Abstract
Receptor-like kinases (RLKs) in plants are a large superfamily of proteins that are structurally similar. RLKs are involved in a diverse array of plant responses including development, growth, hormone perception and the response to pathogens. Current studies have focused attention on plant receptor-like kinases as an important class of sentinels acting in plant defense responses. RLKs have been identified that act in both broad-spectrum, elicitor-initiated defense responses and as dominant resistance (R) genes in race-specific pathogen defense. Most defense-related RLKs are of the leucine-rich repeat (LRR) subclass although new data are highlighting other classes of RLKs as important players in defense responses. As our understanding of RLK structure, activation and signaling has expanded, the role of the ubiquitin/proteasome system in the regulation of these receptors has emerged as a central theme.
Introduction
Receptor-like kinases (RLKs) consist of a large superfamily of proteins in plants that are structurally similar. They consist of an extracellular region, a single membrane spanning domain and an intracellular kinase domain. Plant RLKs have evolved a diverse complement of extracellular domains (more than 20 have been identified) including leucine-rich repeats (LRR), self-incompatibility (S) domains, epidermal growth factor repeats and lectin domains (Cock et al. 2002; Shiu and Bleeker, 2001). It has been proposed that the diversity of the extracellular domains in plant RLKs reflects their need to evolve rapidly in order to defend against an ever-changing population of ligands produced by pathogens (Shiu and Bleeker, 2001). The ability to recognize both general elicitors and specific pathogens through resistance (R) gene mediated resistance is integral to plant defense. Recent work has highlighted the fact that both elicitor perception in the plant innate immune response and R-gene mediated pathogen specific responses are mediated in many cases by plant RLKs. Most of the RLKs identified as being involved in plant defense are of the LRR-RLK class including the rice Xa21 protein and the Arabidopsis FLS2 and EFR receptors. However, the recent identification in rice of a lysine-motif (LysM) receptor kinase involved in the recognition of the fungal elicitor chitin (Kaku et al. 2006) and a lectin receptor kinase (LecRK) involved disease resistance indicates that other RLK classes may also play important or overlapping roles in plant defense and pathogen recognition.
Studies of RLK structural motifs have given clues to their regulation—particularly by ubiquitylation—and members of the ubiquitin system have recently begun to be implicated as major regulators of plant defense (Zeng et al. 2006). This review will focus on our current understanding of the receptor-like kinases involved in plant defense; describing both RLKs involved in elicitor-mediated or innate immunity and those involved in R-gene-mediated defense. In addition, recent work pointing to the regulation of RLKs, components of their signaling pathways and induced defense proteins by ubiquitylation will be explored.
RLKs in General Elicitor Recognition
Plants have the ability to recognize invariant pathogen associated molecular patterns (PAMPs) that are characteristic of microbial and fungal organisms but are not found in the host plant. These molecular cues, also termed general elicitors of plant defense, are essential for the pathogen and, upon receptor-mediated perception, they betray the invader to the plant's surveillance system (Nürnberger et al. 2004). The ability to recognize general elicitors and rapidly initiate defense responses is integral to the basal resistance of plants to most potential pathogens. The identification of receptors involved in the perception of various elicitors and an understanding of their structure and mode of action has been central in shaping our knowledge of elicitor-induced defenses in plant-pathogen interactions (Nürnberger and Kemmerling, 2006).
The first RLK identified involved in elicitor perception was the FLAGELLIN SENSITIVE 2 (FLS2) protein, an RLK with a leucine-rich repeat domain (LRR) that is involved in the perception of flg22, the conserved N-terminal domain of bacterial flagellin (Gómez-Gómez and Boller, 2002). In addition to the extracellular LRR domain, FLS2 has a single membrane spanning domain and a cytoplasmic serine/threonine protein kinase domain. FLS2 appears to play a role in plants similar to that of the TLR5 protein in mammalian systems (Gómez-Gómez and Boller, 2002). Flagellin is a highly conserved PAMP that is recognized by multiple plant species as well as animals. FLS2 was first identified by map-based cloning (Gómez-Gómez and Boller, 2000) and was shown to be expressed in nearly all plant tissues, including flowers, leaves, stems and roots (Gómez-Gómez and Boller, 2002). More recent work using GFP-tagged FLS2 has shown that the receptor is specifically localized to the cell periphery, particularly in the membranes of cells that could serve as a point of entry for pathogens, such as stomatal cells (Robatzek et al. 2006).
Another LRR-RLK that was recently identified as the receptor for a bacterial PAMP is the elongation factor Tu receptor (EFR). EFR is a surface receptor for elf18, a PAMP consisting of the first 18 amino acids of bacterial elongation factor Tu (EF-Tu) a highly conserved protein in bacteria (Zipfel et al. 2006). The EFR receptor belongs to the subfamily LRR-XII and has a domain structure and function similar to that of FLS2. EFR has an extracellular domain comprised of 21 copies of a 24-residue LRR, a transmembrane domain and a cytoplasmic serine/threonine kinase domain. Because the domain structures of EFR and FLS2 were so similar and both elicitors triggered a similar set of responses, Zipfel et al. (2006) investigated whether there was interference or interaction between the two perception systems. When Arabidopsis plants were challenged simultaneously with both flg22 (the conserved portion of bacterial flagellin) and EF-Tu, the plant response was similar to that elicited by flg22 alone, although there was a slight additive effect (Zipfel et al. 2006). Microarray analysis showed a clear correlation between the gene expression patterns in Arabidopsis after treatment with flg22 or EF-Tu. The authors suggest that the lack of differential gene expression between the two treatments may indicate that the plant is recognizing the pathogen-derived elicitor but that the induced response may be generic in nature (Zipfel et al. 2006).
Systemin is an 18 amino acid peptide that acts as an elicitor of defense responses in the Solanaceae and was first isolated in tomato (Ryan and Pearce, 2003). It is released at wound sites in plants (i.e. sites of insect or herbivore feeding) and is perceived by a membrane bound receptor. The systemin cell-surface receptor (SR160) is an LRR-RLK and amino acid analysis of the receptor showed that it contained a signal sequence, a leucine zipper, 25 LRR repeats with a 68 amino acid island between LRR 21 and 22, a transmembrane domain, and a serine/threonine protein kinase domain (Scheer and Ryan, 2002; Boller, 2005). Interestingly, SR160 showed greater than 90% homology to BRASSINOLIDE-INSENSITIVE 1 (BRI1), an Arabidopsis LRR-RLK involved in the perception of the steroid hormone brassinolide (Szekeres, 2003; Boller, 2005). SR160 has been clearly shown to be the tomato BRI1 ortholog (Montoya et al. 2002) but there is debate about whether SR160 and BRI1 are the same receptor in other plant species, particularly in Arabidopsis (Boller, 2005).
Recent work by Huffaker et al. (2006) characterized a new peptide elicitor from Arabidopsis,
Chitin fragments are well characterized fungal PAMPs and have been shown to produce strong defense responses in plants (Stacey and Shibuya, 1997; Shibuya and Minami, 2001). However, the search for a receptor involved in the perception of chitin has been challenging. Recent work by Kaku et al. (2006) characterized a
RLKs and R-Gene Mediated Resistance
In addition to elicitor recognition in broad spectrum defense, RLKs also participate in the recognition of pathogen avirulence factors (Avr genes) produced by specific strains of plant pathogens (Lee et al. 2006). The LRR-RLK genes Xa21 and Xa3/Xa26 have been shown to provide resistance against
The
The ERECTA RLK has recently been implicated in plant resistance to bacterial wilt, caused by
Resistance to Fusarium Oxysporum 1 (RFO1) is an RLK identified in the resistance response to the fungal pathogen
Recently a B-lectin RLK, termed Pi-d2, was identified as an R gene conferring resistance to the fungal pathogen
Receptor Kinase Regulation and Ubiquitin—Tag You're it
While our knowledge of how pathogens and elicitors are perceived by RLKs is increasing rapidly, less is known about how RLKs and their activated signaling pathways are regulated during the infection process. The most predominant theme that is emerging regarding RLK regulation is that of ubiqitination as a means of targeting receptors for degradation in order to mitigate the plant immune response. In brief, the ubiquitin/proteasome system (UPS) targets proteins for degradation by “tagging” proteins with ubiquitin, a 76 amino acid protein (Fig. 1; Dreher and Callis, 2007). Ubiquitin is attached to proteins by three distinct enzymes: the E1 ubiquitin-activating enzymes, E2 ubiquitin-conjugating enzymes and the E3 ubiquitin ligases. An important and universal feature of the ubiquitin pathway is the hierarchical structure of these three enzymes in eukaryotic genomes (Zeng, 2006). In the Arabidopsis genome there are 2 E1 enzymes, 41 E2s and more than 1200 E3s (Zeng et al. 2006). The diversity of E3 ligases highlights the importance of this enzyme in determining the substrate specificity of the ubiquitin system (Ciechanover A, 1998). There are four families of ubiquitin ligases: HECT (homologous to E6-AP COOH terminus), SCF, APC (anaphase promoting complex) and RING/U-box (really interesting new gene) that are distinguished based on their subunit composition and mechanism of action (Fig. 1; Zeng et al. 2006). Recent experimental work has highlighted the importance of the ubiquitin-proteasome system in plant defense.
The ubiquitin/26S proteasome pathway. The pathway begins with the activation of ubiquitin by ATP and the subsequent linkage via thioester bond to an E1. This is followed by the transfer of ubiquitin to an E2 enzyme. From here the pathway branches, depending on the type of E3 ligase to which the ubiquitin moiety will be transferred. RING/U-box, APC and SCF-type E3 ligases will facilitate the direct transfer of ubiquitin from the E2 to the target substrate. In the case of HECT-domain E3 ligases, the E2 will first transfer the ubiquitin to the HECT-E3 and then the HECT-E3 will transfer the ubiquitin to the target substrate. In both cases, after a chain of multiple ubiquitin moieties is attached to the target substrate it will be recognized by the 26S proteasome and degraded.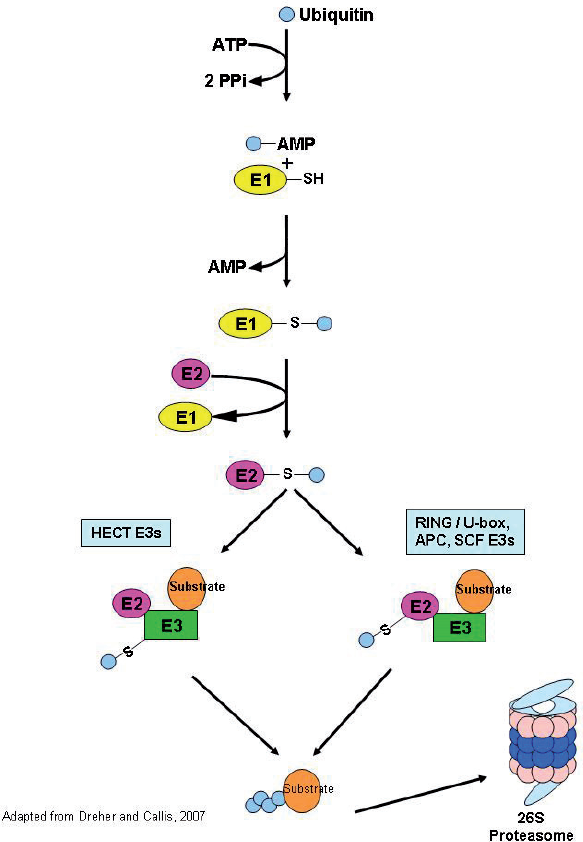
Initial structural analysis of the FLS2 protein revealed the presence of a PEST-like motif. PEST regions are rich in the amino acids proline (P), glutamic acid (E), serine (S) and threonine (T) and are typically present in proteins with short intracellular half-lives i.e. <2 hours (Rogers et al. 1986). Mutations of the PEST-like motif in FLS2 lead to a reduction in downstream signaling, suggesting that FLS2 is degraded after interacting with the ligand (Robatzek et al. 2006; Robatzek, 2007). Recent work with GFP-labeled FLS2 has shown that the FLS2-flg22 complex is internalized and degraded after binding to flg22 (Robatzek et al. 2006). Using fluorescent microscopy, Robatzek et al. (2006) captured the movement of vesicles carrying the labeled FLS2 into the cell and away from the plasma membrane. It was also shown that endogenous levels of FLS2 at the membrane were restored within hours. The restoration is a result of de novo synthesis of the receptor rather recycling as experiments showed that the addition of cyclohexamide blocked the return of FLS2 to normal levels. In addition, treatment of FLS2 with a kinase inhibitor blocked the endocytic process. Besides the presence of a PEST domain in FLS2, microarray analysis of flg22 induced genes showed an increase in the overall expression of E3 ligases, further supporting the idea that FLS2 is ubiquitinated and degraded (Robatzek et al. 2006).
The Xa21 RLK was also recently shown to interact with the ubiquitin ligase XB3 (Wang et al. 2006). These experiments by Wang et al. (2006) were the first to show a direct interaction between an E3 ligase, XB3 and an R protein, Xa21. The XB3 protein contains an ankyrin repeat domain that interacts with XA21's intracellular kinase domain and a RING motif involved in its E3 ligase activity. XB3 acts as a substrate for XA21's serine/threonine kinase activity and this interaction has been demonstrated
Several other E3 ligases have been identified as involved in plant defense. In response to the elicitor chitin, the RING-finger-type E3 ligases ATL2 and ATL6 in Arabidopsis and EL5 in rice were identified as being rapidly induced (Salinas-Mondragón et al. 1999; Serrano and Guzman, 2004; Takai et al. 2002). The tomato ortholog of Arabidopsis ATL6, LeATL6, has recently been shown to respond to cell wall protein fraction elicitor from the biocontrol agent
Conclusions and Future Directions
Our understanding of plant defense responses, both elicitor-mediated and cultivar-specific R-gene-mediated, has rapidly evolved over the past few years. It is clear that RLKs play an important role in both types of defense responses though only a few RLKs involved in defense have been described. Given the large number of RLKs in plant genomes (>400 in Arabidopsis) it is likely that other RLKs are yet to be discovered that play a role in defense. In both plant and mammalian systems, RLKs are capable of forming heterodimers (Cock et al. 2002; Johnson and Ingram, 2005) though those best characterized in plant defense (FLS2, XA21) are homodimers. It will be interesting to see if RLKs identified in future studies are able to form heterodimers and subsequently stimulate novel defense responses. Another intriguing possibility is that several RLK/ligand complexes, when activated simultaneously, may be able to act in an additive manner initiating specific signaling pathways depending on the combinations of RLKs and ligands present. Currently there is also limited information on the precise components of the signaling pathways activated by RLKs. Further characterization both of the RLKs initiating defense responses and their associated signaling molecules will be essential for a complete understanding of the defense response and how these signaling pathways may interact with one another.
Recent work (Bhat et al. 2005) has shown that lipid rafts or microdomains form in the plasma membrane at points of pathogen entry. In addition, Shahollari et al. (2004) have shown that RLKs containing LRR motifs are concentrated in plasma membrane microdomains and specific signaling proteins are then recruited to these sites. It is possible that RLKs acting in plant defense responses may also be grouped into microdomains upon ligand binding and that particular signaling components associated with a pathogen or elicitor would then be enlisted to begin a unique signal transduction cascade leading to defense. The formation of microdomains would also provide a convenient site for regulatory molecules to efficiently interact with activated RLKs. Consequently, future studies focusing on the formation of microdomains will be significant in deepening our understanding of RLK regulation and in the identification of associated signaling components.
The ubiquitin-proteasome system is emerging as an important regulatory player in plant-pathogen interactions. In particular E3 ligases appear to be key players in the regulation of RLK activity; either by regulating their turnover at the plasma membrane via ubiquitination and targeting them to the proteasome or in the case of XB3 (Wang et al. 2006) by binding at a site in the cytoplasmic region of the RLK thereby stabilizing the receptor at the membrane. In addition to interacting with RLKs in pathogen responses, E3's also appear to be important factors in the regulation of RLK-induced defense-related proteins (Hong et al. 2007) suggesting several avenues of regulation in the defense response. Since there are over 1200 E3 ligases in the Arabidopsis genome, the possibility that multiple E3 ligases are involved in the response to a single elicitor or avirulence gene is an important question to be addressed. The data of Robatzek et al. (2006) indicate that multiple E3s may be modulating interactions in the RLK-initiated defense response to flagellin suggesting a complex interplay between receptors, their signaling components and regulatory proteins. A priority for future studies will be the determination of which RLKs and E3s specifically interact, identification of their associated signal transduction components, and further insight into the different levels at which the ubiquitin system regulates the response to various pathogens and elicitors.
Footnotes
Acknowledgements
We thank Chip Goff for critical reading of this manuscript. This work was supported by a grant from the National Institutes of Health (NIGMS grant no. 1 R15 GM073630-01) to K.M.R. and by The University of Alabama.
