Abstract
Background:
Antibiotic resistance (ABR) poses a critical global health challenge, necessitating its surveillance across both human and animal health sectors. This study evaluated ABR in bacteria harboured in reared inland fishes sold in Accra and the pond water from which they originated.
Method:
The study was cross-sectional, involving fishes and water sampled from 80 ponds. The gastrointestinal organs of the fishes were homogenised and cultured for bacteria, as were the water samples. The bacteria were identified using matrix-assisted laser desorption/ionisation time of flight mass spectrometry (MALDI-TOF-MS). Antimicrobial susceptibility test was done using the Kirby-Bauer method. Multidrug-resistant (MDR) bacteria were selected for further testing. The double disc diffusion method was used to detect extended-spectrum beta-lactamase (ESBL) production in isolates that were resistant to third-generation cephalosporins. Whole genome sequencing was performed on the ESBL-positive isolates using the Illumina Miseq platform.
Results:
In total, 39 different bacterial species, with their individual numbers totalling 391, were isolated. The bacteria were predominantly
Conclusion:
The fishes and the pond water were contaminated with a diverse range of bacteria, mainly
Introduction
Antibiotic resistance (ABR) poses a significant and urgent global health threat and was implicated in the deaths of 4.95 million individuals in 2019. 1 If effective measures are not taken to address antibiotic resistance by 2050, 10 million people could die annually, and the gross domestic product (GDP) could decline by 2% to 3.5%, resulting in a total cost of US$100 trillion. 2 Sub-Saharan Africa, which is disproportionately burdened with antibiotic resistance, is expected to experience a relatively higher impact, with a GDP reduction of US$2895 billion, equivalent to 20% of the region’s total economic output.1,2
Aquaculture is a significant contributor to the problem of antibiotic resistance due to its extensive use of antibiotics.3,4 Concerningly, around 70% to 80% of the antibiotics utilised in aquaculture are released into water systems; fish, being ineffective at metabolising these substances, predominantly excrete them into the environment.3,4 Moreover, antibiotics present in fish feed have the potential to persist in aquatic environments for extended periods, rapidly spreading through water systems and exerting selective pressure on ecosystems, thereby driving the development of antibiotic resistance. 5
To help tackle the antibiotic resistance menace efficiently, it is important to continuously conduct surveillance, targeting interphases that hold high relevance regarding the selection and transmission of antibiotic-resistant organisms among wildlife, humans, and the environment, such as aquaculture ecosystems. However, most antibiotic resistance surveillance studies, particularly, those conducted in sub-Saharan Africa, have mainly focused on humans and rarely on animals and the environment.6 -14 The few that have been conducted on animals have usually been on poultry, cattle, and pigs, with very little attention given to the aquaculture sector.6,7,15 -20 In Ghana, the minimal antibiotic resistance surveillance efforts in the aquaculture sector have mainly been on phenotypic characterisation of antibiotic resistance.15,16,19,20 However, data regarding genotypic characterisation of antibiotic resistance in Ghanaian aquaculture settings are generally sparse. 20 Also, on the whole, there is a paucity of data on the patterns and trends of antibiotic resistance in the aquaculture environment, making data on antibiotic resistance surveillance less comprehensive. To help fill this important knowledge gap, this study evaluated antibiotic resistance in bacteria harboured in reared fishes sold in Accra and the pond water from which they originated, employing a combination of phenotypic and genotypic techniques.
Materials and Methods
Study design and site
This was a cross-sectional study in which farm-reared fishes and pond water samples from which the fishes originated were collected, via random sampling. The farmers raised their fishes in intensive farming system, including earthen pond, concrete tanks, plastic tank, and tarpaulin system at Nii Ashley, Gbentaana, Mammomo, Adjringanor, Manhee, Amanfrom, Maledjor, Frafraha, and New Legon, all of which are situated in the Adenta District of the Greater Accra Region of Ghana (Figure 1). The Adenta Municipal Assembly, which has Adenta as its Central Business District, can be found 10 kilometres to the northeast of Accra. Its precise coordinates are 5°43’ north latitude and 0°09’ west longitude. Approximately 928.4 square kilometres of land make up the municipality. In the east and north, it shares borders with the Ashaiman Municipal Assembly and the Kpong Akatamanso District Assembly. In the west and south, it shares boundaries with the La Nkwantanang Municipal Assembly.
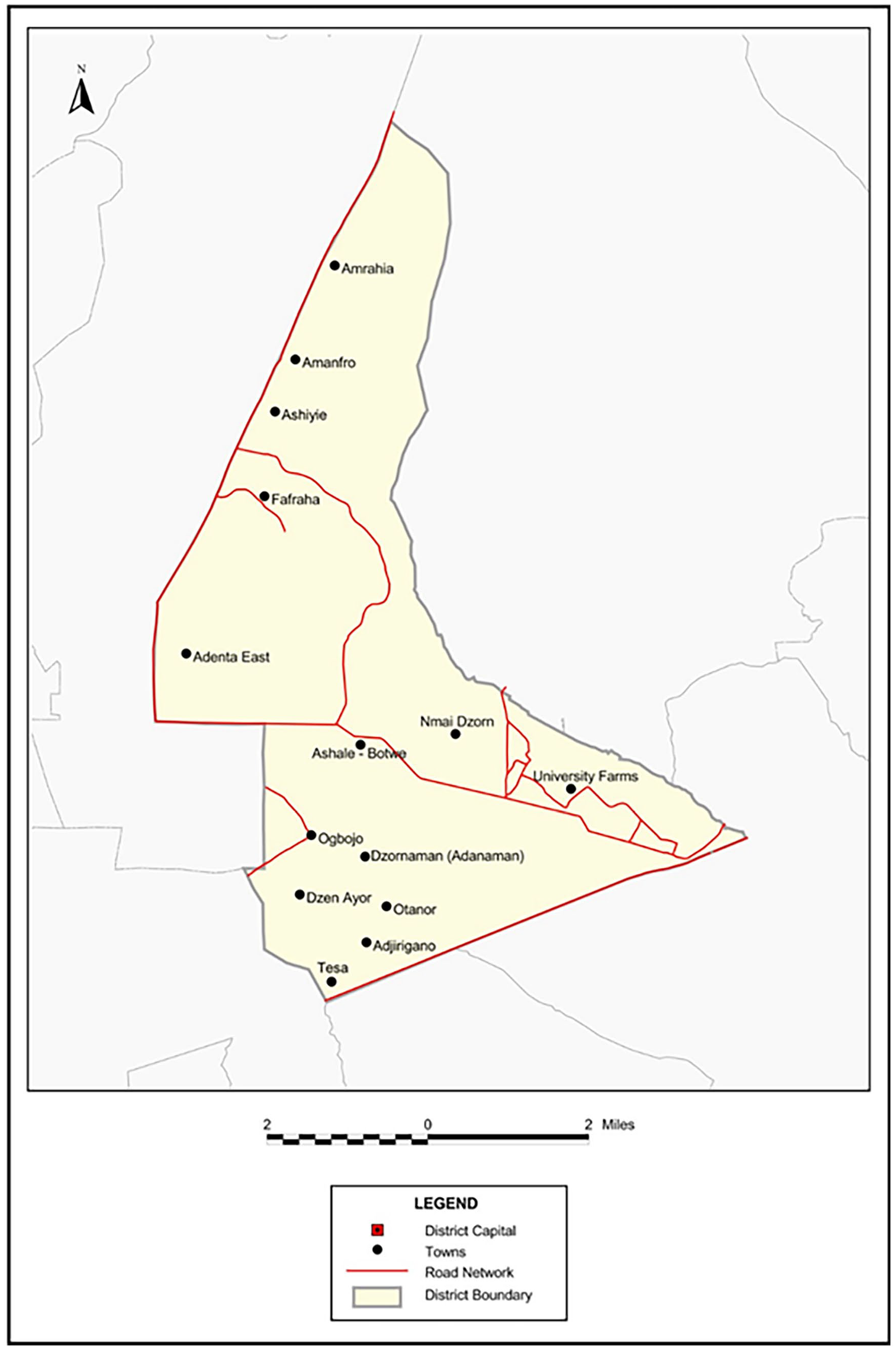
A map of the Adenta Municipality (Source: Ghana Statistical Service, GIS).
Sample collection
The procedures carried out on the fish samples were in accordance with the animal welfare policy of the Veterinary Directorate of the Ministry of Food and Agriculture in Ghana, as well as the sets of animal care and use recommendations. 21 The water and the farmed fishes were both aseptically sampled with the cooperation of the farms’ management teams, and the sampling procedures were carried out in accordance with EPA 22 recommendations. On each day of sampling, the collection of samples was carried out between the hours of 7:00 a.m. and 11:00 a.m. First-hand information on the management techniques followed by the farmers, including the use of antibiotics, was obtained. In total, fishes and pond water from 56 catfish ponds and 24 tilapia ponds were sampled across 12 fish farms. For each pond, five healthy-looking fish were randomly collected from scoop net and casting catches into uniquely-labelled sterile zip-lock bags, following aseptic procedures. Also, 500 ml water samples (in five 100 ml volumes) were collected at five different well-spaced points of the originating ponds using sterile bottles, at a depth of 50 cm below the surface of the water. These samples were transported on ice to the research laboratory of the Department of Medical Microbiology, University of Ghana Medical School.
Laboratory analysis
Bacterial isolation and identification
For each pond, the individual water samples were put together and completely mixed by turning the container slowly and carefully about ten times. A 10 ml portion of each mixed water was placed in a beaker and filled up to the 100 ml mark. The external surface of each fish sample was sterilised with 70% ethanol, and an aseptic incision made across the peritoneal cavity of the fish using a sterile scalpel blade. For the fish from each pond, the intestines and gastric-related organs were carefully removed and separated from the non-gastrointestinal organs, after which the gastrointestinal parts were completely homogenised. This homogenised content (10%) was added to 90% of buffered peptone water and incubated at 37°C for 18 to 24 hours.
After incubation, a loopful of the homogenised mixture was inoculated on blood agar, MacConkey agar, and thiosulfate-citrate-bile salts-sucrose agar (Oxoid, Basingstoke, Hants, UK). For modified semisolid Rappaport Vassiliadis agar, three drops of the mixture were placed on a plate and incubated at 37°C for 18 to 24 hours. From the mixed-growth culture, using the aseptic technique, each group of isolates was obtained using parallel overlapping strokes of the streak method of inoculation. The entire process was repeated for each distinct representative bacterial colony observed in the mixed-growth culture plate, except the modified semisolid Rappaport Vassiliadis agar which was placed on xylose lysine deoxycholate agar and incubated for 18 to 24 hours at 37°C.
Isolates obtained after using the streaking method were identified using Gram staining, and their identities were confirmed using matrix-assisted laser desorption/ionisation time of flight mass spectrometry (MALDI -TOF-MS).
Antimicrobial susceptibility testing
The disc diffusion technique was used to determine the susceptibility of the bacterial isolates using the following antibiotics: ciprofloxacin (5 μg), gentamicin (10 μg), tetracycline (30 μg), meropenem (10 μg), ertapenem (10 μg), chloramphenicol (30 μg), and cefotaxime (30 μg) (Oxoid, Basingstoke, Hants, UK). The criteria of the Clinical Laboratory Standard Institute (CLSI) were followed while conducting the test and interpreting its results. 23
A 0.5 McFarland standard suspension of each bacterial isolate was produced with sterile normal saline and used to inoculate the surface of a sterile Mueller-Hinton agar plate, after which antibiotic discs were carefully placed on the surfaces of the plates. The agar plates were then incubated at 37°C for 16 to 20 hours. After that, diameter measurements of the zones of inhibition were recorded using a pair of callipers, and converted from millimetres to percentages using the CLSI guidelines for specific pathogens and antibiotics. 23
The double disc diffusion method was employed in detecting extended-spectrum beta-lactamase (ESBL) production in isolates that were resistant to third-generation cephalosporins, using cefotaxime (30 μg) and amoxicillin-clavulanate (20/10 μg) or ceftazidime (30 μg) and amoxicillin-clavulanate (20/10 μg) antibiotic discs (Oxoid, Basingstoke, Hants, UK).
Whole-genome sequence analysis
ESBL-producing isolates were sequenced at the Noguchi Memorial Institute for Medical Research, University of Ghana. DNA of each freshly cultured isolate was extracted using the Qiagen DNA MiniAmp kit, following the manufacturer’s instructions. Extracted DNA was quantified using a Qubit 4.0 Fluorometer assay kit (Thermo Fisher Scientific, MA), followed by library preparation with Illumina DNA prep, per the manufacturer’s instructions. DNA concentration between 100 and 500 ng in 30 µl was used as starting material for library preparation. The Beaded-Linked Transposomes (BLT), which included fragmenting and tagging the DNA with adapter sequences, were used to tag the DNA. Following this procedure, a post-tagmentation clean-up was carried out to halt tagmentation and wash the adapter-tagged DNA prior to PCR amplification. The tagmented DNA was subsequently amplified by PCR, and paired index adapters and sequences needed for cluster formation on the flow-cell during sequencing were also added. The amplified libraries were then cleaned up using a post-PCR clean-up procedure to get rid of extra unbound index adapters. The quality and concentration of fragmented DNA were assessed with the 2100 bioanalyser system (Agilent) and qPCR (Kapa Sybr Fast qPCR kit), respectively. Using the fragment sizes generated by the bioanalyser, and the concentration of individual libraries from the qPCR, libraries were then normalised, pooled, and loaded on Illumina 2 × 300 cycle cartridges for sequencing on the Miseq platform (Illumina Inc., San Diego, CA).
Raw sequenced reads were quality-controlled, and adapters were trimmed using FASTQC (http://www.bioinformaticsbabraham.ac.uk/projects/fastqc/) and Trimmomatic (http://www.usadellab.org/cms/index.php?page=trimmomatic), respectively, with a minimum quality set at Q20.24,25 Trimmed reads were
Assembled files were uploaded to ResFinder (https://cge.food.dtu.dk/servicesResFinder/), a tool available on the Centre for Genomic Epidemiology (CGE) (http://www.genomicepidemiology.org/) platform, to detect resistance genes present in the sequenced isolates using an identity threshold of 90% and a minimum length of 60%. Prediction of bacterial species and acquired virulence genes were done using KmerFinder (https://cge.food.dtu.dk/services/KmerFinder/) and VirulenceFinder (https://cge.food.dtu.dk/servicesVirulenceFinder/), respectively. Plasmids and mobile genetic elements were predicted using Plasmidfinder (https://cge.food.dtu.dk/services/PlasmidFinder/) and MGEfinder (https://cge.food.dtu.dk/services/MobileElementFinder/), respectively. The sequence types of the isolates were predicted using MLSTFinder (https://cge.food.dtu.dk/services/MLST/). Distinctively, predicting serotypes of the
Ethical approval
Ethical approval was obtained from the Ethical and Protocol Review Committee of the University of Ghana’s College of Health Sciences (ID: CHS-Et/M.2-P 4.6/2021-2022).
Results
Characteristics of the sampled ponds
Table 1 summarises the pond characteristics. The ponds on the 12 farms were mainly used in rearing catfish (70.0%,
Pond characteristics.

Five fish were randomly sampled from each pond.
Distribution of the isolated bacteria in the fishes and pond water
In total, 391 bacteria (of 39 different species) were isolated from the fishes and pond water. In the fish, a total of 201 bacteria, of 38 different species, were isolated. Conversely, the bacterial isolates from the pond water were 190 in total, from 34 species. As a composite, the top five bacteria were
Distribution of the bacteria harboured in the fishes and pond water.

Antibiotic resistance of the bacterial isolates
The resistance of all the isolated bacteria to the tested antibiotics, stratified by sample type, are displayed in Figure 2, and in Tables 3 to 5, the distributions of the antibiotic resistance rates across the individual bacteria are presented. Across each of the fishes, pond water, and fishes and pond water combined, the antibiotic resistance rates were highest for tetracycline (31% to 42%), cefotaxime (29% to 35%), and chloramphenicol (19%). No resistance was recorded against either meropenem or ertapenem.
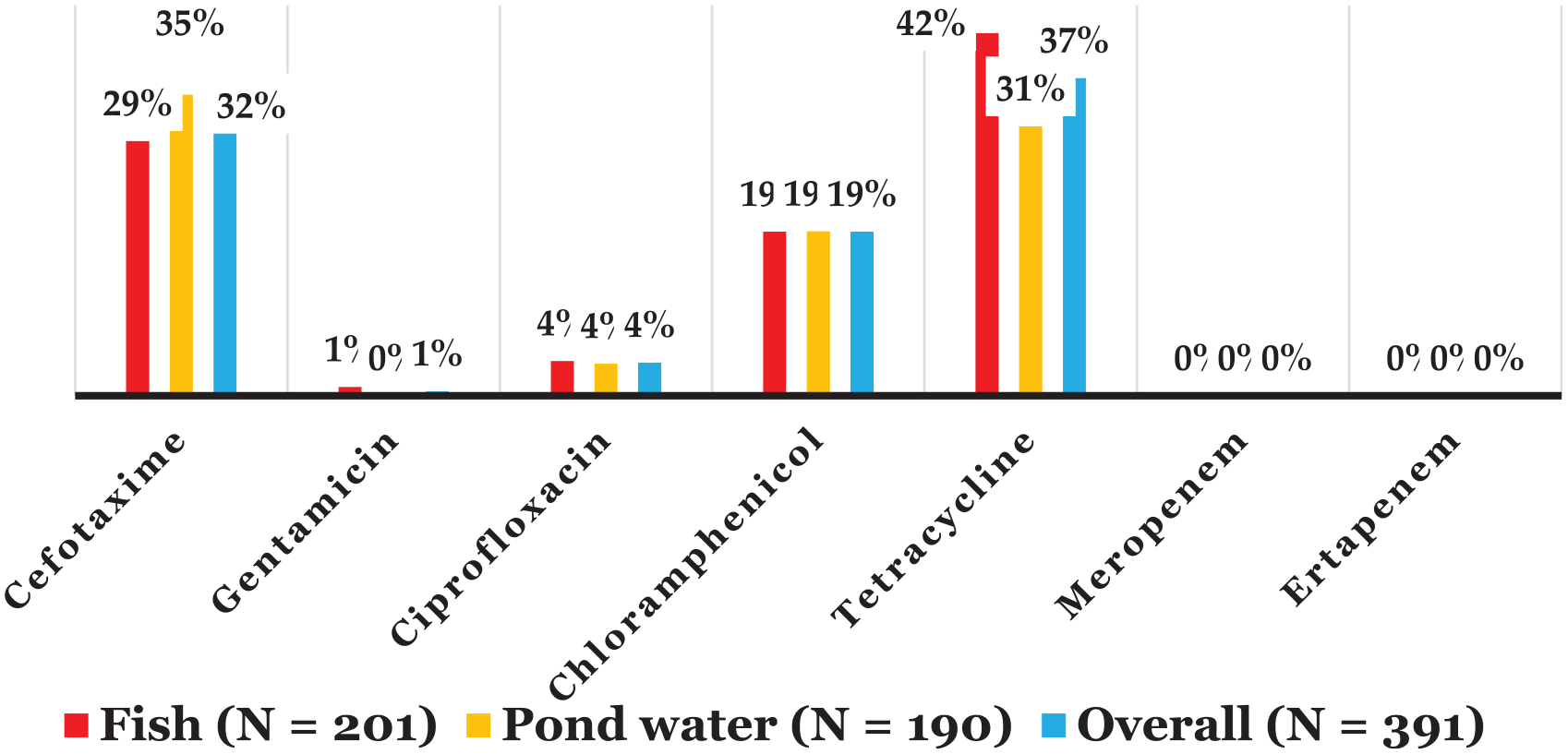
Antibiotic resistance patterns of the bacterial isolates.
Antibiotic resistance among the bacteria isolated from both the fishes and pond water.

CTX, Cefotaxime; GM, Gentamicin; CIP = , Ciprofloxacin; C, Chloramphenicol; TE, Tetracycline; MEM, Meropenem; ERT, Ertapenem.
Antibiotic resistance among the bacteria isolated from the fishes.

CTX, Cefotaxime; GM = Gentamicin; CIP, Ciprofloxacin; C, Chloramphenicol; TE, Tetracycline; MEM, Meropenem; ERT, Ertapenem.
Antibiotic resistance among the bacteria isolated from the pond water.

CTX, Cefotaxime; GM, Gentamicin; CIP, Ciprofloxacin; C, Chloramphenicol; TE, Tetracycline; MEM, Meropenem; ERT, Ertapenem.
Regarding
In
For
Resistance rates in
For
Regarding
In the combined fishes and pond water,
In the case of
Furthermore, in
Moreover,
In terms of the prevalence of multidrug resistant (MDR) and ESBL-producing bacteria isolated, 13.6% and 1.3% of the total bacteria were MDR and ESBL producers, respectively (Table 6).
MDR and ESBL prevalence among the bacteria.

Categorically, 13.4% and 1.0% of the bacteria isolated from the fishes were MDR and ESBL producers, respectively. Similarly, bacteria isolated from the pond water recorded 13.7% and 1.6% for MDR and ESBL-producing bacteria, respectively. Among all the bacteria isolated, 20.6% and 7.4% of
Molecular epidemiology of the ESBL-producing isolates (sequence types, serotypes, resistance genes, and virulence genes)
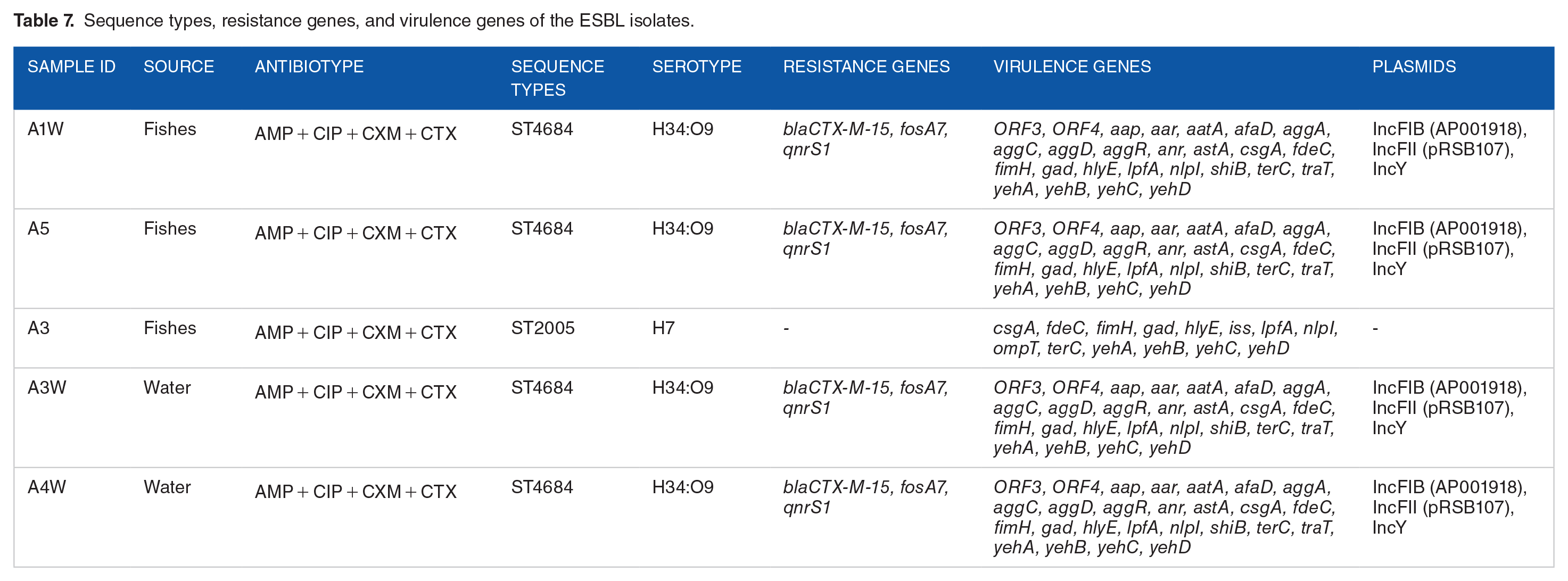
Table 7 shows the molecular epidemiology of the ESBL-producing isolates. These bacterial isolates (A1W, A5, A3, A3W, A4W), which were from the fishes and pond water, exhibited resistance to ampicillin, ciprofloxacin, cefuroxime, and cefotaxime, with specific genetic profiles including resistance genes
Sequence types, resistance genes, and virulence genes of the ESBL isolates.
Discussion
The goal of this study was to evaluate antibiotic resistance in bacteria harboured in reared fishes in Accra and the pond water from which they originated, focusing on distribution of the harboured bacteria and their antibiotic resistance patterns, as well as their ESBL phenotypes, genotypes, and sequence types. The rich bacterial diversity in the fishes and pond water could possibly be accounted for by the different types of fish holding facilities used, namely concrete, earthen, plastic, and tarpaulin. It could also be attributed to the bioaccumulation of microorganisms within the fish.
19
This process allows for the concentration of bacteria and other microorganisms in the tissues of the fish, often in the gut, as well as their presence in the surrounding pond water. This bioaccumulation is particularly significant when considering the diet and feeding habits of the fishes – fishes consume various materials, including plankton, known to harbour high levels of bacteria. Notably,
Essentially, the highlighted concerns about the microbial safety of reared fishes are especially validated by similar observations on frequent occurrence of
To buttress the concerns about the microbial safety of the fishes and the potential risks they pose,
Also,
Among the bacterial isolates,
Other bacterial species, such as
It is evident that the distribution and prevalence of bacterial species in aquaculture systems can vary depending on the specific environment, culture practices, and geographical location. Factors such as water source, feeding regimes, water quality management, and biosecurity measures can significantly influence the microbial ecology of aquaculture systems. Therefore, it is essential to consider these factors when interpreting and extrapolating results from different studies.
The presence of other pathogenic bacteria, such as
Furthermore, the implications of antibiotic resistance in aquaculture cannot be overlooked. The presence of antibiotic-resistant bacteria, such as
It is worth noting that this study focused on a specific aquaculture system and a limited set of bacterial species. Further research is needed to explore the microbial ecology of different aquaculture systems, including marine, brackish, and freshwater environments, and to investigate a broader range of bacterial species. Additionally, the application of advanced molecular techniques, such as metagenomics and metatranscriptomics, can provide a more comprehensive understanding of the microbial community dynamics and functional interactions within aquaculture environments.
Generally, the antibiotic resistance rates were low to moderate, and were highest for tetracycline (31% to 42%), cefotaxime (29% to 35%), and chloramphenicol (19%), while no resistance was recorded against either meropenem or ertapenem. These results align with the global concern of widespread resistance to these antibiotics in various bacterial species. The high levels of resistance to tetracycline are consistent with the findings reported by Adinortey et al. 19 and Ginovyan et al. 33 in fish farms in Ghana and Armenia, respectively. The substantial resistance to cefotaxime highlights the need for careful consideration when using these antibiotics in aquaculture.
The prevalence variations between the fishes and pond water highlight the need for a more targeted investigation into the factors driving resistance in specific bacterial strains. This finding aligns with previous studies by Adinortey et al.
19
and Ginovyan et al.,
33
which reported high levels of antibiotic resistance in coliform bacteria, including
This study provides evidence on the need to inculcate the One Health approach in managing antibiotic resistance in aquaculture. It can be seen from the results that the prevalence of antibiotic-resistant pathogens, antibiotic resistance genes, and virulence genes, in the gastrointestinal of the sampled fishes and the water within which the fishes were kept are determining factors for exacerbating AMR in the environment, in humans after their ingestion of improperly cooked infected fish, and that which is transmitted between and among fishes. However, a comprehensive study including the presence of the different type of antibiotics present in the soil, pond, and gut of fishes will be necessary to buttress this study, and to establish convincing evidence for the need for using One Health approach in tackling this aquaculture menace.
In line with the observed antibiotic resistance traits highlighted above, various antibiotic resistance-conferring genes were observed in this study, as has been the case of aquatic sediments in previous reports, showcasing the intricate nature of the issue.
34
Notably, this study identified the resistance genes
Mechanisms of quinolone resistance frequently involve plasmid-encoded
Two distinct serotypes, H7 and H34:O9, were identified in this study, with both serotypes present in fish samples, while only H34:O9 was found in the water samples. Serotype differentiation, based on specific flagellar (H) and somatic (O) antigens, reveals the diversity of bacterial strains in these environments. Specifically, serotype H34:O9 expresses the 34th flagellar antigen and the ninth somatic antigen, whereas serotype H7 only expresses the seventh flagellar antigen.
47
Although there is limited information on H34:O9 specifically, serogroup O9 was classified by Navarro et al.
48
as commensal strains containing diarrhoeagenic
The identified prevalence rates of antibiotic-resistant bacteria in aquatic environments have potential implications for public health. Also, the transfer of resistance genes from aquatic bacteria to human pathogens, through the food chain or waterborne routes, could contribute to the dissemination of antibiotic resistance in clinical settings. Furthermore, the discernible differences in resistance patterns between the fishes and pond water suggest that the aquatic habitat plays a role in shaping the antibiotic resistance landscape. Moreover, the slightly elevated prevalence of MDR in fishes compared to the overall sample underscores the potential influence of ecological factors on resistance dynamics within specific niches. In addition, the observed patterns of antibiotic resistance in the fish and pond water bacteria underscore the need for prudent antibiotic use in aquaculture. Plus, the resistance of bacteria to certain antibiotics, such as cefotaxime, highlights the potential challenges in disease management within this aquatic ecosystem. Besides, the effectiveness of gentamicin against a high percentage of bacteria isolated from both the fishes and pond water is promising and emphasises the importance of selecting antibiotics carefully in aquaculture practices.
Overall, this study has revealed a highly diverse microbial composition in aquaculture systems in Accra, Ghana, as well as patterns of antibiotic resistance genotypes and phenotypes. The presence of resistance genes and virulence factors suggests a complex interplay between genetic traits and selective pressures in the aquatic environment. It is noted, though, that the whole-genome sequence analysis was limited to the five ESBL-producing isolates recovered from the fishes and pond water in this study, all of which happened to be
Conclusions
The fishes and the pond water were contaminated with a broad range of bacteria, mainly
Further research is needed to explore the microbial ecology of different aquaculture systems, including marine, brackish, and freshwater environments, and to investigate a broader range of bacterial species. Additionally, the application of advanced molecular techniques, such as metagenomics and metatranscriptomics, can provide a more comprehensive understanding of the microbial community dynamics and functional interactions within aquaculture environments. Furthermore, it is recommended that regulatory bodies enforce microbial food safety polices in the fish production chain and sensitise consumers on the need to cook their fish well to destroy contaminating bacteria.
Footnotes
Acknowledgements
We are grateful to the technical staff of the Department of Medical Microbiology, University of Ghana Medical School (UGMS), as well as those of the Bacteriology Department, Noguchi Memorial Institute for Medical Research (NMIMR), University of Ghana. We also appreciate the funding support from the SeQ-Africa (FF RGR2 FF25) Project which enabled the sequencing of the ESBL isolates.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Whole genome sequencing of the ESBL isolates was supported by the SeqAfrica Project (FF RGR2 FF25), funded by the Department of Health and Social Care’s Fleming Fund using UK Aid. The views expressed in this publication are those of the authors and not necessarily those of the UK Department of Health and Social Care or its Management Agent, Mott MacDonald.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Conceptualisation, N.T.K.D.D.; methodology, N.T.K.D.D., F.C.N.K., F.N.N.N.-M., B.E., B.K.A.T., A.D.D., P.B.T.-Q., and E.S.D.; software, B.K.A.T. and P.H.; validation, N.T.K.D.D., B.E., F.C.N.K., A.D.D., P.B.T.-Q., and E.S.D.; formal analysis, N.T.K.D.D., F.C.N.K., F.N.N.N.-M., B.E., B.K.A.T., A.O., and E.S.D.; investigation, N.T.K.D.D., F.C.N.K., F.N.N.N.-M., B.K.A.T., B.E., and A.O.; resources, N.T.K.D.D., F.C.N.K., F.N.N.N.-M., D.E.K.K., A.D.D., P.B.T.-Q., B.E., and E.S.D.; data curation, F.C.N.K., F.N.N.N.-M., B.E., B.K.A.T., D.E.K.K., A.O., P.H.; writing—original draft preparation, N.T.K.D.D., F.C.N.K., B.E., F.N.N.N.-M., P.H., A.D.D., and E.S.D.; writing—review and editing, N.T.K.D.D., F.C.N.K., F.N.N.N.-M., B.E., B.K.A.T., D.E.K.K., A.O., P.H., A.D.D., P.B.T.-Q., and E.S.D.; visualisation, N.T.K.D.D., F.C.N.K., F.N.N.N.-M., D.E.K.K., A.D.D., P.B.T.-Q., B.E., and E.S.D.; supervision, N.T.K.D.D., B.E., and E.S.D.; project administration, N.T.K.D.D., A.D.D., P.B.T.-Q., and E.S.D.; funding acquisition, B.E. All authors have read and agreed to the published version of the manuscript.”
