Abstract
In diploid organisms, half of the chromosomes in each cell come from the father and half from the mother. Through previous studies, it was found that the paternal chromosome and the maternal chromosome can be regulated and expressed independently, leading to the emergence of allele specific expression (ASE). In this study, we analyzed the differential expression of alleles in the high-altitude population and the normal population based on the RNA sequencing data. Through gene cluster analysis and protein interaction network analysis, we found some changes occurred at the gene level, and some negative effects. During the study, we realized that the calmodulin homology domain may have a certain correlation with long-term survival at high altitude. The plateau environment is characterized by hypoxia, low air pressure, strong ultraviolet radiation, and low temperature. Accordingly, the genetic changes in the process of adaptation are mainly reflected in these characteristics. High altitude generation living is also highly related to cancer, immune disease, cardiovascular disease, neurological disease, endocrine disease, and other diseases. Therefore, the medical system in high altitude areas should pay more attention to these diseases.
Introduction
The structural variation and specific expression of alleles are ubiquitous in many organisms. The allele specific expression (ASE) refers to the difference in the expression amount of male and female alleles. 1 ASE is one of the important genetic factors that lead to phenotypic variation and can be used to identify differences in gene regulatory factors. ASE indicates the role of non-coding RNA, DNA methylation, and histone modification mechanisms. The precise mechanism by which allele specific gene expression occurs is still unclear. The study of eQTLs for expression traits indicates that ASE typically reflects genetic polymorphism through cis regulation, 2 while trans genetic regulation or epigenetic mechanisms are relatively rare.2,3 It is generally believed that cis regulatory polymorphism is the main source of phenotypic differences and is associated with many diseases.4 -6 ASE plays an important role in regulating gene expression, and studies have found that the bias of allele expression in the same cell is often fixed. 7 According to different allelic bias, the phenotype reflected in the organism will also be different. 8 As in mammals, ASE is highly associated with epigenetic blunting on the X chromosome. 9 Some studies have also shown that ASE plays an important role in heterosis, because gene variation usually causes gene expression variation.10 -12 With the development of RNA sequencing (RNA-seq), people have been able to obtain basically unbiased and relatively complete deep sequencing data of the whole transcriptome, which effectively improves the accuracy of detecting single nucleotide polymorphisms (SNPs) in the genome, and makes the identification of parental alleles in heterozygotes and the analysis of ASE more reliable. 13 According to previous studies, in order to adapt to the natural conditions in high altitude areas, such as low atmospheric pressure, low oxygen, strong ultraviolet ray, and other factors, the people living in the plateau have undergone a certain survival of the fittest at the genetic level, and the genetic modification that is relatively different from the mainland population has been screened. 14 This kind of gene modification not only improves the adaptability of people living at high altitude, but also causes certain side effects, such as high correlation with cancer, immune disease, cardiovascular disease, neurological disease, endocrine disease, and other diseases.15,16 Imprinting phenomenon is a special case of ASE, which refers to some genes always expressing specific alleles from a certain parent.17 -19 In humans, the absence or improper expression of imprinted genes can cause many clinical syndromes, while in adults, the disorder of imprinting is closely related to the occurrence of tumors. In vitro manipulation and cultivation make imprinted genes more susceptible to epigenetic changes. 20
In the present study, we analyzed the RNA-seq data of human umbilical vein endothelial cells (HUVECs) from the high-altitude population and the normal population, we found that there were some differences of ASE. Combined with genomics data, we researched the function and expression of these genes, discovered the evolution made by human beings in the process of adaptation, and recognized the reasons for the high incidence rate of some diseases in some high-altitude areas.
Method
Datasets
The RNA-seq data of plateau generation residents were obtained from The National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov, access number: GSE145774) and The National Geophysical Data Center (https://ngdc.cncb.ac.cn, project number: CRA002025), 21 Control group RNA-seq data from NCBI (http://www.ncbi.nlm.nih.gov, access number: GSE131681). 22 The RNA-seq data of HUVECs cells were selected for analysis in the above two groups of data.
RNA-seq data processing
For the RNA-seq data used in this experiment, we first use hisat2 23 to blast, and the data after blast use htseq 24 software to count the reads. And then, use R to merge, integrate, and annotate the data, and output the count matrix file. Use DESeq2 25 software to analyze the output matrix data to obtain basic difference expression data. Next, we use the aScan 26 software to process each group and obtain the ASE genes of each group of data for subsequent analysis and processing. The flowchart of data processing is shown in Figure 1.
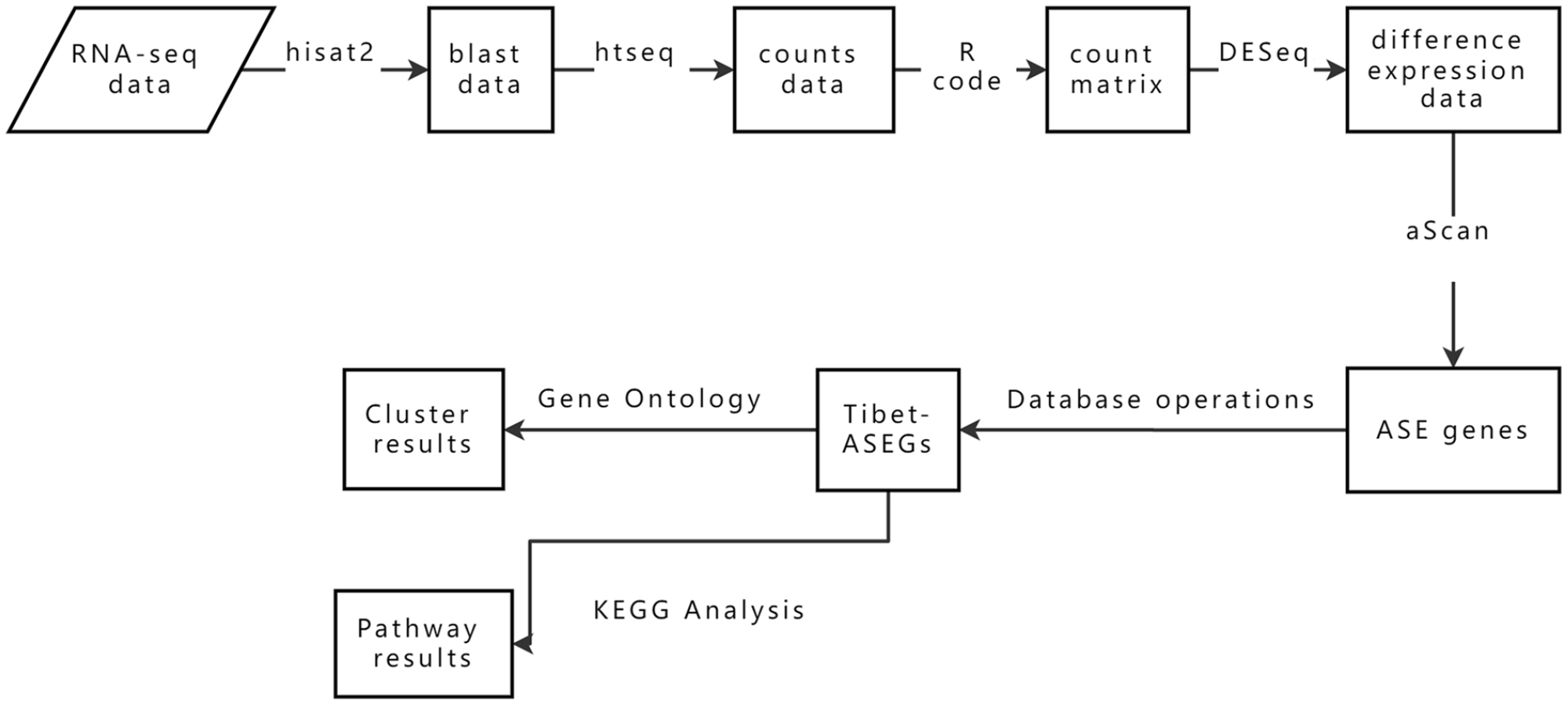
The flowchart of data processing.
Result
Pretreatment and analysis of RNA-seq data
In this study, we selected the RNA-seq data of the ethnic population 21 and the normal population 22 in Tibet for analysis. We used hisat2 23 to align the original sequencing data with the human genome data at first, sorted and ordered the sam files, counted the results by htseq, 24 merged and filtered the datasets, processed, and analyzed the ASE data by the DESeq2. 25 During the processing, the above software uses default parameters. The analysis results are shown in Figure 2.

The expression value of control set and treat set. The first three groups are the control group, and the last nine groups are the experimental group. It can be seen that there is a significant difference in expression levels between the experimental group and the control group.
Identification of ASE genes lists
Through the above preliminary analysis, we have basically confirmed that there are ASE at the gene level of both groups. To further study the differences of genes, we explored the two groups of RNA-seq data using aScan, 26 and got the ASE genes lists (ASEGs). There are 178 genes of ASE in Tibet, and 748 genes of ASE in the normal population. After getting the data, we calculated the P value and fdr of the ASEGs in Tibet population again (P < .05), and obtained a gene group with significant differences in each group’s data, a total of 49 genes (Tibet-ASEGs). There were significant differences in the specific expression of some genes in Tibet-ASEGs, including the gene PKM, it has been confirmed highly correlated with aerobic glycolysis, 27 energy metabolism, 28 and cancer. 29 The gene CCND has been confirmed that it is highly correlated with the cell cycle 30 and cervical cancer. 31 And the gene RHOB is highly correlated with cell proliferation, actin dynamics and cell cycle, 32 and also highly correlated with lung cancer 33 and breast cancer. 34 In summary, it can be seen that the typical genes specifically expressed by the people living at high altitude are all related to high altitude adaptation, such as enhanced energy metabolism and faster cell proliferation. At the same time, it has also brought a series of related diseases common to Tibetan people, such as high altitude erythrocytosis, lung cancer, and breast cancer.
Tibet-ASEGs clustering and Gene Ontology analysis
We performed gene clustering and Gene Ontology analysis on the Tibet-ASEGs, 35 and the clustering results have been shown in Supplemental Table 1, which are highly correlated with protein kinase binding, calmodulin homology domains, Ubl conjugation, ATP binding, and actin filament binding. According to the present research results, calmodulin homeodomain plays an important role in the composition of cytoskeleton proteins and maintaining actin binding proteins. 36 According to the results of functional annotation (Supplemental Table 2, Figure 3), Tibet-ASEGs are mainly related to the following functions: Oxytocin signaling pathway, positive regulation of phosphoprotein phosphatase activity, protein phosphatase activator activity, cell adhesion, and phosphoprotein, etc. According to previous studies, the non-g/g genotypes of anp32d in the phosphoprotein family are associated with chronic high altitude reaction. 37 It can be found from this study that the clustering results of Tibet-ASEGs are all related to cell structure, muscle activity, human metabolism, and altitude reaction.

Functional annotation chart.
Tibet-ASEGs protein interaction network analysis
We used the KEGG database to study the pathway (Figure 4) of Tibet-ASEGs related proteins. 38 According to the processing results of KEGG, it is found that the proteins expressed by Tibet-ASEGs are mainly related to Pathways in cancer (colorectal cancer, breast cancer, pancreatic cancer, and gastric cancer), oxytocin signaling pathway, human cytomegalovirus infection, cellular senescence other tissue system pathways, and are also related to some diseases, such as cancer, catecholaminergic polymorphic ventricular tachycardia, and immune diseases. It can be seen that the protein interaction network of Tibet-ASEGs is mainly reflected in metabolism, environmental adaptation, and common diseases at high altitude, which reflects the compromise and sacrifice made by human beings in the process of adapting to high altitude.

The pathway of metabolism involved by the Tibet-ASEGs, the Tibet-ASEGs involved are marked in red within the pathway.
Discussion
This study confirmed that the long-term residence in the plateau environment has a certain impact on the allele specific expression of related genes
According to this study, it can be confirmed that the allele expression of PKM, RhoB, ccnd, and other genes will be significantly biased when people live at high altitude for a long time. At the same time, these genes are highly correlated with body metabolism and body functions, indicating that there is a certain compensatory relationship at the genetic level in the process of human adaptation to high altitude.
The ASE of 16 genes in the metabolism pathway, such as PKM, DGKH, and BCKDHA, also indicates that in the process of adaptation, the human body changes some pathways instead of some genes, thus ensuring the gradual adaptation of the human body.
Calmodulin homology domain has a certain correlation with long-term high-altitude reaction
At present, there is no specific study showing that the calmodulin homology domain has a correlation with altitude reaction. According to the results of go analysis, we found that the clusters related to the calmodulin homology domain have high reliability and relatively high enrichment scores. This indicates that there are significant differences in the ASE of calmodulin homology domain related genes in the population living at high altitude for generations. According to previous studies, calmodulin homeodomain often plays an important role in composing cytoskeleton proteins and maintaining actin binding proteins. In high altitude areas, due to the influence of external factors such as strong ultraviolet rays, hypoxia, and low air pressure, the overall strength of cell structure will also change, and important cell functions such as cell movement and contraction will also be affected accordingly. Therefore, we infer that with the adaptation and evolution of human beings, calmodulin homology domain related genes also gradually appear specific expression.
Living on the plateau for generations mainly affects metabolism and body energy supply
According to the analysis results of KEGG, among the genes specifically expressed by residents at high altitude, 16 genes are in the metabolic pathway. Combined with the results of aScan analysis, PKM, ANPEP, and GANAB with significant differences in ASE are highly related to energy metabolism, aerobic glycolysis, etc. On the basis of Go clustering results, gene clustering related to ATP binding has a high enrichment score, of which 27 genes are related to ATP binding. It can be found that the main problem faced to human beings in the hypoxic environment is the supply of body energy. Therefore, a large number of genes related to energy supply and metabolism are differentially expressed in alleles.
Long term living on the plateau also has a certain impact on the occurrence of various diseases
From this study, we can draw the following conclusions: human beings have undergone a series of changes at the genetic level in the process of adapting to high altitude. On the one hand, some of them have significant effects on improving the survival rate of residents at high altitude. On the other hand, the differential expression of some genes also brings side effects of some diseases. From the above analysis results, it can be found that some ASE genes promote metabolism and maintain cell processes, but also increase the probability of people suffering from cancer, immune diseases, cardiovascular diseases, neurological diseases, endocrine diseases, and other diseases. Combined with the previous research results, it can be found that the cancer mortality of people at high altitude is highly related to altitude, 39 chronic altitude reaction often leads to immune system disorder and immune diseases,40,41 high altitude has a certain positive correlation effect on cardiovascular disease and nervous system disease. 42 All these indicate a series of compromises and concessions made in the evolutionary process at the level of human genes.
Limitations of this study
In this study, due to limitations in funding, experimenters, and ancillary facilities, we did not conduct relevant experiments, but directly selected some publicly available datasets for analysis and research. To ensure that the cell lines analyzed in the control group and experimental group are consistent, we have relatively few datasets to choose, which have some limitations. Therefore, some results may be affected to some extent.
Conclusion
According to our research, living on high plateaus for a long time can also cause adaptive changes in human genes. This change is reflected in many aspects. On the one hand, people are more adaptable to live in high altitude, low pressure, and hypoxia environment, and on the other hand, the incidence rate of some diseases also increases. In practical work environments, if it is necessary to work at high altitudes, genetic screening can be carried out first to select populations more suitable for high-altitude areas, thereby improving work efficiency and reducing cost consumption.
Supplemental Material
sj-csv-1-evb-10.1177_11769343241257344 – Supplemental material for Study on Allele Specific Expression of Long-Term Residents in High Altitude Areas
Supplemental material, sj-csv-1-evb-10.1177_11769343241257344 for Study on Allele Specific Expression of Long-Term Residents in High Altitude Areas by Chao He, Bin Zhu, Wenwen Gao, Qianjin Wu and Changshui Zhang in Evolutionary Bioinformatics
Supplemental Material
sj-csv-2-evb-10.1177_11769343241257344 – Supplemental material for Study on Allele Specific Expression of Long-Term Residents in High Altitude Areas
Supplemental material, sj-csv-2-evb-10.1177_11769343241257344 for Study on Allele Specific Expression of Long-Term Residents in High Altitude Areas by Chao He, Bin Zhu, Wenwen Gao, Qianjin Wu and Changshui Zhang in Evolutionary Bioinformatics
Supplemental Material
sj-csv-3-evb-10.1177_11769343241257344 – Supplemental material for Study on Allele Specific Expression of Long-Term Residents in High Altitude Areas
Supplemental material, sj-csv-3-evb-10.1177_11769343241257344 for Study on Allele Specific Expression of Long-Term Residents in High Altitude Areas by Chao He, Bin Zhu, Wenwen Gao, Qianjin Wu and Changshui Zhang in Evolutionary Bioinformatics
Footnotes
Acknowledgements
First and foremost, I would like to show my deepest gratitude to my supervisor, Dr. Zhang Changshui, a respectable, responsible, and resourceful scholar, who has provided me with valuable guidance in every stage of the writing of this thesis. Without his enlightening instruction, impressive kindness, and patience, I could not have completed my paper. His keen and vigorous academic observation enlightens me not only in this paper but also in my future study. I shall extend my thanks to Mrs. Liu for all her kindness and help. I would also like to thank all my teachers who have helped me to develop the fundamental and essential academic competence. My sincere appreciation also goes to the teachers and students from The General Hospital of Tibet Military Region, who participated this study with great cooperation. Last but not least, I would like to thank all the authors of this article, especially my project leader Zhu Bin, for their encouragement and support.
Statement on Author’s Contribution
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by Chao He, Bin Zhu, and Wenwen Gao. The first draft of the manuscript was written by Chao He and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Central Guidance Local Science and Technology Development Fund Project (Construction of Science and Technology Innovation Base) (XZ202201YD0004C).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
