Abstract
Providencia rettgeri has increasingly been responsible for several infections, including urinary tract, post-burn wounds, neonatal sepsis, and others. The emergence of drug-resistant isolates of P rettgeri, accompanied by intrinsic and acquired antibiotic resistance, has exacerbated the challenge of treating such infections, necessitating the development of novel therapeutics. Hypothetical proteins (HPs) form a major portion of cellular proteins and can be targeted by these novel therapeutics. In this study, 410 HPs from a pan-drug-resistant (PDR) P rettgeri strain (MRSN845308) were functionally annotated and characterized by physicochemical properties, localization, virulence, essentiality, druggability, and functionality. Among 410 HPs, the VirulentPred 2.0 tool and VICMpred combinedly predicted 33 HPs as virulent, whereas 48 HPs were highly interacting proteins based on the STRING v12 database. BlastKOALA and eggNOG-mapper v2.1.12 predicted 13 HPs involved in several metabolic pathways like Riboflavin metabolism and Lipopolysaccharide biosynthesis. Overall, 83 HPs were selected as primary drug targets; however, only 80 remained after nonhomology searching and essentiality analysis. In addition, all were detected as novel drug targets according to DrugBank 5.1.12. Considering the potential of membrane and extracellular proteins, 29 HPs (extracellular, outer, and inner membrane) were selected based on the combined prediction from PSORTb v3.0.3, CELLO v.2.5, BUSCA, SOSUIGramN, and PSLpred. According to the prevalence of those HPs in different strains of P rettgeri sequences in National Center for Biotechnology Information Identical Protein Groups (NCBI-IPG), 5 HPs were selected as final drug targets. In addition, 5 other HPs annotated as transporter proteins were also added to the list. As no crystal structures of our targets are present, 3-dimensional structures of selected HPs were predicted by the AlphaFold Server powered by AlphaFold 3. Our findings might facilitate a better understanding of the mechanism of virulence and pathogenesis, and up-to-date annotations can make uncharacterized HPs easy to identify as targets for novel therapeutics.
Introduction
Providencia rettgeri is a Gram-negative opportunistic bacterium belonging to the genus “Providencia.” It is now classified under the Morganellaceae family, although it previously belonged to the Enterobacteriaceae family. Like some members of the Enterobacteriaceae family, such as Klebsiella pneumoniae, Acenetobacter baumannii, and Serratia marcescens, P rettgeri is commonly involved in hospital-acquired infections.1,2 Providencia rettgeri mainly causes travelers’ diarrhea and catheter-related urinary tract infections.3,4 Recently, it has also been reported in several other infections, including post-burn wound infections, meningitis, and neonatal sepsis.5 -7 The increasing involvement in different types of infections is particularly a problem to public health as drug-resistance is nothing unfamiliar to P rettgeri. It has intrinsic resistance to colistin and tigecycline as well as first-generation cephalosporins making it insensitive to many antibiotic treatments.8,9
Other than intrinsic resistance, the acquisition of antibiotic resistance has also been well-documented in P rettgeri. In 2000, carbapenem-resistant P rettgeri was first identified in western Japan. Since then, various antibiotic-resistance genes carrying P rettgeri strains have been identified worldwide. 10 A P rettgeri isolate harboring the blaNDM-1 gene was reported in Israel in 2013. 11 These isolates, carrying resistance genes, have been involved in several outbreaks in recent years. Providencia rettgeri strain co-producing New Delhi metallo-β-lactamase-1 (NDM-1) carbapenemase and Pseudomonas Extended Resistance-1 (PER-1) extended-spectrum β-lactamase (ESBL) was documented for causing nosocomial outbreaks during 2016 to 2017 in South Korea. 9 Similarly, in 2020, an outbreak mediated by multidrug-resistant (MDR) P rettgeri has come into concern in South-eastern Nigeria. 12
The transfer of resistance through plasmids can lead to more frequent outbreaks of P rettgeri as plasmids carrying antibiotic-resistant genes are frequently detected in P rettgeri. The presence of plasmids carrying blaIMP-11 or blaIMP-70 was detected by sequencing P rettgeri isolates. 10 In Japan, unique plasmids carrying the blaIMP-70 and blaCTX-M-253 genes were found in MDR P rettgeri strains, as reported in 2022. 13 The availability of plasmid loaded with antibiotic-resistance genes elevates the risk of MDR P rettgeri turning into pan-drug-resistant (PDR) under persistent selective pressure resulting from the widespread use of antibiotics. Recently, a PDR P rettgeri MRSN845308 was noted to cause severe fatality in a COVID-19 patient. 8 The strain’s extensive resistance to all classes of antibiotics resulted in multiorgan failure, ultimately leading to the patient’s death on hospital day 66. If the emergence of such PDR P rettgeri is left unnoticed, the current trend of drug failure in the treatment of P rettgeri is unlikely to change.
Hence, a study was designed with the aim of identifying novel therapeutic targets for the treatment of PDR P rettgeri infections. New drugs that are effective against PDR isolates would likely be functional against MDR and Extensively Drug-Resistant (XDR) isolates. Therefore, the best practice in designing drugs or finding drug targets would be taking PDR isolates as target pathogens. Besides, most practices to unearth drug targets are based on analysis of defined or known proteins leaving a significant portion of targets unexplored. To our concern, no study thus far has explored the possibility that the hypothetical proteins (HPs) of P rettgeri can potentially be the targets of developing and already developed drugs. Considering the potential, we have explored the HPs of the recently sequenced PDR P rettgeri strain MRSN845308 for drug targets. In this study, we employed a range of computational algorithms to deduce the potential functions of HPs from the PDR P rettgeri strain MRSN845308. Our analysis includes in silico assessments of these HPs, focusing on both the molecular and structural levels, aiming to identify promising therapeutic targets for drug design and development. Through computational methodologies, we sought to elucidate the roles and characteristics of these proteins, paving the way for targeted approaches in combating antibiotic resistance and advancing antimicrobial strategies.
Materials and Methods
Sequence retrieval
The complete genome of P rettgeri strain MRSN845308 was retrieved on April 10, 2024, from the National Center for Biotechnology Information (NCBI) (http://www.ncbi.nlm.nih.gov/) with accession number JAEEFV000000000.1. According to the Average Nucleotide Identity (ANI)-based NCBI taxonomy check, the strain is attributed to Providencia hangzhouensis. The sequences of HPs were mined using the keyword “hypothetical protein” by comparing with the reference (GCF_016512695.1) annotation (referenced on December 2, 2023). Their Uniprot accession IDs were retrieved using the Uniprot database. Proteins with more than 100 amino acids were considered for further analysis (Figure 1).

Flowchart of methodology.
Physicochemical properties of hypothetical proteins
The physicochemical characteristics of all the HPs were investigated using Expasy’s ProtParam server (https://web.expasy.org/protparam/). 14 This analysis facilitated the determination of key parameters, including molecular weight, isoelectric point, 15 instability index, 16 aliphatic index, and grand average of hydropathicity (GRAVY). 17 The GRAVY score, calculated as the sum of hydropathy values divided by the number of residues in the sequence, indicates the overall hydrophobicity of a peptide or protein.
Cellular and subcellular localization and protein classification
The topology of membrane proteins was predicted using TOPCONS, a web server designed for consensus prediction of membrane protein topology (https://topcons.cbr.su.se/pred/). 18 In addition, CCTOP (https://cctop.ttk.hu), 19 TMHMM v.2.0 (https://services.healthtech.dtu.dk/services/TMHMM-2.0/), 20 and the SignalP 5.0 server (https://services.healthtech.dtu.dk/services/SignalP-5.0/) 21 were used to identify the cellular locations of these proteins.
To determine the subcellular localization of the HPs, we employed PSLpred (https://webs.iiitd.edu.in/raghava/pslpred/submit.html), which integrates Position-Specific Iterated BLAST (PSI-BLAST) and 3 SVM (Support Vector Machine) modules based on residue compositions, dipeptides, and physicochemical properties, 22 BUSCA (https://busca.biocomp.unibo.it), 23 CELLO v.2.5 (http://cello.life.nctu.edu.tw), 24 SOSUIGramN (https://harrier.nagahama-i-bio.ac.jp/sosui/sosuigramn/sosuigramn_submit.html), 25 and PSORTb version 3.0.3 (https://www.psort.org/psortb/). 26
Functional domain/motif prediction
Functional domain prediction aims to identify conserved segments within proteins, as these domains often represent functional units. In the case of the HPs, we used several publicly available databases for domain prediction, including ScanProsite (https://prosite.expasy.org/scanprosite/), 27 NCBI-CDD (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi), 28 InterProScan 5 (https://www.ebi.ac.uk/interpro/search/sequence/), 29 and PFP-FunDSeqE (http://www.csbio.sjtu.edu.cn/bioinf/PFP-FunDSeqE/). 30
Function prediction
Accurately annotating protein function is essential for comprehending molecular-level life processes. In our study, we predicted the Gene Ontology (GO) terms, specifically Molecular Function and Biological Process, associated with the HPs. With this aim, Argot2 (Annotation Retrieval of Gene Ontology Terms) (https://www.medcomp.medicina.unipd.it/Argot2/) 31 and eggNOG-mapper v2.1.12 (http://eggnog-mapper.embl.de) were used. 32
Pathway analysis of hypothetical proteins
The HPs were analyzed using the tool BlastKOALA (https://www.kegg.jp/blastkoala/) from the Kyoto Encyclopedia of Genes and Genomes (KEGG) database to predict the pathways regulated by these proteins. 33
Interactional network analysis of hypothetical proteins
We conducted a protein interaction network analysis to identify HPs with significant protein-protein interactions (PPIs) using the STRING database version 12 (https://string-db.org). 34
Virulence protein prediction
Identifying virulent proteins within bacterial sequences is valuable for assessing pathogenic potential and elucidating the intricate mechanisms of virulence in pathogenesis. To achieve the desired goal, we employed the VirulentPred 2.0 tool (https://bioinfo.icgeb.res.in/virulent2/index.html/) and VICMpred (https://webs.iiitd.edu.in/raghava/vicmpred/index.html) to identify virulence factors among the HPs.35,36
Nonhomology analysis by Basic Local Alignment Search Tool for proteins human proteome
In our analysis, HPs from P rettgeri present in the metabolic pathways, proteins with higher PPIs, and proteins involved in virulence were compared with the proteome of Homo sapiens (human; taxid 9606) using BLASTp (Basic Local Alignment Search Tool for proteins) (https://blast.ncbi.nlm.nih.gov/Blast.cgi). It allows us to identify similarities and differences in protein sequences between the bacterial HPs and the human proteome. Proteins that showed BLAST hits with an expectation value (E-value) less than 0.001 were considered homologous to the human proteome. Conversely, proteins without significant BLAST hits (E-value ⩾ 0.001) were deemed specific to P rettgeri, suggesting potential species-specific functions or roles. 37
Essentiality analysis
Essentiality analysis for the selected nonhomologous HPs was performed using BLASTp against a Database of Essential Genes (DEG; http://origin.tubic.org/deg/public/index.php/blast/bacteria). 38
Druggability analysis
We assessed the druggability of nonhomologous HPs by investigating their potential to interact with known drug candidates. Each nonhomologous HP underwent a homology search using the DrugBank 5.1.12 Target collection database (https://go.drugbank.com/structures/search/bonds/sequence#results). 39
Homologs of known antibiotic-resistance determinants
The Comprehensive Antibiotic Resistance Database (CARD) (https://card.mcmaster.ca/analyze/blast) was used to search for homologs of known antibiotic-resistance determinants of HPs. 40
Abundance of hypothetical proteins
To identify and prioritize potential drug targets among HPs, we conducted an abundance analysis using the NCBI Identical Protein Groups database (IPG) (https://www.ncbi.nlm.nih.gov/ipg). 41
Structural modeling of functionally important hypothetical proteins
In our study, protein structural prediction was conducted using the AlphaFold Server powered by AlphaFold 3 (https://alphafoldserver.com). 42
Results
Selection of hypothetical proteins
Using P rettgeri strain MRSN845308 from the NCBI Genome database, 573 proteins were found to be HPs containing specific Uniprot accession IDs. Among all the HPs, 410 proteins were found to be longer than 100 amino acids and were therefore considered suitable for further analysis.
Functional domain/motif and physicochemical characteristics prediction
Based on the NCBI-CDD database, we identified 111 proteins with conserved domains (Supplementary File 1.1). InterProScan shows 67 proteins have functional domains (Supplementary File 1.2). ScanProsite, which matches proteins against the Prosite collection of motifs, identified 44 functional regions, and others showed no hit. PFP-FunDSeqE predicted protein fold pattern with functional domain for all the proteins, as shown in Figure 2 (Supplementary File 2). Physicochemical characteristics showed that 183 proteins were found to be basic (pI ⩾ 7), whereas 227 proteins were found to be acidic (pI < 7). The GRAVY, amino acid composition, molecular weight, instability index, and aliphatic index were also calculated; the details are given in Supplementary File 3.

Domain/motif prediction by PFP-FunDSeqE.
Cellular and subcellular localization of hypothetical proteins
Several bioinformatics tools were used to determine the presence of transmembrane helices. TOPCONS predicted 23 proteins with transmembrane helices. Specifically, the TMHMM algorithm, known for its accuracy in distinguishing between soluble and membrane proteins using a Hidden Markov Model, identified 108 HPs containing transmembrane helices. CCTOP identified 99 proteins with transmembrane segments, and SignalP identified 82 HPs containing transmembrane residues (Supplementary File 4).
For subcellular localization, we employed PSORTb, CELLO v.2.5, BUSCA, SOSUIGramN, and PSLpred tools. By taking consensus from all, we have identified 281 sequences in the cytoplasm, 45 proteins in the extracellular space, 63 as inner membrane proteins, 6 as outer membrane proteins, and 15 predicted to be in the periplasmic space (Supplementary File 5).
Functional classification of hypothetical proteins
For functional annotation at the cellular, molecular, and biological levels, Argot2 was used. It predicted biological processes of 372 HPs (Figure 3), cellular functions of 366 HPs (Figure 4), and molecular functions of 406 HPs (Figure 5). More than 24 broad classes of molecular functions were identified at the molecular level. The most significant group of proteins was predicted to be involved in transferase activity, followed by DNA, ATP, Ion, other binding proteins, and hydrolase activity. In biological processes, 17 broad categories were identified, with most proteins primarily associated with DNA-associated metabolic processes, transportation, signal transduction, folding, biosynthesis, cell division, and stress resistance. With respect to cellular components, 15 broad functional categories were identified, as presented in Figure 3 (Supplementary File 6.1).

Figure representing the Gene Ontology of hypothetical proteins: biological process.

Figure representing the Gene Ontology of hypothetical proteins: cellular components.
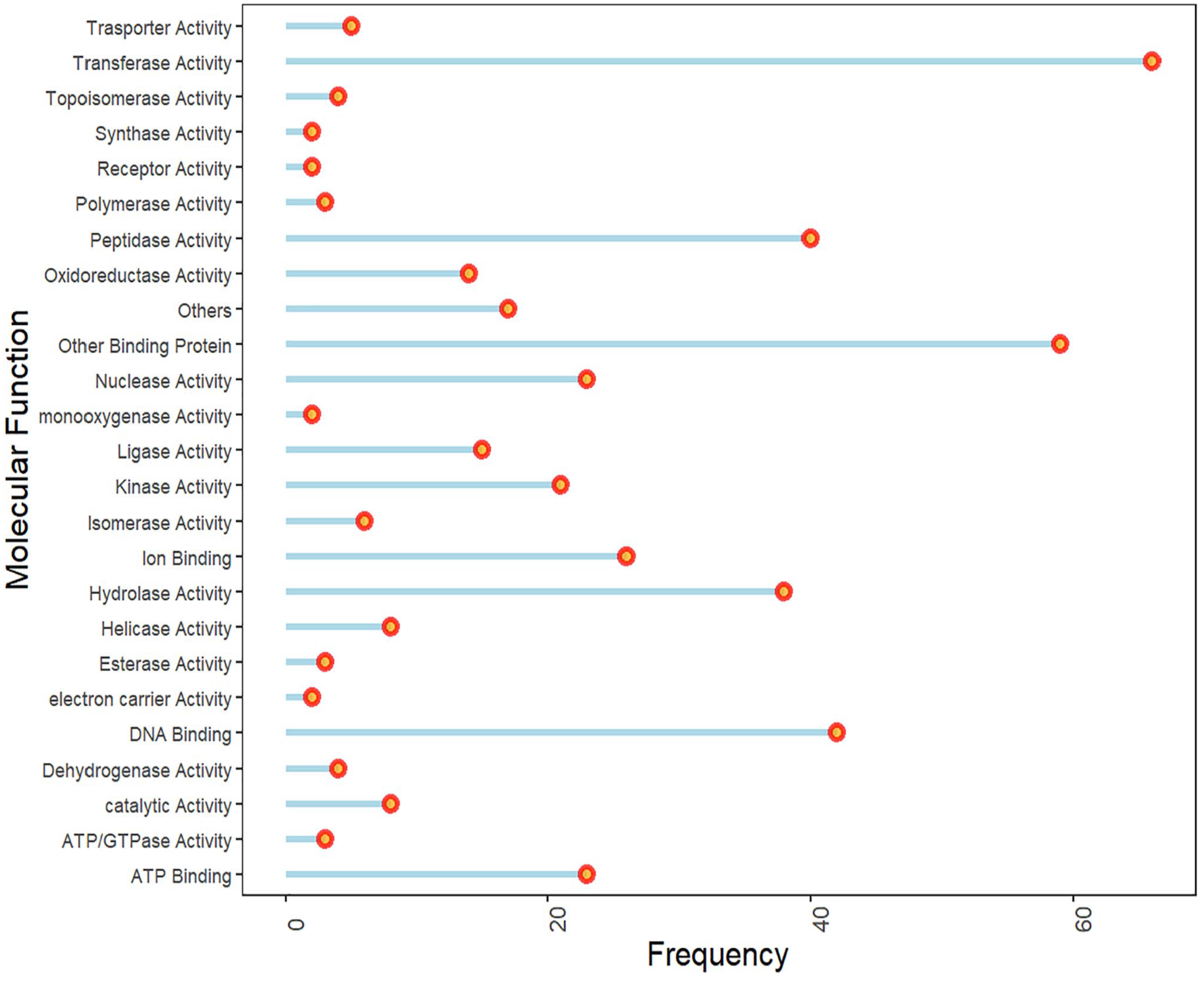
Figure representing the Gene Ontology of hypothetical proteins: molecular function.
Pathway analysis of functionally characterized proteins
The functional roles of HPs in various pathways were predicted using the tool BlastKOALA and KO (KEGG ORTHOLOGY) from the KEGG database and with eggNOG-mapper v2.1.12 to elucidate their regulatory functions (Supplementary File 6.2). Our analysis identified 13 proteins involved in different metabolic pathways (Table 1).
Hypothetical proteins interacting with functionally important proteins.

Protein-protein interaction analysis
Proteins exhibiting highly interactive functional associations were investigated using the STRING database, identifying 48 proteins with strong interacting relationships at a confidence level exceeding 0.400. Among them, 4 highest interacting clusters were shown in Figure 6 along with their details provided in Table 1 (Supplementary File 7).

Protein-protein interaction of hypothetical proteins.
Virulence factor analysis of hypothetical proteins
To identify virulent proteins associated with bacterial pathogenesis, VirulentPred 2.0 tool and VICMpred were used. Based on the prediction from both tools, 33 HPs were classified as potential virulence factors (Supplementary File 8).
Selection of hypothetical proteins as putative drug targets
A total of 83 HPs, including 13 HPs associated with metabolic pathways, 33 HPs associated with virulence proteins, and 48 HPs associated with PPIs were primarily selected as drug targets. Among them, 3 proteins were duplicated in both pathway and PPI, 7 duplicated in virulent and PPI, and 1 duplicated in virulent and pathway. Homology to humans was predicted using BLASTp against the human proteome (taxid: 9606) available in the NCBI database. Among the 83 HPs, only 2 showed homology with the reference proteome (E-value < 0.001). 43
Essentiality and druggability analysis
Among the 81 nonhomologous HPs identified, 80 were determined to be essential for pathogen survival based on a similarity search conducted in the DEG database (Supplementary File 9). The 80 essential nonhomologous HPs were then assessed for druggability through a similarity search against targets listed in the DrugBank database. On manual inspection, it was found that none of the 80 essential nonhomologous HPs exhibited similarity to a target in the DrugBank database (E-value = 0.00001), classifying them as novel drug targets (Supplementary File 9).
Homologs of known antibiotic-resistance determinants and their abundance
According to the CARD database, 66 of the total 80 essential nonhomologous novel HPs showed similarity to different antibiotic-resistance determinants. In addition, the abundance of those 80 proteins in different strains of P rettgeri based on NCBI-IPG database searching has been recorded in Supplementary File 9.
Structural modeling of functionally important hypothetical proteins
The identification and structural characterization of HPs are essential for selecting potential targets in drug design. 44 Among the 80 essential novel nonhomologous HPs analyzed, 29 proteins were primarily selected based on their subcellular locations (extracellular, outer membrane, and inner membrane). Subsequently, from these 29 proteins, the top 5 most abundant HPs (MBJ9971398.1, MBJ9971402.1, MBJ9971395.1, MBJ9971147.1, and MBJ9973330.1) were identified from the NCBI-IPG database and their tertiary structures were modeled using AlphaFold Server powered by AlphaFold 3. In addition, 5 other HPs (MBJ9971209.1, MBJ9972277.1, MBJ9972284.1, MBJ9973148.1, and MBJ9973050.1) involved in various transporter activities were also selected for structure prediction (Table 2 and Figure 7). The details of their physicochemical characteristics are provided in Table 3.
Hypothetical proteins annotated as transporter by Argot2.

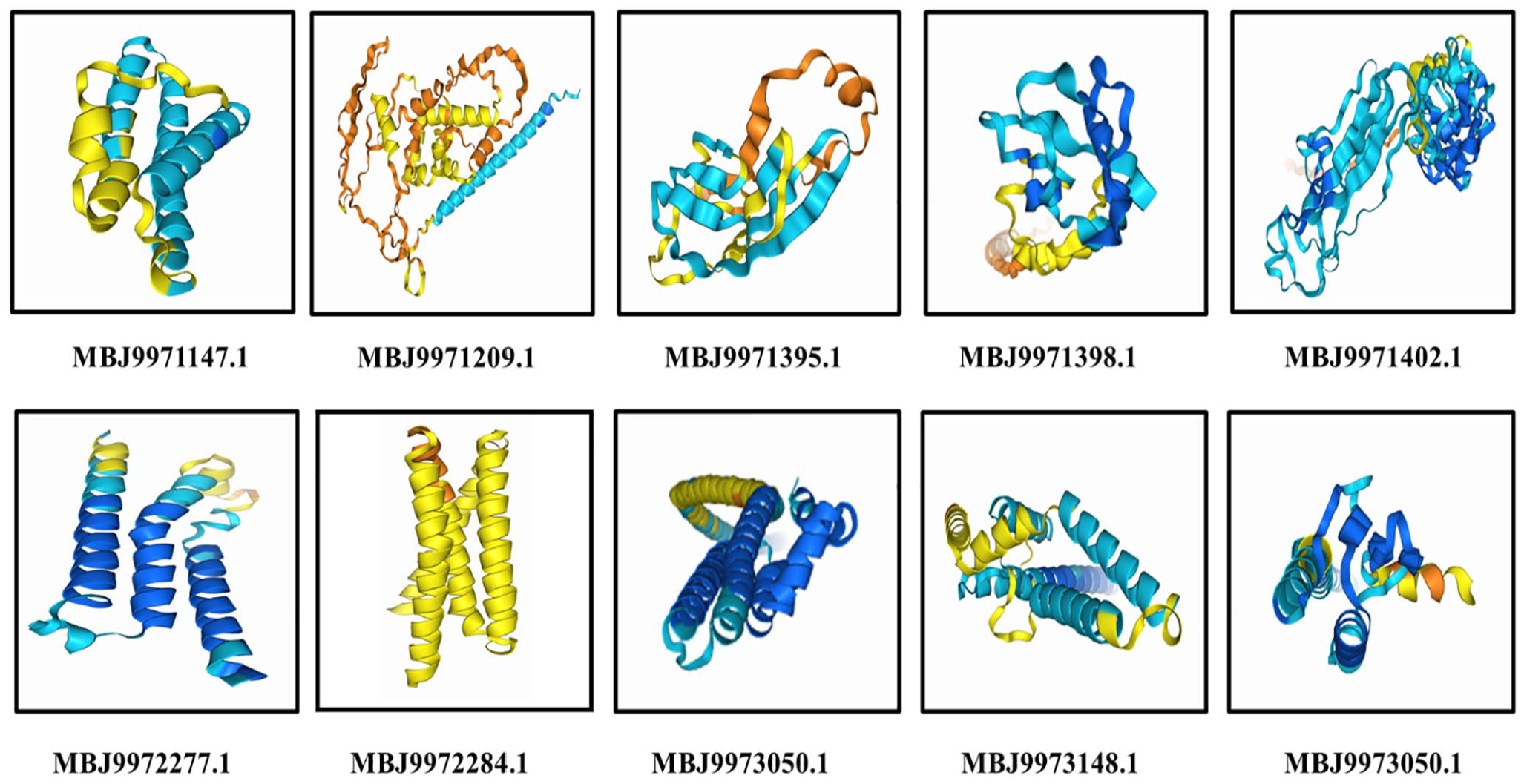
The 3D structure of the 10 target proteins are modeled by AlphaFold Server powered by AlphaFold 3.
Physicochemical characteristics of the selected drug targets.

Discussion
In this study, our target was to characterize HPs of a PDR P rettgeri strain MRSN845308, with an emphasis on identifying potential drug targets. The sequence of the selected strain was obtained from the NCBI Genome database. The genome of the strain encodes 4,405 proteins, including 3832 (87%) defined proteins and 573 (13%) uncharacterized (hypothetical) proteins. In a study conducted by Wang et al, 45 the HP was around 30% in a strain of P rettgeri annotated by Prokka. The lower percentage of HPs in the selected strain is due to the use of the Prokaryotic Genome Annotation Pipeline (PGAP) by NCBI, which has up-to-date and vast databases for annotation. 44 Hypothetical proteins shorter than 100 amino acids, categorized as pseudogenes, were omitted from this analysis to enhance the accuracy of functional predictions. Identifying functional domains can be invaluable in selecting drug targets. Proteins identified as bacteriophage proteins, including viral coat and capsid proteins, were excluded from the downstream analysis to ensure that the results are specific to bacterial proteins. The functional or conserved domains were explored by the NCBI-CDD database, InterProScan, PFP-FunDSeqE, and ScanProsite.
The cellular, molecular, and biological roles of the HPs were predicted by Argot 2. This website predicted cellular functions, biological processes, and molecular functions of 410 HPs. There was a high prevalence of binding proteins that attach to DNA, RNA, protein, ions, or other metabolites. DNA-binding proteins play an important role in survival and stress tolerance and may also be responsible for virulence.46,47 Metal-binding proteins can serve as cofactors in a range of functions, such as controlling metabolism, regulating signal transport, and maintaining metal homeostasis. 48 Five HPs were annotated as transporter proteins. Transporter proteins that act as efflux pumps can help bacteria prevent the entry of antibiotics. 49 Some proteins were identified as outer membrane porins, which are crucial for the survival of Gram-negative bacteria. Several antibiotics enter the cell through the pores formed by porins, and by regulating these pores, bacteria can acquire drug-resistance. Their presence is also crucial to maintain membrane integrity. 50 Some HPs function as enzymes contributing to virulence and antibiotic resistance. They also play key roles in metabolic pathways, the synthesis of cell organelles, and bacterial multiplication, among other cellular processes.51 -53 We also found several proteins characterized as toxins, inhibitors, and immunogens. These proteins can play a significant role in bacterial pathogenesis.
Pathway analysis provides additional insights into the potential roles of these proteins within biological processes. A pathway analysis based on predictions from the KEGG database and eggNOG-mapper (v2.1.12) revealed that 13 HPs are involved in different metabolic pathways. Two important metabolic pathways involve riboflavin metabolism and lipopolysaccharide biosynthesis, affected by MBJ9973193.1 and MBJ9973676.1, respectively. In Riboflavin metabolism, MBJ9973193.1 HP may play a vital role in the synthesis or utilization of riboflavin (Vitamin B2), which is essential for cellular metabolism. In Lipopolysaccharide biosynthesis, the identified MBJ9973676.1 may be involved in the biosynthesis of lipopolysaccharides, crucial components of Gram-negative bacterial outer membranes. Nonhomology searching suggests that the 2 HPs associated with these pathways share no homology to human proteins. Hence, they can be targeted by drugs without affecting metabolic pathways in humans.
Protein-protein interaction analysis revealed that 48 proteins had strong interacting relationships with a confidence level above 0.400. This threshold sets a minimum required interaction score, ensuring that only interactions with higher confidence are included in the predicted network. Lower scores mean more interaction but also higher false positive results. The HP MBJ9971369.1 was found to interact with the type III secretion (T3SS) systems, suggesting a role in virulence. The T3SSs are protein transport nanomachines that play a role in Gram-negative bacterial virulence and pathogenesis. 54 Owing to its role in host-pathogen interaction, the T3SS has been an attractive target for the development of new antibacterial therapies. Hence, MBJ9971369.1 can also be a potential target for drugs attacking T3SS systems. 43 Other HPs with a high number of interactions belong to families such as signal transduction response regulators, the tRNA endonuclease-like domain superfamily, protein secretion and bacterial export proteins, DNA integration, and transmembrane transport proteins. Biological networks serve as pivotal starting points for studying and addressing numerous human diseases. 55 Gram-negative pathogens often undergo evolutionary adaptations to enhance their virulence within the host environment, manifesting in alterations to traits such as motility, cell adhesion, and evasion of host immune responses. A consensus sequence analysis conducted by VICMpred and VirulentPred classified 43 (7.05% of HPs) proteins as potential virulence factors. These virulent proteins are involved in bacterial pathogenesis, including responses to stress and signaling pathways, as well as cellular functions such as enzymatic activity and transportation. The targeting of virulence factors has been increasingly recognized as a promising therapeutic approach against bacterial pathogenesis, offering an alternative to conventional antibiotics or as adjuncts to enhance host immune responses. 56
Gram-negative pathogens often undergo evolutionary adaptations to enhance their virulence within the host environment, manifesting in alterations to traits such as motility, cell adhesion, and evasion of host immune responses. A consensus sequence analysis conducted by VICMpred and VirulentPred classified 33 proteins (8.1%) as potential virulence factors. A slightly higher percentage (18.02%) of virulence proteins were classified from HPs of Orientia tsutsugamushi in a study by Imam et al. 57 These virulent proteins are involved in bacterial pathogenesis, including responses to stress and signaling pathways, as well as cellular functions such as enzymatic activity and transportation. The targeting of virulence factors has been increasingly recognized as a promising therapeutic approach against bacterial pathogenesis, offering an alternative to conventional antibiotics or as adjuncts to enhance host immune responses. 56
A total of 83 HPs from interaction, virulence, and pathway analysis was our primary drug target. Targets similar to human proteins may induce immunogenic reactions and, hence, often are eliminated from the lists of potential drug targets. Among the 83 HPs, only 2 showed homology with human proteins and thus were not considered in further analysis. Necessarily, the drug targets should be essential to the target pathogens, and as such, the essentiality of the 81 HPs was determined by the DEG database. A total of 80 HPs were found to be essential for pathogen survival. The HPs that match any target with similar biological functions from the list of nonhomologous HPs in the DrugBank database indicate potential druggability. Alternatively, if no match is found, the target is considered a “Novel target,” suggesting a unique opportunity for drug discovery and development. 58 All were detected as novel targets, which suggest that a large number of HPs as novel drug targets are still unexplored.
The localization of HPs in the cell is also an important factor when considering drug accessibility. Although intracellular drug targets may be less accessible, the extracellular targets are more exposed and have more potential to serve as drug aims. Besides, the extracellular proteins are often critical components of the biofilm matrix, and few attempts have been taken to target them for antimicrobial drugs.59 -61 Membrane proteins (inner or outer) represent a substantial proportion of current drug targets due to their roles in signal transduction, transport, bacterial pathogenicity, and drug-resistance. 62 Extracellular and outer membrane proteins have an extra advantage as being viable targets for vaccine development. 63 Considering the potential of membrane and extracellular proteins, 29 HPs (extracellular, outer, and inner membrane) were selected based on the combined prediction from PSORTb, CELLO v.2.5, BUSCA, SOSUIGramN, and PSLpred. The prevalence of HPs in the P rettgeri population is also an important consideration for drug target evaluation. The targeted HPs abundance was compared with the mean abundance of the selected HPs using the NCBI-IPG database. The NCBI-IPG database is a resource within the NCBI that aims to simplify protein searches by grouping together protein sequences that are 100% identical. Based on the high abundance in NCBI-IPG, the top 5 HPs were selected as drug targets that are available in most P rettgeri isolates. The MBJ9971398.1 was the most prevalent one and was present in 216 strains, followed by MBJ9971402.1 (189 strains), MBJ9971395.1 (150 strains), MBJ9971147.1 (119 strains), and MBJ9973330.1 (111 strains). These highly abundant HPs can be targeted for developing broad-spectrum drugs that will be effective against most strains of P rettgeri.
Transporter proteins can be potential drug targets and have been targeted by antibacterial vaccines and antibiotics.64,65 However, 5 HPs annotated as transporter proteins by Argot 2 were not found in HPs associated with virulence, metabolic pathways, and PPI. Considering their potential as drug targets, the HPs annotated as transporter proteins were also added to the list. The 5 transporter proteins, including MBJ9973148.1, MBJ9972277.1, MBJ9972284.1, MBJ9971209.1, and MBJ9973050.1, were present in 42, 15, 10, 2, and 2 strains, respectively. Although these transporter proteins are not that abundant, they can be targeted for developing strain-specific drugs. As the selected HPs have no crystal structure, determining their 3-dimensional (3D) structure is required to understand their function and suitability as targets for drug design. Accordingly, their 3D structures were modeled using the AlphaFold Server powered by AlphaFold 3. The recently developed artificial intelligence (AI)-based AlphaFold 3 can predict the structure and interactions of proteins and other molecules with unprecedented accuracy. 42 All these analyses in our work might help facilitate narrowing down targets and accelerate the drug screening process against P rettgeri infections.
Conclusions
This study facilitates the prompt identification of unknown functions associated with HPs using a variety of bioinformatics web tools and databases. The rigorous functional and spatial characterization of HPs in this study might unveil promising targets for therapeutic intervention. The 3D modeling of these targets will help identify the optimal binding sites that can be targeted for drug design by molecular docking and simulation. Virtual screening against these target proteins might be effective in the discovery of novel therapeutic compounds against P rettgeri. Further studies are, hence, required to address the therapeutic potential of these HPs and validate their suitability as drug targets through experimental approaches.
Supplemental Material
sj-xlsx-1-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-1-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-10-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-10-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-11-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-11-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-2-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-2-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-3-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-3-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-4-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-4-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-5-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-5-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-6-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-6-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-7-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-7-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-8-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-8-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Supplemental Material
sj-xlsx-9-bbi-10.1177_11779322241280580 – Supplemental material for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets
Supplemental material, sj-xlsx-9-bbi-10.1177_11779322241280580 for Identification and Functional Annotation of Hypothetical Proteins of Pan-Drug-Resistant Providencia rettgeri Strain MRSN845308 Toward Designing Antimicrobial Drug Targets by Dipta Chandra Pal, Tasnimul Arabi Anik, Atiq Abrar Rahman and S M Mahfujur Rahman in Bioinformatics and Biology Insights
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
DCP conceived and designed the experiment. DCP and TAA conducted the primary investigation and literature review. DCP, TAA, AAR, and SMMR performed data collection. DCP and TAA conducted data validation, formal analysis, and interpreted the results. DCP, TAA, AAR, and SMMR wrote the manuscript. DCP and TAA primarily revised and edited the manuscript. All authors reviewed and approved the final manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
