Abstract
The epigenetic silencing of tumor suppressor genes (TSGs) is a common finding in several solid and hematological tumors involving various epigenetic readers and writers leading to enhanced cell proliferation and defective apoptosis. Thymoquinone (TQ), the major biologically active compound of black seed oil, has demonstrated anticancer activities in various tumors by targeting several pathways. However, its effects on the epigenetic code of cancer cells are largely unknown. In the present study, we performed RNA sequencing to investigate the anticancer mechanisms of TQ-treated T-cell acute lymphoblastic leukemia cell line (Jurkat cells) and examined gene expression using different tools. We found that many key epigenetic players, including ubiquitin-like containing plant homeodomain (PHD) and really interesting new gene (RING) finger domains 1 (
Introduction
Genetic abnormalities constitute the basis for the etiology of several tumors. However, in addition to genetic alterations, epigenetic modifications, including DNA methylation and
Materials and Methods
Cell culture and treatment
T-cell ALL JK cell line and MDA-MB-468 cell line, a human epithelial breast cancer cell line, were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). Cells were maintained in RPMI1640 media for JK and Dulbecco’s modified Eagle medium (DMEM) (UFC-Biotech, Riyadh, Saudi Arabia) for MDA-MB-468 supplemented with 15% (v/v) Fetal bovine serum (FBS), penicillin (100 IU/mL), and streptomycin (100 µg/mL). All cell lines were maintained in a humidified incubator containing 5% CO₂ at 37°C. For all treatments, a 10-mM solution of TQ (Sigma-Aldrich, Louis, MO, USA) was prepared in 10% dimethyl sulfoxide (DMSO) (Millipore, Molsheim, France) and appropriate working concentrations were prepared with cell culture medium. The final concentration of DMSO was always less than 0.1% in both control and treated conditions.
Cell proliferation assay
The effect of TQ on cell proliferation was analyzed by a colorimetric cell proliferation assay using WST-1 Cell Proliferation Reagent Kit (Sigma-Aldrich, catalog no. 11644807001). Briefly, the cells were seeded in 96-multiwell plates at a density of 104 cells/well (counted using Scepter 2.0 Handheld Automated Cell Counter, Millipore, Billerica, MA, USA; catalog no. PHCC20040) for MDA-MB-468 cells or 4 × 104/well for JK cells. After 24 h of incubation, the cells were exposed to different concentrations of TQ for the desired times. Cell proliferation rate was evaluated through a rapid WST-1 reagent. After incubation for the above-mentioned time, 10 µL of the WST-1 solution were added and cells were incubated for an additional 3 h at 37°C. Finally, the absorbance was read at 450 nm with a microplate ELISA (enzyme-linked immunosorbent assay) reader (ELx800, BioTek, USA) and the results were analyzed using the Gen5 software (BioTek, Winooski,Vermont). The reaction is based on the cleavage of the tetrazolium salt WST-1 to formazan by cellular mitochondrial dehydrogenases. The quantity of formazan dye in the medium is directly proportional to the number of viable metabolically active cells. The percentage of cell viability was calculated by assuming that control (untreated) samples are 100% viable.
RNA-seq and differentially expressed gene analysis
RNA-seq was carried out as described elsewhere. 34 Jurkat cells were treated with 20 µM TQ for 24 h, as this concentration was the nearest value to the half maximal inhibitory concentration (IC50), in triplicates. 10 Total RNA was extracted using RNeasy kit (Qiagen, Valencia, CA, USA). RNA quality was assessed using Bioanalyzer RNA integrity number (RIN score > 7.0). Then, the sequencing libraries were generated (TruSeq Stranded mRNA Sample Preparation Kits; Illumina, San Diego, CA, USA) from 2.5 µg of total RNA from each of the 3 biological replicates). Single-end deep sequencing (50 bp long) was performed using Illumina HiSeq 2000 System. The adaptor sequences were removed, and the low-quality base call was filtered by FASTX-Toolkit. The obtained filtered short sequencing reads were mapped to the human genome using TopHat2 and the subsequent gene expression values were quantified using Subreads package Feature Counts function. The DGE analysis was further performed from the gene expression values after identifying the library size and appropriate data set dispersion. Differential gene expression was tested using R/Bioconductor package edgeR. Differential gene expression is determined as the Log2 Fold-Change (Log2FC) with a false discovery rate (FDR) ⩽ 0.05 (Log2FC ⩾ 0.5 or ⩽–0.5). Gene ontology (GO) analysis and BIOCARTA pathway analysis were then performed by setting all the GO terms and BIOCARTA pathway genes as background genes. Overrepresented GO terms or pathways are determined by enrichment score (EASE ⩽ 0.1, gene count ⩾ 2). The heatmap of the genes responsible for epigenetic regulation was plotted using the web tool ClustVis, as per the mentioned algorithm.
Apoptosis assay
To study the apoptosis, JK cells were seeded in 96-well plates at a density of 4 × 104 cells/well, grown for 24 h, and exposed to TQ at different concentrations for 24 h. Cell apoptosis rate was assessed using the Annexin V Binding Guava Nexin Assay by capillary cytometry (Guava Easycyte Plus HP system, with absolute cell count and 6 parameters) following the manufacturer’s recommendations (Guava Technologies, Inc, Hayward, CA, USA). Guava Nexin Assay utilizes Annexin V-PE.
Real-time reverse transcription polymerase chain reaction analysis
Real-time reverse transcription polymerase chain reaction (RT-PCR) analysis was described elsewhere. 35 Briefly, the cells were treated with different concentrations of TQ for 24 h. Then, total RNA was purified and subjected to reverse transcription using Oligo(dt) (Sigma, Steinheim, Germany) and Superscript II reverse transcriptase (Invitrogen, Carlsbad, CA, USA). Quantitative real-time PCR was performed using the LightCycler 480 SYBR Green I Master Kit (Roche Diagnostics, Indianapolis, Indiana, USA) and the Mastercycler Realplex apparatus (Eppendorf, Montesson, France). The results were normalized with Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) mRNA. The sequences of the primers for PCR amplification were UHRF1 (sense: 5′-GTCGAATCATCTTCGTGGAC-3′, antisense: 5′-AGTACCACCTCGCTGGCAT-3′), DNMT1 (sense: 5′-GCACAAACTGACCTGCTTCA-3′, antisense: 5′-GGCCTTTTCACCTCCATCAA-3′), HDAC1 (sense: 5′-GACAAGGCCACCCAATGAAG-3′, antisense: 5′-GCTTGCTGTACTCCGACATG-3′), G9a (sense: 5v-GGAGAAGTGACCCTGACGAA-3′, antisense: 5′-CCTCTTCCTCCTCCTCCTCT-3′), and GAPDH (sense: 5′-GGTGAAGGTCGGA-GTCAAC-3′, antisense: 5′AGAGTTAAAAGC-AGCCCTGGTG-3′). Amplicons were size controlled on agarose gel, and purity was assessed by analysis of the melting curves at the end of the RT-PCR reaction.
Statistical analysis
All data were presented as mean ± SEM of triplicates done in the same experiment or an average of at least 3 separate experiments. The differences between control and treated conditions were analyzed by Student t test (2-tailed) using GraphPad Prism 6 (GraphPad Software, San Diego, CA, USA), and the significant differences were indicated as
Results
Pro-apoptotic effects of TQ involve modulations of several writer and reader enzymes implicated in epigenetic process
We first analyzed gene expression in JK cells incubated for 24 h with 20 µM of TQ, a concentration approximately corresponding to its half-maximal activity.
10
RNA-seq was done using next-generation sequencing, as described in the “Materials and Methods” section. The data obtained from RNA-seq showed that epigenetic integrator
Downregulated genes involved in epigenetic pathways in TQ-treated Jurkat cells as compared with untreated cells.
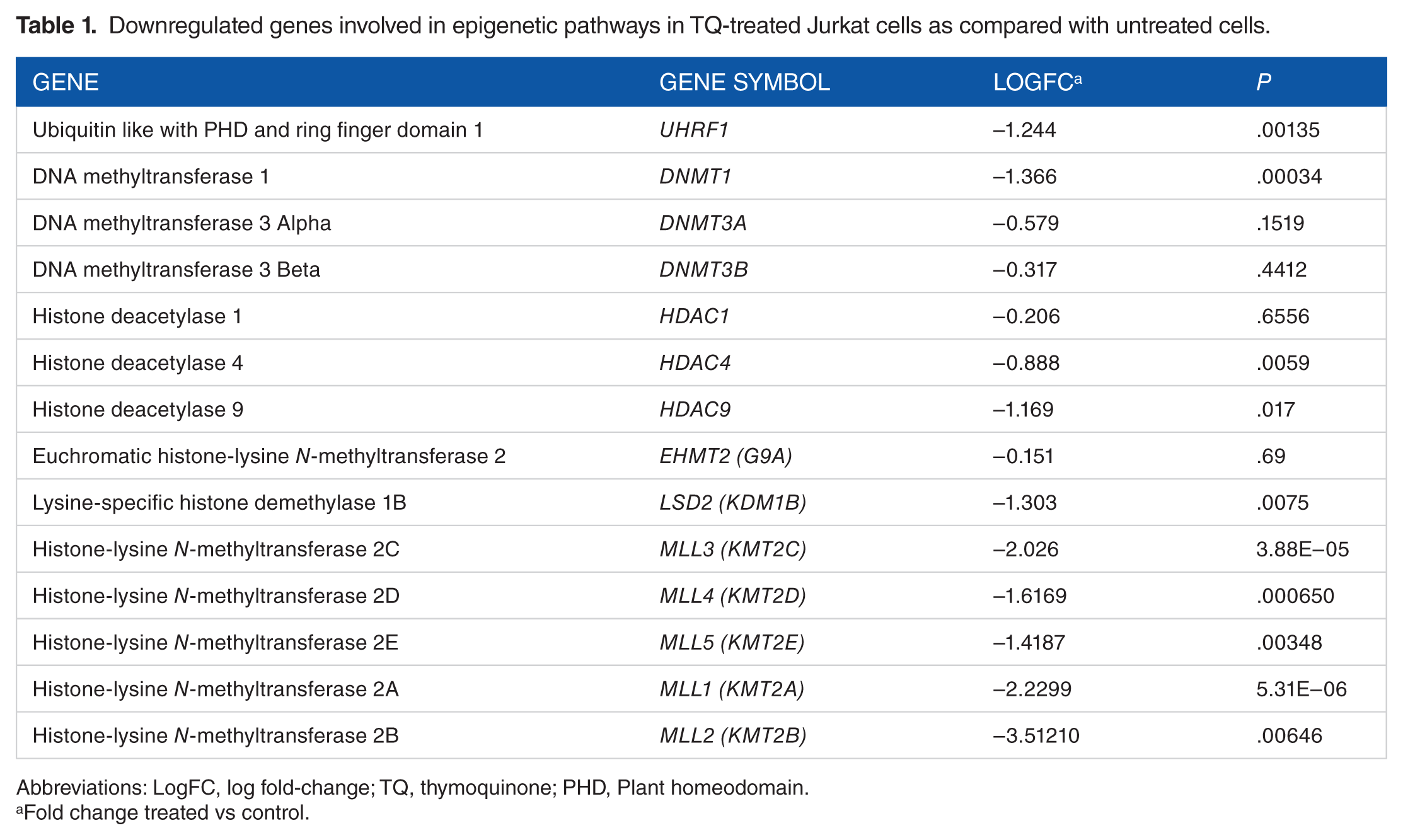
Abbreviations: LogFC, log fold-change; TQ, thymoquinone; PHD, Plant homeodomain.
Fold change treated vs control.
Upregulated tumor suppressor genes in TQ-treated Jurkat cells as compared with untreated cells.

Abbreviations: LogFC, log fold-change; TQ, thymoquinone.
Fold change treated vs control.
Upregulated pro-apoptotic genes in TQ-treated Jurkat cells as compared with untreated cells.

Abbreviations: LogFC, log fold-change; TQ, thymoquinone.
Fold change treated vs control.
These results suggest that the TQ-induced upregulation of TSGs leading to apoptosis in JK cells involves epigenetic mechanisms. The heatmap presented in Figure 1 gives an overall overview of the expression of the modulated genes with respect to both Log2FC and “

Heatmap of significantly altered Differentially expressed genes (DEGs) showing the corresponding change in expression. The intensity of the color varies with the LogFC from −1 to +2 in TQ-treated Jurkat cells as compared with untreated cells. LogFC indicates log fold-change; TQ, thymoquinone.

Different gene interactions of the most significantly deregulated Differentially expressed genes (DEGs) are represented as strings depicting cell death and tumor suppressor genes in TQ-treated Jurkat cells as compared with untreated cells. TQ indicates thymoquinone.
TQ decreased mRNA expression of UHRF1, DNMT1, HDAC1, and G9a in cancer cells
Our previous study showed that TQ-induced apoptosis in JK cells is associated with downregulation of the protein expression of UHRF1, DNMT1, and HDAC1. 10 Thus, we studied the effect of 5 and 10 µM TQ for 24 h on cell viability and mRNA expression of UHRF1, DNMT1, HDAC1, and G9a in JK (Figure 3) and MDA-MB-468 cells (Figure 4), as a model of solid tumor, using RT-qPCR. We found that mRNA expression of target genes was significantly decreased in a dose-dependent manner in JK (Figure 3A) and MDA-MB-468 cells (Figure 4A) treated with TQ compared with control. Under the same conditions, this effect was associated with a significant decrease in cell viability in JK (Figure 3B) and MDA-MB-468 cells (Figure 4B), as well as with a significant increase in apoptosis in JK cells (Supplemental Figure 1), suggesting a significant role for these epigenetic regulators in the anti-proliferative and pro-apoptotic effects of TQ in cancer cells.

Effect of thymoquinone on cell viability and mRNA expression of UHRF1, DNMT1, HDAC1, and G9a in Jurkat cells. Cells were exposed to increasing concentrations of TQ for 24 h. (A) The histograms show the quantification data of mRNA expression for UHRF1, DNMT1, HDAC1, and G9a as assessed by real-time PCR. (B) Cell viability rate was assessed by WST-1 assay. Values are shown as mean ± SEM (n = 3). *

Effect of thymoquinone on cell viability and mRNA expression of UHRF1, DNMT1, HDAC1, and G9a in MDA-MB-468 cells. Cells were exposed to increasing concentrations of TQ for 24 h. (A) The histograms show the quantification data of mRNA expression of UHRF1, DNMT1, HDAC1, and G9a as assessed by real-time PCR. (B) Cell viability rate was assessed by WST-1 assay. Values are shown as mean ± SEM (n = 3). **
Discussion
The epigenetic silencing of TSGs is a common characteristic in human cancer cells and is considered one of the main mechanisms involved in the regulation of TSGs.3,36,37 This process is mainly ensured through a coordinated dialogue between DNA methylation and histone post-translational modifications, such as acetylation and methylation. In cancer cells, promoters of several key TSGs are hypermethylated by the DNMT1 enzyme, and histone proteins are deacetylated and/or hypermethylated by HDACs and HMTs, respectively. Consequently, those TSGs are inhibited with the subsequent defect in apoptosis.38–41 Therefore, many drugs have been developed that act as inhibitors of DNMT, HDACs, and HMTs leading to the re-expression of TSGs.42–46 In the present study, we evaluated the effect of TQ on the epigenetic code of JK cells, an established cell line for acute T-cell leukemia. By analyzing the data obtained from next-generation sequencing, we found that many key epigenetic players were downregulated in TQ-treated JK cells, including the epigenetic integrator
The promoter of TSG
Next-generation sequencing data were confirmed using RT-qPCR that showed that TQ significantly decreased the expression of
In the present study, we also evaluated the effect of TQ on MDA-MB-468, a human epithelial breast cancer cell line characterized as triple-negative/basal-A mammary carcinoma, as a model of solid tumor. Similar to our results in JK cells, TQ was able to significantly decrease the cell viability and the expression of
Several studies have shown the presence of a coordinated crosstalk between DNA methylation and histone methylation in cancer cells leading to gene expression inhibition.68–70 One of the most important histone modifications, having inhibitory effects on the expression of TSGs, is H3K9me2 or H3K9me3, catalyzed by G9a.71,72 In the present study, we found, for the first time, that TQ significantly decreased the expression of G9a in both cancer cell lines from blood and solid tumors, suggesting that TQ-induced inhibition of G9a is key in the reactivation of TSG and induction of apoptosis. Indeed, inhibition of G9a in breast tumor cell lines using 5-aza-2′-deoxycytidine was reported to be sufficient to induce a reactivation of 2 TSGs,
Conclusions
The present study suggests that the inhibitory effects of TQ on both blood and solid tumors involve epigenetic mechanisms leading to the upregulation of several TSGs with subsequent apoptosis. These results also suggest that TQ could be used as an epigenetic drug that targets both DNA methylation and histone post-translational modifications, which could be a promising strategy for the epigenetic therapy of both types of tumors.
Supplemental Material
Supplemental_data – Supplemental material for Thymoquinone-Induced Reactivation of Tumor Suppressor Genes in Cancer Cells Involves Epigenetic Mechanisms
Supplemental material, Supplemental_data for Thymoquinone-Induced Reactivation of Tumor Suppressor Genes in Cancer Cells Involves Epigenetic Mechanisms by Shahad A Qadi, Mohammed A Hassan, Ryan A Sheikh, Othman AS Baothman, Mazin A Zamzami, Hani Choudhry, Abdulrahman Labeed Al-Malki, Ashwag Albukhari and Mahmoud Alhosin in Epigenetics Insights
Footnotes
Acknowledgements
The data and material presented in this manuscript have neither been published before nor have been submitted for publication to another scientific journal or are being considered for publication elsewhere. All the co-authors have read this manuscript and approved it for submission.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors would like to acknowledge the support provided by King Abdulaziz City for Science and Technology (KACST), Grant no. 1-17-01-009-0010.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MA and AA designed the project and wrote the paper. SAQ, MAH, and RAS performed research and analyzed data. OASB, MAZ, ALA and HC helped with experimental design, data interpretation, and drafting of the paper.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
