Abstract
Background:
Previous studies have shown that the 5-year survival rates of patients with nasopharyngeal carcinoma (NPC) were still not ideal despite great improvement in NPC treatments. To achieve individualized treatment of NPC, we have been looking for novel models to predict the prognosis of patients with NPC. The objective of this study was to use a novel deep learning network structural model to predict the prognosis of patients with NPC and to compare it with the traditional PET-CT model combining metabolic parameters and clinical factors.
Methods:
A total of 173 patients were admitted to 2 institutions between July 2014 and April 2020 for the retrospective study; each received a PET-CT scan before treatment. The least absolute shrinkage and selection operator (LASSO) was employed to select some features, including SUVpeak-P, T3, age, stage II, MTV-P, N1, stage III and pathological type, which were associated with overall survival (OS) of patients. We constructed 2 survival prediction models: an improved optimized adaptive multimodal task (a 3D Coordinate Attention Convolutional Autoencoder and an uncertainty-based jointly Optimizing Cox Model, CACA-UOCM for short) and a clinical model. The predictive power of these models was assessed using the Harrell Consistency Index (C index). Overall survival of patients with NPC was compared by Kaplan–Meier and Log-rank tests.
Results:
The results showed that CACA-UOCM model could estimate OS (C index, 0.779 for training, 0.774 for validation, and 0.819 for testing) and divide patients into low and high mortality risk groups, which were significantly associated with OS (P < .001). However, the C-index of the model based only on clinical variables was only 0.42.
Conclusions:
The deep learning network model based on 18F-FDG PET/CT can serve as a reliable and powerful predictive tool for NPC and provide therapeutic strategies for individual treatment.
Background
One of the most frequent malignant head and neck tumors is nasopharyngeal carcinoma (NPC), particularly in Southeast Asia and China. 1 Radiotherapy is the main treatment for stage I-II NPC, and concurrent chemoradiotherapy is used for stage III-IV NPC. The 5-year survival rate is about 80%. 2 Nevertheless, the latest revised AJCC 8th edition staging for NPC is far from optimal, because clinical outcomes from the same treatment vary widely even among patients with tumor of identical TNM stage and treatment strategies. Therefore, predictors other than the TNM stage are needed to predict the prognosis. Some quantitative parameters derived from PET/CT imaging such as maximum standard uptake value (SUVmax), peak SUV (SUVpeak), metabolic tumor volume (MTV), and total lesion glycolysis (TLG), showed prognostic potential. For example, SUVmax of primary tumors is an effective biomarker for predicting overall survival (OS) and event-free survival (EFS) in patients of NPC. However, Chang et al 3 showed different results. They argue that SUVmax is a limited parameter and does not represent the whole tumor. Many studies have shown that a higher MTV or TLG is associated with a higher risk of adverse events or survival. However, Shi et al 4 proved that MTV or TLG could not predict EFS or OS, suggesting that traditional 18F-FDG PET-CT metabolic parameters are controversial for predicting the prognosis of patients with NPC. The reasons of the variation in findings were due to different quantitative characteristics of patients, the threshold of traditional 18F-FDG PET-CT metabolic parameters, and the failure of traditional metabolic parameters to fully describe the heterogeneity of tumors. As can be seen, a survival prediction model for NPC patients can be created using the metabolic parameters of conventional 18F-FDG PET-CT, but its predictive power is constrained. As a result, we require a new indicator or model to more accurately predict the prognosis of patients with NPC.
Deep learning is used to discover the internal structures and levels of representation of sample data. Its ultimate goal is for machines to have the analytical learning capabilities of humans and to recognize data such as text, images, and sound. New evidence suggests that deep learning-based information extracted from medical images has made commensurate progress in predicting survival outcomes.5-8 Particularly, convolutional neural networks (CNNs) have been applied to many survivals prediction studies and have achieved good performance.9-12 Zhu et al 13 developed CNN to predict the OS of lung cancer patients and conducted model training on pathological images of lung cancer, with a consistency index of 0.63. Byun et al 14 constructed the random survival forest model (RSF), DeepSurv model, and Cox proportional hazards model (CPH) to predict RFS and tumor-specific survival (CSS) in patients with nonmetastatic clear cell RCC. Harrel’s C-indices of RFS and CSS in the test dataset were 0.794, 0.789, 0.802 and 0.831, 0.790, and 0.834 for CPH, RSF, and DeepSurv, respectively. Therefore, DeepSurv is superior in predicting RFS and CSS in non-metastatic clear cell RCC patients compare with CPH and RSF. Noteworthy, deep learning technology has been maturely implemented in the automated identification of structures and lesions on CT or magnetic resonance (MR) images in many cancers including NPC. 15 Recent developments in machine learning and artificial intelligence have shown the potential of 18FDG-PET/CT to improve the accuracy of reading anatomical and metabolic characterization of malignancies.16,17 However, the majority of existing studies about 18FDG-PET/CT have focused on diagnosis and staging, even though some research studies aimed at prognostication have mostly relied on manual image segmentation rather than deep learning.18,19
To overcome the error of manual image segmentation and the difference of threshold defined by traditional metabolic parameters, the objective of the study was to establish a prognostic model of the deep learning (CACA-UOCM) network structure to predict the prognosis of patients with NPC. Furthermore, its predictive value was compared with a traditional prognostic model of clinical parameters.
Methods
Patients
A total of 173 patients with NPC admitted to Shandong Cancer Hospital and Weifang People’s Hospital from July 2014 to April 2020 were collected. Inclusion criteria: histologically confirmed NPC, pretreatment PET/CT data, and cervical lymph node metastasis. Exclusion criteria: distant metastasis before treatment; have other tumors; incomplete clinical data are not available. The protocol of this retrospective study was approved by the Ethics Committee of Shandong Institute of Cancer Prevention and Treatment (2019GGX101057) and Weifang People’s Hospital (2019034).
PET/CT acquisition
All patients fasted for at least 8 hours before receiving an 18F-FDG PET-CT scan and checking blood glucose levels. The scans were performed using 2 sets of advanced PET/CT scanners. Shandong Cancer Hospital was equipped with a Discovery Lightspeed PET-CT and a Minitrace cyclotron. Its spiral CT component has a peak voltage of 140 kV, 80 mA, a pitch of 2:1, a thickness of 4.25 mm, and a speed of 0.8s/r. 18F-FDG MBq (5.55-7.40/kg) was injected intravenously with a purity of >95%, and radioactive drugs were used. Two-dimensional pattern images were collected on PET/CT on Discovery LS 1 hour later. The other one is the introduction of BioGRAPH-64-True Point PET/CT produced by SIEMENS in Germany for the PET-CT Center of Weifang People’s Hospital. This device combines excellent PET and 64 spiral CT, 2 mature medical imaging technologies. This design, which is equipped with a unique Siemens high-resolution design, doubles the detection signal-to-noise ratio, significantly reducing image blur and distortion, giving it a consistent view spatial resolution of 2 mm, and pushing the limits of its ability to detect small lesions.
PET/CT Analysis
Gross tumor volume (GTV-P) and lymph node (GTV-N) of the primary tumor were manually segmented on each PET/CT by the same experienced investigator (radiation oncologist), referring to 2 nuclear radiologists’ reports. Region of interest (ROI) was drawn on the primary tumor and metastatic cervical lymph nodes. All patients’ PET and CT images were imported into the MEDEX workstation (Philips Health Care), and the ROI of the lesions was calculated manually using the 40% SUVmax threshold. The metabolic and volume parameters SUVmax, SUVpeak, MTV, and TLG were measured by 3-dimensional measurement software.
Treatment and Follow-up Method
All patients were treated with 1.8 to 2.0 Gy per fraction with 5 daily fractions per week for 6 to 7 weeks. The cumulative radiation dose of the primary tumor was 50 to 76 Gy, and the cumulative dose of bilateral lymph node irradiation was 50 to 70 Gy. The chemotherapy regimen was mainly platinum therapy. Follow-up included head and neck examination, nasopharyngeal endoscopy, chest X-ray examination, abdominal B-ultrasound examination, and whole-body bone scan. When recurrence or metastasis is found, re-staging and treatment plans should be made. Regular follow-up was performed every 3 months for the first 3 years, every 6 months for the next 3 years, and then annually thereafter.
Model Based on Clinical Factor and Conventional PET-CT Metabolic Parameters
A training set of patients and a test set of patients were separated. Adasyn algorithm was used to over-sample the training data so that the ratio of surviving patients and dead patients was 1:1, and the test data was the original data. Preliminary model construction included 6 clinical characteristics (gender, age, T stage, N stage, AJCC stage, pathological type) and 8 conventional PET-CT metabolic parameters (SUVmax-P, SUVpeak-P, MTV-P, TLG-P, SUVmax-N, SUVpeak-N, VMTV N, TLG-N). Figure 1A shows the coefficient estimates for various hyperparameters alpha in the training queue. With C-index as the evaluation standard, the optimal hyperparameter alpha was selected after 5 times of cross verification, as shown in Figure 1B. The orange line segment was the hyperparameter corresponding to the optimal consistency score, and the Lasso coefficient corresponding to the selected variable was as shown in Figure 2. Figure 3 shows the estimated results for the horizontal risk coefficients for each parameter, and Table 1 shows the coefficients and P values for each parameter. Cox regression fitting was performed again for the selected variables. Variables were removed from the model if P > .1, and multivariate Cox analysis was used to identify the significant prognostic model parameters and standardized coefficients.
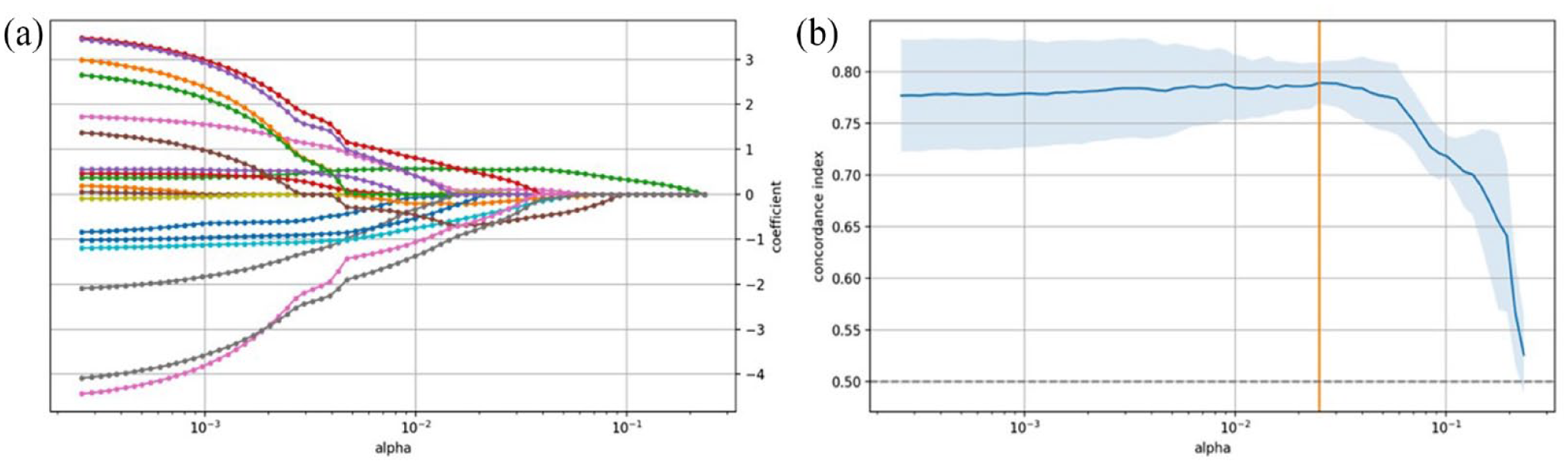
In the training cohort, given different hyperparameter alpha, the coefficient estimates are shown in A. With C index as the evaluation standard, the optimal hyperparameter alpha is selected after 5 times of cross-validation, as shown in B.

The orange line segment was the hyperparameter corresponding to the optimal consistency score and the Lasso coefficient corresponding to the selected variable.

The estimation results of the level risk coefficient of traditional PET-CT metabolic parameters.
Coefficient estimates.

The value and P value of risk coefficient of each parameter level are shown in Table 3.
Abbreviations: Staging_1 = Clinical stage II, Staging_2 = Clinical stage III.
Model Based on Deep Learning Network
Patients were enrolled and randomly assigned to test, validation, and training groups. CACA-UOCM takes CT clip-out according to the ROI area as the input. First, input the down-sample layer in the network for 2 consecutive 3D convolutions and normalized activation, and then enter the convolution layer for maximum pooling and repeat the operation of the down-sample layer. After the 3D coordinate attention is input by the autoencoder, the size of the features in the input directions of the other 2 dimensions is first 1 by the 3 average attention autoencoders, and its size remains unchanged. Then the outputs of the 3 directions of X, Y, and Z are concatenated and entered into the 1D convolutional layer. It is divided into distinct features in 3 directions for convolution and activation after the activation function. The 3D coordinate attention encoder and output features from the original input are multiplied by the output features to complete the process. The data output from the encoder is fed into a decoder where the feature graph is up-sampled using 2×2×2 trilinear interpolation to match the size of the previous block. With the addition of the coordinate attention mechanism, the decoder is down-sampled using convolution, just like the down-sampling layer. The final output of the decoder is a reconstructed image of the same size as the input image. Extract the data output from the encoder, perform the adaptive average pooling operation, and aggregate the input information into the 1-dimensional feature vector marked as the intermediate feature as the input of the survival prediction task (the network structure and flow chart are shown in Figure 4).
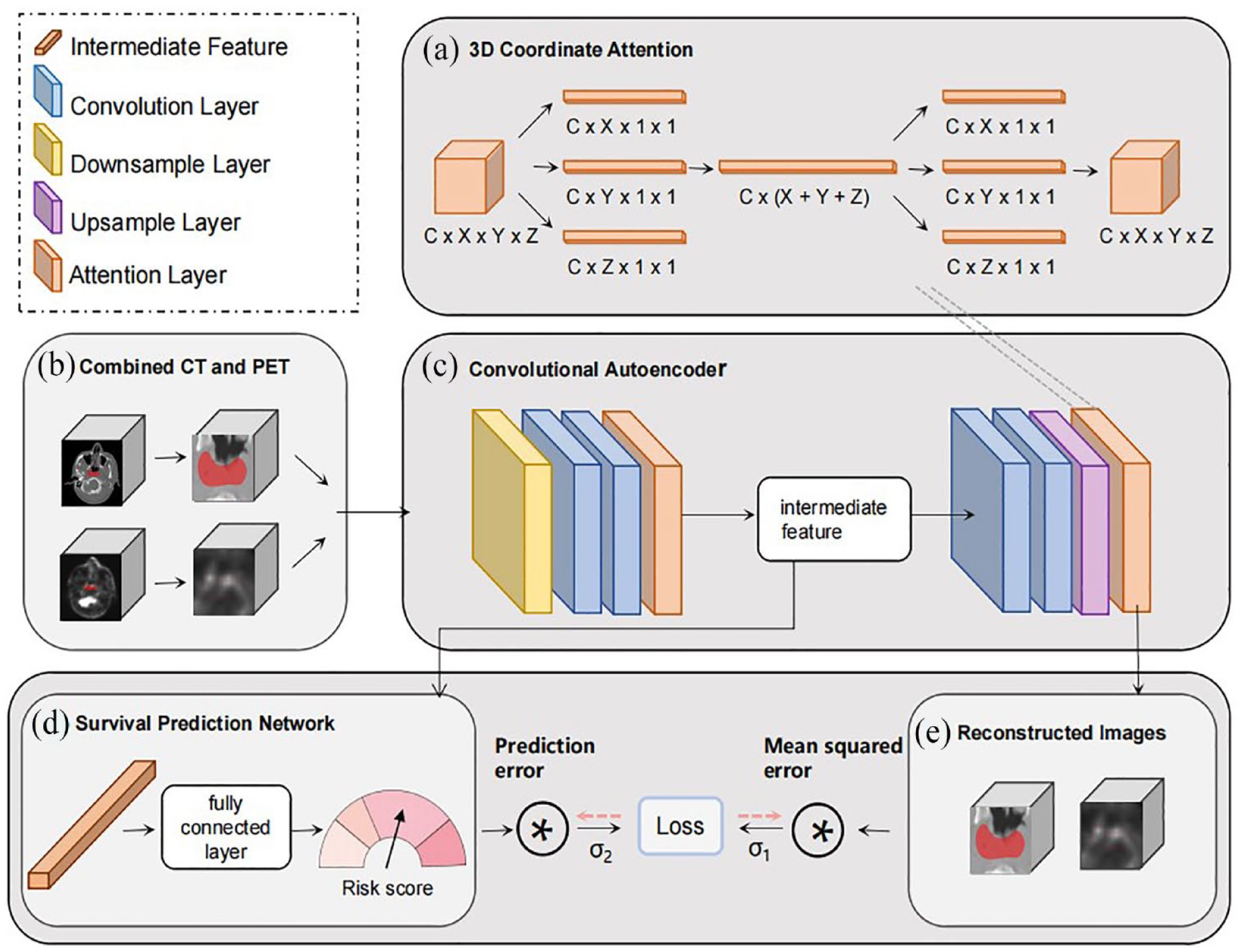
The network structure and flow chart.
Statistical Analysis
OS is defined as the time between patient’s pathological diagnosis and the patient’s death from any cause. Kaplan–Meier was used for the statistical description of survival data, and different groups were plotted separately, and a log-rank nonparametric test was performed to compare whether survival curves of different groups were the same. Cox proportional risk regression model was used to estimate survival function, establish a direct relationship model between influencing factors and survival time, and calculate the risk ratio of influencing factors. The degree of conformity between model prediction results and actual observation results is measured using the C index, which assesses the accuracy of the model prediction. Built on the Torchtuples package used to train the PyTorch model. The optimizer uses Adam + Warm Restart, and batchSize is 32.
Missing values were first filled in during the preprocessing of the data, followed by the mean value filling in for continuous variables, the mode filling in for classified variables, and the classified variables being set as dummy variables. After removing outliers, the mean values of normalized continuous variables were divided by their variance. Variables that had correlation coefficients of more than 0.9 were eliminated after calculating the correlation coefficients between them. The results use averages and ranges to represent quantitative variables, while numbers are used to classify the results. Python is used to implement the model’s predictive values, and code is used to calculate the conformance index. The critical level of 5% considered the results to be statistically significant (P < .05).
Results
Patient characteristic
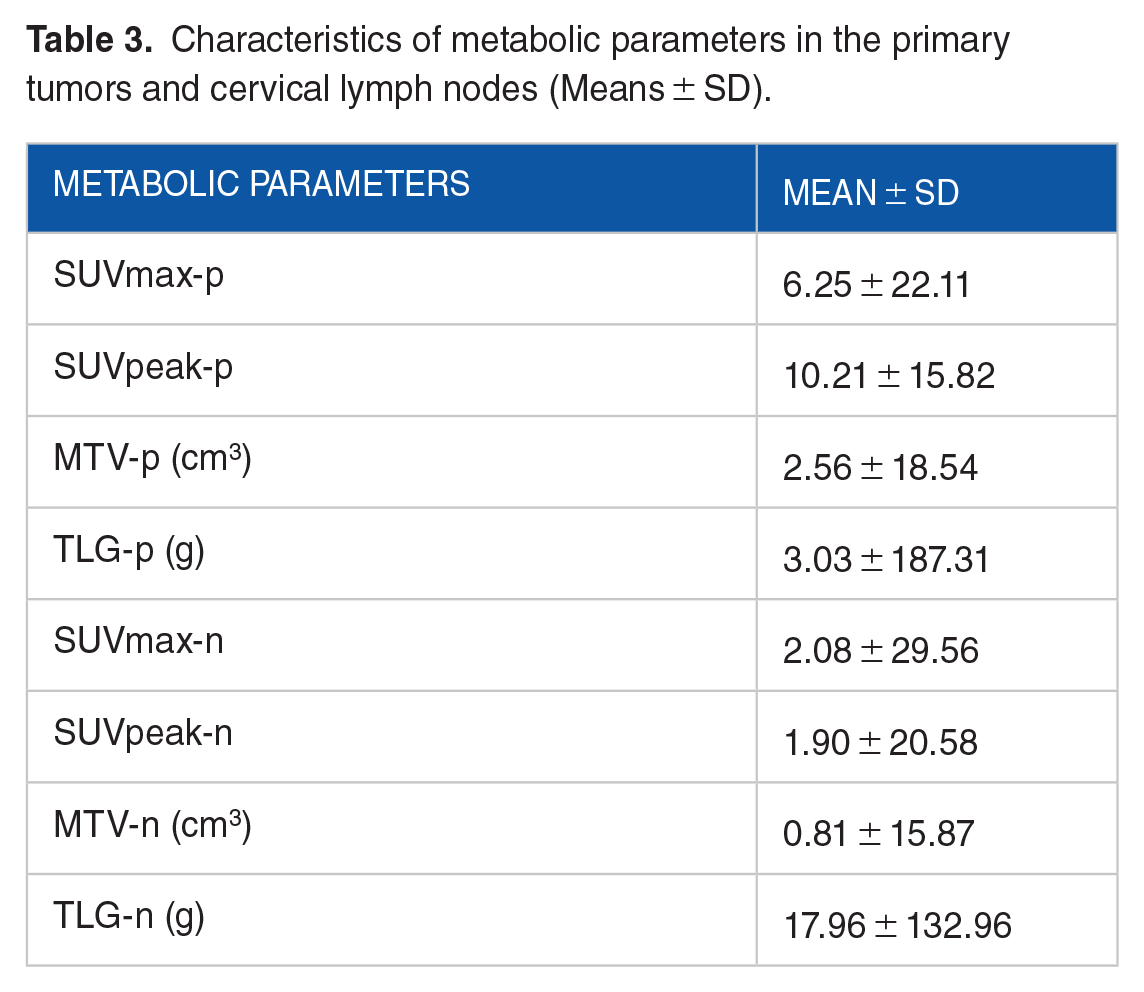
In the study, patients comprised 124 males (71.7%) and 49 females (28.3%); 37 patients were ⩾60 years old, and 136 patients were <60 years old. The median follow-up time was 31.4 months (range 0.5-115.2 months). A total of 18 deaths (10.4%) occurred in the follow-up period. The mean OS was 38.8 months (95% CI: 35.0, 42.9; Figure 5). Table 2 lists the patient’s characteristics. The mean value of SUV max-P was 14.18 cm3 (2.01, 48.14), and SUVmax-N was 15.82 cm3 (1.65, 94.51). The patient’s baseline characteristics and PET-CT traditional metabolic parameters of the primary site and cervical lymph node range are summarized in Tables 2 and 3.

Function curve of overall survival.
Patient characteristics.

Characteristics of metabolic parameters in the primary tumors and cervical lymph nodes (Means
Prediction of OS using models based on clinical factors and conventional 18F-FDG PET-CT metabolic parameters
One hundred and seventy-three patients with standard according to the proportion of 4:1 was divided into a training set (138 people) and a test set, 20 in the training set, a total of 125 patients survived, with 13 cases of death.
The significant parameters retained after re-fitting in COX analysis are shown in Figure 6. The coefficients of SUVpeak-P, T-3, Staging-1, age, MTV-P, N-1, Staging-2, and pathology were −0.44, 1.04, −1.91, −1.15, 0.48, 0.91, −2.11, and −0.78, respectively, with P ⩽ .05 (Staging -1 represents stage II and staging-2 represents stage III). Table 4 shows the specific values. The C index of the test is 0.42. In addition, the mean risk coefficient predicted by the training cohort model (0.5) was divided into a survival group (low-risk group) and a death group (high-risk group). Death (high risk, risk score > 0.5) and survival (low risk, risk score ⩽ 0.5) were represented by the above and below means, respectively. Kaplan–Meier curves showed no statistical difference between high-risk and low-risk groups (Figure 7A, P = .098). The survival model based on clinical factors and conventional 18F-FDG PET-CT metabolic parameters has no significant clinical value in predicting the survival status of NPC patients.
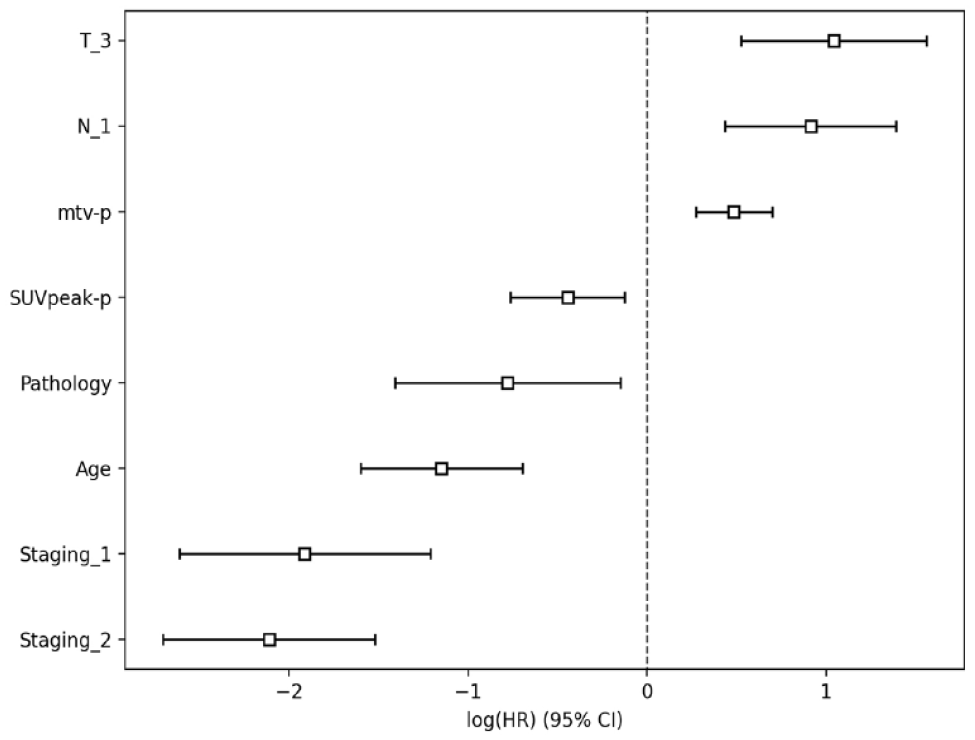
The important parameters retained after re-fitting in COX analysis.

The Kaplan–Meier curves of the high-low risk group grouped by clinical model were not statistically significant (A, P = .098). Kaplan–Meier curves of the test cohort and validation cohort grouped by the deep network model showed statistical significance (B, P = .0041; C, P = .0024).
Coefficient estimates.

The selected variables were fitted by Cox regression again. If P > .1, the variables were deleted from the model, and multivariate Cox analysis was used to determine the significant parameters and standardized coefficients of the prognostic model. The significant coefficients and P values retained after re-fitting in COX analysis are shown in Table 3.
Abbreviations: Staging_1 = Clinical stage II, Staging_2 = Clinical stage III.
Prediction of OS using models based on deep learning network model
A total of 173 patients with cervical lymph node metastasis were enrolled and randomly divided into 35 test sets, 35 validation sets, and 103 training sets. CACA-UOCM takes CT clip-out according to the ROI area as the input.
Based on CACA-UOCM, we changed the input from 1-channel 3D CT image to 3-channel 3D input, with each channel being CT, PET, and graphing Mask respectively. The final output of the decoder was a 3-channel reconstructed image consistent with the number of input channels so that the 3D coordinate attention convolution encoder could adapt multi-mode image data. In OS, the C index of training cohort (n = 103) was 0.779 (0.669, 0.859), 0.819 (0.425, 0.978), and 0.774 (0.550, 0.956). The CACA-UOCM network structure in Table 5 has obtained the test queue (n = 35) and verification queue (n = 35) respectively. Additionally, based on the mean value of the risk coefficient predicted by the model, the test cohort and training cohort was split into high-risk and low-risk groups (0.5). The mean value of the high-risk group was greater than (risk score > 0.5), and the mean value of the low-risk group was less than (risk score ⩽ 0.5). The log-rank test yielded p values of 0.0041 and 0.0024 (P < .05), respectively. It can be seen that the difference in survival risk between the high-risk and low-risk groups of NPC patients is statistically significant (Figure 7B and C).
Five-fold cross-validation performance prediction model.

Discussion
Instead of using the image segmentation method, we used GTV reconstruction to train our model, which retains the original high-quality intensity values and facilitates better image information acquisition.
Accurate prediction of prognosis is key to risk stratification and management of NPC patients. To develop and validate the prognostic value of deep learning network structure for NPC, we carried out this study. Our results show that the CACA-UOCM network structure has better predictive value than the current TNM staging system, traditional 18F-FDG PET-CT metabolic parameters, and age. The study outcomes suggest that it may someday serve as a novel and practical tool for the diagnosis and treatment of NPC.
Medical images can be used to characterize tumor heterogeneity, characterize a large amount of noninvasive information, and assess the prognosis of cancer patients.21-23 In particular, the field of “radiomics,” which extracts texture features from medical images, has been widely used in the prognosis assessment of cancer patients.22,24-30 However, due to the lack of unified image acquisition and imaging algorithm standards in medical imaging equipment, features based on gray values such as histogram and texture analysis are affected. At the same time, the premise of feature extraction is the accurate delineation of the target volume. For tumors with fuzzy boundaries, subjective influence is greater, which reduces the stability of features based on size, shape, and boundary. 25 In-depth research and analysis-based artificial intelligence are currently being actively investigated as a prognostic tool in tumor radiotherapy to predict clinical outcomes.7,15,20,31-37
The survival of esophageal cancer has been successfully predicted using this network. 15 First, CACA is built upon an autoencoder structure with 3D coordinate attention layers, capturing deep and potential information from medical images. The advantages of down-sampling and up-sampling make our model focus on the target region and improve its expressiveness. Second, a joint optimization Cox model based on uncertainty was designed, which carried out the joint optimization survival prediction task for CACA. Cox proportional hazard regression will oversee the task of predicting survival to model the interaction between patient characteristics and clinical outcomes and to predict a patient’s credible hazard ratio. In UOCM, we design a specific loss function based on the relative weight of uncertainty in the data to improve the performance of the survival prediction network structure. Experimental results show that the consistency index of this method is high, and the network structure is conducive to the construction of a model to predict the survival of NPC patients. In addition, the test set data were divided into a death group (high-risk group) and a survival group (low-risk group) according to the mean risk coefficient predicted by the model. The P values of the survival curves of the test cohort and training cohort were all less than .05. It can be seen that the CACA-UOCM network structure is conducive to the classification of NPC patients into high-risk and low-risk groups and the formulation of treatment plans.
We found that the prognostic factors of age, stage, pathological type, SUV peak-P, and MTV-P were significant, whereas the traditional 18F-FDG PET-CT metabolic parameters of cervical metastatic lymph nodes were not significant for predicting the survival of NPC patients. Ab Hamid Siti-Azrin et al 38 found that the vital factors that changed survival rate and time were age (P = .041) and stage (P = .002), which was consistent with our research results. However, compared with SUV peak, SUVmax, TLG, and MTV survival prediction are more valuable in patients with NPC. 39 Our study also confirmed the important value of MTV-P (P < .005) in the survival of NPC patients. Huang Yecai et al 40 suggested that SUVmax, MTV, and TLG (SUV fixed value 2.5) before primary tumor treatment may be independent factors affecting the prognosis of patients with NPC. Metastatic lymph nodes’ SUVmax, MTV, and TLG did not change significantly. This is consistent with our conclusion that the metabolic parameters of traditional 18F-FDG PET-CT cervical metastatic lymph nodes are not significant for predicting survival in patients with NPC. Our study suggests that clinical data and routine 18F-FDG PET-CT metabolic parameters alone are insufficient to predict survival model conformance indicators in patients with NPC. Models that can predict survival are still limited. Recently, Zhao et al 41 studied 420 patients with rNPC who underwent PET/CT imaging and followed up overall survival (OS). They constructed multi-modality deep learning signatures from PET and CT images with a light-weighted deep convolutional neural network EfficienetNet-lite0 and survival loss DeepSurvLoss. It is concluded that PET-CT-based deep learning features show satisfactory prognostic performance in rNPC patients. Therefore, the CACA-UOCM network structure established has more predictive value than the clinical data model for survival prediction of patients with NPC.
Our study has limitations. First, it was a retrospective study with small sample size, requiring prospective, multicenter validation. Second, data from 2 different hospitals as well as data from various scanners at various facilities may differ. Third, we only used PET-CT before treatment for imaging. In the future, OS predictions can be further updated by combining treatment and posttreatment information.
Conclusion
Based on the above studies, we established 2 prognostic survival models, one based on clinical factor and conventional 18F-FDG PET-CT metabolic parameters model, and the other based on deep learning network structure model. The C-index of clinical combined with traditional 18F-FDG PET-CT metabolic parameters was 0.42, and the C-index of the deep network model training group was 0.779, demonstrating that the structure of the deep learning network based on 18F-FDG PET-CT is a reliable and powerful prognostic tool, and can be used as an indicator to guide the personalized treatment of NPC.
Footnotes
Acknowledgements
We thank the patients who participated in this study, their families, and the staff members at the study sites who cared for them.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the Shandong Natural Science Foundation (ZR2021LSW002).
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MH conceived this project. ZCL, FRH, and ZF collected and prepared the samples. ZCL and SY collected the data. ZCL and ZRL analyzed and interpreted the data. ZCL, XCD, and XBZ wrote the manuscript. All authors read and approved the final manuscript.
Ethics Approval and Consent to Participate
The study complied with the principles set forth in the Declaration of Helsinki. It was approved by the Ethics Committee of Shandong Institute of Cancer Prevention and Treatment (2019GGX101057) and Weifang People’s Hospital (2019034). Written informed consent was obtained from each patient.
Consent for Publication
Acknowledgements Informed consent was obtained from all patients for being included in the study.
Availability of Data and Materials
The data is available from the corresponding author upon reasonable request.
