Abstract
Aim:
This study aimed to characterize ESBL-producing E. coli isolates obtained from apparently healthy captive wild birds from selected zoological gardens in Nigeria.
Materials and Methods:
A total of 121 freshly deposited faecal samples were collected from captive wild birds in seven major zoological gardens and pooled into 50 samples. The samples were screened for the presence of E. coli. The isolates obtained were tested against a panel of antibiotics and screened for ESBL production using the double disc synergy test (DDST). Primer-specific PCR was used to detect the carriage of ESBL genes (blaCTX-M, blaTEM and blaSHV) by the isolates.
Results:
A total of 26 ESBL-producing E. coli isolates (52%, n = 26/50) were obtained from the pooled faecal samples of captive wild birds. The highest resistance rate to antibiotics was observed with amoxicillin-clavulanate (88.5%), while the lowest resistance rate was observed with fosfomycin (3.8%). The isolates had Multiple Antibiotic Resistance Index (MARI) values ranging from 0.1 to 0.8 across the tested antibiotics. Approximately 65.4% of the isolates carried blaCTX-M, while blaTEM and blaSHV were detected in 15.4% and 34.6% of the isolates, respectively.
Conclusion:
This study revealed a significant prevalence of ESBL-producing E. coli in captive wild birds in Nigeria and highlighted the need to institute control measures in zoological gardens to prevent the transmission of antibiotic resistance.
Summary of Points
• A high prevalence of ESBL-producing E. coli (52%) was found in captive wild birds in Nigeria.
• A high resistance rate was found in amoxicillin-clavulanate and tetracycline.
• The frequency of ESBL genes detected was blaCTX-M (65.4%), blaSHV (34.6%) and blaTEM (15.4%).
• Adequate biosecurity measures are needed in zoological gardens and animal sanctuaries to prevent the transmission of AMR.
Introduction
Antimicrobial resistance (AMR) is a significant global health concern due to the widespread use of antibiotics.1 -3 Even though AMR is a global issue, the situation has become more catastrophic in developing countries, such as Nigeria, 4 due to unrestricted access to and indiscriminate use of antibiotics, a lack of national AMR surveillance programmes and antimicrobial regulation policies. 5 Multidrug-resistant bacteria, notably those resistant to β-lactam antibiotics, have further reduced the number of available therapeutic options. This contributes to treatment failure, extended hospital stays for patients and the need for additional antimicrobials, which are typically more costly and have more adverse effects on patients.6,7 Extended Spectrum β-Lactamase (ESBL)-producing bacteria have become more widespread worldwide due to the broad application of β-lactam antibiotics in both human and veterinary medicine, particularly in commensal Enterobacteriaceae.8,9
Escherichia coli is an opportunistic bacteria found in the intestines of mammals, including humans and animals. This bacterium is commonly exposed to antibiotics, which may contribute to its pathogenic potential. E. coli is considered a good bio-indicator for observational research in antibiotic resistance because it may readily acquire and transfer AMR genes.10 -12 Thus, research on the AMR of E. coli is currently ongoing for strains that are present in meat, the environment, and wild and companion animals.13 -16 Animals and humans can share antibiotic resistance genes (ARGs) and/or host microorganisms. Therefore, its control requires a coordinated approach based on One Health.
An increasing number of studies have revealed the essential role of wild animals, especially birds, as carriers and reservoirs of bacteria resistant to antibiotics.17,18 Alonso et al 19 and Poeta et al 20 revealed the presence of ESBL-producing Enterobacteriaceae in at least 80 species of wildlife that are primarily wild birds. E. coli is the most common bacteria identified among the Enterobacteriaceae family. However, relatively few reports from Africa exist.21,22 Zoological gardens may serve as human reservoirs for ESBL-producing bacteria and their genes, because of their popularity and capacity to house exotic animals sharing close proximity to people.23,24
Zoological gardens have recently evolved to concentrate on species conservation and animal welfare. However, some continue to prioritize other objectives, such as animal feeding, to engage visitors and manage the zoo. Zoo animals maintained in captivity can act as reservoirs for pathogenic bacteria resistant to antibiotics, including strains of pathogenic E. coli, 25 Mycobacterium spp., 26 Salmonella spp., 26 Yersinia spp. 26 and Campylobacter spp. 27 Research indicates that visitors to zoological gardens may be exposed to antibiotic-resistant E. coli strains carrying antibiotic resistance genes (ARGs). These environments are considered reservoirs for ARGs and sources of resistance transmission to humans and other animals. 28
In Nigeria, zoological gardens frequently house and nurture wild birds to attract visitors. Captive wild birds harbouring antibiotic-resistant E. coli significantly threaten human health and the environment. Zoological garden animals from Belgium, China, Japan, Israel and the Czech Republic have previously been shown to carry ESBL- and AmpC-producing Enterobacteriaceae in their faeces, with prevalence rates ranging from 11% to 32%.24,29 -31 To the best of our knowledge, very few studies have investigated ARG-mediated E. coli antibiotic resistance in captive wild birds housed in Nigerian zoological gardens, making it difficult to institute public measures via the One Health approach to control the spread of antimicrobial resistance. This study aimed to characterize ESBL-producing E. coli isolates obtained from apparently healthy captive wild birds from different zoological gardens in Nigeria.
Materials and Methods
Ethical Statement
The University of Ibadan Animal Care and Use Research Ethics Committee (UI-ACUREC/031-0423/13) gave ethical approval for this study. The directors of each zoological garden granted formal written and physical authorization for the collection of bird samples.
Sample Collection
The samples were collected from seven major zoological gardens and sanctuaries in Southwest Nigeria in May 2023. The selected zoological gardens and sanctuaries were Greenfingers Wildlife Conservation (GWC) and Omu Resort Zoo (OMU) located in Lagos State, Zoological Park, Federal University of Agriculture, Abeokuta (FUNAAB) and Olusegun Obasanjo Presidential Library Wildlife Park (OOPL), located in Ogun State, Zoological Garden, University of Ibadan (UI) located in Oyo State, Biological Garden, Obafemi Awolowo University (OAU), Osun State and Wildlife Park, Federal University of Technology, Akure (FUTA) located in Ondo State (Figure 1). The sampled birds are typically housed either individually or in groups. A total of 121 freshly deposited faecal (either swab or freshly voided) samples were collected. The samples were transported in a cold chain to the Environmental Microbiology and Biotechnology Laboratory, Department of Microbiology, University of Ibadan, Nigeria, for microbiological analysis. Table 1 shows the distribution of collected samples across the zoological gardens.

Geographical location of selected zoological gardens in Southwest Nigeria.
Distribution of samples collected in major zoological gardens in Southwest Nigeria.

Abbreviations: FUNAAB: Zoological Park, Federal University of Agriculture, Abeokuta; FUTA: Wildlife Park, Federal University of Technology, Akure; GWC: Greenfingers Wildlife Conservation; OAU: Biological Garden, Obafemi Awolowo University; OMU: Omu Resort Zoo; OOPL: Olusegun Obasanjo Presidential Library Wildlife Park; UI: Zoological Garden, University of Ibadan.
Isolation and Characterization of E. coli
The faecal samples (n = 121) were pooled into 50 samples. The samples from the same bird species and zoological garden were pooled and treated as one sample. This was done according to the method of Aho. 32 The samples were pooled in order to achieve high cost efficiency and a suitable statistical power for the prevalence estimation. Streak plate method was used to isolate E. coli from the pooled faecal samples. The tips of the swab sticks were cut aseptically, inoculated into the previously prepared nutrient broth (Oxoid, UK) and incubated overnight at 35°C ± 2°C. A loopful of the incubated broth was inoculated onto prepared MacConkey agar (Oxoid, UK) plates and incubated at 35°C ± 2°C for 24 hours. Lactose-fermenting colonies appearing pink on the media were selected and inoculated on Eosin Methylene Blue (Oxoid, UK). Colonies showing green-metallic sheen appearance on the plates presumptive of E. coli were picked, further subcultured on nutrient agar (Oxoid, UK) and stored on agar slants for further studies. Genomic DNA of suspected E. coli isolates was extracted using the boiling lysis method according to Janezic et al, 33 for the performance of a PCR aiming to detect uidA, a housekeeping gene in E. coli. The detection of the gene was performed according to the method of Janezic et al. 33
ESBL Phenotyping of Isolates and Antibiotic Susceptibility Testing
The double disc synergy test (DDST) was used for the phenotypic detection of ESBL production in the confirmed E. coli isolates. Isolates that phenotypically displayed ESBL production were selected for antibiotic susceptibility testing. The susceptibility of the isolates to a panel of nine selected antibiotics was assessed using the Kirby–Bauer disc diffusion technique. 34 The tested antibiotics were ciprofloxacin (30 µg), tetracycline (30 μg), amoxicillin-clavulanate (30 µg), trimethoprim-sulfamethoxazole (1.25/23.75 μg), gentamicin (10 μg), azithromycin (15 μg), chloramphenicol (30 µg), cefpodoxime (30 µg) and fosfomycin (200 µg). The choice of antibiotics and interpretation of the data were performed according to the CLSI34,35 guidelines. Organisms showing resistance to ≥ 3 classes of antibiotics were termed multidrug-resistant (MDR), according to the method of Magiorakos et al 36 .
Detection of ESBL Genes (blaCTX-M, blaTEM and blaSHV)
Multiplex PCR was used for the amplification of the blaSHV and blaTEM genes as described by Maynard et al 37 using the following PCR conditions: initial denaturation step at 94°C for 5 minutes, denaturation at 94°C for 30 seconds, primer annealing at 50°C for 30 seconds, extension at 72°C for 90 seconds (30 cycles) and terminal extension at 72°C for 10 minutes. The amplification of blaCTX-M was performed via monoplex PCR, as reported by Mendonça et al, 38 with the following conditions: initial denaturation step at 94°C for 5 minutes, denaturation at 94°C for 30 seconds, primer annealing at 56°C for 1 minute, extension at 72°C for 60 seconds (30 cycles) and terminal extension at 72°C for 10 minutes. Amplicons were resolved on 1% agarose gel electrophoresis. A multidrug-resistant E. coli ALC08 isolated from abattoir leachate carrying blaCTX-M, blaSHV and blaTEM was used as the positive control. 39
Multiple Antibiotic Resistance Index (MARI) Evaluation
The multiple antibiotic resistance index (MARI) was used to assess the proportion of antibiotics to which an organism demonstrates resistance among those tested. 40

where n = number of antibiotics to which the isolate showed resistance.
t = total number of antibiotics tested against the isolate.
Statistical Analysis
The data were sorted and analysed via Microsoft Excel. A descriptive analysis was performed on the data presented in Figures and Tables. Exploratory data analysis was performed on R to understand outlier and distribution discrepancies in our data. The frequency distribution of MARI was represented with a histogram and box plot. The chi-square test was used to test the statistical significance of resistance patterns across antibiotics and sources (animal species or zoological gardens). Pearson correlation was used to examine the correlation between MARI and the resistance profile of selected antibiotics. Cluster Analysis was performed using hierarchical clustering (complete linkage method and Euclidean method) to reveal the clusters of ESBL-producing Escherichia coli strains that have similar resistance profiles across multiple antibiotics. The clusters were visualized with a dendrogram. P-value <.05 was considered statistically significant in all the statistical analyses. All R codes used in this study are available as a Supplementary File included in this manuscript (Supplementary File 1).
Results
Distribution of ESBL-producing E. coli in Selected Zoos
A total of 26 ESBL-producing E. coli was identified from a pool of 50 samples from captive wild birds in Southwest Nigeria, accounting for a 52% prevalence rate (Table 2). Ostriches (15.4%), African grey parrots (11.5%), crowned cranes (11.5%) and mallard ducks (11.5%) housed the highest percentage of ESBL-producing E. coli among the sampled zoological birds.
Characteristics of ESBL-producing E. coli isolates to selected antibiotics.

Abbreviations: AUG: Amoxicillin-clavulanate; AZM: Azithromycin; CPD: Cefpodoxime; CHL: Chloramphenicol; CIP: Ciprofloxacin; FOS: Fosfomycin; GEN: Gentamicin; SXT: Trimethoprim-Sulfamethoxazole; TET: Tetracycline; R: resistant; S: sensitive; I: intermediate; MARI: Multiple Antibiotic Resistance Index.
Resistance and Antibiotypes of ESBL-producing E. coli to Selected Antibiotics
Amoxicillin-clavulanate (88.5%) and tetracycline (61.5%), were the antibiotics the ESBL-producing E. coli, mostly resisted (Figure 2 and Supplementary File 2 (Table 1)). In contrast, a few of the ESBL-producing E. coli (3.8%) were resistant to fosfomycin. The antibiotypes of the ESBL-producing E. coli are presented in Table 2.

Resistance of isolated ESBL-producing E. coli to the tested antibiotics.
Fifty-four percent of the ESBL-producing E. coli were resistant to ≥ 3 classes of antibiotics and termed MDR, whereas the remaining 46% were resistant to ≤ 2 classes of antibiotics. Resistance to Amoxicillin-clavulanate was detected in all the multidrug resistance cases of ESBL-producing E. coli in this study (Supplementary File 2 (Table 2)). There was a statistically significant difference between the resistance patterns and the studied antibiotics (P < 0.05; Table 3). However, there was no statistically significant difference among the resistance patterns, surveyed birds (P = .6293), and selected zoological gardens (P = 0.8772; Table 3).
Antibiotics, captive wild birds and zoological gardens influencing the resistance patterns.

χ2: Chi-square; df: degrees of freedom.
Statistically significant.
Relationship Between MARI and Selected Antibiotics
The Multiple Antibiotic Resistance Index (MARI) value of the ESBL-producing E. coli ranged from 0.1 to 0.8 (Table 2). Approximately 76.9% (20/26) of the ESBL-producing E. coli had MARI between 0.2 and 0.8. The highest MARI was recorded for birds such as hens (0.8), crown cranes (0.7), and ostriches (0.7) housed in OOPL, FUTA and OAU zoological gardens. Although the frequency of distribution of MARI was not normally distributed (Supplementary File 2 (Figures 1-3)), MARI had a positive correlation across all the tested antibiotics (Figure 3). Gentamicin (0.76) and ciprofloxacin (0.72) had the highest positive correlation to MARI, while fosfomycin (0.09) had the lowest positive correlation (Figure 3).

Correlation between MARI and resistance profile to selected antibiotics.
Cluster analysis revealed the similarity of each of the ESBL-producing E. coli based on their resistance profile (Figure 4). ESBL-producing E. coli with the same resistance profile (MARI) clustered around each to form a distinct clade in a tree distance. A total of 17 isolates (63%) had a similar MARI and formed different clusters.
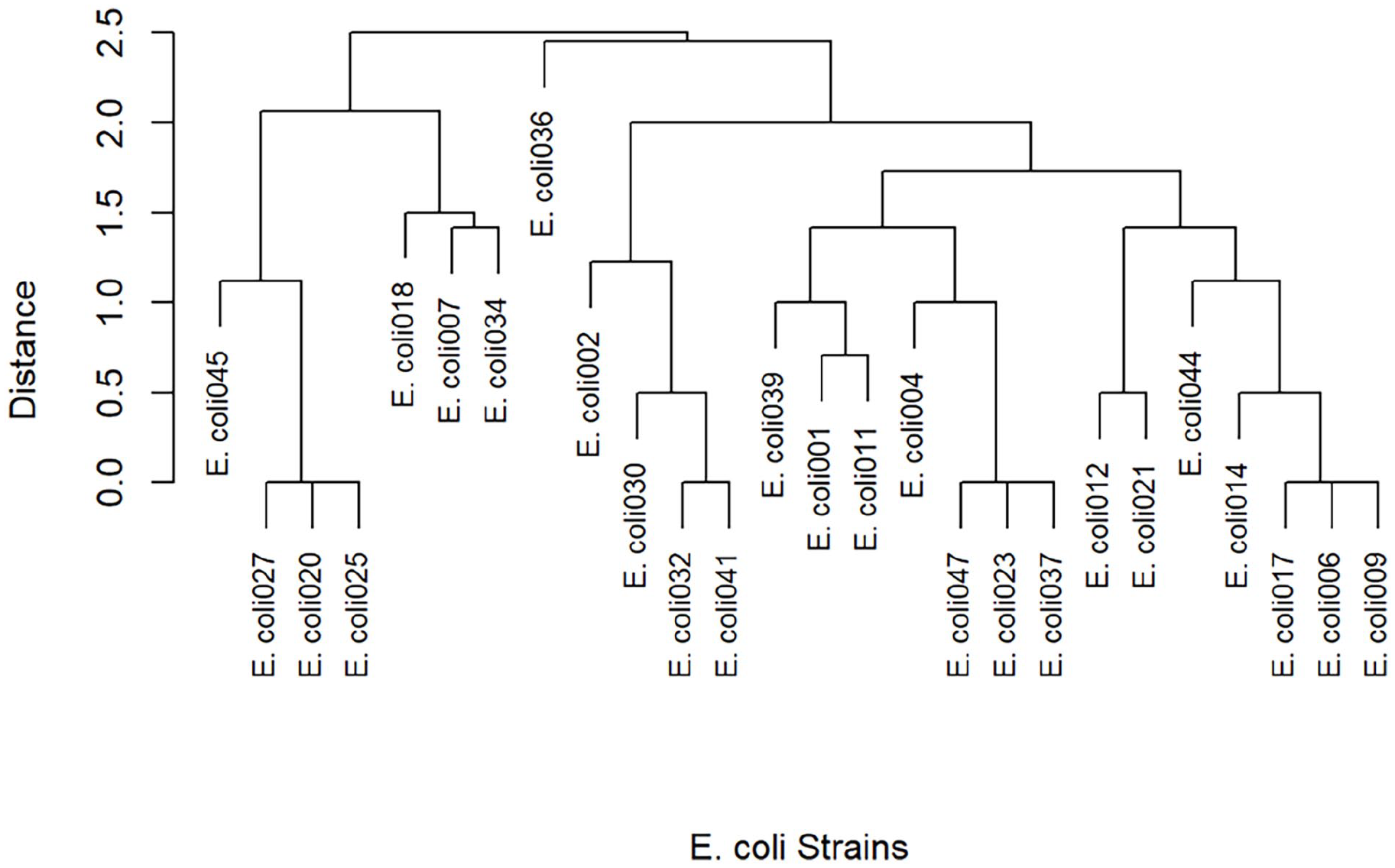
Hierarchical clustering dendrogram of the ESBL-producing E. coli.
Detection of ESBL Genes in the ESBL-producing E. coli
The three ESBL genes targeted were detected in the ESBL-producing E. coli, albeit at different percentages. The most predominant of the ESBL genes was blaCTX-M (62%), followed by blaSHV (34.6%), with the least being blaTEM which was detected in 15% of the ESBL producers isolated (Figure 5).
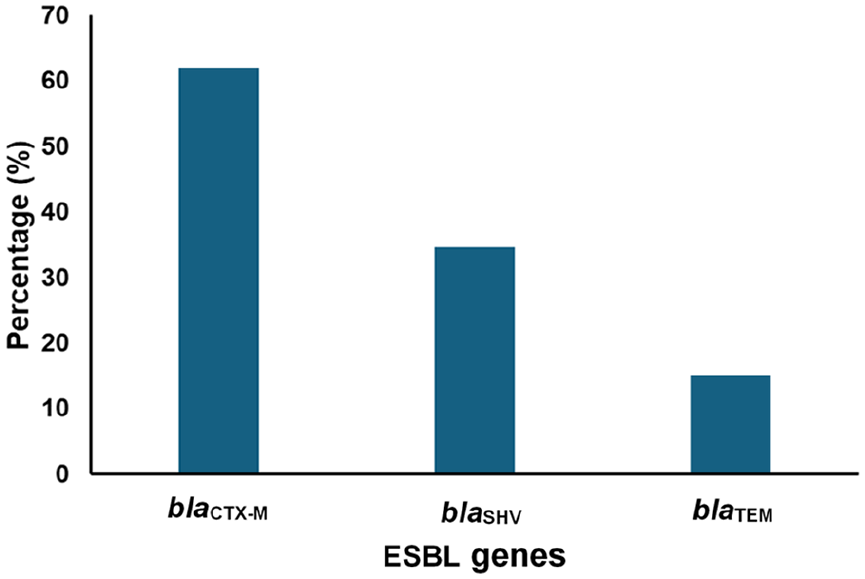
Detection of ESBL genes in ESBL-producing E. coli isolated from the faecal samples of captive wild birds in Southwest Nigeria.
Captive wild birds from GWC: African grey parrot (blaCTX-M and blaSHV), hen (blaCTX-M and blaTEM), and peacock (blaSHV and blaTEM)), FUNAAB: crowned crane (blaCTX-M and blaSHV), FUTA: crowned crane (blaCTX-M and blaSHV), and OMU: palm-nut vulture (blaCTX-M and blaTEM) harboured at least two ESBL genes (Table 4).
Distribution of ESBL genes in ESBL-producing E. coli from the faecal samples of captive wild birds in Southwest Nigeria.

Discussion
This study provides insights into the occurrence of ESBL-producing E. coli in captive wild birds in major zoological gardens in southwestern Nigeria. This is significant considering the number of zoo workers working directly with these animals as well as the diverse population of tourists and regular visitors interacting with this environment. 41 Our study recorded 52% ESBL-producing E. coli in major zoological gardens in Southwest Nigeria. We, therefore, reported a higher prevalence of ESBL-producing E. coli in captive wild birds when compared with studies performed in other locations such as Turkey (4.3% 42 and 2.7% 43 ) and Cameroon (0% 44 ). The high prevalence of ESBL-producing E. coli in our study corroborates the high prevalence reported by Fashae et al 18 and Albrechtova et al. 45 A high prevalence of ESBL-producing E. coli was found in ostriches (15.4%), African grey parrots (11.5%), crowned cranes (11.5%), and mallard ducks (11.5%), among other zoological birds included in this study. This is likely attributable to the exposure of these birds to substantial quantities of antibiotics, potentially stemming from human or animal activities, as well as the distinct microbiomes of captive wild birds. These two factors were identified by Jorquera et al 46 , to cause an increase in AMR in captive wild animals. Other factors such as feed supplies, water sources, bird-to-bird contact, zoo workers and rodents can also contribute to the high prevalence rate. The birds were not exposed to any agricultural or environmental wastes as they were all in cages. But they are in close proximity to one another and are fed with feed in which the antibiotic residue is unknown. During the sampling, some rodents were spotted in the bird cages. There is a high possibility of the rodents contaminating the feed of the birds with antibiotic-resistant bacteria and transmitting them throughout the zoos. There is no strict biosecurity or hygiene practices by the zoo workers. Individuals use a single clothing and boots to serve different animals in different cages. There is a high chance of carrying and transmitting different antibiotic-resistant bacteria to naïve birds through their clothing and boots. It is important to increase the biosecurity of the zoo in order to prevent the spread of resistant bacteria as the zoo workers and the visitors are at high risk of ingesting ESBL-producing E. coli.
Our study recorded high resistance of the ESBL-producing E. coli to amoxicillin-clavulanate (88.5%) and tetracycline (61.5%). Both antibiotics are reported to be among the primary antibiotics utilized in veterinary practices in Nigeria, 47 which might have contributed to the increased resistance rate recorded for them in our study. There might be the possibility of AmpC β-lactamases producer strains among the isolates that were resistant to amoxicillin-clavulanate, which has contributed to the high resistance. AmpC β-lactamase was the first E. coli enzyme reported to destroy penicillin, and exposure to any β-lactams can activate a cascade of events that lead to high AmpC β-lactamase production and resistance. Further study is required to confirm the presence of AmpC β-lactamases producers among our isolates.
The isolates also exhibited resistance to several other commonly-used antibiotics, such as trimethoprim/sulfamethoxazole, chloramphenicol, cefpodoxime and azithromycin, indicating the high multidrug resistance nature of the isolated ESBL-producing E. coli. Our study showed a significant difference between resistance patterns of ESBL-producing E. coli and the selected antibiotics. However, there was no statistically significant difference between the resistance patterns, birds surveyed and/or selected zoological gardens. The indiscriminate use of antibiotics in Nigeria may have contributed to this difference, as reported by previous studies.48 -50
We recorded 76.9% (20/26) of the ESBL-producing E. coli having MARI value between 0.2 and 0.8, and a positive correlation was seen between MARI and the tested antibiotics, which ascertained the high index. Krumperman 51 reported that isolates with MARI ⩾ 0.2 constitute high-risk contamination sources with high antibiotic usage. This suggests that antibiotics are used indiscriminately in each of the sampled zoos, or the horizontal transfer of genes encoding antibiotic resistance could have occurred. The emergence of AMR and means of transmission can also occur simultaneously, which our cluster analysis supported, as 17 (63%) of the ESBL-producing E. coli isolates had a similar MARI and formed clusters. Our finding is supported by the results of Nelly et al 44 in Cameroon. However, Albrechtova et al, 45 who reported a low resistance level in Tai National Park in Cote d’Ivoire, disagreed with this finding. Variations in sample size and location peculiarities may be contributing factors to the disparity.
The major ESBL gene carried by the isolated E. coli was blaCTX-M (62%), followed by blaSHV (34.6%) and blaTEM (15%). Previous studies conducted on poultry birds had similar trends to our study. For instance, Nelly et al 44 reported a high prevalence of blaCTX-M (75.0%), blaSHV (12.5%) and blaTEM (6.3%) in healthy broilers in Cameroon, while Lemlem et al 49 reported the occurrence of ESBL genes in chickens and the farm environment in Malaysia. However, the study of Sakin et al 42 reported only blaCTX-M in caged wild birds in Turkey. CTX-M enzymes are encoded by blaCTX-M, and they have become the most widely occurring β-lactamase in humans and animals. The first case was reported in the 1980s. 52 The distribution of blaCTX-M is attributed to its dissemination via plasmids within bacterial species, which has caused the spread of a silent pandemic (AMR) worldwide. The paucity of data for captive animals showed a need to conduct further genomic studies in the zoological environments in Nigeria, a foundation this study has provided.
Limitation
Despite the great importance of this study, it has some limitations that need to be discussed. We acknowledge the small number of samples available per bird, which might limit the species-level comparison of our analysis. The number of samples collected for each bird was small because of a limited number of zoological gardens and wildlife parks available in the study region, the knowledge and attitude of zoo owners to our study, the birds’ International Union for Conservation of Nature (IUCN) status and the lack of research funding for the study. Having a larger number of samples per bird will further increase our understanding of which bird harbours the highest antibiotic resistance genes and their impact in sharing the genes to the environment. As this is a preliminary study that looks into the occurrence and spread of AMR in captive wild birds, we hope to include a large number of samples in our future study.
Conclusion
This is the first study to investigate the role of captive wild birds in the spread of AMR in Southwest Nigeria. We opined that captive wild birds in Nigeria harbour a significant detection of the ESBL-producing E. coli and ESBL genes, as a result of either indiscriminate use of antibiotics by the zoos or horizontal transfer of antibiotic resistance genes. The study provides a basis for understanding the role of captive wild birds in the spread of AMR. This highlights the need to institute adequate control measures in zoological gardens and animal sanctuaries to prevent the transmission of antibiotic resistance to visitors, workers, other wild animals and the environment. An adequate surveillance system undertaken through the lens of One Health is needed to provide a holistic situation report of AMR in captive wildlife settings.
Supplemental Material
sj-docx-1-ehi-10.1177_11786302251329300 – Supplemental material for Molecular and Epidemiological Characterization of ESBL-producing Escherichia coli from Captive Wild Birds in Zoological Gardens in Nigeria
Supplemental material, sj-docx-1-ehi-10.1177_11786302251329300 for Molecular and Epidemiological Characterization of ESBL-producing Escherichia coli from Captive Wild Birds in Zoological Gardens in Nigeria by Ridwan Olamilekan Adesola, Adetolase Azizat Bakre, Abimbola Olumide Adekanmbi, Bamidele Nyemike Ogunro, Oluwaseun Adeolu Ogundijo, Abdulafees Hamzat, Delower Hossain, Moses Aimanosi Aribana and Luqman Adeola Balogun in Environmental Health Insights
Footnotes
Acknowledgements
We thank all the people who assisted in sample collection and provided support during the project. We are also grateful to the Morris Animal Foundation for providing a seed financial support to ROA for the project.
Ethical Statement and Consent to Participate
The University of Ibadan Animal Care and Use Research Ethics Committee (UI-ACUREC/031-0423/13) gave ethical approval for this study. The Directors of each zoological garden also gave physical and written permission to collect samples from the birds.
Author Contributions
AOA, ROA, and AAB conceptualized the idea. AOA, ROA, AAB, BNO, OAO, AH, and MAA participated in the sample collection and experiment. ROA, AOA, DH, AH and OAO wrote the initial draft of the manuscript. LAB, ROA, AH, and DH performed data analysis, and designed the figures. All authors reviewed the final draft and approved it for publication.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by a grant from the Morris Animal Foundation.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Data in this study will be made available upon request from the first and corresponding authors.
Supplementary Material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
