Abstract
High-throughput screening (HTS) has become an indispensable tool for the pharmaceutical industry and for biomedical research. A high degree of automation allows for experiments in the range of a few hundred up to several hundred thousand to be performed in close succession. The basis for such screens are molecular libraries, that is, microtiter plates with solubilized reagents such as siRNAs, shRNAs, miRNA inhibitors or mimics, and sgRNAs, or small compounds, that is, drugs. These reagents are typically condensed to provide enough material for covering several screens. Library plates thus need to be serially diluted before they can be used as assay plates. This process, however, leads to an explosion in the number of plates and samples to be tracked. Here, we present SAVANAH, the first tool to effectively manage molecular screening libraries across dilution series. It conveniently links (connects) sample information from the library to experimental results from the assay plates. All results can be exported to the R statistical environment or piped into HiTSeekR (http://hitseekr.compbio.sdu.dk) for comprehensive follow-up analyses. In summary, SAVANAH supports the HTS community in managing and analyzing HTS experiments with an emphasis on serially diluted molecular libraries.
Keywords
Introduction
High-throughput screening (HTS) is a key technology in functional genomics and drug (target) research. 1 Thanks to a high degree of automation and the use of robotics, researchers may nowadays perform experiments on a large (>1000) to ultralarge scale (>100,000). HTS was first applied by the pharmaceutical industry for small-compound screens, but is now also an important tool in functional genomics 2 and drug target discovery. 3 The basis for these screens is formed by molecular libraries, that is, sets of plates containing the samples or reagents to be tested on a particular cell line and under specific experimental conditions. For instance, a genome-wide siRNA screen spans more than 200 × 96-well or more than 50 × 384-well microtiter plates. Molecular libraries are typically manufactured in a highly condensed form to allow performing several screens with the available material. Hypothetically, the solubilized reagents in a library plate can simply be resuspended, allowing for small amounts to be extracted, producing a single set of assay plates. However, the quality of subsequent screens might be affected, since reagents will slowly degrade with each freeze/thaw cycle. Consequently, library plates need to be diluted in as few freeze/thaw cycles as possible. Assay plates obtained in this way can then be thawed and directly used for a screen when needed. However, fully diluting an entire library would lead to a large number of plates that typically have to be stored at −80°C, posing a storage problem. Therefore, library plates are typically diluted in several steps ( Fig. 1 ). At first, each library plate is diluted and samples are divided between a number of master plates. These sets of master plates, each representing the full library, are then stored in a freezer, except for one set, which is processed further. Again, the samples are diluted and transferred to a number of so-called mother plates. Only a single set of mother plates is in turn processed further and diluted into daughter plates. Master, mother, and daughter plates are thus technically identical and only differ in the concentration of the sample, where only daughter plates are appropriately diluted for screening. Each set of daughter plates, also called assay plates, covers an entire screen. This process enables conservation of a large portion of the screening library in a concentrated form, while reducing freeze/thaw cycles to a minimum.
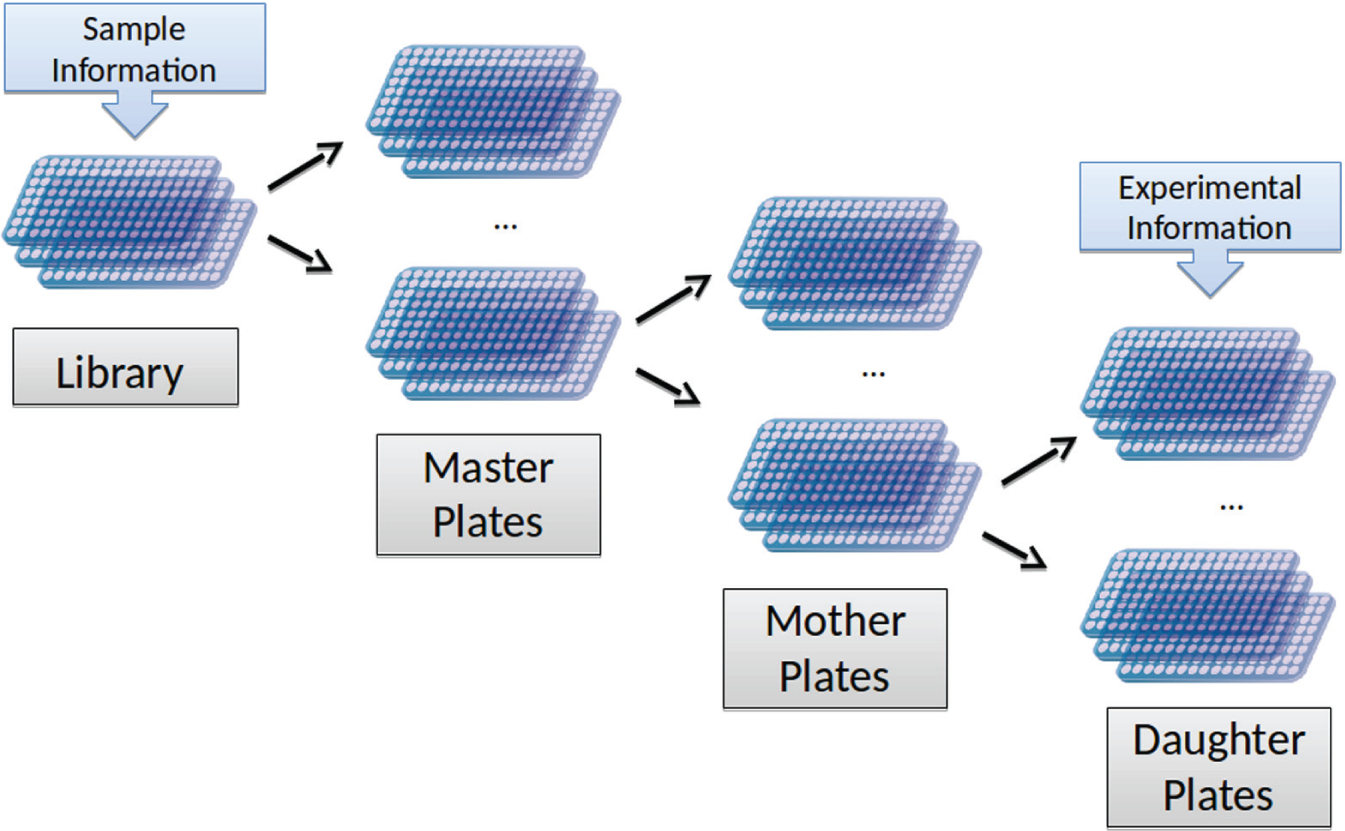
HTS libraries are typically delivered in a highly concentrated form. To keep the number of freeze/thaw cycles low and to minimize the freezer space needed for the entire library, plates are diluted stepwise via master and mother plates, down to daughter plates, which can finally be used as assay plates in a screen.
Depending on the actual concentrations, a screening library can be used for dozens of screens. A more comprehensive overview of this procedure can be found in Entzeroth et al. 4 Over time, the thousands of samples in a library are distributed across hundreds or thousands of plates. This demands a systematic solution for sample management. Based on our experience with HTS in cancer stem cell research, 5 we have developed Sample Management and Visual Analysis of High-Throughput Screens (SAVANAH), a free web application that addresses common issues in HTS library sample management.
Requirements
We identified a number of requirements, listed in the following as R1–R28, that an HTS sample management system should optimally fulfill. These requirements are the basis for a comprehensive comparison of SAVANAH with two existing applications in Table 1 .
Feature Comparison between SAVANAH and Comparable Open-Source Solutions for Managing High-Throughput Screening Data.
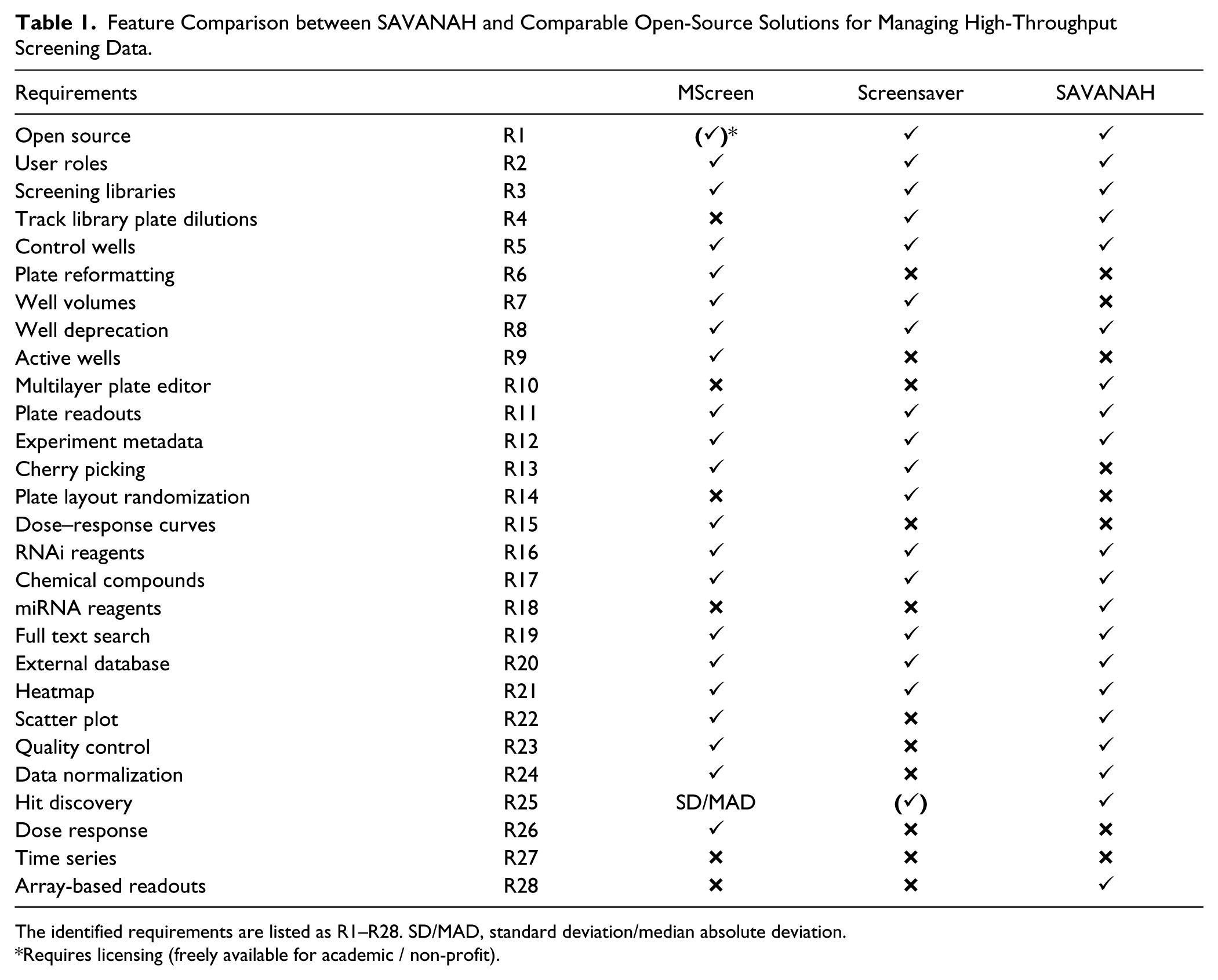
The identified requirements are listed as R1–R28. SD/MAD, standard deviation/median absolute deviation.
Requires licensing (freely available for academic / non-profit).
A sample management system for HTS should be open source (R1) to allow for adapting the system to the particular requirements of individual screening facilities. Moreover, access should be restricted based on different user roles (R2). The system should be able to import and manage sample information associated with molecular screening libraries (R3). The multitude of plates representing different dilution stages of a library requires an efficient plate tracking system. Consequently, an HTS sample management system should model the entire library dilution process in a user-friendly way (R4). In addition, the system should keep track of empty wells in the library plates that are reserved for controls (R5). Occasionally, it might be necessary to allow the user to transfer samples to a different plate format, for example, from 96- to 384-well plates (R6). Apart from monitoring the well volumes (R7), it would be advantageous to avoid considering inactive or deprecated samples. The system should thus allow for the corresponding well to be flagged as deprecated (R8) or even automatically assess the activity of all samples across different screens (R9). The sample information for all sets of diluted plates is identical, which suggests that entering and keeping this information in the system redundantly should be avoided (R10). Finally, only the fully diluted daughter plates are actually used as assay plates in a screen. For those plates, the sample information of each well needs to be linked to data originating from one or several experimental readouts (R11). Apart from information about an individual sample, experiment parameters need to be tracked that are often shared across a batch or an entire screen (R12). This includes, for instance, reagents that have been used or the cell lines that have been tested. Secondary screening can be guided by so-called cherry picking (R13), where the most promising hits are transferred in silico to a new assay plate. Ideally, an automated mechanism for randomizing the sample position is included to avoid clustering similar reagents (R14). The system should cover typical use cases like RNAi screens (R16), small-compound screens (R17), and miRNA inhibitor and mimics screens (R18). All samples should be searchable (R19) and linked to external databases (R20) to provide additional information to the user. Data should be visualized (R21 and R22) and subjected to quality control (R23). After appropriate normalization to counter known sources of variation (R24), hit discovery (R25) can be performed to filter for active samples that show an effect. In a typical screen, each sample is considered an independent experiment. There are, however, other types of HTS with related experiments, such as dose–response (R26) or time-series (R27) experiments, which would benefit from dedicated features. Finally, HTS readout data could be followed up in a secondary readout with another technology, such as reverse-phase protein arrays (RPPAs), suggesting that support be included for linking such data (R28).
State of the Art
Most bioinformatics software tools for HTS focus on statistical data analysis and lack capabilities to address the sample management issues. Furthermore, vendor-provided software for HTS systems typically focuses on plate logistics required to drive the robotic platform. Beckman-Coulter (http://www.beckman.com/liquid-handling-and-robotics/liquid-handling-software), as an example, provides tools for liquid handling and plate transfer (SAMIex), normalization and hit picking (PowerPack), and data reporting (DART). DART facilitates easy integration with third-party laboratory information management systems (LIMS), which leaves a gap in the efficient library handling in large-scale screening projects. To close this gap, powerful commercial solutions have emerged, such as, for instance, IDBS ActivityBase (https://www.idbs.com/en/platform-products/activitybase/). However, the costs of such a system are usually prohibitive in academic research. In the past, few freely available LIMS have been developed to accommodate small-compound and RNAi screens. OpenBIS 6 is a general-purpose LIMS that includes support for high-content screening, where the focus is mainly on image acquisition and processing. In HTS, however, only numeric readouts are produced. One system accommodating this type of data is Screensaver, 7 which has been developed at the Harvard Medical School to manage siRNA, as well as compound screens. The system keeps track of screening libraries, including the remaining volume of the wells. Reagents are linked to external databases to provide additional information. Furthermore, Screensaver provides cherry picking, which allows promising hits to be selected and assembled for secondary screening. Plates can be randomized and wells can be flagged as deprecated to exclude them from future screens. Normalization and hit detection are not supported and have to be performed externally. However, the results of such an analysis can subsequently be uploaded back into the system. An alternative to Screensaver is MScreen, which has been developed at the University of Michigan. 8 MScreen provides similar features and in addition supports dose–response curves, quality control, normalization, and hit detection. However, neither MScreen nor Screensaver supports miRNA mimics or inhibitor libraries. In both systems, all sample information is uploaded via externally defined files. While it is an appropriate strategy to upload information about a screening library, which is only necessary once, it would be advantageous if users were offered a convenient and interactive interface to design smaller experiments manually. This is of particular interest for CRISPR/Cas9 screens, which have emerged as the new state of the art for genome editing. While newly developed tools such as CRISPRdirect 9 can be used to easily generate custom guide RNAs to direct the Cas9 nuclease, additional challenges in managing a rising number of custom experimental designs are not met. The lack of a system that supports small customized experiments, as well as large-scale HTS, motivated us to develop SAVANAH. See Table 1 for a feature comparison between SAVANAH, Screensaver, and MScreen.
Material and Methods
Implementation
SAVANAH was built using Grails (http://grails.org), a Java/Groovy web application framework that enables rapid development of robust applications. Grails is supported by a plethora of open-source plug-ins, which deliver high-quality solutions for nontrivial web application tasks, for example, database search (Compass and Apache Lucene, http://www.compass-project.org/), spreadsheet import and export (Apache POI, https://poi.apache.org/), and user/security management (SpringSecurity, http://projects.spring.io/spring-security/).
Data export to R in general and to HiTSeekR in particular is realized through a set of REST web services. Here, data retrieval is facilitated through simple HTTP requests by the client, which are replied to by the server that sends experimental data in the lightweight Java Serial Object Notation (JSON) format. The efficient conversion between database content and JSON objects on the server side is facilitated by the Jackson library (https://github.com/FasterXML/jackson). JSON is converted into R data structures via the RJSONIO package (https://cran.r-project.org/web/packages/RJSONIO/index.html).
Results
SAVANAH aims to simplify HTS data management for screening data, supporting both small experiments involving manually created plates and large-scale HTS. To achieve the latter, SAVANAH offers a set of features for managing screening libraries efficiently.
Plate Layouts
In many cases, experiments involve several replicate plates to accommodate technical and biological variation. To avoid redundancy, SAVANAH thus separates the so-called plate layouts holding the sample information from the actual plates. This has the advantage that changes applied to the layout will automatically be propagated to the respective plates. To avoid long and cryptic sample names, sample information is divided in several categories shown in Figure 2 . Each property is modeled as a distinct layer of the plate layout, which provides a clear data structure to the system.
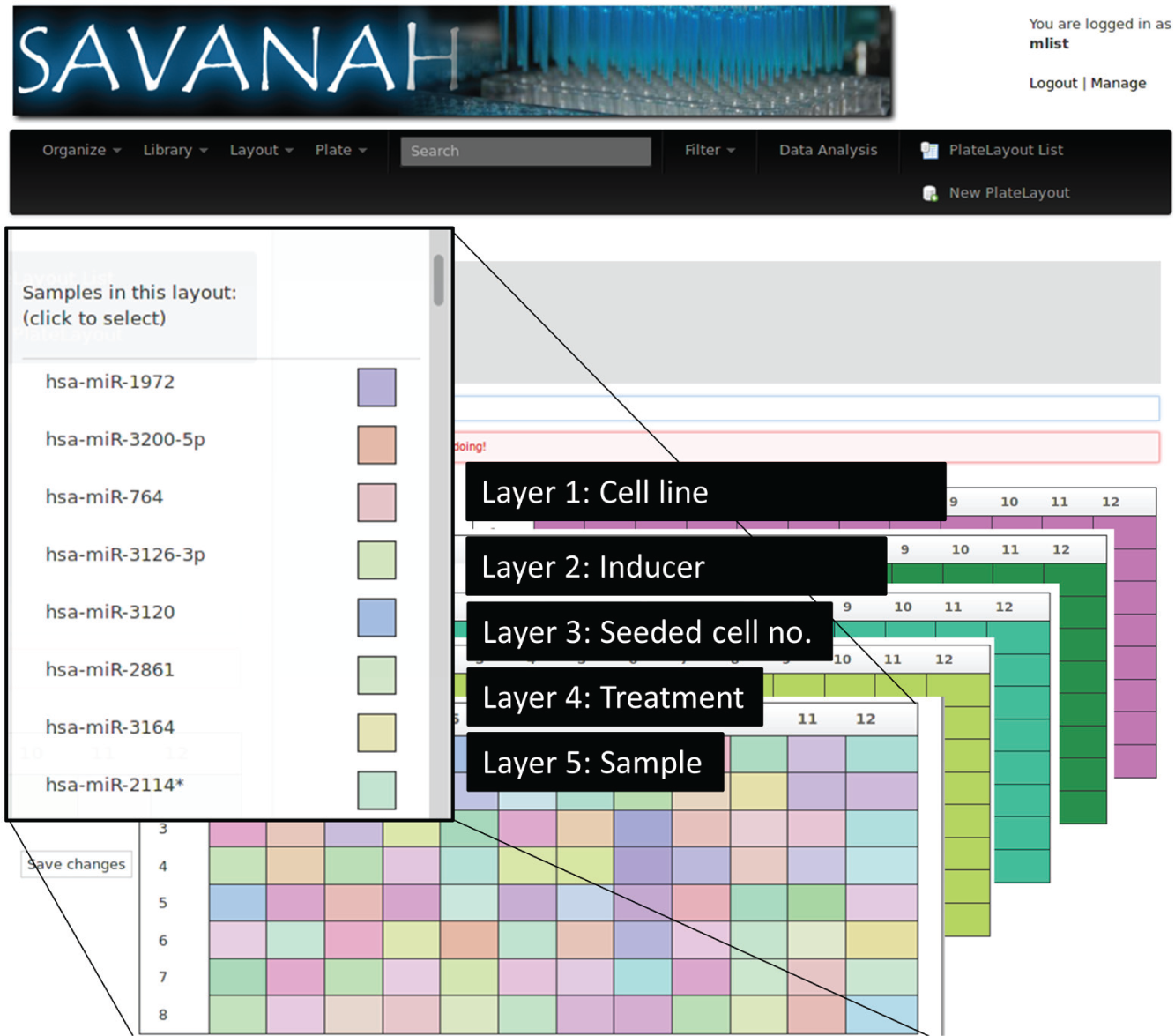
Plate layouts in SAVANAH capture sample information in several layers and can be conveniently edited by the user. This multilayer approach avoids the use of complex and ambiguous sample names. Each layer can be edited interactively by the user.
Plates and Readout Import
In contrast to plate layouts, which hold sample information, plate objects identify physical plates by properties such as their plate type or barcode. Moreover, plates can be linked to measurements, where readout data can be uploaded for each plate individually or as a batch import. For the latter, the user is required to add all readout files to a zip file in which each file is named after the unique barcodes of the plate. SAVANAH accepts XLSX-, comma-, tab-, and semicolon-separated files as input for readouts. During the import, the columns of the input file can be mapped to the expected properties, such as the well position and signal value. The uploaded readout data can subsequently be visualized as a scatter plot or a heatmap ( Fig. 3 ).
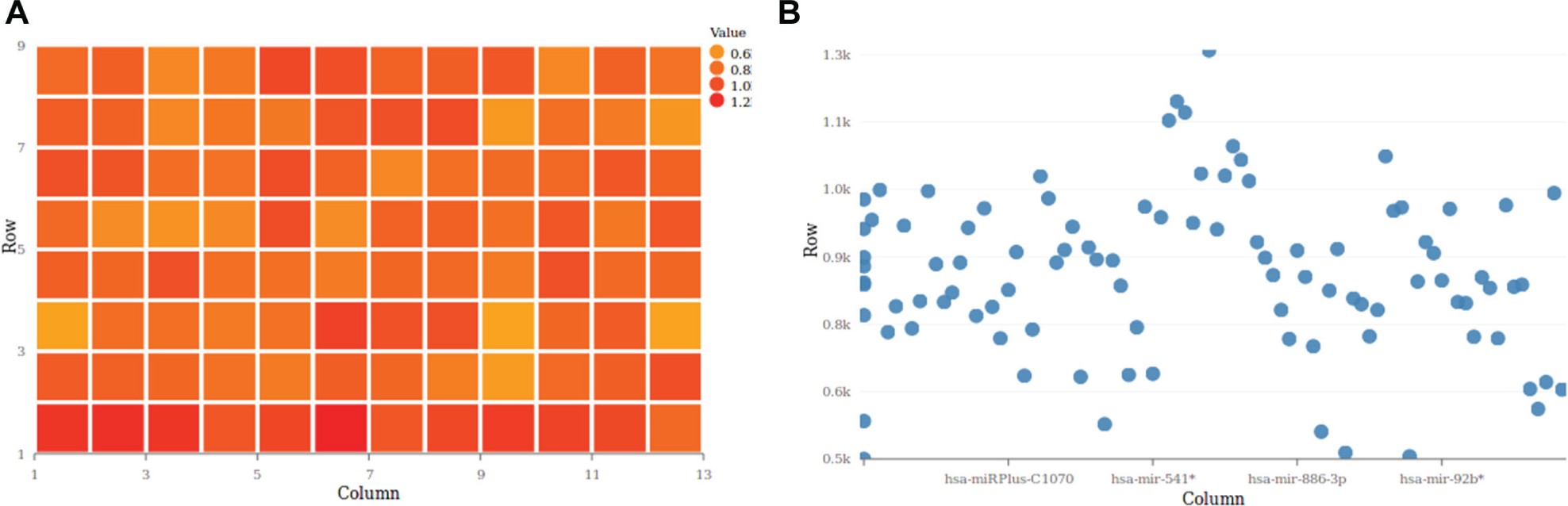
SAVANAH visualizes readout data in two ways: (
Projects and Experiments
To keep a growing number of plate layouts organized, users are encouraged to group them into experiments, which in turn can be grouped into projects. First, this will make it easier to apply the search function efficiently. Moreover, a filter is available in the navigation menu. Here, a specific experiment and/or project can be selected by the user. Whenever the user opens a list view, for example, a list of plate layouts or a list of plates, only items that are associated with the selected experiment and/or project will be displayed.
Screening Libraries
A screening library comprises a set of library plates with hundreds or thousands of molecules to be tested. For each of them, information about the plate and well location, as well as sample and product identifiers, needs to be added to the system. To this end, a tab-separated file can be uploaded. SAVANAH expects this file to have a specific header, which is further described in the user manual (http://nanocan.github.io/SAVANAH/). SAVANAH will process this information and create a library together with the corresponding library plates. To avoid multiple database entries for the same sample, the system will check if an existing entry can be used by looking up the sample name and sample accession in the database.
Library Dilutions
Prior to use, library plates are serially diluted via master and mother plates to daughter plates, which are also called assay plates. Consequently, a large number of library plate copies need to be tracked. To avoid burdening the user with creating and linking these plates individually, SAVANAH introduces the concept of library dilutions. Each library dilution represents a full copy of the original library and can be of either “Master,” “Mother,” or “Daughter” type. In addition, diluted libraries of type “Mother” and “Daughter” keep track of their respective source.
Platform Integration
Data analysis: Follow-up data analysis, including normalization, quality control, and hit detection, is supported through export functionality to R or HiTSeekR (http://hitseekr.compbio.sdu.dk). 10 HiTSeekR is a dedicated web application suited for the analysis of various types of HTS data ranging from quality control, normalization, and hit discovery to a systems biology evaluation.
Functional genomics: Several tools are available to modify existing cell lines to exhibit a certain trait, such as the controlled (over-)expression of a particular gene. The effect of such modifications can then be characterized systematically using HTS. To keep track of the numerous modified cell line samples, we linked SAVANAH to the LIMS OpenLabFramework, 11 which is tailored toward functional genomics to enable researchers to conveniently identify the location of the cell lines tested in a particular HTS experiment.
Reverse-phase protein arrays: The number of assays that can be performed in parallel in an HTS setup is limited. A promising strategy to gain more information is thus to transfer the samples after the primary HTS readout onto RPPAs for a secondary readout. RPPAs require only small sample volumes, allowing several identical array copies to be printed and each of them to be interrogated with a different antibody. In this way, HTS experiments can be augmented with additional information about the expression levels of proteins of interest, their phosphorylation status, and so forth. 12 This also adds another level of complexity to sample tracking, which we address by integrating SAVANAH with MIRACLE, 13 a web application tailored toward sample management and analysis of RPPA experiments. Both web applications can utilize a shared database, allowing users of MIRACLE to directly utilize plate layout and sample information from SAVANAH, as well as plate readout data in the subsequent array spotting and data analysis.
Use Cases
SAVANAH is implemented for a wide range of screen types: (1) gene-targeted screens facilitated by siRNA-, shRNA-, or sgRNA-mediated silencing; (2) gene-activating screens facilitated by sgRNA screens; (3) small-compound screens; and (4) miRNA inhibiting or mimicking screens. SAVANAH supports both large and small screen experiments, which we describe in two typical use cases.
Large-scale screen: In a use case for a large-scale experiment, a researcher is typically equipped with an already prepared library covering several microtiter plates, as well as a tabular file describing the content of each well. SAVANAH enables users to upload such a library file to the system to generate an in silico representation of the library (
Small-scale screen: To perform a small-scale screen, for example, for hit validation of a larger primary screen, users can create an experiment with a series of new (empty) plates. Each plate is opened in the plate layout editor, where the user selects one or several wells to manually add samples and controls previously entered into the system. Moreover, additional sample information pertaining to the used cell line or the applied treatment, for instance, can be added. Finally, readout data can be uploaded and linked to each plate to facilitate further analysis. As the main advantage, the user avoids ambiguity in sample naming and can quickly identify all experiments and plates related to a condition or sample of interest.
Discussion
A major challenge for any screening center regularly performing HTS is the continuously growing number of plates and sample information that need to be tracked efficiently. To make this elaborate task easier, SAVANAH bundles diluted copies of library plates as abstract entities and handles the bulk of the plate setup automatically. This separates SAVANAH from similar tools, such as Screensaver or MScreen. Another unique feature is the distinction between plate layouts and actual plates, which makes it possible to edit the sample information for all replicates at once. Finally, the interactive plate layout editor makes sample information accessible and editable in a convenient way. In addition to HTS, SAVANAH is ideally suited for designing and managing small customized experiments. This also makes the system attractive for laboratories that do not perform HTS, but deal with a large number of custom plate designs. The multilevel sample management may at first appear as a burden, since users have to tediously add sample information before designing plate layouts and experiments. However, once this process is complete, sample properties can be applied quickly and efficiently, while the data model ensures consistency and efficient information retrieval. To allow users to familiarize themselves with SAVANAH, we offer a user manual (http://nanocan.github.io/SAVANAH), as well as an online demo (http://nanocan.org/savanah/demo). The link to HiTSeekR for direct follow-up data processing and systems biology analyses is a clear advantage compared with platforms, such as Screensaver, which currently require manual data export. Finally, the integration of SAVANAH with MIRACLE uniquely enables researchers to combine two complementing high-throughput technologies, namely, HTS and RPPAs, to maximize the information gain of costly screening experiments.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Lundbeckfonden grant for the NanoCAN Center of Excellence in Nanomedicine, the Region Syddanmarks ph.d.-pulje and Forskningspulje, the Fonden Til Laegevidenskabens Fremme, the DAWN-2020 project financed by the Rektorpuljen SDU2020 program, and the MIO project of the OUH Frontlinjepuljen. Furthermore, MPE acknowledges the Grosserer M. Brogaard og Hustrus Fond and the Aase og Ejnar Danielsens Fond for their financial support.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
