Abstract
Despite widespread vaccination, pertussis resurgence persists due to waning immunity and emerging resistant strains in
Introduction
Pertussis, commonly known as whooping cough, is a highly contagious respiratory infection caused by the Gram-negative coccobacillus
Despite the availability of vaccination programs, pertussis remains a major global public health concern. According to the World Health Organization, pertussis exhibits cyclical epidemic peaks every 2 to 5 years, with multiple large-scale outbreaks reported over the past decade. 2 Although reported cases declined markedly during the COVID-19 pandemic, a substantial resurgence has been observed in the post-pandemic period, reaching alarming levels worldwide.3,4
Vaccination has long been the cornerstone of pertussis prevention and has substantially reduced disease incidence worldwide. The shift from DTwP to DTaP vaccines was driven by the latter’s improved safety profile and tolerability due to reduced reactogenicity, making DTaP preferable for pediatric use. 5 Waning immunity, antigenic variations in circulating strains, and diminished natural boosting from subclinical infections have been associated with the resurgence of pertussis in highly vaccinated populations.5,6 Moreover, DTaP vaccines, which target a limited set of antigens, exhibit limitations that may restrict the breadth and durability of the immune response.7,8 Together, these limitations underscore a critical gap, highlighting the urgent need for next-generation vaccines with enhanced and broader protective efficacy.
The persistence and resurgence of pertussis are further illuminated by the genetic evolution of
Pangenomics, a subset of this paradigm, examines the complete genomic repertoire across strains to pinpoint conserved elements, such as those involved in essential metabolic processes or signaling pathways, ensuring broad applicability irrespective of strain-specific variations.15,16 By integrating multiple genomes, this approach enables the identification of the core genome and conserved genes shared across all strains, as well as the non-core genome; classified into accessory and singleton genomes; that contributes to phenotypic variability and adaptation. 17
This study addresses these challenges by employing pangenomics reverse vaccinology—a sophisticated, data-driven framework—to identify novel vaccine candidates from
Materials and Methods
This

Schematic representation of the pangenomics-driven reverse vaccinology pipeline applied to prioritize targets in
Pangenome Analysis
Prior to initiating the pangenome analysis, a series of preparatory steps are required, starting with the acquisition of genome data. Ten sequencing reads of
Identification of Host and Pathogen Metabolic Pathways
Through a systematic workflow (Figure 1), this study represents the first computational and subtractive genomics analysis of various metabolic pathways to identify potential vaccine targets in
Identifying the Non-Homologues and Unique Proteins
This step was performed to eliminate proteins from common pathways which serve to minimize potential side effects, the perturbation of the protein function and avoid inducing an immune response against the microbiota. To execute this step, 2-step comparisons were conducted between human and pathogen proteomes to identify non-homologous proteins of
Identification of Essential Proteins for Bordetella pertussis
After identifying all non-homologous and unique proteins, we further filtered them based on their essentiality, specifically, targeting proteins critical for
Prioritization of Vaccine Targets
Subcellular Localization
The subcellular localization prediction and secretion screening was performed on the non-homologous and essential proteins identified in
Physicochemical Parameters
Initially, information regarding non-homologous, unique, and essential proteins was evaluated to prioritize suitable vaccine targets; however, to minimize the time required for vaccine testing and development, it is preferable to evaluate these vaccine targets by several molecular and structural criteria. 31 This involved calculating the molecular weight, which must be less than 110 kDa, and the GRAVY score, which should be less than or equal to zero. These 2 parameters facilitate the purification of targeted proteins. 32 The stability index, which must be less than or equal to 45, indicates protein stability. For pI, there is no critical value, but an extreme pI value could cause issues; thus, the pI should range between 5 and 9 and differ from human pH. These calculations were performed using the ProtParam (https://web.expasy.org/protparam/) computational web tool. 33 The next criteria involved employing TMHMM (v2.0) (https://services.healthtech.dtu.dk/services/TMHMM-2.0/), which used to predict the number of the transmembrane helices of the proteins. The selection was made if a protein has 1 or 0 transmembrane helices.34,35
Prediction of Antigenicity and Immunogenicity
Proteins classified as PSE or secreted are those that first interact with the immune system
36
; thus, we predicted their ability to elicit a strong immune response by evaluating their antigenicity using VaxiJen (v2.0) (https://www.ddg-pharmfac.net/vaxijen/VaxiJen/VaxiJen.html). VaxiJen employs to its alignment-independent approach, which evaluates protein antigenicity based on the physicochemical properties of amino acids rather than sequence similarity. Proteins with a score > 0.4 were considered probable antigens with significant immunogenic potential.37,38 Subsequently, we proceeded to predict of potentially immunogenic epitopes in a given protein sequence, that may significantly reduce wet lab effort needed to discover the epitopes required for the design of vaccines and for immunodiagnostics. The goal of B cell epitope prediction was to identify immunogenic epitopes on potential antigens, encompassing both linear (continuous) and discontinuous (conformational) categories, capable of binding to B cell receptors, thereby activating B lymphocytes and eliciting a humoral immune response characterized by the secretion of neutralizing antibodies.39,40 Linear B-cell epitopes were predicted employing BepiPred (v3.0) (https://services.healthtech.dtu.dk/services/BepiPred-3.0/) with a threshold of 0.5 and a minimum residue length of 7 amino acids.
41
For conformational epitopes, which necessitate a 3D protein structure model, the Protein Data Bank (PDB) (https://www.rcsb.org/) was initially queried to ascertain the availability of structures for the target proteins; in cases where no structure existed, a homology model was generated using SWISS-MODEL (https://swissmodel.expasy.org/).
42
The corresponding .
Results
Analysis Pangenomics
Using Roary_plots for visualization of pangenomics analysis results give an overview of
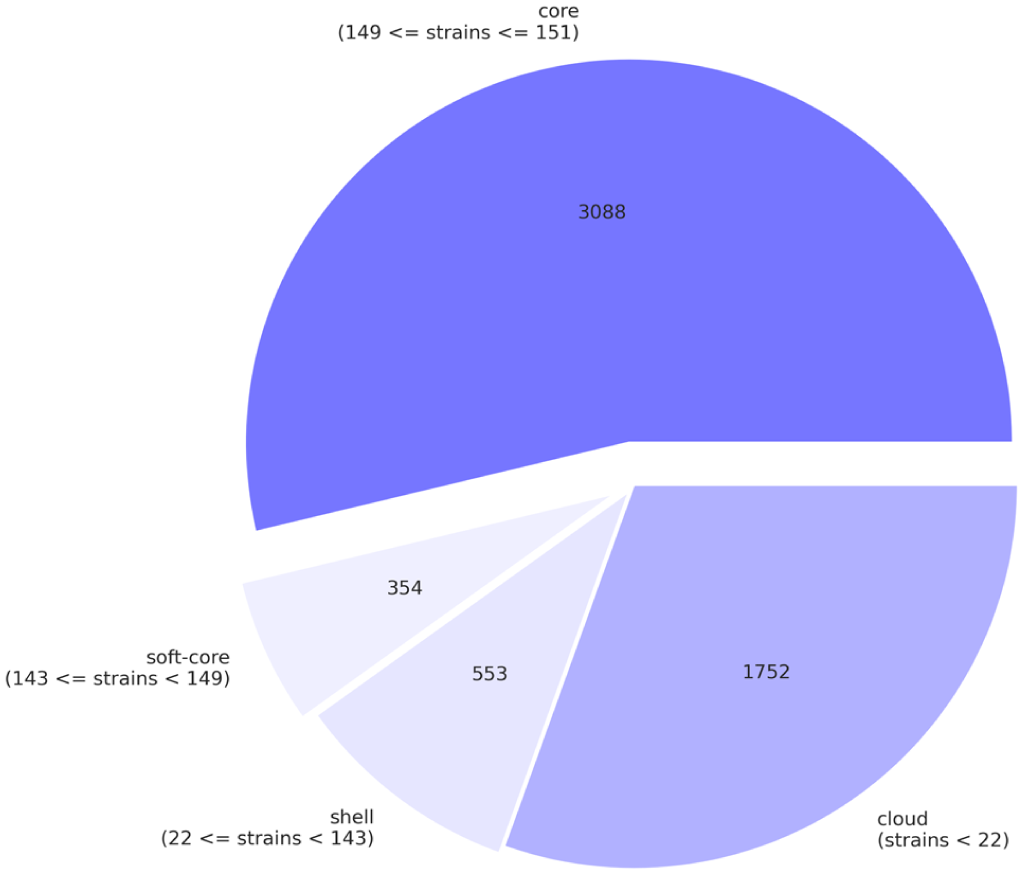
Distribution of 5747 genes into the core (core + soft-core) and the non-core (shell + cloud) genome categories.
Identifying the Non-Homologues and Unique Proteins
Proteins identified as human homologs were excluded in the initial step, as such targets could adversely affect host metabolism. Unique metabolic pathways specific to
Identification of Essential Proteins for Bordetella pertussis
Being non-human and involved in unique metabolic pathways are not the sole criteria for selecting favorable vaccine targets. Identifying proteins that regulate key factors, such as nutrient uptake, growth, secretion systems, virulence, and pathogenicity, is of great importance for disrupting pathogen functions and survival.27,44 Such proteins are considered essential for the pathogen. However, not all essential proteins are non-homologous. Therefore, pathogen proteins that are both non-homologous and essential represent more attractive vaccine targets.
45
The non-homologous proteins identified through resistance gene analysis were further screened based on their essentiality. A total of 63
Non-Homologous and Essential
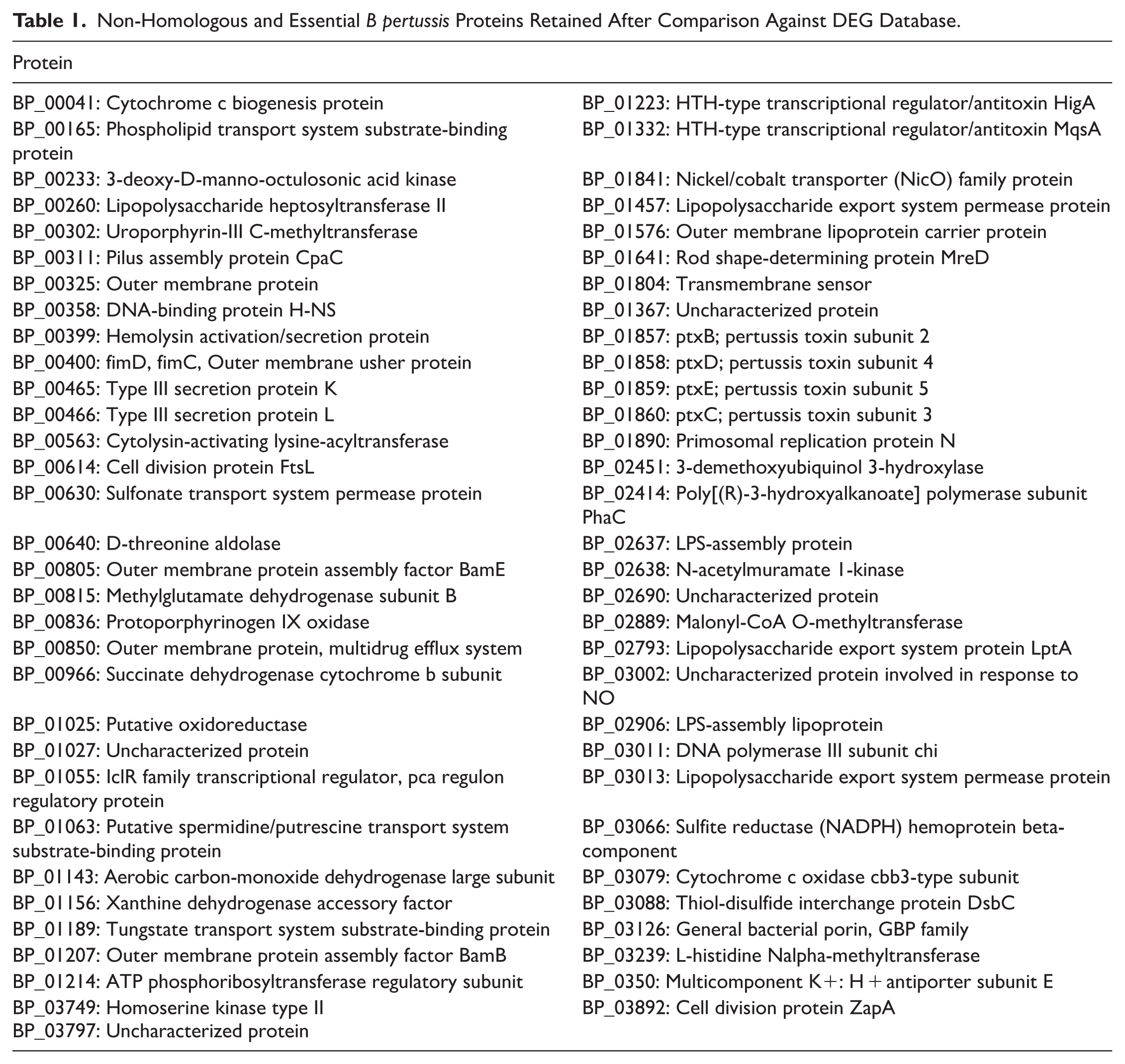

Prioritization of Vaccine Targets
Sub-Cellular Localization
Subcellular localization is a key functional attribute of a protein, as many cellular processes are compartmentalized; thus, predicting the localization of unknown proteins can offer valuable insights into their roles and assist in selecting candidates for further study. 31 In bacteria, all proteins are synthesized in the cytoplasm, where many remain to perform specific functions, while others contain export signals that direct them to different cellular compartments. In Gram-negative bacteria, compartments include the cytoplasmic membrane, periplasm, inner and outer membranes, and extracellular space. 28 Although most proteins are confined to a single compartment, some span multiple locations.
The subcellular localization of 63 non-homologous essential proteins of

A multi-selection screening pipeline for the characterization and prioritization of 63 essential proteins using subcellular localization, signal peptide, transmembrane topology, physicochemical, and antigenicity prediction tools (Made using Napkin).
Subcellular localization is a critical factor in identifying effective drug and vaccine targets; proteins localized in the cytoplasm or membrane are often considered for drug development, while those secreted or exposed on the bacterial surface are ideal candidates for vaccines. 46 The next step involves refining these candidates further through multilevel analysis of physicochemical parameters.
Physicochemical Parameters
We initially screened the secreted or extracellularly exposed proteins of
Analysis of Physicochemical Properties (Molecular Mass, pI, Stability, GRAVY), Subcellular Localization, Transmembrane Helix Number, Signal Peptide Prediction, and Antigenicity.

Prediction of Antigenicity and Immunogenicity
Antigenicity
The proteins classified as secreted or PSE were analyzed using VaxiJen software. This analysis revealed that all 11 proteins have a probability above the threshold of 0.4, indicating that they are immunogenic and capable of generating antibodies, thus making them strong candidates for vaccine targets (Figure 3 and Table 2).
Immunogenicity
Peptide vaccines are more convenient and safer than conventional vaccines, as they include only immunogenic epitopes rather than the full 3-dimensional protein structure. B-cell epitopes are antigenic determinants that are recognized and bound by receptors on the surface of B lymphocytes. A total of 103 linear B-cell epitopes were identified across the 11 examined proteins of
Number of Predicted Linear and Conformational Epitopes for the 11 Candidate Targets.

Discussion
Face to decline of vaccine efficacy and increase of pertussis cases reported around the world, it was suggested the necessary of finding novel therapeutic strategy to improve the vaccines and to combat this resurgence. The purpose of this study was to identifying new vaccine targets against
The pangenomics approach, which is applied in this study, successfully reduced the number of candidate proteins from 1010 to 11. These proteins were selected as vaccine targets for treating
Two pertussis toxin (Ptx) subunits, S2 (BP_01857) and S5 (BP_01859), are widely recognized as major virulence factors expressed by
Two LPS assembling proteins (BP_02637 and BP_02906) are highly appeared to be the novel vaccine targets, replacing LPS, which are the major component of DTwP and the main responsible for the reactogenicity. 47 Assembling proteins, such as LptD, whose immunogenicity was demonstrated in a study done by Dorji, showed that mice vaccinated with LptD alone exhibited significantly lower bacterial loads compared with those vaccinated solely with DTaP. 48
The outer membrane protein assembly factors BamE (BP_00805) and BamB (BP_01207) are components of the β-barrel assembly machinery (BAM) complex. This complex facilitates the insertion and folding of β-barrel proteins into the outer membrane (OM) of Gram-negative bacteria. 52 Targeting the BAM complex holds potential not only for vaccine development but also for developing potentiators that enhance OM permeability. The immunogenicity of BamB was demonstrated in vivo by Dorji, who showed that mice vaccinated with BamB alone exhibited significantly lower bacterial loads compared with those vaccinated solely with DTaP. 48 However, the immunogenicity of BamE remains to be fully evaluated and warrants further investigation.
Outer membrane protein (BP_03126) belonging to the general bacterial porin (GBP) family are found in the outer membrane of Gram-negative bacteria, which is responsible for antibiotics permeability.
53
Among the proteins of this family in
Hemolysin activation/secretion protein, FhaC (BP_00399),is an outer membrane protein that acts as a transporter in the 2-partner secretion pathway (TPS). Its essentiality has been confirmed in the secretion of filamentous hemagglutinin (FHA). 54 Furthermore, Xu and Hu have noted that secretion systems have changed under vaccine pressure, and they suggested that targeting FhaC is an innovative approach. 55 Therefore, it is suggested that the immunogenicity of FhaC needs to be demonstrated.
The outer membrane usher protein, FimC (BP_00400), is predicted to be an integral outer membrane (OM) β-barrel usher protein involved in fimbriae (Fim2/3) assembly.
56
Despite the critical role of FimC in fimbriae assembly and its potential as a novel vaccine target to disrupt fimbriae assembly and reduce
The outer membrane protein (BP_00850), part of the multidrug efflux system, is a relevant element within the microbial repertoire that contributes to multidrug resistance (MDR) and the failure of anti-infectious therapies
57
; thus, it is considered an attractive target for the development of new vaccines. It has been reported that the multidrug efflux system of
Protein involved in the biogenesis of cytochrome c (BP_00041), an essential component of bacterial energy metabolism, is critical for cellular function. The entire system appears to operate on the outer surface of the cytoplasmic membrane, 59 which limits its potential as a direct vaccine target, but it could be exploited for antimicrobial strategies and used as a potential drug target.
Conclusions
Unlike classical vaccinology methods, which rely on in vitro bacterial cultivation for antigen identification, reverse vaccinology leverages genomic analysis to predict vaccine targets more rapidly and cost-effectively, while enhancing specificity and minimizing risks of autoimmune reactions. These predicted targets subsequently undergo immunogenicity assessments to precisely characterize the elicited immune responses, thereby facilitating advanced testing of potential candidates. In this study, 160
Supplemental Material
sj-docx-1-bbi-10.1177_11779322261442522 – Supplemental material for Pangenomics-Driven Reverse Vaccinology for the Discovery of New Vaccine Candidates Against Bordetella pertussis
Supplemental material, sj-docx-1-bbi-10.1177_11779322261442522 for Pangenomics-Driven Reverse Vaccinology for the Discovery of New Vaccine Candidates Against Bordetella pertussis by Majid Abdelouahab and Diawara Idrissa in Bioinformatics and Biology Insights
Supplemental Material
sj-docx-2-bbi-10.1177_11779322261442522 – Supplemental material for Pangenomics-Driven Reverse Vaccinology for the Discovery of New Vaccine Candidates Against Bordetella pertussis
Supplemental material, sj-docx-2-bbi-10.1177_11779322261442522 for Pangenomics-Driven Reverse Vaccinology for the Discovery of New Vaccine Candidates Against Bordetella pertussis by Majid Abdelouahab and Diawara Idrissa in Bioinformatics and Biology Insights
Supplemental Material
sj-docx-3-bbi-10.1177_11779322261442522 – Supplemental material for Pangenomics-Driven Reverse Vaccinology for the Discovery of New Vaccine Candidates Against Bordetella pertussis
Supplemental material, sj-docx-3-bbi-10.1177_11779322261442522 for Pangenomics-Driven Reverse Vaccinology for the Discovery of New Vaccine Candidates Against Bordetella pertussis by Majid Abdelouahab and Diawara Idrissa in Bioinformatics and Biology Insights
Footnotes
Consent and Ethics
Not applicable (this is an in silico study only).
Author Contributions
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
