehensive approach combining multicenter, multiomics data with our local cohort. This systematic and comprehensive methodology aims to explore the potential of CTSL as a biomarker in LUAD. Previous studies have been limited to single-omic analyses and lack validation with clinical samples from their respective centers. Our integrated approach provides a more holistic view of the implications of CTSL in LUAD.
Methods
Data processing
This research began with acquiring RNA sequencing (RNA-seq) data, mutation details, and clinical information for Lung Adenocarcinoma (TCGA-LUAD) and pan-cancer cohorts, sourced from the UCSC Xena database (https://xenabrowser.net/datapages/). To further substantiate our findings, LUAD datasets from the GEO database (http://www.ncbi.nlm.nih.gov/geo/), namely GSE50081, GSE31210, and GSE68465, were collected. These datasets are listed in Supplemental Table S1. To convert Ensembl IDs to SYMBOL IDs within the RNA-seq datasets, we employed the R packages clusterProfiler and org.Hs.eg.db (version 3.17.0). 16 For the microarray data, batch effects were adjusted through the ComBat function within the sva R package. 17 After obtaining these datasets, we retained the LUAD samples only. Additionally, only samples with both gene expression and follow-up information were included in subsequent analyses.
Statistical analysis
Prognostic analysis in this study involved calculating Hazard Ratios (HR) using Kaplan–Meier (KM) analysis, coupled with time-dependent Receiver Operating Characteristic (ROC) curve assessments. The “survival” R package was employed for KM analysis and Cox regression modeling. The “survivalROC” R package was used to determine the Area Under the ROC Curves (AUC). 18 Specifically, Cox regression models were built using the coxph function from this R package to investigate the association between gene expression and clinical outcomes. The significance of the Kaplan–Meier survival curves was determined through the log-rank test.
In assessing the relationships between variables, Spearman correlation analysis was employed to calculate correlation coefficients. When comparing two continuous variables, we utilized either the Wilcoxon rank-sum test based on the distribution characteristics of the data.
Immunohistochemistry (IHC)
In this investigation, LUAD tissue microarray (TMA) were obtained from Shanghai Zhuoli Biotech Company. The TMA, identified as ZL-Luc601, contained 31 validated LUAD tissue samples, each accompanied by relevant clinical data. The acquisition of these samples was approved by the Ethics Committee of Shanghai Zhuoli Biotech Company (approval number SHLLS-BA-22101102), with written informed consent acquired from each participant for the use of their samples in this study.
For the immunohistochemistry protocol, detailed methodologies are outlined in our previously published paper, 19 with a summary provided herein. Antigen retrieval was performed on tissue slices or TMAs using a citrate antigen retrieval solution. This was followed by blocking the endogenous peroxidase activity with a 1% H2O2 solution and applying nonimmune goat serum for blocking. Tissue slices were incubated overnight at 4°C with the primary antibody, followed by a 30-minute room temperature incubation with a biotinylated secondary antibody. This was succeeded by a 15-minute room temperature incubation with streptavidin-conjugated horseradish peroxidase (HRP). The HRP activity was visualized using diaminobenzidine tetrahydrochloride (DAB), and nuclear staining was conducted using haematoxylin. The scoring of immunoreactive cells was categorized as 0 (0%), 1 (1-10%), 2 (11-50%), and 3 (>50%), while staining intensity was visually scored and graded as 0 (negative), 1 (weak), 2 (moderate), and 3 (strong). The specific antibody employed in this study was the anti-CTSL antibody (Immnoway, YT5124).
Functional enrichment analysis
We explored the relationship between CTSL expression and the expression profiles of all mRNAs, utilizing the Spearman correlation method for analysis. To pinpoint functional enrichment and establish connections with Hallmark pathways, the “clusterProfiler” 16 R package was employed. This method enabled us to detect and highlight biologically significant pathways that may be associated with CTSL expression.
Assessing CTSL dependency
Our investigation into the dependency of various tumor cell lines on CTSL was conducted through the Dependency Map (DepMap) portal, specifically leveraging CRISPR data (https://depmap.org/portal/). The analysis encompassed two main modules within the DepMap resource: “Perturbation Effects” and “Predictability.” These modules were critical in providing a comprehensive analysis of gene dependency and co-dependency. Key to this analysis was the utilization of the “Achilles_gene_effect.csv” and “sample_info.csv” files from the DepMap dataset. These files provided the necessary data for a thorough assessment of CTSL’s role across different tumor cell lines.
The landscape of mutation
In our genomic analysis, we quantified all base substitutions, insertions, and deletions within the coding regions of targeted genes. The top 10 genes exhibiting frequent mutations were identified as driver genes using the “maftools” R package.
20
Corresponding to the expression levels of CTSL, we delineated the mutation landscapes for these genes across two distinct sample groups, considering a
Immunotherapy response evaluation
We utilized the online tool “BEST” (https://rookieutopia.com/app_direct/BEST/) to forecast patient responses to immune checkpoint blockade (ICB) therapy. 22 The transcriptomic expression profiles were compared among patient groups with different levels of CTSL expression and varied immunotherapy outcomes. For the immunotherapy cohort, samples were stratified according to optimal CTSL expression cut-off values for KM survival analysis.
Results
Multicenter data indicate that CTSL is an unfavorable prognostic factor in LUAD
This study, outlined in Figure 1, involved the collection of multiomic data from the TCGA and three large-sample LUAD datasets from the GEO, specifically from Japan (GSE31210), Canada (GSE50081), and the United States (GSE68465). Our analysis identified CTSL as a potential biomarker for LUAD. Subsequent validation through immunohistochemical analyses in a local cohort, multi-omics evaluations, and pan-cancer assessments confirmed that CTSL was an adverse prognostic factor in LUAD.
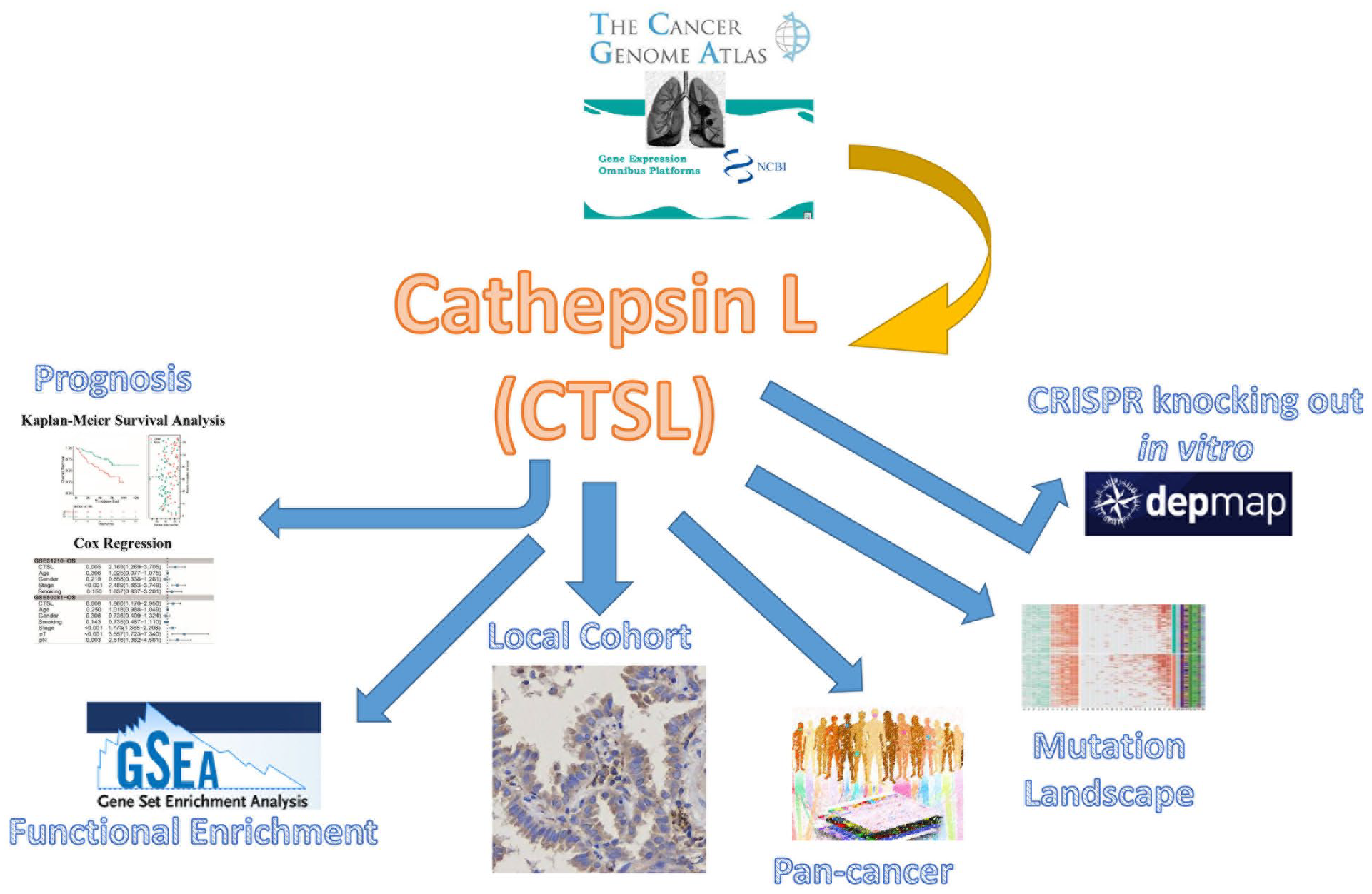
Flowchart.
Initially, to assess the prognostic significance of CTSL in LUAD, KM survival analysis was conducted across multicenter cohorts. Considering various prognostic endpoints (OS, progression-free interval [PFI], disease-free survival [DFS], and progression-free survival [PFS]), groups with higher CTSL expression had poorer prognoses in LUAD patients than those with lower CTSL expression (Figure 2A–G,
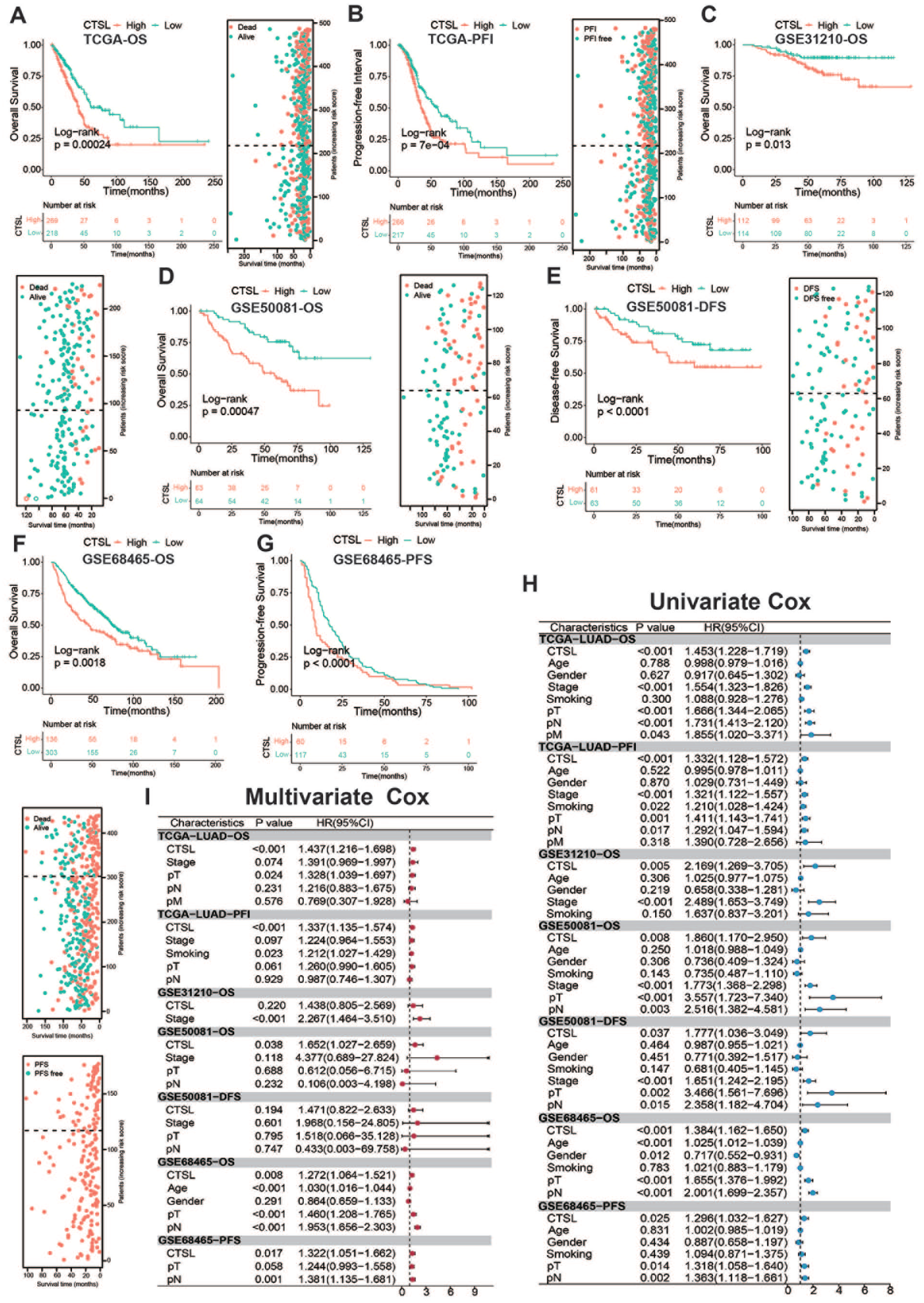
CTSL is an unfavorable biomarker in LUAD: (A–G) Kaplan–Meier survival analyses for CTSL in different LUAD datasets, (H) univariate Cox regression, and (I) multivariate Cox regression assessing CTSL in multi-center datasets.
IHC of CTSL in the local cohort
To substantiate the findings obtained from public datasets, we performed an IHC assay on our local cohort. This step was pivotal for confirming the prognostic significance of CTSL in LUAD at the protein expression level using a tissue microarray. Our findings, as depicted in Figure 3A, revealed a predominant cytoplasmic localization of CTSL. Notably, elevated CTSL expression was correlated with advanced disease stage (Figure 3A). KM survival analysis further revealed that high CTSL expression was indicative of an unfavorable prognosis in terms of OS (Figure 3B). Time-dependent ROC analysis, as shown in Figure 3C, revealed that the CTSL had a significant ability to predict third- and fifth-year survival (3-year AUC = 0.85, 5-year AUC = 0.91). This in-depth analysis highlights the crucial role of CTSL as a prognostic marker for survival outcomes in LUAD patients.
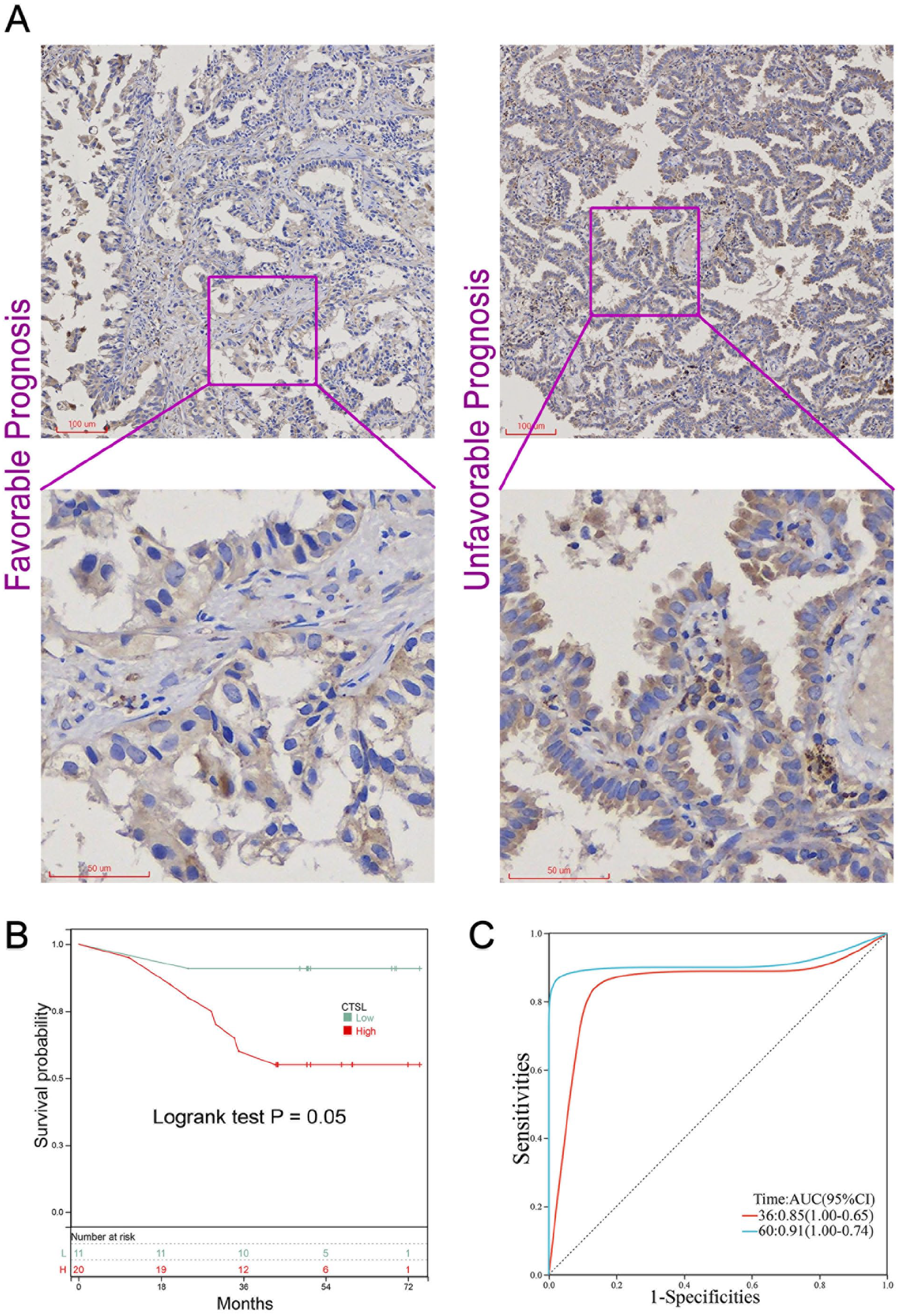
IHC of CTSL in local LUAD cohort: (A) representative staining images of CTSL in patients with varying prognoses within the LUAD cohort, (B) Kaplan–Meier survival curves depicting OS in LUAD patients, categorized into low (
CTSL as a prognostic factor in multiple solid tumors
Our research initially highlighted the significant prognostic role of CTSL in LUAD. To explore whether CTSL’s predictive capability extends beyond LUAD, we performed a comprehensive pan-cancer analysis. Using the TCGA database, we examined 32 different cancer types. Our findings revealed increased CTSL expression in various cancers, including uveal melanoma (UVM), skin cutaneous melanoma (SKCM), kidney renal papillary cell carcinoma (KIRP), sarcoma (SARC), and diffuse large B-cell lymphoma (DLBC). Conversely, a reduction in CTSL expression was noted in Esophageal Carcinoma (ESCA), Rectum Adenocarcinoma (READ), Colon Adenocarcinoma (COAD), Stomach Adenocarcinoma (STAD), Cervical Squamous Cell Carcinoma and Endocervical Adenocarcinoma (CESC) (Figure 4A). Univariate Cox regression analysis (Figure 4B) identified CTSL as a risk factor for several tumors, which may affect OS, PFI, or both, including glioblastoma multiforme (GBM), ESCA, head and neck squamous cell carcinoma (HNSC), low-grade glioma (LGG), liver hepatocellular carcinoma (LIHC), lung squamous cell carcinoma (LUSC), STAD, kidney chromophobe (KICH), and prostate adenocarcinoma (PRAD). Interestingly, CTSL appeared to act as a protective factor in UVM and kidney renal clear cell carcinoma (KIRC). These observations suggest that the role of CTSL is not only related to LUAD prognosis but also extends to other solid tumors, highlighting its broader predictive significance.
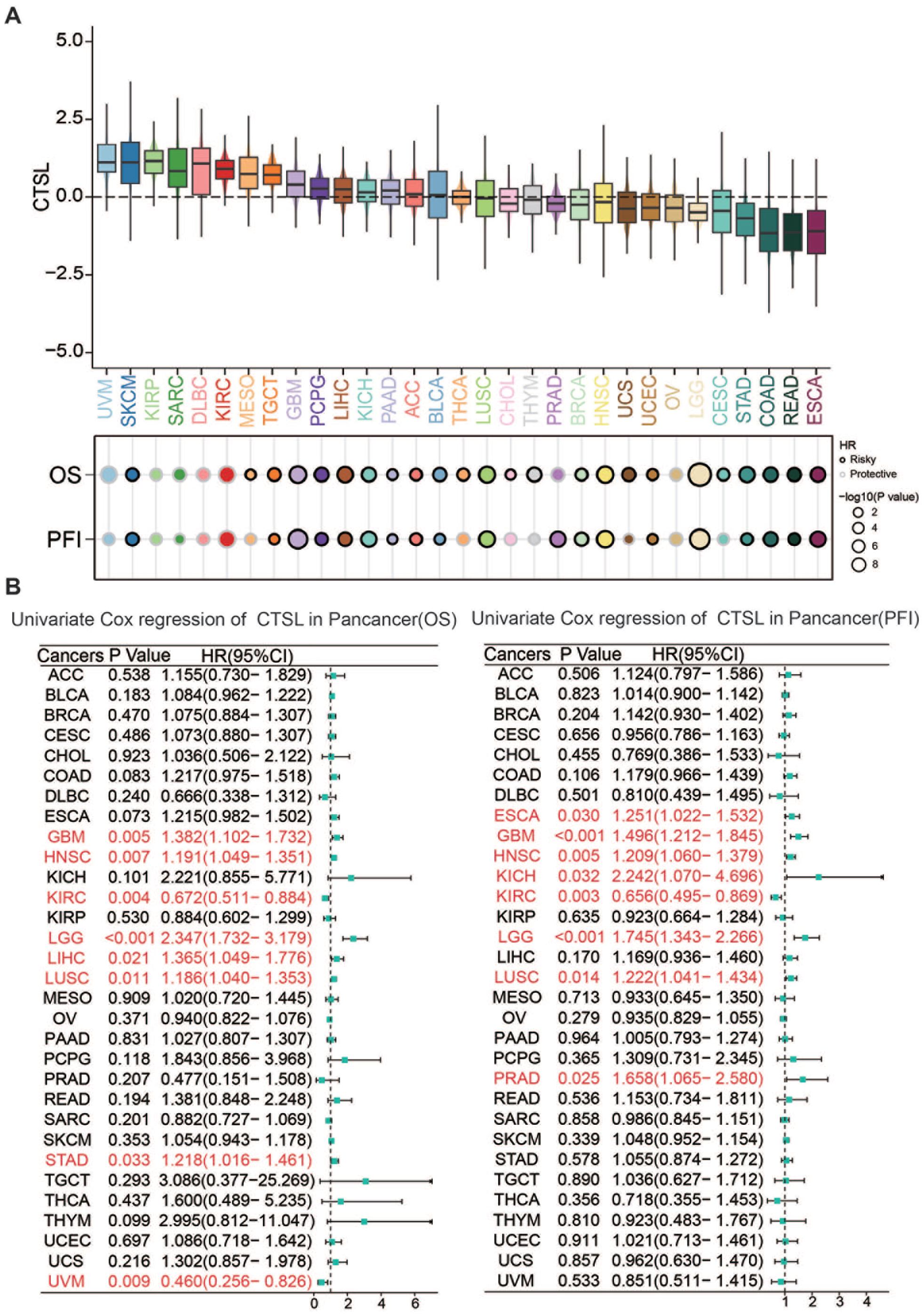
Pan-cancer analysis of CTSL: (A) expression levels of CTSL in TCGA pan-cancer and (B) univariate Cox regression assessing CTSL expression in PFI and univariate Cox regression assessing CTSL expression in OS across different tumor types.
Knocking out CTSL inhibits LUAD cell viability
To elucidate the functional impact of CTSL in vitro, we used CRISPR/Cas9 datasets for the targeted knockout of CTSL in various cell lines. Analysis conducted with DepMap revealed that negative scores on the gene effect scale (dependency scores) are indicative of reduced cell proliferation and survival subsequent to the depletion of a specific gene. As depicted in Figure 5A, the knockout of CTSL manifested differential effects across cell lines, with a majority displaying dependency scores below zero. This trend suggests a predominant inhibitory influence on cellular growth (Supplemental Table S3). In particular, the LUAD cell line NCIH1573 showed the most pronounced dependency (Figure 5B). NCIH1573 is a type of adherent cell that originates from a 35-year-old female who was diagnosed with stage 4 LUAD. It is derived from tumor cells of soft tissue prior to radiation therapy. Furthermore, Figure 5C shows that the top ten lung cancer cell lines exhibited the most significant sensitivity to CTSL depletion, underscoring their vulnerability to its absence.

Knocking out CTSL displays negative dependency score: (A) knocking out CTSL by CRISPR/Cas9 in different human cancer cell lines, (B) identification of the top 10 cancer cell lines exhibiting heightened sensitivity to CTSL knockout, and (C) identification of the top 10 lung cancer cell lines exhibiting heightened sensitivity to CTSL knockout.
Functional enrichment analysis of CTSL
Given the prognostic importance of CTSL in cancer, along with its in vitro oncogenic effects, our research aimed to explore its underlying biological functions. We employed GSEA focused on hallmark pathways to characterize the activity profile of CTSL. Figure 6 shows that CTSL predominantly activated several pathways critically associated with tumor progression, such as angiogenesis, hedgehog signaling, TGF-beta signaling, epithelial‒mesenchymal transition, and hypoxia. Concurrently, CTSL was found to suppress a number of classical pathways, including apoptosis, mTORC signaling, DNA repair, and those related to the cell cycle. This dual role of CTSL, both in the activation and suppression of key pathways, offers a strategic direction for our subsequent mechanistic investigations into the role of CTSL in cancer biology.
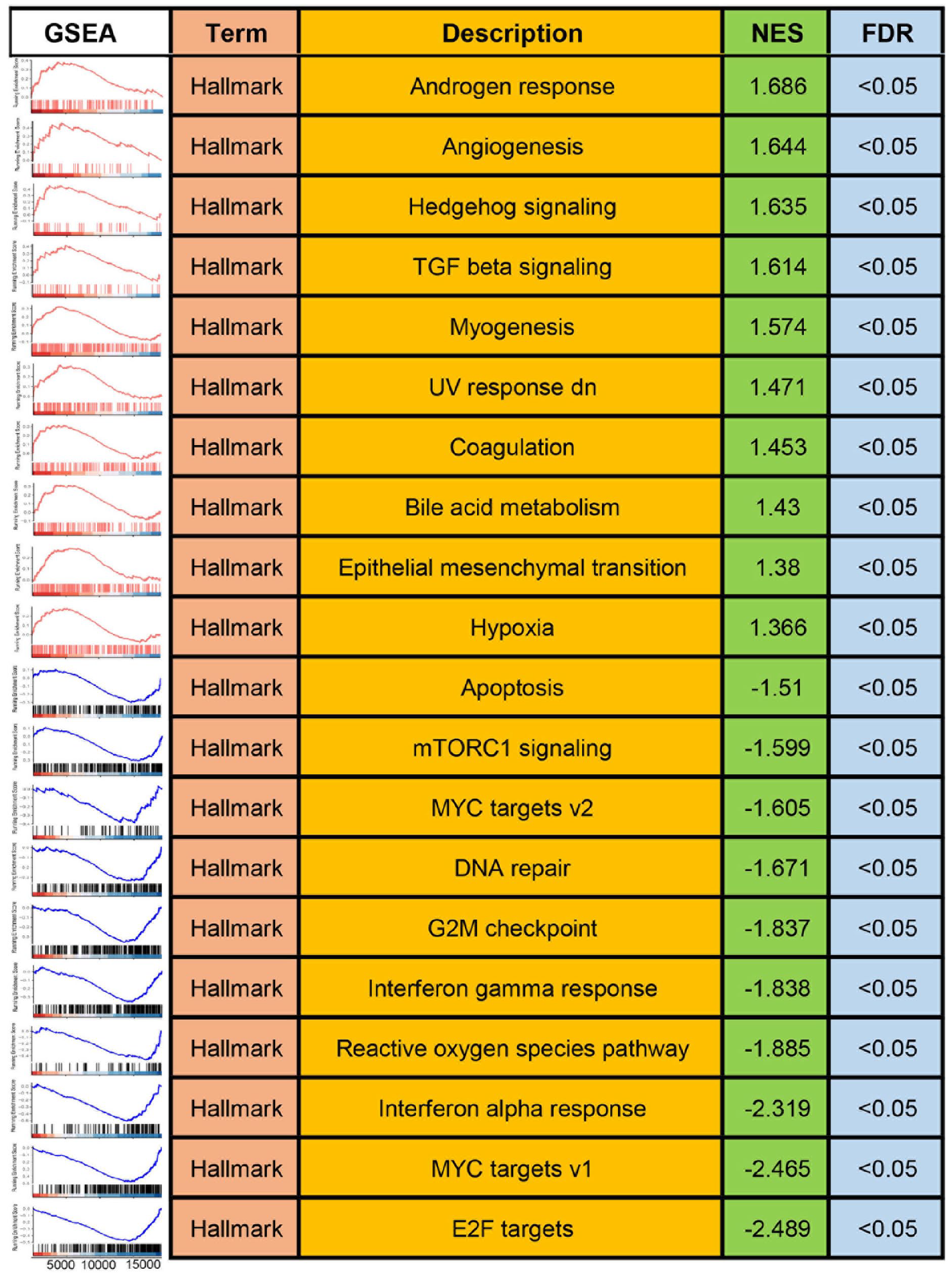
Functional enrichment analysis.
Role of CTSL in the genomic landscape of LUAD mutations
While our preceding analyses focused primarily on the transcriptome, we explored the role of CTSL at the mutation level in LUAD (Figure 7). Within the TCGA-LUAD cohort, the top ten genes exhibiting high mutation rates in tumor samples included TP53, TTN, MUC16, APOB, ASXL3, HMCN1, CNTNAP5, PKHD1, NF1, and SCEP1 (Figure 7A). Notably, TP53, TTN, and MUC16 had mutation frequencies of 49%, 45%, and 39%, respectively. Differential analysis based on CTSL expression levels revealed that genes such as TP53, APOB, and FRAS1 presented a relatively high mutation frequency in the high-CTSL-expressing group (Figure 7B). With respect to copy number variations (CNVs), the most commonly amplified genomic regions were at 1q22-Amp and 1q21.3-Amp, affecting up to 80% of the samples. Additionally, the most frequent CNV losses were 9q21.3-Del, 9q23-Del, and 13q12.11-Del, with frequencies of 60%, 59%, and 58%, respectively. However, no significant differences were observed between the high and low CTSL expression groups in terms of CNVs (Figure 7B). These findings suggest that CTSL may be more closely associated with gene mutations than with changes at the CNV level in LUAD.

Mutation landscape: (A) heatmap presents an integrated genomic landscape of high and low CTSL expression and (B) mutational differences in FMGs of CTSL expression subgroups.
CTSL is associated with the immunotherapy response
Immune checkpoint inhibitors (ICIs) have become a pivotal treatment modality for LUAD. Following the identification of CTSL as a prognostic factor in LUAD, we explored its association with the immunotherapy response. In the anti-PD-1/PD-L1 cohorts of Cho and Kim, both NSCLC patients, elevated CTSL expression was correlated with improved outcomes. Specifically, patients in the responder group presented increased CTSL expression levels, and the AUC for the ability of CTSL to predict treatment response exceeded 0.7 (Figure 8A and B). Furthermore, similar trends were observed in other cancer cohorts treated with immune checkpoint inhibitors; high CTSL expression was linked to better outcomes. Although the
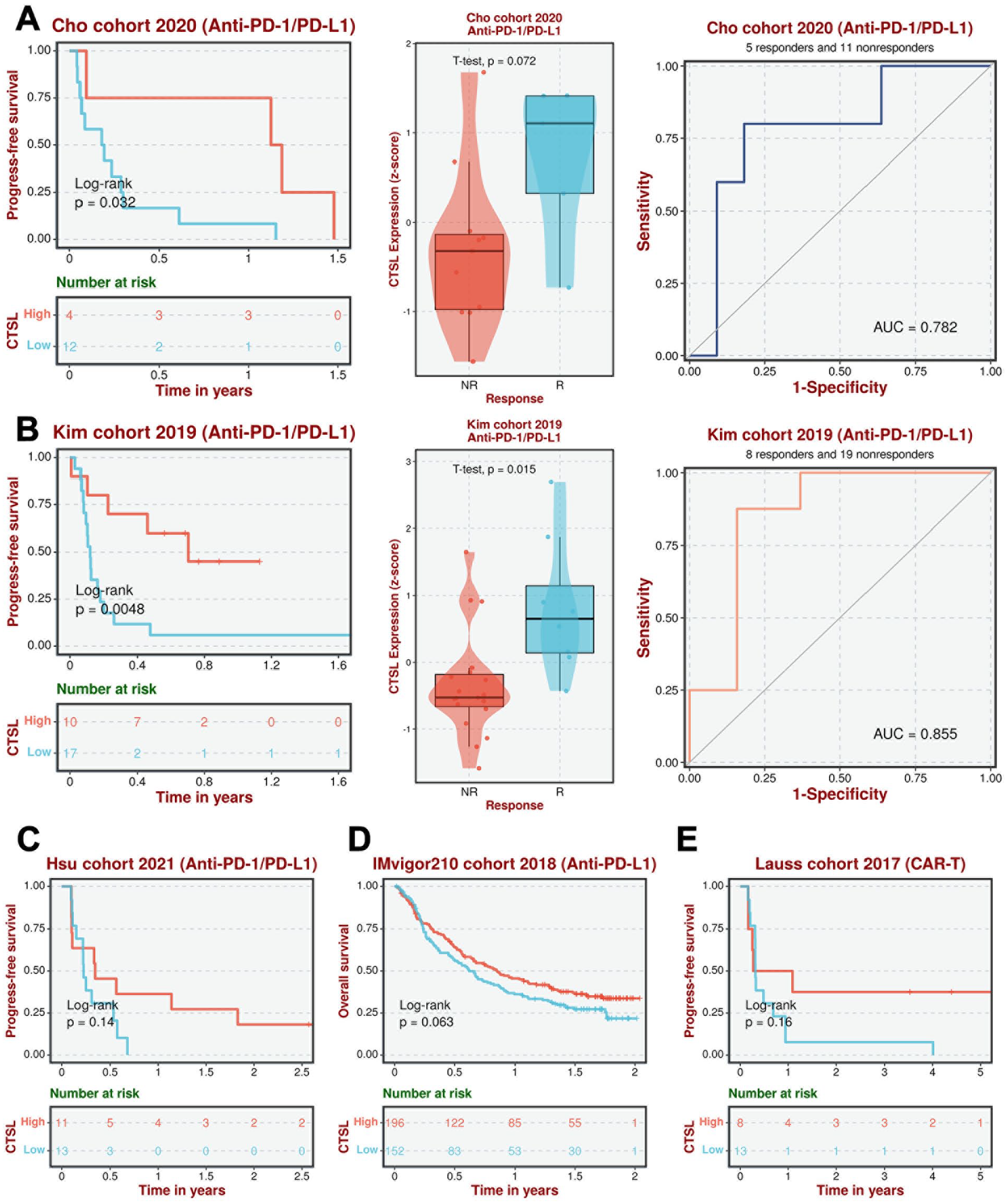
CTSL is a biomarker for therapy response in LUAD: (A and B) therapy response analysis of CTSL in LUAD anti-PD-1/PD-L1 cohorts and (C–E) therapy response analysis of CTSL in ICI cohorts.
Discussion
Lung cancer continues to be the leading cause of cancer-related deaths worldwide, despite significant advancements in diagnosis and treatment modalities.23,24 LUAD, the most common form of lung cancer, is characterized by substantial morphological heterogeneity and comprises tumor cells across a broad spectrum of histological subtypes.25,26 Through comprehensive bioinformatics analysis of big data, we identified candidate genes correlated with poor prognosis in patients with LUAD. This approach enables the prediction of target genes that may play crucial roles in the molecular progression of lung cancer, thus guiding the implementation of gene therapy in clinical settings. Capitalizing on these extensive data repositories, our study identified CTSL as a promising biomarker for LUAD.
CTSL, a ubiquitously expressed lysosomal endopeptidase, functions primarily in the terminal degradation of intracellular and endocytosed proteins.24,25 Recent studies have highlighted the role of CTSL in the entry mechanism of SARS-CoV-2 into host cells, in which it cleaves the receptor-bound viral spike protein, facilitating viral activation and infection. 27 Furthermore, in the realm of cancer research, CTSL has been the subject of several studies. Its upregulation has been observed in various cancers, including ovarian, breast, prostate, lung, gastric, pancreatic, and colon cancers.10,11,14,28 -30 Notably, increasing evidence suggests a correlation between CTSL expression and cancer grade and stage. Its potential role in advancing tumor progression and metastatic aggressiveness has also been proposed. Previous research has demonstrated a significant association between increased CTSL expression and reduced OS in patients with lung cancer and its contribution to chemotherapy resistance in vitro.12,31 Additionally, during tumor immunotherapy, elevated CTSL expression has been identified as an indicator of poor response. 15 However, these findings have offered a somewhat one-dimensional view of CTSL in LUAD, lacking comprehensive validation through multiomics, multicenter, and local cohort validation. Our current research endeavors to fill this gap, providing a more holistic understanding of the role of CTSL in LUAD.
Our study revealed that CTSL serves as a detrimental prognostic factor not only in the TCGA-LUAD cohort, which predominantly consists of American patients, but also in Japanese, Canadian, and our local Chinese cohorts. This uniformity in the role of CTSL across various ethnic backgrounds highlights its potential as a universal biomarker for LUAD prognosis. Interestingly, while CTSL serves as a negative prognostic factor in surgical intervention cohorts, it acts as a protective factor in immunotherapy cohorts. These results corroborate those of previous studies. This study revealed that CTSL is especially differentially expressed in myeloid cells. CTSL-related genes are involved mostly in complement and coagulation cascades, immune response activation, cytokine-mediated signaling pathways, etc., according to functional analysis. Moreover, the CTSL high-expression group had higher StromalScore, ImmuneScore, and ESTIMATEScore, indicating enrichment of immune components and increased immune activity in the population. 15 The interaction between PD-1 and PD-L1 leads to the suppression of T cells, causing tumor immune escape, which can be blocked by anti-PD-1/PD-L-1. 32 One hypothesis is that CTSL enhances the immune response, which is equivalent to enhancing the function of anti-PD-1/PD-L-1. Moreover, high-CTSL patients tend to have a greater tumor mutation burden (TMB), and a previous study suggested that patients with a higher TMB might respond better to anti-PD1/PD-L1 therapy. 33 This observation suggests that, at an early stage, high CTSL expression predicts worse outcomes than low CTSL expression does; however, patients with high CTSL expression postsurgery might benefit more from prioritizing immunotherapy. This discovery could facilitate the development of more personalized treatment strategies for patients.
Moreover, CTSL also emerged as an indicator of poor prognosis in a range of solid tumors in our pancancer analysis. Acting as a cysteine protease, CTSL might be involved in a series of common pathways that are not confined to LUAD. We further explored the expression pattern of CTSL among various cancer types. Next, we explored the influence of CTSL on tumor cells. The knockout of CTSL not only affects lung cancer cell proliferation but also inhibits the growth of various cancer cell lines within the DepMap project. Remarkably, the dependency scores for the NCIH1573 LUAD cell line were among the highest. The functional enrichment analysis in our study suggested that the oncogenic effect of CTSL in LUAD may be attributed to the activation of pathways such as angiogenesis, Hedgehog signaling, TGF-beta signaling, epithelial‒mesenchymal transition, and hypoxia. These pathways reveal that CTSL might affect cell proliferation, invasion and metastasis. The generation of blood vessels is also involved in tumor growth. In tumor cells, CTSL can be largely produced and augment the invasive/metastatic potential of cancer cells through direct degradative proteolysis of several components of the extracellular matrix (ECM) and basement membrane. In the nucleus, CTSL-processed CCAAT-displacement protein/cut homeobox (CDP/Cux) exhibits enhanced DNA-binding properties, which in turn confers a replicative and metastatic advantage. CDP/Cux promotes proliferation by accelerating the S phase of the cell cycle. 13 These findings offer a comprehensive perspective on the role of CTSL in the context of LUAD.
Overall, we can view the comprehensive potential therapeutic implications of targeting CTSL in lung adenocarcinoma treatment. It is involved in several aspects of tumor development, such as invasion and proliferation, while it also acts in modulating the immune response and tumor microenvironment. Thus, breakthroughs in both tumor biology and targeted therapy are needed. All these potential therapeutic implications may propel personalized therapy to distinguish CTSL levels among patients.
Limitation
It is important to acknowledge some limitations in our current study. For example, the analysis results derived from DepMap lack validation through in vivo and in vitro assays. Additionally, elucidation of the underlying mechanisms of CTSL in LUAD warrants further investigation in future research endeavors. In the future, further research on the molecular biology and immune profile of CTSL is needed. We hope to validate our conclusions clinically based on a large-scale study.
Conclusion
Overall, our results align with and build upon those of previous studies, establishing a foundation for further research to explore the broader implications of CTSL in different cancer types.
Supplemental Material
sj-xlsx-1-cix-10.1177_11769351241307492 – Supplemental material for Cathepsin L in Lung Adenocarcinoma: Prognostic Significance and Immunotherapy Response Through a Multi Omics Perspective
Supplemental material, sj-xlsx-1-cix-10.1177_11769351241307492 for Cathepsin L in Lung Adenocarcinoma: Prognostic Significance and Immunotherapy Response Through a Multi Omics Perspective by Jianming Lu, Jiaqi Liang, Gang Xiao, Zitao He, Guifang Yu, Le Zhang, Chao Cai, Gao Yi and Jianjiang Xie in Cancer Informatics
Footnotes
Acknowledgements
Abbreviations
AUC area under the curve
CDP CCAAT-displacement protein
CESC cervical squamous cell carcinoma and endocervical adenocarcinoma
CNA copy number alteration
CNV copy number variation
COAD colon adenocarcinoma
CRISPR clustered regularly interspaced short palindromic repeats
CTSL Cathepsin L
Cux cut homeobox
DAB diaminobenzidine tetrahydrochloride
DepMap dependency map
DFS disease free survival
DLBC diffuse large B-cell lymphoma
ESCA esophageal carcinoma
GBM glioblastoma multiforme
GISTIC Genomic identification of significant targets in cancer
HNSC head and neck squamous cell carcinoma
HR hazard ratio
HRP horseradish peroxidase temperature
ICB immune checkpoint blockade
ICI immune checkpoint inhibitor
IHC immunohistochemistry
KICH kidney chromophobe
KIRC kidney renal clear cell carcinoma
KIRP kidney renal papillary carcinoma
KM analysis Kaplan–Meier analysis
LGG lower grade glioma
LIHC liver hepatocellular carcinoma
LUAD lung adenocarcinoma
LUSC lung squamous cell carcinoma
OS overall survival
PFI progression-free interval
PFS progression-free survival
PRAD prostate adenocarcinoma
READ rectum adenocarcinoma
ROC receiver operating characteristic
SARC sarcoma
SKCM skin cutaneous melanoma
STAD stomach adenocarcinoma
TGF-β transforming growth factor β
TMA tissue microarray
TMB tumor mutation burden
UVM uveal melanoma
Author Contributions
Chao Cai, Jianjiang Xie and Gao Yi were instrumental in formulating the concept and design of this study. The task of gathering and analyzing the public datasets was carried out by Jianming Lu, Le Zhang, Jiaqi Liang, and Guifang Yu. The draft of the manuscript was prepared by Jianming Lu, Jiaqi Liang, and Gang Xiao. Additionally, the immunohistochemistry work was adeptly completed by Jianming Lu, Zitao He and Jiaqi Liang. All authors made significant contributions to the development of the article and have given their approval for the final version to be submitted.
Availability of Data and Materials
The datasets analyzed during the current study are available in public databases. This RNA sequencing (RNA-seq) data, mutation details, and clinical information for Lung Adenocarcinoma (TCGA-LUAD) and pan-cancer cohorts, sourced from the UCSC Xena database (https://xenabrowser.net/datapages/). To further substantiate our findings, LUAD datasets from the GEO database (![]() ), namely GSE50081, GSE31210, and GSE68465, were collected. For any further inquiries, please contact the corresponding author.
), namely GSE50081, GSE31210, and GSE68465, were collected. For any further inquiries, please contact the corresponding author.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The article is supported by Guangdong Basic and Applied Basic Research Foundation (2020A1515011290) and the Guangzhou Municipal Science and Technology Project (202201010053, 202201010787).
Ethics Approval
LUAD tissue microarray (TMA) were obtained from Shanghai Zhuoli Biotech Company. The acquisition of these samples was approved by the Ethics Committee of Shanghai Zhuoli Biotech Company (approval number SHLLS-BA-22101102), with written informed consent acquired from each participant for the use of their samples in this study.
Consent for Participate
The written informed consent has been acquired from each participant.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
