Abstract
Narciclasine is an alkaloid belonging to the Amaryllidaceae family which has been reported to have many beneficial properties. Especially its anticancer properties have been widely reported. Here, we have focused on its potential use in suppressing the inflammatory response in sepsis using in silico methods. Lipopolysaccharide (LPS) is an endotoxin which is present in the outer membrane of gram-negative bacteria and is a crucial player in the pathogenesis of gram-negative sepsis. Activation of toll-like receptor 4 (TLR4) signaling by LPS is an important event in the pathogenesis of gram-negative sepsis. This initiates a downstream signaling pathway comprising of several adaptor proteins such as toll/interleukin-1 receptor domain-containing adapter protein (TIRAP), myeloid differentiation primary response protein 88 (MyD88), interleukin-1 receptor–associated kinase (IRAK)-1, IRAK-4, interferon regulatory factor 3 (IRF-3), tumor necrosis factor receptor–associated factor 6 (TRAF-6) leading to nuclear factor kappa B (NF-κβ) activation resulting in elevated production of inflammatory cytokines such as tumor necrosis factor alpha (TNF-α) and interleukin (IL)-6. S100 calcium binding proteins A8/A9 (S100A8/A9) have been found to be an agonist of TLR4, and it amplifies the inflammatory response in sepsis. Molecular docking studies of narciclasine with target proteins associated with the LPS-TLR4 pathway showed that it has good binding affinity and stable interactions with the targets studied. Molecular dynamics (MD) simulation studies over 100 ns showed that most of the ligand-target complexes were stable. The structures of all the targets except TRAF-6 were retrieved from the Protein Data Bank (PDB) database. Homology modeling was done to predict the 3-dimensional structure of TRAF-6. MD simulation of narciclasine-TRAF-6 complex showed that the structure is stable. Metapocket was used for active site prediction in the target proteins. Toxicity analysis by admetSAR revealed that narciclasine was readily biodegradable and exhibited minimum toxicity. These results indicate that narciclasine has effective anti-inflammatory properties which could be useful in suppressing the inflammatory response in sepsis.
Keywords
Introduction
Sepsis can be defined as a life-threatening organ dysfunction that occurs due to dysregulated host response to infection. 1 Despite recent advancements in antibiotics and critical care, it is one of the major causes of death in hospitalized patients affecting around 18 million people worldwide annually. 2 It draws huge costs every year and still therapeutic interventions are far from satisfactory. 3 Blood culture is the gold standard for confirmation of diagnosis. Antibiotics and supportive care is currently the mainstay of treatment for sepsis. Blood culture results takes time to come and the administration of broad spectrum antibiotics is common which has led to the evolution of more multi-drug-resistant strains. A good number of culture negative cases turn out to be actually positive as in many cases no specific organism can be identified in culture. 4 These culture negative cases account for high mortality and thus alternate strategies such as the use of anti-inflammatory drugs for sepsis could be therapeutically beneficial.
After infection, the initiation of host inflammatory response is through the recognition of pathogen-associated molecular patterns (PAMPs) in microbes by pattern recognition receptors on phagocytes. Phagocytes could also be activated by endogenous danger signals known as damage-associated molecular patterns (DAMPs) or alarmins. Among them, S100 calcium binding protein A8/A9 (S100A8/A9) complex which is also known as calprotectin are important endogenous DAMPs. They are found to be released from phagocytes during inflammatory stimuli and have proinflammatory properties and amplify the inflammatory response. The S100A8/A9 proteins are found to be endogenous activators of TLR4 and are found to enhance lethal endotoxin induced shock. 5
Among sepsis cases, gram-negative sepsis is quite prevalent which accounts for high mortality every year. 6 Lipopolysaccharide (LPS) is an endotoxin present in the outer membrane of gram-negative bacteria and is considered to induce shock and organ failure in gram-negative sepsis. 7 The LPS is sensed through a lipopolysaccharide-binding protein (LBP) complex and then signaling happens through the TLR4-MD2 complex. The downstream intracellular signaling involves several key adaptor proteins such as myeloid differentiation primary response protein 88 (MyD88), toll/interleukin-1 receptor domain-containing adapter protein (TIRAP), interleukin-1 receptor–associated kinase 1 (IRAK-1), IRAK-4, interferon regulatory factor 3 (IRF-3), and tumor necrosis factor receptor–associated factor 6 (TRAF-6). This downstream signaling pathway leads to the activation and translocation of NF-κβ dimer to the nucleus where it induces the transcription of various target genes including those for production of inflammatory cytokines. 8 The excessive production of inflammatory cytokines during sepsis may worsen the outcome causing capillary leakage, tissue injury, and multiple organ failure. 2 Septic patients had higher levels of plasma tumor necrosis factor alpha (TNF-α) and interleukin (IL)-1β which correlated with the severity of illness. 9 Cytokines such as TNF-α and IL-1β peak within the first few hours of sepsis, thus providing a narrow therapeutic window.
Narciclasine is an isocarbostyril alkaloid belonging to the Amaryllidaceae family. Narciclasine was found to be the main bioactive component in Haemanthus coccineus extracts and is found to be responsible for its anti-inflammatory actions. We have earlier reported the therapeutic potential of narciclasine in neonatal rats with sepsis. 10 Here, in this study, we have done molecular docking of narciclasine with target proteins of the LPS-TLR4 pathway followed by molecular dynamics (MD) simulation of selected docked complexes for 100 ns. We found that narciclasine showed good binding affinity and stable interactions with most of the selected targets. The drug likeness of narciclasine was evaluated using Lipinski filter and quantitative estimate of drug likeness (QED). Toxicity analysis was done using admetSAR.
Material and Methods
The 3-dimensional (3D) structure of target proteins viz, TLR4, CD14, MD2, calprotectin, MyD88, IRAK-1, IRAK-4, TIRAP, IRF-3, NF-κβp50, NF-κβp65, TNF-α, and IL-6 were retrieved from the Protein Data Bank (PDB) database. The chemical structure of narciclasine (ligand) CID 72376 was retrieved from PubChem database. The 3D structure of TRAF-6 was predicted by homology modeling and validated using PDBsum server and ERRAT analysis. Active-binding sites were identified using Metapocket 2.0 server. Docking was done using AutoDock, and the docked complexes were visualized using Discovery Studio Visualizer 2016. Molecular dynamics simulation was performed using GROMACS. The drug likeness was determined using Lipinski filter and QED analysis. Toxicity of narciclasine was analyzed using admetSAR. The overall methodology followed is depicted as a flowchart in Figure 1.

Flowchart showing the methodology followed for this study. CD14 indicates cluster of differentiation 14; IL-6, interleukin 6; IRAK, interleukin-1 receptor–associated kinase; IRF-3, interferon regulatory factor-3; MD2, myeloid differentiation factor 2; MyD88, myeloid differentiation primary response protein 88; NF-κβ, nuclear factor kappa B; PDB, Protein Data Bank; TIRAP, TIR domain-containing adaptor protein; TLR4, toll-like receptor 4; TNF-α, tumor necrosis factor alpha.
Ligand retrieval
The 3D structure of narciclasine (CID 72376) was retrieved from the PubChem database (Figure 2). 11 The ligand was energy minimized using Marvin software.

Structure of narciclasine (A) 2D (B) 3D structure (ball and stick representation).
Retrieval of target protein structures
The structures of following target proteins were downloaded from the Research Collaboratory for Structural Bioinformatics (RCSB) PDB (http://rcsb.org) which is an archive containing all the PDB data. 12 They are TLR4 (3FXI, DOI: 10.2210/pdb3FXI/pdb), calprotectin (1XK4, DOI: 10.2210/pdb1XK4/pdb), MD2 (2E56, DOI:10.2210/pdb2E56/pdb), CD14 (4GLP, DOI: 10.2210/pdb4GLP/pdb), MyD88 (4DOM, DOI: 10.2210/pdb4DOM/pdb), IRAK-1 (6BFN, DOI: 10.2210/pdb6BFN/pdb), IRAK-4 (2NRU, DOI: 10.2210/pdb2NRU/pdb), TIRAP (4FZ5, DOI: 10.2210/pdb4FZ5/pdb), IRF-3 (2O6G, DOI: 10.2210/pdb2O6G/pdb), NF-κβ p50 (1NFK, DOI: 10.2210/pdb1NFK/pdb), NF-κβ p65 (1NFI, DOI: 10.2210/pdb1NFI/pdb), TNF-α (2AZ5, DOI: 10.2210/pdb2AZ5/pdb), and IL-6 (1ALU, DOI: 10.2210/pdb1ALU/pdb).
Template identification and homology modeling of human tumor necrosis factor receptor–associated factor 6
Homology modeling is used to determine the 3D structure of a given protein sequence based mainly on its alignment to one or more proteins of known structure (templates). Homology modeling was done to generate the 3D structure of TRAF-6 (Uniprot ID: Q9Y4K3).
For protein modeling, Iterative Threading Assembly Refinement (I-TASSER) was used. I-TASSER is a hierarchical protocol for automated protein structure prediction and structure-based function annotation. 13 I-TASSER modeling begins from the structure templates recognized by LOMETS from the PDB library. The top 10 template-query alignments generated by LOMETS are mentioned in Table 1.
Templates used.
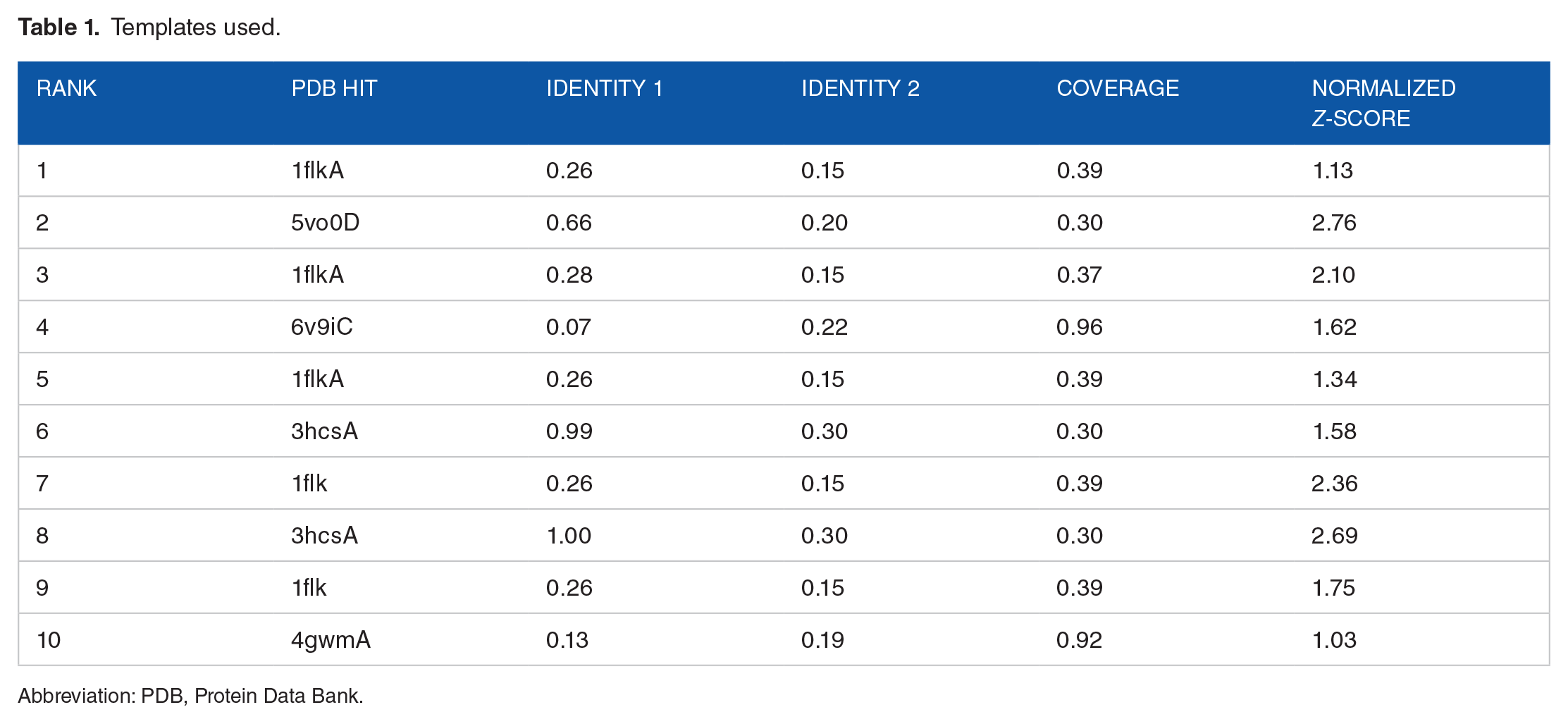
Abbreviation: PDB, Protein Data Bank.
Templates of high significance were measured by the Z-score, which is the difference between the raw and average scores in the unit of standard deviation. Z-score > 1 indicate a good alignment. The normalized Z-score of the threading alignments indicated a good alignment.
Structural details of target protein model
The final model selected by I-TASSER is based on the pair-wise structure similarity. C-score implies the confidence of each model and is calculated based on the significance of threading alignments and the convergence parameters of the structure assembly simulations. Usually, the C-score is in the range of [−5, 2] and a C-score of a higher value means a model with higher confidence or vice versa.
Model validation
The predicted TRAF-6 model was validated using Ramachandran Plot (RC plot) and ERRAT. 14 The RC plot was generated using PDBsum (which used PROCHECK for the plot generation). 15 The RC plot shows the torsional angles—phi (φ) and psi (ψ)—of the residues (amino acids) in the structure. The various regions on the plot are represented by the different coloring. The core regions signifying the most favorable phi-psi combinations are the darkest areas (here shown in red). Ideally, it is expected to have more than 90% of the residues in these “core” (favored) regions. The percentage of residues in the “core” region is one of the better guides to stereo chemical quality. Also, residues in the disallowed region should ideally be less than or equal to 0.2%.
ERRAT analyses
ERRAT is to analyze the statistics of nonbonded interactions between different atom types. The value of the error function vs position of a 9-residue sliding window is plotted.
Active site prediction
Before performing docking analysis, active site prediction of the target proteins (TLR4, CD14, MD2, calprotectin, MyD88, IRAK-1, IRAK-4, TIRAP, IRF-3, TRAF-6, NF-κβp50, NF-κβp65, TNF-α, and IL-6) was done using Metapocket 2.0 server. 16 The top 3 major binding pockets were retrieved for analysis of active binding residues and for comparison of the docking results (see Supplementary Information).
Docking analysis
Drugs generally work by interacting with protein receptors. The binding affinity of narciclasine with various targets was found using protein-ligand docking. The 3D structure of target proteins was downloaded from the RCSB PDB (http://rcsb.org). The chemical structure of narciclasine (CID 72376) was obtained from PubChem compound database. 11 Marvin software was used for energy minimization of the ligand. AutoDock 4.0 (autodock.scripps.edu/) tool was used for Docking. AutoDock uses the Lamarckian genetic algorithm for obtaining the interaction regions of residues in feasible sites of binding. The ligand was docked against the active sites of targeted proteins. The best lowest energy state of the receptor-ligand complexes was identified.
Molecular dynamics simulation studies
Molecular dynamics simulation of the protein-ligand complexes (narciclasine with TLR4, calprotectin, TIRAP, MyD88, IRF-3, and IRAK-1) was performed using GROMACS (2022.3). GROMACS is a very fast program for MD simulation. 17 From the docking results, the ligand-target complex structure was obtained and subjected to MD simulation (100 ns). Molecular dynamics simulation trajectories included protein-ligand complex root-mean square deviation (RMSD), protein-RMSD, and ligand-RMSD. Visualization of these MD simulation results was done with XMGRACE, plot-analyzing GUI software.
Toxicity analysis
For toxicity analysis, we have used the database “admetSAR.” 18 This gives simple manually curated data for wide variety of chemicals associated with known Absorption, Distribution, Metabolism, Excretion, and Toxicity (ADMET) profiles. Main features involved in toxicity analysis are drug-induced toxicity, drug-induced liver injury, rat acute toxicity, skin sensitivity, genomic toxicity, carcinogens, rodent (human, rat, mouse, hamster, etc) animals’ toxicity, biodegradability, and bioconcentration factors.
Analysis of drug likeness of narciclasine
The drug-likeness prediction was done using Lipinski filter according to which an orally active drug should comply with at least 4 of the 5 mentioned criteria for drug likeness. They are molecular mass, hydrogen bond donor and acceptor, Log P, and molar refractive index. 19
In Lipinski analysis, poor absorption or permeation is implied when the molecule has more than 5H-bond donors, more than 10H-bond acceptors, molecular weight more than 500, and Log P > 5. In addition to the 4 rules defined by Lipinski, the extended rule of 5 takes rotatable bond counts into account. This implementation considers a rotatable bond count greater than 10 as rule violation. 20 The Konstanz Information Miner (KNIME) was used to find the pass or fail for Lipinski rule of 5. 21 The physicochemical properties were predicted by SWISSADME. 22
In addition to the Lipinski rule of 5, QED was estimated using RDKit (Open-Source Cheminformatics Software). The SMILES format of narciclasine compound was downloaded from PubChem (https://pubchem.ncbi.nlm.nih.gov/) and given as input to RDKit for calculating QED.
The properties of narciclasine with respect to prediction of adsorption, distribution, metabolism, excretion, and toxicity were analyzed using admetSAR. 18
Results
Homology modeling—structural details of the target protein model
The final model selected by I-TASSER is based on the pair-wise structure similarity. The model has an estimated TM-score = 0.55 ± 0.15 and estimated RMSD = 10.5 ± 4.6 Å calculated based on C-score which is −1.33 and protein length following the correlation observed between these qualities, as shown in the Table 2.
Quality of model.
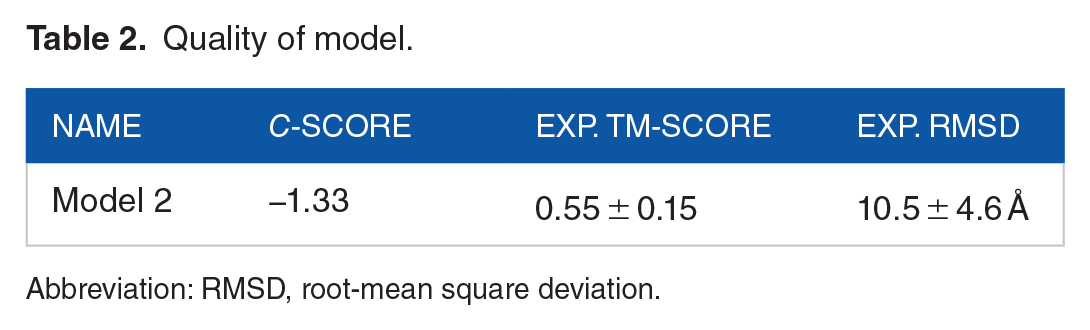

Abbreviation: RMSD, root-mean square deviation.
The 3D image of the final protein model as visualized with Discovery Studio is shown in Figure 3.

The 3D protein structure image of TRAF-6 predicted by homology modeling, viewed in Discovery Studio. TRAF-6 indicates tumor necrosis factor receptor–associated factor 6.
Model validation—Ramachandran plot and ERRAT
The quality of the 3D model was analyzed by the RC plot using PROCHECK software. The RC plot for the selected final model is shown in Figure 4. The RC plot statistics is shown in Supplementary Information file (Table S1).
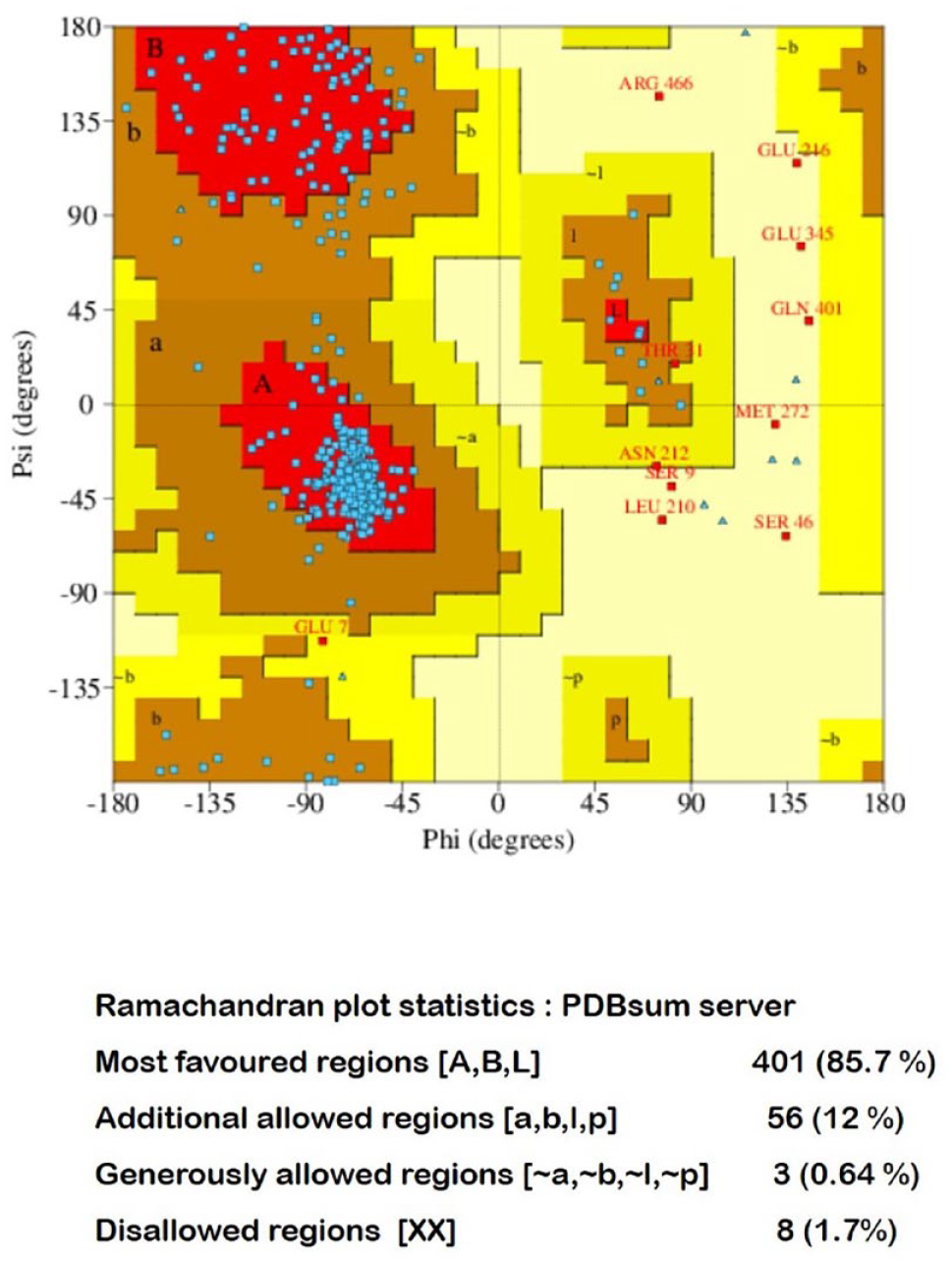
Ramachandran plot for the model using PDBsum (PROCHECK). PDB indicates Protein Data Bank.
The generated model of TRAF-6 was visualized using Discovery Studio Visualizer 2016. The RC plot for the generated model showed that 85.7% of residues to be in the most favored regions, whereas 11.96% were in the allowed region. This shows that the predicted model is of good quality.
ERRAT analyses
ERRAT is used to assess the overall quality factor for nonbonded atomic interactions where higher scores point to higher quality. The generally accepted range is more than 50% for a high-quality model. For the current 3D model structure, 74.708% of an overall quality was observed (Figures S1 and S2 in Supplementary Information). The plot generated indicates the confidence and overall quality of the model.
Simulation studies of narciclasine-tumor necrosis factor receptor–associated factor 6 complex
The energy minimization was done, and potential energy for TRAF-6 is shown in Figure 5A. The Epot obtained for TRAF-6 was −5.18651 × 106 kJ/mol. The plot in Figure 5A shows that complex TRAF-6 was energy minimized and is stable in potential energy. After energy minimization, equilibration was done and temperature reached to a target value of 300 K and remained stable over the equilibration. The average temperature obtained for TRAF-6 was 299.994 K (Figure 5B). The protein TRAF-6 achieved stability in 1000 ps (1 ns) of approximately 300 K. The temperature equilibration stabilizes the temperature of the system. The pressure stabilization should also be done. The pressure value fluctuates widely over the course of the 1000-ps equilibration phase and over the equilibration. The average pressure value obtained for TRAF-6 was −2.30652 bar which show that the structure is equilibrated (Figure 5C). Thus, the energy minimization and equilibration (temperature and pressure) process ensures the structure is stable or we can say that the system is well equilibrated. The final simulation was done using GROMACS. The changes in structure were checked in RMSD and radius of gyration. After analyzing the RMSD plot (Figure 5D), we can say that over the period of 1 ns (1000 p s), protein TRAF-6 is showing its RMSD in the required range (0.1–0.7 nm). Also, the graph may go lower in RMSD if studied for longer timeframe. Figure 5E shows that the radius of gyration is relatively stable for the protein TRAF-6, as graph is quite stable and does not show much variation.

Simulation of TRAF-6 protein. (A) Potential energy graph obtained from energy minimization using GROMACS. (B) Temperature graph obtained after temperature equilibration. (C) Pressure equilibration graph obtained after equilibration. (D) RMSD plot obtained for protein (relative to backbone). (E) Radius of Gyration plot obtained for protein. RMSD indicates root-mean square deviation; TRAF-6, tumor necrosis factor receptor–associated factor 6.
Docking analysis of narciclasine
AutoDock 4.0 was used in this study, and the docked complexes were visualized using Discovery Studio Visualizer 2016. The docking result is shown in Table 3.
Docking result for narciclasine with targets—TLR4, MD2, CD14, MyD88, TIRAP, IRAK-1, IRAK-4, IRF3, NF-κβp50, NF-κβp65, IL-6, TNF-α, calprotectin, and TRAF-6..

Abbreviations: CD14 indicates cluster of differentiation 14; IL-6, interleukin 6; IRAK, interleukin-1 receptor—associated kinase; IRF-3, interferon regulatory factor 3; MD2, myeloid differentiation factor 2; MyD88, myeloid differentiation primary response protein 88; NF-κβ, nuclear factor kappa B; TIRAP, TIR domain-containing adaptor protein; TLR4, toll-like receptor 4; TNF-α, tumor necrosis factor alpha; TRAF-6, tumor necrosis factor receptor–associated factor 6.
The prominent binding sites were predicted using Metapocket 2.0 server (see Supplementary Information file). Docking of narciclasine with target compounds of the LPS-TLR4 pathway was studied with respect to the following parameters: (1) interacting amino acids, (2) ligand and protein molecules involved in H bonding, (3) predicted docking and binding energy, (4) inhibition constant, (5) ligand efficiency, (6) sum of van der Waals energy, hydrogen bond energy, and desolvation energy (Vdw_hb_desolv_energy), (7) electrostatic energy, (8) total internal energy, (9) torsional energy, and (10) unbound energy.
The 3D interaction image between narciclasine and the various target compounds shows the hydrogen bonds involved in the docking interactions. In the protein-ligand interactions, the hydrogen bonds in association with other noncovalent interactions such as van der Waals forces, ionic interactions, and hydrophobic interactions play an important role. The docked complexes were visualized by Discovery Studio Visualizer to display various interactions involved in the ligand-target docking (Figures 6 to 8).

Visualization of docked complexes. Figure shows the 2D interaction images and 3D interaction images for TLR4, MD2, CD14, TIRAP, and TRAF-6 with narciclasine using Discovery Studio (dashed green line in 3D interaction images shows the hydrogen bonds). The molecular surface model complex, secondary structure model complex, and stick model complex showing interaction between narciclasine and targets. CD14 indicates cluster of differentiation 14; MD2, myeloid differentiation factor 2; TIRAP, TIR domain-containing adaptor protein; TLR4, toll-like receptor 4; TRAF-6, tumor necrosis factor receptor–associated factor 6.

Visualization of docked complexes. Figure shows the 2D interaction images and 3D interaction images for MyD88, IL-6, IRAK-1, IRAK-4, and IRF-3 with narciclasine using Discovery Studio (dashed green line in 3D interaction images shows the hydrogen bonds). The molecular surface model complex, secondary structure model complex, and stick model complex showing interaction between narciclasine and targets. IL-6 indicates interleukin 6; IRAK, interleukin-1 receptor−associated kinase; IRF-3, interferon regulatory factor-3; MyD88, myeloid differentiation primary response protein 88.

Visualization of docked complexes. The molecular surface model complex, secondary structure model complex, and stick model complex showing interaction between narciclasine and targets—calprotectin, TNF-α, NF-κβp50, and NF-κβp65 using AutoDock tools. NF-κβ indicates nuclear factor kappa B; TNF-α, tumor necrosis factor alpha.
The H-bond receptor surface, hydrophobic receptor surface, ionizability receptor surface, and solvent accessibility receptor surface showing interaction between the targets with narciclasine using Discovery Studio are depicted in Figures 9 and 10.

Visualization of docked complexes. Aromatic receptor surface, charge receptor surface, H-bond receptor surface, hydrophobic receptor surface, ionizability receptor surface, and solvent accessibility receptor surface showing interaction between targets—TLR4, MD2, CD14, TIRAP, and TRAF-6 with narciclasine, using Discovery Studio. CD14 indicates cluster of differentiation 14; MD2, myeloid differentiation factor 2; TIRAP, TIR domain-containing adaptor protein; TLR4, toll-like receptor 4; TRAF-6, tumor necrosis factor receptor–associated factor 6.

Visualization of docked complexes. Aromatic receptor surface, charge receptor surface, H-bond receptor surface, hydrophobic receptor surface, ionizability receptor surface, and solvent accessibility receptor surface showing interaction between targets—MyD88, IRAK-1, IRAK-4, IRF-3, and IL-6 with narciclasine, using Discovery Studio. IL-6 indicates interleukin 6; IRAK, interleukin-1 receptor–associated kinase; IRF-3, interferon regulatory factor-3; MyD88, myeloid differentiation primary response protein 88.
Molecular dynamics simulation studies
Molecular dynamics simulation has been performed to find the stability of the ligand narciclasine with the targets (TLR4, TIRAP, MyD88, IRAK-1, IRF-3, and calprotectin) through the analysis of a variety of trajectories that accurately predict the stability of the ligands bound to the protein during the simulation. They include ligand-target complex RMSD, protein-RMSD, and ligand-RMSD. The RMSD plot of the ligand-target complex shows the stability of the ligand inside the binding pocket of target over the course of a 100-ns simulation. The RMSD trajectories of the docked complexes formed by narciclasine with TLR4, TIRAP, MyD88, and IRF-3 were found to be stable after 10 ns (Figures 11 and 12). The narciclasine-calprotectin (1XK4) complex (Figure 12B) was found to be only partially stable with irregular fluctuations throughout the simulation process. The narciclasine-IRAK-1 protein-ligand interactions were stable (Figure 12C). However, narciclasine-IRAK-1 complex (Figure S8 in Supplementary Information) was found to be not stable. Figures 11 and 12 denote the multiple trajectories of the simulation process that run for 100 ns.

Analysis of RMSD trajectories for the ligand (narciclasine)-target protein complexes throughout 100 ns MD simulation. (A) TLR4 (protein-ligand RMSD and complex RMSD). (B) TIRAP (protein-ligand RMSD and complex RMSD). (C) MyD88 (protein-ligand RMSD and complex RMSD). MyD88 indicates myeloid differentiation primary response protein 88; RMSD, root-mean square deviation; TIRAP, TIR domain-containing adaptor protein; TLR4, toll-like receptor 4.

Analysis of RMSD trajectories for the ligand (narciclasine)-target protein complexes throughout 100 ns MD simulation. (A) IRF3 (protein-ligand RMSD and complex RMSD). (B) Calprotectin (protein-ligand RMSD and complex RMSD). (C) IRAK-1 (protein-ligand RMSD). IRF-3 indicates interferon regulatory factor; RMSD, root-mean square deviation; IRAK-1, interleukin-1 receptor–associated kinase 1.
Testing of drug likeness of narciclasine
The drug likeness of narciclasine was analyzed by Lipinski filter and also by QED. The results generated after the Lipinski filter analysis, and the physicochemical properties as predicted by SWISSADME are mentioned in Table 4. Findings were found to be in agreement with the Lipinski rule of 5and passed through the filtering analysis.
Physiochemical properties as predicted by SWISSADME.

Quantitative estimate of drug-likeness analysis showed that narciclasine had a score of 0.43 which suggests a reasonable degree of drug likeness. 23 The absorption, distribution, metabolism, and excretion (ADME) report revealed the solubility and absorption efficiency of narciclasine in blood and various organs of the body (Table 5).
ADME report.

Abbreviation: ADME, absorption, distribution, metabolism, and excretion.
Toxicity analysis was done using admetSAR. 18 The results are shown in Table 6.
Toxicity analysis of narciclasine using admetSAR.

Abbreviation: AMES, Bacterial reverse mutation test; hERG, human ether-a-go-go-related gene.
The detailed report of toxicity analysis along with description can be found in the Supplementary Information file (Table S2).
Discussion
Despite advancements in medical care, sepsis still remains a serious clinical problem accounting for high mortality in the intensive care units worldwide.24,25 The plants belonging to Amaryllidaceae family possess wide range of biological activities and have been the subject of pharmacologic investigations. 26 Narciclasine is an isocarbostyril alkaloid present in H coccineus extracts of the Amaryllidaceae family. Our previous study on neonatal rats with sepsis showed that narciclasine could effectively suppress the excessive inflammatory response and improve the outcome. 10 Here, we found using molecular docking that narciclasine could effectively bind with various targets of the LPS-TLR4 pathway associated with gram-negative bacterial sepsis. All of these targets studied are interconnected with one another and have a specific role in contributing to the aberrant inflammatory response during sepsis.
Despite several underlying reasons for the onset of sepsis, bacteremia is considered a major factor that influences the outcome. 27 Recent studies have shown that gram-negative bacteremia was significantly higher in the patients with septic shock in intensive care unit (ICUs). 28 Gram-negative bacteria have a much stronger cell membrane than gram-positive bacteria, thus making them more difficult to treat.
The endotoxin LPS is an important component of the outer membrane of gram-negative bacteria and is a crucial player in the pathogenesis of bacterial sepsis. 29 Toll-like receptor 4 plays a crucial role in the detection of LPS of gram-negative bacteria. The activation of TLR4 signaling pathways by LPS is an important event in the pathogenesis of gram-negative bacterial sepsis. 29 Thus, as TLR4 occupies a fundamental role in the pathogenesis of gram-negative sepsis. It is a target of choice for developing novel antisepsis therapies. In our study, we found that narciclasine forms stable complexes with TLR4 and the low binding energy of −6.71 kcal/mole shows that narciclasine could bind effectively with TLR4. It was also evident from the MD simulation studies that RMSD graph (TLR4-narciclasine complex) over 100 ns is stable and does not show much variation (Figure 11A).
The S100A8 and S100A9 proteins are present in the cells of the myelomonocytic lineage and are predominantly present in the cytoplasm of neutrophils and monocytes. When they are activated, the S100A8/A9 heterodimer get translocated to the membrane and other cytoskeletal structures. During inflammatory conditions, its levels are found to be elevated in circulation. 30 Plasma calprotectin was found to be useful early marker of bacterial infection in critically ill patients. 31 Calprotectin levels were found to be higher in serum of patients with bacterial sepsis than the healthy controls. 32 The S100A8/A9 complex acts as an endogenous activator of TLR4 and amplifies the inflammatory responses. Thus, we proposed that calprotectin could be a potential target for antisepsis therapies. The docking result of calprotectin (1XK4) with narciclasine showed a binding energy of −6.56 kcal/mol. The low binding energy shows stable interaction between calprotectin and narciclasine. However, MD simulation studies over 100 ns had continuous variations implying that the narciclasine-calprotectin complex is only partially stable (Figure 12B).
The recognition of LPS by innate immune cells is crucial for providing protection to the host against gram-negative bacteria. This involves activation of several adaptor proteins which is initiated by the recognition of LPS by LBP which is an acute phase protein in the circulating bloodstream. 29 This then binds to a glycosylphosphatidylinositol (GPI)-linked protein—CD14, which is present on the cell surface of phagocytes. Lipopolysaccharide then gets transferred to MD2 which is present in the extracellular portion of TLR4. 33 In this study, we found that narciclasine formed stable complexes with CD14 and MD2 which was evident from the low binding energy of −6.34 and −6.82 kcal/mole, respectively. The binding of LPS to the MD2-TLR4 complex causes TLR4 dimerization and activates the signaling cascade comprising of the toll-interleukin receptor (TIR) domain-containing adaptor molecules such as MyD88 and TIRAP. 33 In our study, molecular docking results of narciclasine with MyD88 and TIRAP showed a low binding energy of −8.52 and −8.08 kcal/mole, respectively, which reveals that narciclasine is showing good activity with MyD88 and TIRAP and exhibits stable interactions. The RMSD graphs (for TIRAP-narciclasine and MyD88-narciclasine complex) over 100 ns were stable and does not show much variation (Figure 11B and C).
Myeloid differentiation primary response protein 88 associates with the cytoplasmic portion of TLR and recruits IRAK-1 and IRAK-4. In TLR4 signaling, another adaptor protein TIRAP is needed to recruit MyD88 to the receptor. After IRAK-1 interacts with MyD88, it is phosphorylated by activated IRAK-4 which subsequently gets associated with TRAF-6. 33 In our study, molecular docking results of narciclasine with IRAK-1, IRAK-4, and TRAF-6 showed a very low binding energy of −8.57, −8.35, and −7.10 kcal/mole, respectively. This shows that narciclasine exhibits stable interactions with these targets. Interferon regulatory factor 3 is another transcription factor activated downstream in the TLR4 pathway and has been found to have an important role in the induction of interferon beta (IFN-β) in sepsis. 34 The RMSD graph for IRF3-narciclasine complex over 100 ns was stable and does not show much variation (Figure 12A). The narciclasine-IRAK-1 protein-ligand interactions were stable (Figure 12C), but the complex was not stable (Supplemental Figure S8). This is followed by the activation of inhibitor of nuclear factor-κB (IκB) kinase (IKK) complex. The activation of IKKs leads to the phosphorylation and degradation of Iκβ that subsequently leads to the activation of NF-κβ signaling pathways. This causes the release and nuclear translocation of multiple NF-κβ dimers, although the main target of Iκβα is the p65/p50 heterodimer. 35 The nuclear translocation of NF-κBp65/p50 dimer results in the transcription of several target genes such as those for inflammatory cytokines. The LPS-TLR4 pathway associated with the immunopathogenesis of gram-negative sepsis is depicted in Figure 13.

The LPS-TLR4 pathway associated with the pathogenesis of gram-negative sepsis. The potential of narciclasine to bind to various targets of the pathway as evidenced from our docking results is also represented in the figure. CD14 indicates cluster of differentiation 14; CP: calprotectin; IL-6, interleukin 6; IRAK, interleukin-1 receptor–associated kinase; IRF-3, interferon regulatory factor-3; LBP, lipopolysaccharide-binding protein; LPS, lipopolysaccharide; MD2, myeloid differentiation factor 2; MyD88, myeloid differentiation primary response protein 88; NC: narciclasine; NF-κβ, nuclear factor kappa B; TIRAP, TIR domain-containing adaptor protein; TLR4, toll-like receptor 4; TNF-α, tumor necrosis factor alpha; TRAF-6, tumor necrosis factor receptor–associated factor 6.
Toll-like receptor–mediated NF-κβ signaling pathway is activated in sepsis and has crucial role in the inflammatory response. 36 Nuclear factor kappa B activation has a fundamental role in inflammatory disorders and sepsis and leads to aberrant outcomes. 37 In response to various stimuli such as the cytokines, the Iκβα dissociates and translocates to the nucleus where it induces various target genes related to apoptosis and inflammation. 38 The suppression of NF-κβ activation has shown to alleviate the inflammatory response and organ damage in sepsis. In our study, the docking result of narciclasine with NF-κβp50 and NF-κβp65 showed a low binding energy of −6.75 and −7.21 kcal/mole, respectively. This implies that narciclasine exhibits stable interaction with NF-κβp50 and NF-κβp65.
The NF-κβ activation and translocation to nucleus lead to the transcription of several genes coding for inflammatory cytokines. High levels of circulating TNF-α and IL-6 were found in patients with septic shock which correlated with disease severity and increased mortality. 39 Most studies have shown that TNF is the chief mediator of the inflammatory response in sepsis and septic shock. It has also been postulated that sepsis treatment in future will be focused on immune modulating therapy against the deleterious effects of cytokines such as TNF. 40 Interleukin 6 is induced by TNF and is secreted after the initial TNF response, but it has a longer half-life than TNF making it a good surrogate marker of localized TNF activity. 41 In our study, the molecular docking results of narciclasine with TNF-α and IL-6 showed a binding energy of −6.98 and −6.77 kcal/mole, respectively. The low binding energy revealed stable interaction between narciclasine and TNF-α and narciclasine with IL-6.
It has been found that S100A8/myeloid-related protein 8 (MRP-8) directly interacts with TLR4/MD2 and induces NF-κβ activation. The S100A8 induces the intracellular translocation of MyD88 and activation of IRAK-1 and NF-κβ that subsequently results in the increased expression of TNF-α. 42 In our study, we observed that narciclasine exhibits stable interactions and has a high probability of activity with target proteins viz, S100A8/A9, MyD88, IRAK-1, and NF-κβ p50/p65. Thus, the suppression of S100A8 by narciclasine could have an impact in the downregulation of other adaptor proteins of the NF-κβ pathway.
Many drugs have failed in clinical trials due to the poor pharmacokinetics properties of drugs.43,44 These properties include ADMET. These are vital in determining the success of any drug for human therapeutic use. The ADMET result showed that narciclasine was readily biodegradable, having low toxicity and is noncarcinogenic. Likewise, the Lipinski analysis and QED results showed that narciclasine showed desirable drug-like criteria to be used in biological systems.
The reason for the high mortality in sepsis is mainly due to the nonspecific treatment for sepsis which at present is limited to supportive care including administration of intravenous fluids, antibiotics, and oxygen. 45 Many agents that have shown promising results in animal models of sepsis have failed to replicate the same impact in clinical trials. Despite initial encouraging results shown by antisepsis therapies using drotrecogin-alfa, corticosteroids, intensive insulin therapy, and vasopressin, their benefits remain uncertain. 46
For long, the major emphasis in treating sepsis was focused on controlling the spread of pathogens. Recently, in the past few decades, the focus has shifted on inhibiting the excessive inflammatory mediators released from immune cells which have been found to be involved in the pathogenesis of septic shock. 47 Thus, drugs which could reduce the excessive inflammatory response could be beneficial in sepsis.
Previously, we have found that narciclasine could effectively suppress the expression of inflammatory cytokines and improve the outcome in Escherichia coli induced sepsis in neonatal rats even at a very low concentration. 10 Thus, the results of this study further reiterates the anti-inflammatory potential of narciclasine which could be beneficial in sepsis. Moreover, narciclasine has shown to inhibit calprotectin-induced cytotoxicity at a 10 fold lower concentration than other alkaloids in the Amaryllidaceae family. 48 This can be a significant advantage in the drug development process as it can alleviate any undue side effects and toxicity issues associated with a higher concentration of any potential drug candidate.
Conclusions
Sepsis is a major cause of mortality worldwide especially among neonates and elderly. Anti-inflammatory drugs could be helpful in reducing the excessive inflammatory response in sepsis. In this study, we have found using in silico tools that narciclasine exhibits stable interactions with most of the target proteins of the LPS-TLR4 pathway. Molecular docking and MD simulation studies showed that most of the ligand-target complexes formed by narciclasine with target proteins of the LPS-TLR4 pathway were stable. This reveals that narciclasine could be effective in suppressing the inflammatory signaling response during sepsis. Furthermore, analysis of the drug likeness of narciclasine by the Lipinski filter and admetSAR indicated that narciclasine showed desirable drug-like criteria. Nevertheless, more preclinical studies on larger animals over a longer period of time need to be done to assess the toxicity and side effects associated with narciclasine before any major conclusion can be drawn.
Supplemental Material
sj-pdf-1-bbi-10.1177_11779322241233436 – Supplemental material for Effectiveness of Narciclasine in Suppressing the Inflammatory Response in Sepsis: Molecular Docking and In Silico Studies
Supplemental material, sj-pdf-1-bbi-10.1177_11779322241233436 for Effectiveness of Narciclasine in Suppressing the Inflammatory Response in Sepsis: Molecular Docking and In Silico Studies by Manoj Kumar Kingsley, Gurugubelli Krishna Rao and Ballambattu Vishnu Bhat in Bioinformatics and Biology Insights
Footnotes
Acknowledgements
The authors thank RASA Life Science Informatics, Pune, India, for their technical support and Mr V. Suresh Kumar from National Institute of Immunology, New Delhi, for his assistance with MD Simulation works.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MKK conceived and designed the study, performed the experiments, analyzed and interpreted the data, and wrote the paper. GKR assisted in performing experiments and data analysis. BVB supervised the study, assisted in the design of the experiments, revising the manuscript, and analysis of data.
Data availability statement
All relevant data supporting the findings of the study are available within the manuscript and its Supplementary Information files.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
