Abstract
Objective:
Host immunogenetics (Human Leukocyte Antigen, HLA) play a critical role in the human immune response to melanoma, influencing both melanoma prevalence and immunotherapy outcomes. Beneficial outcomes hinge on the successful binding of epitopes of melanoma antigens to HLA Class I molecules for an effective engagement of cytotoxic CD8+ lymphocytes and subsequent elimination of the cancerous cell. This study evaluated the binding affinity and immunogenicity of HLA Class I to melanoma tumor antigens to identify alleles best suited to facilitate elimination of melanoma antigens.
Methods:
In this study, we used freely available software tools to determine in silico the binding affinity and immunogenicity of 2462 reported HLA Class I alleles to all linear nonamer epitopes of 11 known antigens expressed in melanoma tumors (TRP2, S100, Tyrosinase, TRP1, PMEL(17), MAGE1, MAGE4, CTA, BAGE, GAGE/SSX2, Melan).
Results:
We identified the following 9 HLA Class I alleles with very high immunogenicity and binding affinity against all 11 melanoma antigens: A*02:14, B*07:10, B*35:10, B*40:10, B*40:12, B*44:10, C*07:11, and C*07:13, and C*07:14.
Conclusion:
These 9 HLA alleles possess the potential to aid in the elimination of melanoma both by themselves and by enhancing the beneficial effect of immune checkpoint inhibitors.
Keywords
Introduction
Melanoma accounts for a high number of skin cancer related deaths and its incidence has been increasing for decades.1,2 Therapeutic advances such as immune checkpoint inhibitors (ICI) have improved melanoma survival rates; however, ICI outcomes partially depend on host immunogenetics (Human Leukocyte Antigen, HLA). 3 Notably, a recent study documented that the population frequency of HLA alleles associated with favorable ICI outcomes are also negatively associated with the population prevalence of melanoma. 4 Burgeoning evidence highlighting the influence of HLA on melanoma prevalence, survival, and treatment response suggests a broad influence of HLA on melanoma risk and protection.3-6
A primary function of HLA involves elimination of foreign antigens including cancer neoantigens. 7 HLA Class I molecules bind and export melanoma antigen-derived peptides to the cell surface where they are presented to CD8+ T lymphocytes to signal destruction of tumor cells. This process depends on (a) the good binding of tumor epitopes with HLA Class I molecule(s) for the formation of epitope-HLA molecule complex, and (b) the successful engagement by this complex of the T-cell receptor (TCR) for subsequent cytotoxic elimination of the tumor cell. However, due to the highly polymorphic nature of HLA, its molecules vary tremendously with respect to peptide binding capability, with the result that any specific combination of 6 HLA Class I alleles carried by an individual subject may offer only limited coverage of the tumor antigens,8-16 particularly in light of immune-escape mechanisms exploited by tumors.17-19 Yet, relatively little is known about binding affinity and immunogenicity of specific HLA alleles with melanoma antigens. Indeed, alleles with high immunogenicity against specific melanoma/cancer antigens would be beneficial in tumor elimination by themselves and by enhancing the effect of immune checkpoint blockade immunotherapy.
In this study we used an in silico approach20-23 to characterize binding affinity and immunogenicity of 2462 HLA alleles to 11 known melanoma antigens, 24 and successfully identified alleles with high immunogenicity and binding affinity against those antigens.
Materials and Methods
Melanoma/cancer antigens
Eleven cancer antigens (Table 1) were used based on their known occurrence in melanoma tumors 24 ; of those 6 are specific to melanoma and 5 are expressed in melanoma as well as in other tumors 20 (labeled [CTA] in Table 1). The amino acid (AA) sequences of the 11 melanoma antigens used were retrieved from the Uniprot Database 25 and are given in the Appendix.
In silico determination of immunogenicity of antigen nonamers with HLA Class I alleles
We determined the immunogenicity between 2462 reported HLA Class I alleles
26
and 11 melanoma antigens (Table 1) using the INeo-Epp method
22
for T-cell receptor (TCR) epitope prediction using theINeo-Epp web tool,
20
downloaded in April 2023, and run locally on a CentOS 6.7 server. For that purpose, we split a given melanoma antigen to all possible linear 9-mer (nonamer) AA residue epitopes using a sliding window approach, as detailed previously
27
(Figure 1) and submitted each epitope to the application together with a specific HLA allele. More specifically, we paired all epitopes with all alleles and obtained for each pair its percentile rank, a measure of binding affinity of the epitope-HLA allele complex; smaller percentile rank indicates higher binding affinity. The application gave as an outcome a TCR predictive score,


Illustration of the sliding nonamer approach used. See text for details.
Number of alleles with allele-antigen immunogenicities above the thresholds mentioned in the text.

Finally, TIMG scores were summed for each allele, across all allele-nonamer combinations, and the fractional ranks of these sums were computed for each allele and antigen. For each antigen, the allele with the top-ranked sum (ie, 100%) was the most potent against the particular antigen.
In silico determination of predicted binding affinity of antigen nonamers with HLA Class I alleles
The determination of nonamer immunogenicity above was obtained using the INeo-Epp tool which employs a variety of T-cell immunogenicity-related measures, in addition to the predicted binding affinity of the antigen to the peptide motif of the HLA Class I molecule. As such, the INeo-Epp tool provides a wide coverage of antigen immunogenicity. However, we also determined the predicted binding affinity for the best INeo-Epp outcomes using a different tool, namely the Immune Epitope Database (IEDB) NetMHCpan (ver. 4.1) tool.21,23 For each nonamer-HLA molecule tested, this tool gives, as an output, the percentile rank of binding affinity of the HLA molecule and the epitope among predicted binding affinities of the same HLA molecule to a large number of different peptides of the same AA length; the smaller the percentile rank, the better the binding affinity. Now, given a protein of N amino acid length and a nonamer, there are N-9 binding affinity predictions, that is, N-9 percentile ranks. Of these predictions, for each melanoma antigen and HLA molecule tested, we retained the lowest percentile rank (LPR) as the best possible binding affinity of the protein-HLA molecule pair.
Results
We identified 2462 reported HLA Class I alleles (Table 2) that possessed high binding affinity to nonamers of 11 known melanoma antigens and yielded HLA-antigen complexes with high immunogenicity (ie, good prediction of engaging cytotoxic CD8+ TCR). We then ranked the alleles according to their immunogenicity strength and identified those with the highest immunogenicity for each melanoma antigen, a total of 9 alleles shown in Table 3. (The results for all alleles are given in Table S1 in Supplemental Material). Remarkably, these 11 alleles were also potent against all other melanoma antigens (Table 4), thus conferring very high protection across the board. The specific nonamers and their highly ranked immunogenicities are shown in Table 5. Finally, Table 6 shows the predicted best binding affinities obtained for the antigen-allele combinations (Table 5) using the NetMHCpan tool. It can be seen that all alleles had excellent predicted binding affinities: all LPR values were <1, a common threshold for a good binding affinity. Thus both INeo-Epp and NetMHCpan prediction tools gave very congruent results.
Alleles with highest immunogenicities (among the 2462 alleles tested) for the different cancer/melanoma antigens.

Fractional ranks of average immunogenicity of alleles in Table 1 across all nonamers. Maximum (best) is 100.
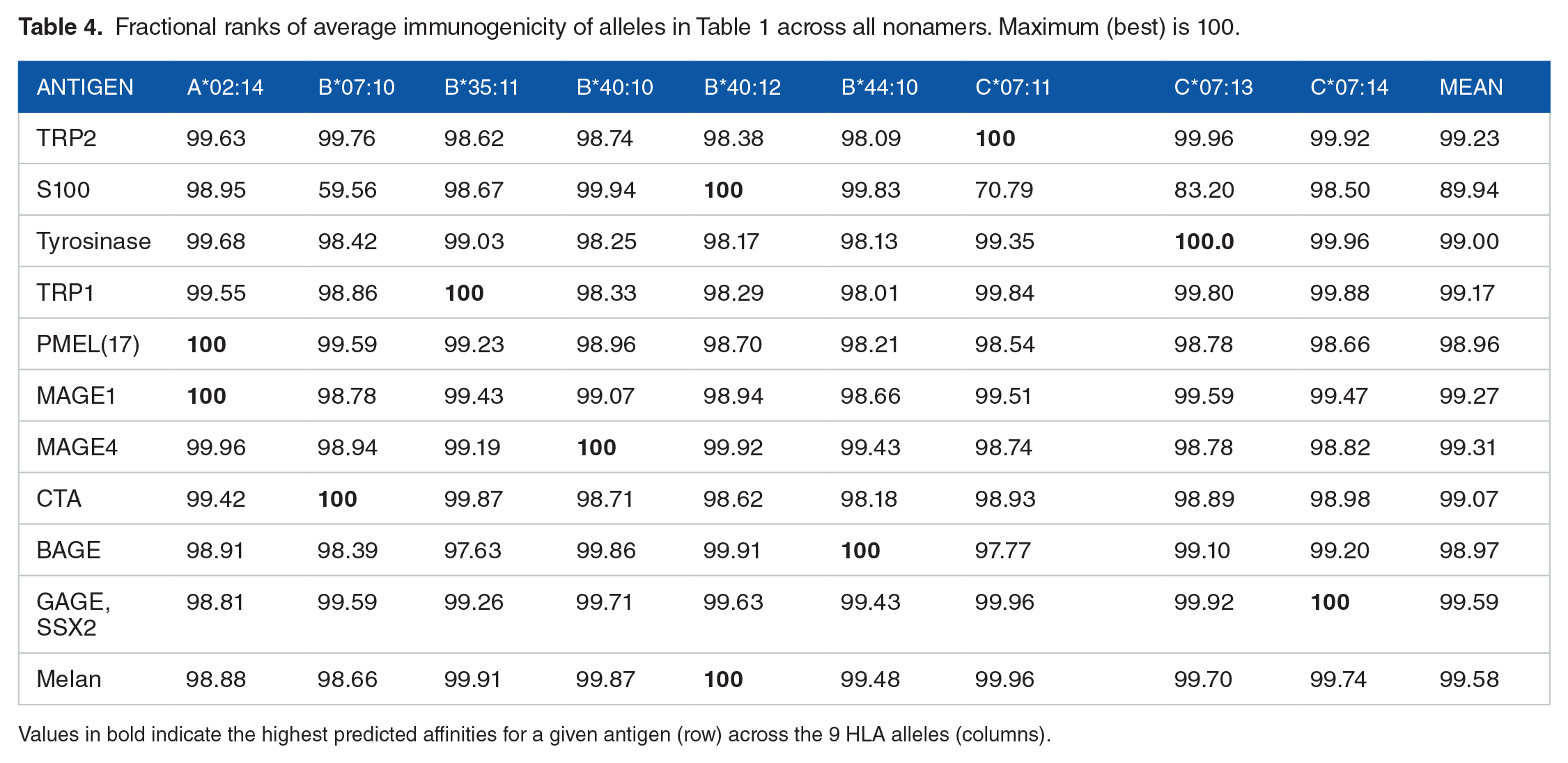
Values in bold indicate the highest predicted affinities for a given antigen (row) across the 9 HLA alleles (columns).
Nonamers of best allele for each melanoma antigen. Begin and End denote the beginning and end of the nonamer within the antigen AA sequence (Table S3).

Predicted best binding affinities (LPR values) using the NetChan tool. The lower the LPR, the higher the predicted affinity.

Values in bold indicate the highest predicted affinities for a given antigen (row) across the 9 HLA alleles (columns).
Discussion
Here we identified in silico 9 HLA Class I alleles with the highest immunogenicity (out of 2462 tested) against 11 melanoma antigens. These alleles, and others with lower but still high immunogenicities against the tested antigens (See Table 1 in Supplemental Material), are excellent candidates as therapeutic anti-melanoma agents, acting both by themselves (inducing the lysis of cancerous cells via engagement of cytotoxic CD8+ T-cells) and by aiding ICI, the success of which has been found to depend on the HLA Class I genetic makeup of the patient. 3
Since HLA is the most highly polymorphic region of the human genome and a particular individual carries only 6 Class I HLA alleles, the likelihood that a given individual may possess a HLA allele that can bind melanoma antigens with sufficient immunogenicity to be substantially effective in tumor cell elimination is limited. Thus, the outcome of ICI would be substantially potentiated if such HLA Class I molecules were present in the tumor cell. Indeed, we have recently proposed 28 that this could be achieved by inducing the biosynthesis of desired HLA Class I molecules in the tumor cells by injecting in the tumor site the suitably LNP-mRNA (i.e. mRNA packaged with lipid nanoparticles, LNP) of those Class I molecules, (“Direct HLA therapy, dHLA” 28 ). In the specific case of melanoma, the treatment would involve the introduction in the tumor site of the LNP-mRNA of the 9 omnipotent alleles in Table 3 using recently improved methods of administration. 29 An additional benefit could be derived indirectly from the well-known immunogenicity of the HLA molecules themselves, that is, the production of antibodies against them leading, for example, to the rejection of a transplanted organ. In the present application, such an effect would actually be beneficial, since it will enhance the elimination of the tumor, which would be rejected as a foreign organ. This treatment could be applied to all forms of solid tumors for which tumor antigens and suitably potent HLA Class I alleles against them have been identified, a feasible objective, as demonstrated in the present study. In this context, it is noteworthy that 5 of the 11 antigens tested, namely the antigens in the Cancer Testis Antigen (CTA) family (Table 3) are also expressed in other tumors24,30-32 (e.g. lung carcinoma, 30 head and neck squamous cell carcinoma 31 ) and, therefore, administration of the 9 alleles discovered here (Tables 4 and 6) would be beneficial to such tumors, in addition to melanoma.
Finally, it is important to distinguish the proposed therapy here from cancer vaccine therapies currently being developed and applied.33-36 In a cancer vaccine, cancer (neo)antigens are suitably administered to induce the production of antibodies against them and thus reduction/elimination of the tumor, engaging antigen presenting cells (APC), the HLA Class II system, CD4+ lymphocytes, B cells, etc. In contrast, in our proposed dHLA application, 28 the mRNA of specific HLA Class I molecules, found to bind with high affinity to cancer antigen fragments (nonamers), are injected directly into the tumor, where the protein of the HLA molecule is synthesized, binds to antigen nonamers, is transported to the cell surface, and engages the cytotoxic CD8+ lymphocytes, killing the cancerous cell. Thus this method does not involve the production of antibodies but depends on the ability of cytotoxic lymphocytes to kill the cancer cell, an ability that, when reduced or absent, can be restored by ICI, hence the prospect of dHLA therapy as a promising aid in ICI success, in addition to its expected effectiveness in early stages of cancer when immune blockade is weak or absent. Given the cost and toxicity of current cancer immunotherapies,37-39 dHLA therapy 28 would be a good addition: the cost should be low (just making the LNP-mRNA of the 9 HLA molecules) and adverse effects minimal, if any, given the local, intratumoral injection of natural molecules already present in humans. In a way, dHLA cancer therapy opens a new avenue in response to the “need for innovation” in cancer immunotherapy requested recently. 40
Limitations
The main limitation is that these effective alleles are to be injected to tumors locally, and not systemically. Thus, they are not suitable for diffuse tumors, multiple metastases, or inaccessible sites.
Conclusions
The 9 HLA alleles identified here to possess high affinity and immunogenicity against 11 common cancer antigens are specific, key molecules for use in direct HLA cancer therapy with expected favorable application to melanoma and other solid tumors, and minimal (if any) adverse effects. Their evaluation in preclinical assessments and clinical trials regarding tumor reduction/elimination remains to be determined with respect to effects of themselves alone and/or administered in combination with ICI.
Supplemental Material
sj-docx-1-cix-10.1177_11769351241274160 – Supplemental material for Nine Human Leukocyte Antigen (HLA) Class I Alleles are Omnipotent Against 11 Antigens Expressed in Melanoma Tumors
Supplemental material, sj-docx-1-cix-10.1177_11769351241274160 for Nine Human Leukocyte Antigen (HLA) Class I Alleles are Omnipotent Against 11 Antigens Expressed in Melanoma Tumors by Apostolos P Georgopoulos, Lisa M James and Matthew Sanders in Cancer Informatics
Footnotes
Appendix
Amino acid sequences of the 11 melanoma antigens analyzed (Table 1).
MSPLWWGFLLSCLGCKILPGAQGQFPRVCMTVDSLVNKECCPRLGAESANVCGSQQGRGQ
CTEVRADTRPWSGPYILRNQDDRELWPRKFFHRTCKCTGNFAGYNCGDCKFGWTGPNCER
KKPPVIRQNIHSLSPQEREQFLGALDLAKKRVHPDYVITTQHWVGLLGPNGTQPQFANCS
VYDFFVWLHYYSVRDTLLGGFFPWLKVYYYRFVIGLRVWQWEVISCKLIKRATTRQP
MSELEKAMVALIDVFHQYSGREGDKHKLKKSELKELINNELSHFLEEIKEQEVVDKVMET
LDNDGDGECDFQEFMAFVAMVTTACHEFFEHE
MLLAVLYCLLWSFQTSAGHFPRACVSSKNLMEKECCPPWSGDRSPCGQLSGRGSCQNILL
SNAPLGPQFPFTGVDDRESWPSVFYNRTCQCSGNFMGFNCGNCKFGFWGPNCTERRLLVR
RNIFDLSAPEKDKFFAYLTLAKHTISSDYVIPIGTYGQMKNGSTPMFNDINIYDLFVWMH
YYVSMDALLGGSEIWRDIDFAHEAPAFLPWHRLFLLRWEQEIQKLTGDENFTIPYWDWRD
AEKCDICTDEYMGGQHPTNPNLLSPASFFSSWQIVCSRLEEYNSHQSLCNGTPEGPLRRN
PGNHDKSRTPRLPSSADVEFCLSLTQYESGSMDKAANFSFRNTLEGFASPLTGIADASQS
SMHNALHIYMNGTMSQVQGSANDPIFLLHHAFVDSIFEQWLRRHRPLQEVYPEANAPIGH
NRESYMVPFIPLYRNGDFFISSKDLGYDYSYLQDSDPDSFQDYIKSYLEQASRIWSWLLG
AAMVGAVLTALLAGLVSLLCRHKRKQLPEEKQPLLMEKEDYHSLYQSHL
MSAPKLLSLGCIFFPLLLFQQARAQFPRQCATVEALRSGMCCPDLSPVSGPGTDRCGSSS
GRGRCEAVTADSRPHSPQYPHDGRDDREVWPLRFFNRTCHCNGNFSGHNCGTCRPGWRGA
ACDQRVLIVRRNLLDLSKEEKNHFVRALDMAKRTTHPLFVIATRRSEEILGPDGNTPQFE
NISIYNYFVWTHYYSVKKTFLGVGQESFGEVDFSHEGPAFLTWHRYHLLRLEKDMQEMLQ
EPSFSLPYWNFATGKNVCDICTDDLMGSRSNFDSTLISPNSVFSQWRVVCDSLEDYDTLG
TLCNSTEDGPIRRNPAGNVARPMVQRLPEPQDVAQCLEVGLFDTPPFYSNSTNSFRNTVE
GYSDPTGKYDPAVRSLHNLAHLFLNGTGGQTHLSPNDPIFVLLHTFTDAVFDEWLRRYNA
DISTFPLENAPIGHNRQYNMVPFWPPVTNTEMFVTAPDNLGYTYEIQWPSREFSVPEIIA
IAVVGALLLVALIFGTASYLIRARRSMDEANQPLLTDQYQCYAEEYEKLQNPNQSVV
MDLVLKRCLLHLAVIGALLAVGATKVPRNQDWLGVSRQLRTKAWNRQLYPEWTEAQRLDC
WRGGQVSLKVSNDGPTLIGANASFSIALNFPGSQKVLPDGQVIWVNNTIINGSQVWGGQP
VYPQETDDACIFPDGGPCPSGSWSQKRSFVYVWKTWGQYWQVLGGPVSGLSIGTGRAMLG
THTMEVTVYHRRGSRSYVPLAHSSSAFTITDQVPFSVSVSQLRALDGGNKHFLRNQPLTF
ALQLHDPSGYLAEADLSYTWDFGDSSGTLISRALVVTHTYLEPGPVTAQVVLQAAIPLTS
CGSSPVPGTTDGHRPTAEAPNTTAGQVPTTEVVGTTPGQAPTAEPSGTTSVQVPTTEVIS
TAPVQMPTAESTGMTPEKVPVSEVMGTTLAEMSTPEATGMTPAEVSIVVLSGTTAAQVTT
TEWVETTARELPIPEPEGPDASSIMSTESITGSLGPLLDGTATLRLVKRQVPLDCVLYRY
GSFSVTLDIVQGIESAEILQAVPSGEGDAFELTVSCQGGLPKEACMEISSPGCQPPAQRL
CQPVLPSPACQLVLHQILKGGSGTYCLNVSLADTNSLAVVSTQLIMPGQEAGLGQVPLIV
GILLVLMAVVLASLIYRRRLMKQDFSVPQLPHSSSHWLRLPRIFCSCPIGENSPLLSGQQ
V
MSLEQRSLHCKPEEALEAQQEALGLVCVQAATSSSSPLVLGTLEEVPTAGSTDPPQSPQG
ASAFPTTINFTRQRQPSEGSSSREEEGPSTSCILESLFRAVITKKVADLVGFLLLKYRAR
EPVTKAEMLESVIKNYKHCFPEIFGKASESLQLVFGIDVKEADPTGHSYVLVTCLGLSYD
GLLGDNQIMPKTGFLIIVLVMIAMEGGHAPEEEIWEELSVMEVYDGREHSAYGEPRKLLT
QDLVQEKYLEYRQVPDSDPARYEFLWGPRALAETSYVKVLEYVIKVSARVRFFFPSLREA
ALREEEEGV
MSSEQKSQHCKPEEGVEAQEEALGLVGAQAPTTEEQEAAVSSSSPLVPGTLEEVPAAESA
GPPQSPQGASALPTTISFTCWRQPNEGSSSQEEEGPSTSPDAESLFREALSNKVDELAHF
LLRKYRAKELVTKAEMLERVIKNYKRCFPVIFGKASESLKMIFGIDVKEVDPASNTYTLV
TCLGLSYDGLLGNNQIFPKTGLLIIVLGTIAMEGDSASEEEIWEELGVMGVYDGREHTVY
GEPRKLLTQDWVQENYLEYRQVPGSNPARYEFLWGPRALAETSYVKVLEHVVRVNARVRI
AYPSLREAALLEEEEGV
MQAEGRGTGGSTGDADGPGGPGIPDGPGGNAGGPGEAGATGGRGPRGAGAARASGPGGGA
PRGPHGGAASGLNGCCRCGARGPESRLLEFYLAMPFATPMEAELARRSLAQDAPPLPVPG
VLLKEFTVSGNILTIRLTAADHRQLQLSISSCLQQLSLLMWITQCFLPVFLAQPPSGQRR
MAARAVFLALSAQLLQARLMKEESPVVSWRLEPEDGTALCFIF
MNGDDAFARRPTVGAQIPEKIQKAFDDIAKYFSKEEWEKMKASEKIFYVYMKRKYEAMTK
LGFKATLPPFMCNKRAEDFQGNDLDNDPNRGNQVERPQMTFGRLQGISPKIMPKKPAEEG
NDSEEVPEASGPQNDGKELCPPGKPTTSEKIHERSGPKRGEHAWTHRLRERKQLVIYEEI
SDPEEDDE
MPREDAHFIYGYPKKGHGHSYTTAEEAAGIGILTVILGVLLLIGCWYCRRRNGYRALMDK
SLHVGTQCALTRRCPQEGFDHRDSKVSLQEKNCEPVVPNAPPAYEKLSAEQSPPPYSP
Acknowledgements
None.
Author Contributions
APG conceived the proposal and analyzed the data. MS performed the INeo-Epp and NetChan analyses. APG and LMJ wrote the paper. APG, LMJ and MS reviewed and approved the paper.
Availability of Data and Materials
All data used for analysis are freely available from public databases, as stated in the Materials and Methods Section of the manuscript.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Partial funding for this study was provided by the University of Minnesota (the Kunin Chair in Women’s Healthy Brain Aging, the Brain and Genomics Fund, and the American Legion Brain Sciences Chair) and the U.S. Department of Veterans Affairs. The sponsors had no role in the current study design, analysis or interpretation, or in the writing of this paper. The contents do not represent the views of the U.S. Department of Veterans Affairs or the United States Government.
Ethical Approval and Consent to Participate
This article simply and solely analyzed publicly available data and does not contain any studies with human participants or animal experiments performed by any of the authors. Hence, ethical approval is not applicable to this study.
Consent for Publication
Not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

