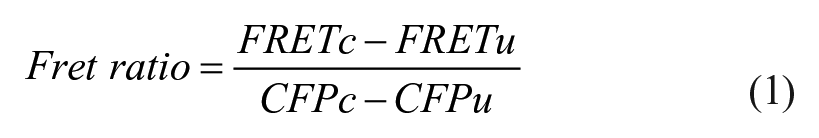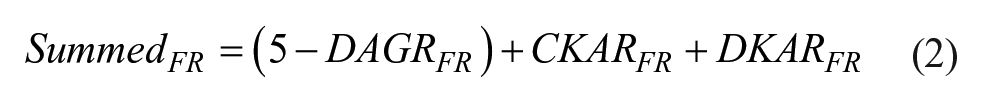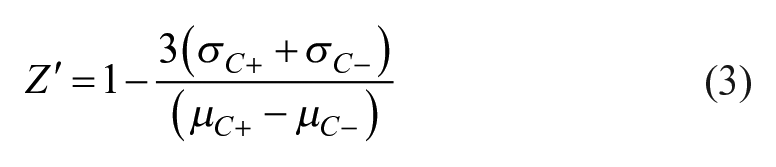Abstract
Genetically encoded sensors based on intramolecular FRET between CFP and YFP are used extensively in cell biology research. Flow cytometry has been shown to offer a means to measure CFP-YFP FRET; we suspected it would provide a unique way to conduct multiplexed measurements from cells expressing different FRET sensors, which is difficult to do with microscopy, and that this could be used for screening. We confirmed that flow cytometry accurately measures FRET signals using cells transiently transfected with an ERK activity reporter, comparing responses measured with imaging and cytometry. We created polyclonal long-term transfectant lines, each expressing a different intramolecular FRET sensor, and devised a way to bar-code four distinct populations of cells. We demonstrated the feasibility of multiplexed measurements and determined that robust multiplexed measurements can be conducted in plate format. To validate the suitability of the method for screening, we measured responses from a plate of bacterial extracts that in unrelated experiments we had determined contained the protein kinase C (PKC)–activating compound teleocidin A-1. The multiplexed assay correctly identifying the teleocidin A-1-containing well. We propose that multiplexed cytometric FRET measurements will be useful for analyzing cellular function and for screening compound collections.
Introduction
Genetically encoded sensors based on Förster resonance energy transfer (FRET) between fluorescent proteins (FPs) have seen wide use in cell biology. 1 There are two basic types of FRET reporter systems: intermolecular and intramolecular sensors. Intramolecular FRET sensors are comprised of a single chimeric protein that includes the two FPs and an engineered sequence that confers responsiveness to a desired stimulus. This results in stoichiometric expression of the two FPs, making the analysis of intramolecular FRET signals much simpler than analysis of intermolecular FRET signals (see, e.g., Violin et al. 2 and Depry and Zhang 3 ). Changes in intramolecular sensor FRET ratio provide a measure of the sensor’s target activity without the need to quantify absolute FRET efficiency, correct for bleed-through of the donor FP into the acceptor FP’s channel, or compensate for differing FP expression levels between cells or in different regions of a cell. Intramolecular FRET sensors have been developed to report a large variety of cellular parameters, such as the activity of various protein kinases, 4 cellular metabolic status,5,6 changes in intracellular and organellar calcium, 7 and caspase activation. 8
Signals from intramolecular FP FRET sensors are commonly monitored using microscopy, which allows time-lapse imaging and provides researchers the ability to resolve the spatial localization of signaling events. However, the method is fairly laborious to implement, largely because the magnitude of FRET ratio changes is typically only in the range of 5%–50%, and there can be a great deal of heterogeneity in the FRET ratio of different cells, due in part to intermolecular interactions between sensor molecules and variability in the status of the parameter the reporter senses. It is particularly difficult to perform measurements with multiple FP FRET sensors using imaging. A number of different strategies to do so have been developed (see Depry et al. 9 for a review). Multiple sensors utilizing the same FP FRET pair can be employed in microscopy if they can be targeted to sufficiently distinct subcellular localizations so that their activity can be discriminated spatially. Alternatively, FP FRET pairs can be used if they can be separated spectrally, which entails the use of FPs other the CFP (cyan fluorescent protein) and YFP (yellow fluorescent protein), or their spectrally related variants, which are used most commonly for FRET. CFP-RFP and YFP-RFP FRET pairs (RFP is red fluorescent protein) have been used, as have CFP-YFP- and OFP-RFP-based reporters (OFP is orange fluorescent protein). Both combinations allow acquisition of signals from two FP FRET constructs. Recently, Woehler reported a sophisticated method involving spectral “fingerprinting” that allowed the acquisition of signals from three FP FRET sensors with overlapping spectra and common spatial localization. 10 In this method, FRET pairs comprised of combinations of three different FPs (e.g., mTurquoise2-cpVenus, mTurquoise2-mCherry, and mCherry-cpVenus) are discriminated based on three-dimensional analysis of excitation–emission spectra. Validation of the method comes from experiments that show that it can follow simultaneously changes in cytosolic calcium, cAMP, and PKA activity. Alternative approaches for multiplexing that have been explored include fluorescence lifetime imaging of a CFP-YFP reporter of intracellular calcium together with a homo-FRET indicator of phosphinositides based on mCherry. 11
Cytometers with lasers capable of exciting CFP can measure CFP-YFP FRET. 12 The method has been used for measuring intermolecular FRET,13,14 and even to sort cells in which intermolecular FRET is occurring, 15 but has not often been used to measure intramolecular FP FRET. Caspase activation has been monitored using genetically encoded intramolecular FP FRET sensors in cytometry (e.g., He et al. 8 ), and Adachi and Tsubata measured FRET signals from a genetically encoded chameleon calcium sensor in B lymphocytes. 16 It is surprising that there has been relatively little use of cytometry to measure intramolecular FP FRET, as it should be technically simpler than measuring FRET in imaging, and because cytometry is an intrinsically high-throughput method, 8 it should allow measurement from many more cells.
We suspected that if cytometry provides a relatively simple means of measuring intramolecular FP FRET signals, it might make it possible to conduct multiplexed measurements of FP FRET in a new and different way. The strategy we envisioned would involve expressing different CFP-YFP-based FRET sensors singly in cell lines, and then labeling the different sensor-expressing cells with marker dyes to “bar-code” them 17 prior to mixing them, stimulating them, and measuring FRET. Provided bar coding does not interfere with FP FRET, the approach should be easier to implement than imaging approaches to multiplexing, as it relies on the inherent ability of cytometry to distinguish populations of cells labeled with different fluorophores. However, because a given cell would express only a single FRET construct, the method would not be useful for measuring the relationship of different parameters in single cells. It would only provide information about population behavior. However, we suspected that this might be extremely useful for certain purposes, including high-throughput screening, where we anticipated that it could enable the development of new kinds of powerful cell-based multiplexed screens.
Here we describe a series of experiments to explore the limitations of measuring intramolecular FP FRET with cytometry. We show that the method is easy to implement, but yields accurate measures of FRET ratios only for cells that express relatively high levels of the FP FRET construct. We demonstrate that it is possible to bar-code cells while preserving FRET ratios and FRET ratio changes, and show that the method can be used for multiplexed screening in plates.
Materials and Methods
Cells, Constructs, and Reagents
TALL-104 cells were obtained from the ATCC and cultured and transfected using nucleofection as described previously.
18
K562 cells were obtained from the ATCC and cultured in Iscove’s modified Eagle’s medium (MEM) supplemented with 10% heat-inactivated serum, glutamine, and penicillin-streptomycin, and were transfected using nucleofection according to the manufacturer’s recommended protocol. CDNAs encoding EKAR,
19
CKAR,
2
DAGR,
2
DKAR,
20
and D3CPV
21
were obtained from Addgene (see
FRET Imaging
Imaging of EKAR FRET in transiently transfected TALL-104 cells was done on a Zeiss Axiovert 200M inverted microscope equipped with a Sutter DG4 light source, a Uniblitz shutter system, and a Hammamatsu ORCA ER CCD camera. Open Lab software from Improvision was used to control hardware and acquire images. After acquisition, images were exported in TIFF format and imported into Slidebook software (Intelligent Imaging Innovations, Denver, CO), where it was background subtracted before ratios were computed. Ratio measurements were made for user-defined regions of interest after setting a threshold on the CFP image to avoid the contribution to the ratio of cell-free areas within the regions of interest (ROIs). Ratio data were exported and analyzed further using Excel.
Cytometry to Measure FRET
Flow cytometric measurement of FRET signals was performed using a BD FACSAria II Special Order cell sorter equipped with 405, 488, and 640 nm lasers. Three channels were collected to characterize FRET signals: CFP (405 nm excitation, 450/50 nm emission, 405 nm laser trigon), FRET (405 nm excitation, 525/50 emission, 405 nm laser trigon), and YFP (488 nm excitation, 530/30 emission, 488 nm laser octagon). For bar coding, singly labeled cells were stained with dyes at a final concentration of 100 ng/mL in normal Ringer’s solution, and then washed twice with medium before use. Unloaded cells were treated similarly. Doubly labeled cells were incubated with both dyes, each at 100 ng/mL. Bar-coding signals were acquired with 488 nm excitation and 575/20 nm emission (488 nm laser trigon) for calcein red-orange, and 640 nm excitation and 660/20 nm emission (647 nm laser trigon) for Far Red DDAO-SE.
Flow data were analyzed offline using FlowJo software. Samples were first gated on forward and side scatter. FRET ratios for the events in the live-cell gate were calculated in FlowJo according to eq 1:
where FRETc and CFPc are the fluorescence intensities of transfected cells acquired at the indicated wavelengths, while FRETu and CFPu are the fluorescence intensities of YFP-negative cells in the case of TALL-104 transient transfection or of untransfected parental K562 cells for long-term K562 cell experiments. For quantification of the FRET ratio, gates were drawn on plots of FRET versus CFP to select cells for inclusion (see
Fig. 2
and
Measuring FRET Ratio in 96-Well Plates
Plate experiments were performed on a BD Fortessa X20 analyzer equipped with a BD HTS sampler. Signals acquired for FRET ratio analysis were CFP, 405 nm excitation, 450/50 emission, 405 nm laser octagon; FRET, 405 nm excitation, 520/15 emission, 405 nm laser octagon; and YFP, 488 nm excitation, 530/30 emission, 488 nm trigon. Cells were bar-coded by loading them with calcein red-orange, Far Red DDAO-SE, or no dye essentially as described above. Calcein red-orange signals were acquired using 561 nm excitation and 585/15 nm emission, 561 nm octagon, and Far Red DDAO-SE signals were acquired with 640 nm excitation and 670/30 nm emission, 640 nm trigon. Samples were prepared in flat-bottomed Greiner half-area 96-well plates. A total of ~50,000–80,000 bar-coded cells in 10 µL of medium were added to wells that were prefilled with 80 µL of medium containing DMSO, 1 µM PMA, or test extracts at a final concentration of 10 µg/mL. After addition of cells, plates were vortexed and then read after 45 min at room temperature. Fifty microliters from each well was acquired at the maximum flow rate of 3 µL/s. Reading a plate took ~45 min. Data were analyzed as described above using FlowJo, and then exported to Microsoft Excel or Igor Pro. Heat maps were generated using online software from Plotly (https://plot.ly/). The heat map labeled “all sensors” in Figure 6C was generated according to eq 2:
where FR indicates FRET ratio. Subtracting the DAGR FRET ratio from the constant resulted in a signal that, like the CKAR and DKAR signals, decreased with activation, and preserved both the magnitude of the FRET ratio changes and their standard deviation. The plate used in Figure 6 to validate the 96-well format of a three-sensor assay and our determination that well A2 contains the known PKC-activator teleocidin 23 are described in a submitted manuscript (Zhao et al., submitted) and in the supplementary information.
Results
FRET Signals from a Genetically Encoded ERK Activity Reporter in Transiently Transfected TALL-104 Cells Are Similar When Measured with Microscopy and Flow Cytometry
Since there have been so few reports using flow cytometry to measure intramolecular CFP-YFP FRET, we first wanted to confirm that cytometry and microscopy yield similar measurements of intramolecular FP FRET signals when compared directly ( Fig. 1 ). We transiently expressed EKAR, a genetically encoded sensor of the activity of the mitogen-activated protein kinase (MAPK) ERK, 19 in TALL-104 human leukemic CTLs, and analyzed FRET using microscopy and flow cytometry. EKAR has been constructed in such a way that when it is phosphorylated by ERK, a conformational change occurs that brings CFP and YFP into closer proximity, resulting in an increase in FRET ratio.
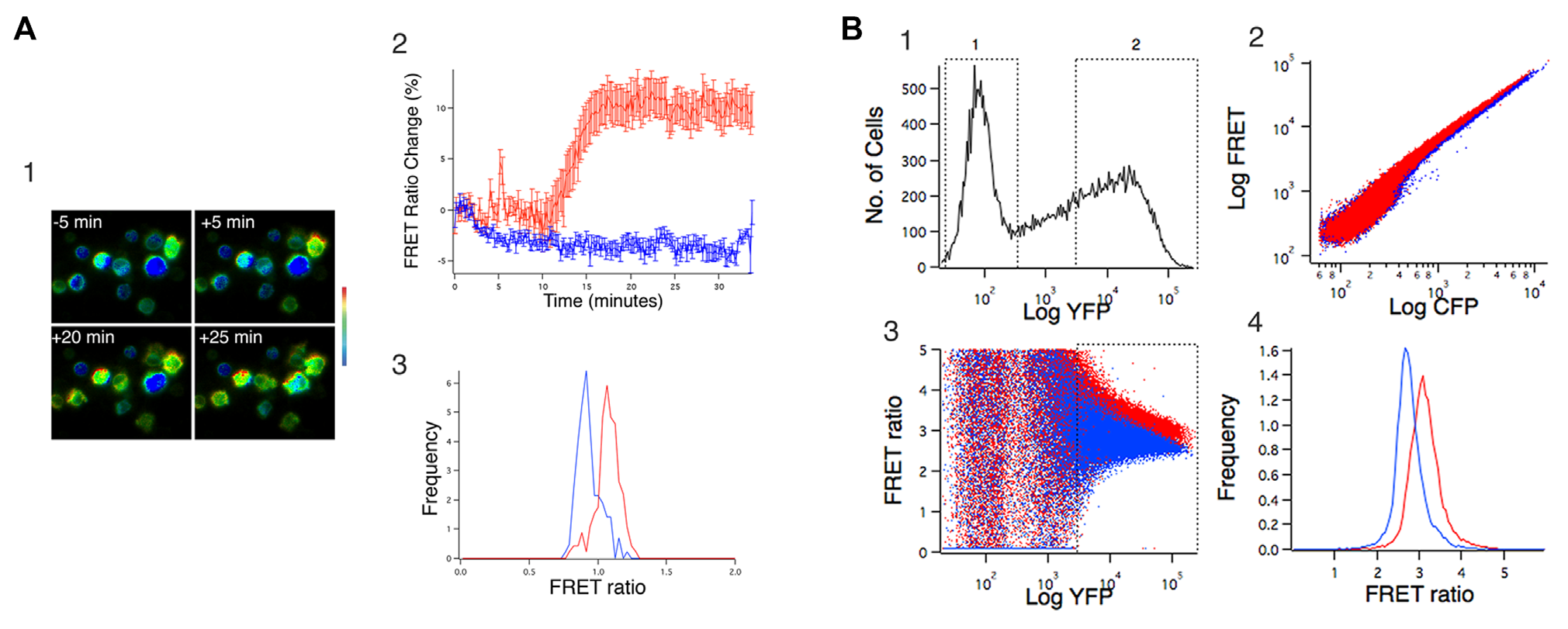
Measuring EKAR FRET in transiently transfected TALL-104 cells with microscopy and flow cytometry. (
A typical result using microscopy to measure EKAR signals is shown in Figure 1A . Figure 1A1 shows pseudocolored ratio images from a cluster of cells at various times before and after stimulation with 50 nM PMA, which we have shown previously activates ERK in TALL-104 cells in a PKC-dependent manner. 24 We stimulated the cells with 50 nM PMA, collecting time-lapse data over the course of 35 min for a field of view (which included the cluster of cells shown in Fig. 1A1 ) from which we were subsequently able to analyze data from 10 highly transfected cells. For these cells (red trace in Fig. 1A2 ), the FRET ratio increased from 0.93 ± 0.1 to 1.02 ± 0.09, an increase of ~10% (mean ± SD). The traces in Figure 1A2 are displayed after normalizing each cell’s FRET ratio to the average of its value for the first 10 frames, a common transformation that allows the percent change in FRET ratio to be estimated easily from visual inspection (see, e.g., Harvey et al. 19 ) and eliminates differences due to different initial FRET ratios. In separate imaging experiments, cells were left untreated for 35 min to serve as a control (blue trace in Fig. 1A2 ). A slight decrease in FRET ratio occurred over time in these cells.
Using microscopy, it is possible to measure changes in FRET ratio over time, providing a kind of internal control for each cell that enhances confidence that the small changes in FRET ratio observed are real. However, measuring FRET from the same cells over time is not possible using cytometry. To more closely approximate the kind of data we expected from cytometry, we performed the following analysis ( Fig. 1A3 ). Prior to the time-lapse imaging shown in Figure 1A1,2 , we first collected signals from 140 unstimulated cells in multiple microscopic fields. These cells displayed resting FRET ratios of 0.95 ± 0.085, with values ranging from 0.75 to 1.21. We also collected images and calculated FRET ratios for 151 cells after stimulation with PMA, again from multiple fields of view. FRET ratios for these cells were 1.07 ± 0.087, ranging from 0.79 to 1.27. Overall, the increase in FRET ratio comparing stimulated cells to unstimulated cells was 12.6%, consistent with the time-lapse imaging results. Quantitatively similar results were obtained in two other experiments.
We next examined the behavior of EKAR in transiently transfected TALL-104 cells using flow cytometry ( Fig. 1B ). We measured three signals, two for calculation of FRET ratios (CFP, 405 excitation/450 emission; FRET, 405 excitation/520 emission) and one to allow us to quantify sensor expression (YFP, 488 excitation/520 emission). A histogram of YFP fluorescence ( Fig. 1B1 ) shows that in this experiment, transient transfection led to high levels of expression in some cells, while other cells did not express the plasmid (gate labeled 1). Plotting FRET versus CFP fluorescence for unstimulated cells (blue) and for cells stimulated with 50 nM PMA (red) revealed a small shift to slightly higher levels of FRET fluorescence for a given level of CFP fluorescence upon stimulation ( Fig. 1B2 ), although this shift is extremely difficult to visualize on the logarithmic plots commonly used in cytometry. We computed the FRET ratio (FRET/CFP) on a cell-by-cell basis after subtracting average cellular autofluorescence, which we measured from untransfected cells in the sample (gate labeled 1 in Fig. 1B1 ). Those cells had comparable fluorescence values to control K562 cells that were not subjected to the nucleofection protocol (not shown). Plots of FRET ratio versus YFP fluorescence ( Fig. 1B3 ) reveal a great deal of scatter in FRET ratio at low levels of YFP fluorescence, suggesting that FRET ratios are poorly defined, likely due to low signals relative to cellular autofluoresence. However, as sensor expression levels increased, scatter in the FRET ratio decreased, appearing to converge to a relatively well-defined value. Figure 1B4 shows histograms of FRET ratio data for unstimulated and PMA-stimulated cells that were gated on YFP fluorescence to be >30 times brighter than the YFP-negative nonexpressing cells (gate labeled 2 in Fig. 1B1 , which is redrawn in Fig. 1B3 , and represents ~41% of the total population for this experiment). FRET ratio did not vary markedly with expression level for these cells. When analyzed this way, PMA stimulation increased FRET ratios from 2.8 ± 0.59 to 3.14 ± 0.39, an increase of 12.1%. Similar results were obtained in two other experiments. This change in FRET ratio, while extremely small, is comparable to what we observed in the imaging experiments. Both measures are in reasonable agreement with what has been reported in the literature for this sensor, which gave ~20% FRET ratio increases in HEK-293 cells stimulated with EGF. 19
Long-Term Expression of FRET Sensors in Leukemic Cells
Aoki et al. have discussed the advantages of stable cell lines for FRET-based measurements, and assessed several strategies for generating them, including retroviral transduction and the use of transposons. 25 We took a different approach, generating long-term lines by nucleofecting cells followed by two rounds of sorting ~2 weeks and ~4 weeks after transfection. Sorted sensor-positive cells were not cloned. These polyclonal lines retained relatively high levels of fluorescence for periods of weeks to months. Note that because cells are not cloned, there might be more variability in their responses. However, a polyclonal population may better represent the diversity of the parental line, which may have advantages for some applications.
We created long-term K562 transfectants expressing six different intramolecular FP FRET sensors: EKAR, CKAR, DKAR, DAGR, JNKAR, and D3CPV. EKAR was described above. CKAR reports the activity of protein kinase C (PKC), and was constructed so that when phosphorylated by PKC, a decrease in FRET ratio occurs.
2
DKAR reports the activity of protein kinase D (PKD), a serine-threonine kinase that is activated physiologically by diacylglycerol binding and PKC activity. When phosphorylated, DKAR FRET ratio decreases.
20
DAGR is comprised of the diacylglycerol-binding C2 domain of PKCΒII. It translocates from cytosol to membrane when DAG or an analog is present, resulting in an increase in the sensor’s effective concentration that generates an increase in FRET ratio in the absence of a conformational change.2,26 JNKAR reports the activity of the stress-activated kinase JNK, and its FRET ratio increases upon phosphorylation.
22
Finally, D3CPV is a calcium sensor.
27
Its FRET ratio increases when intracellular calcium levels rise. Additional information about the different sensors can be found in the
We assessed the behavior of the sensor-expressing long-term transfectants using flow cytometry (
Fig. 2
). The left column of
Figure 2
shows histograms of YFP fluorescence from cells expressing the indicated constructs after gating cells on forward scatter versus side scatter to exclude dead cells and debris. The middle column shows plots of FRET versus CFP fluorescence for unstimulated (blue) and stimulated (red) cells for the live-cell scatter gate. Cells were stimulated with the chemical agents described above. Shown on the plots of FRET versus CFP fluorescence are gates that were used to select events for calculation of the FRET ratio, which is shown in histograms in the right column for stimulated (red) and unstimulated (blue) cells. The gates used include cells that are >30 times brighter than untransfected cells, and also allow us to exclude cells that are clearly outliers from the main population. Outliers are most prominent in the case of EKAR, in which a population of cells with much higher FRET at a given level of CFP expression is evident. A complete outline of the gating strategy used to select cells for calculation of the FRET ratio is shown in
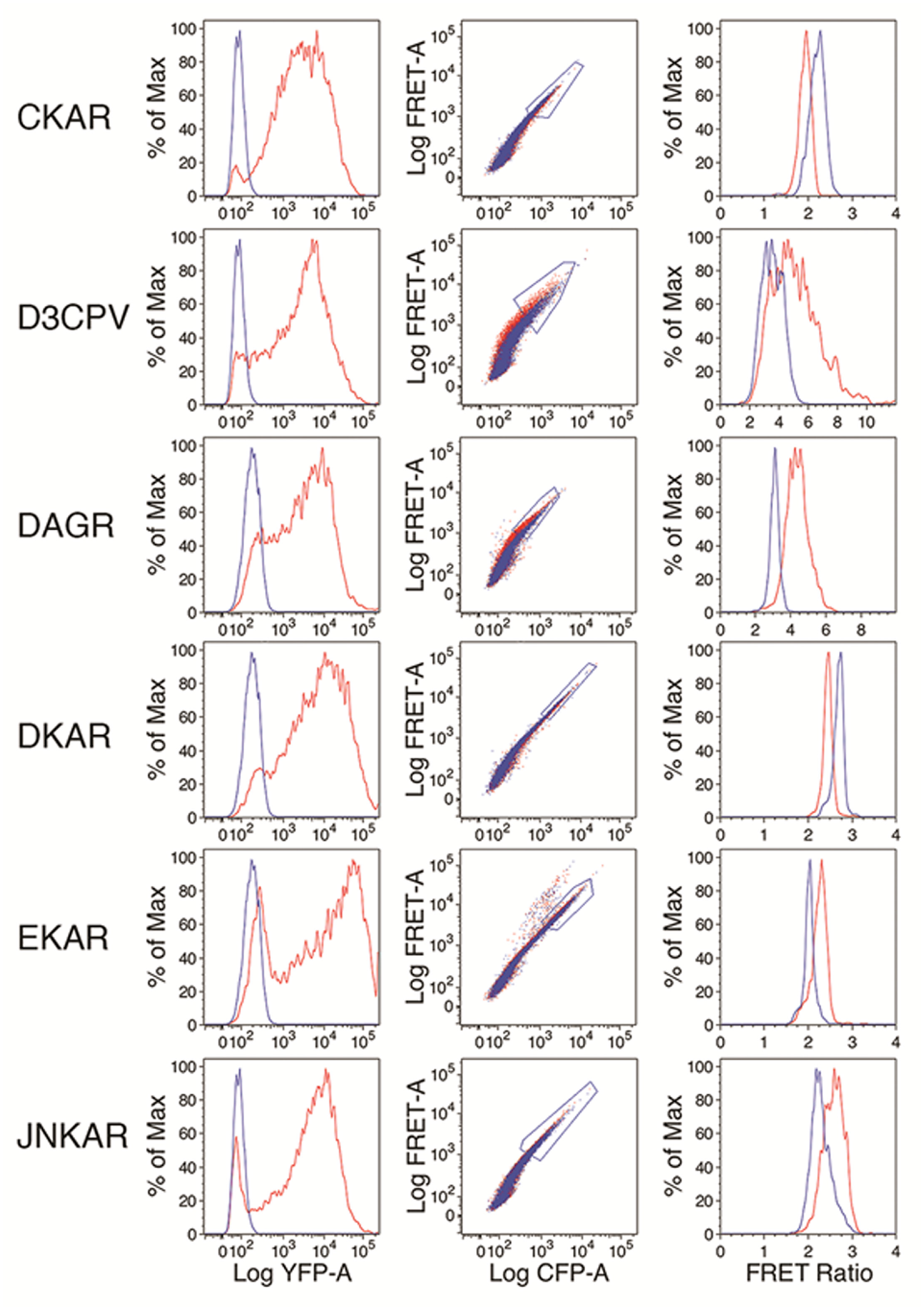
Long-term FRET sensor expression and function in K562 cells. Left column: Histograms of YFP fluorescence for long-term transfectants expressing the indicated FRET sensor are shown in red. The blue trace represents the YFP fluorescence of untransfected K562 cells. Cells were gated on forward and side scatter to eliminate dead/dying cells and debris. Middle column: Plots of FRET fluorescence vs. CFP fluorescence for unstimulated (blue) and stimulated (red) cells from the live-cell forward vs. side scatter gate. In most cases, the unstimulated and stimulated samples overlap sufficiently that only the unstimulated population is discernible. Gates used to select cells to analyze FRET signals further are shown. Right: Histograms of FRET ratio calculated for unstimulated (blue) and stimulated (red) cells from the gates shown in the plots of FRET vs. CFP fluorescence in the middle column. PMA was used to stimulate EKAR, CKAR, DKAR, and DAGR; anisomycin was used to stimulate JNKAR; and thapsigargin was used to stimulate D3CPV.
Bar-Coding Long-Term Transfectants for Multiplexed Measurements of FRET
To implement our multiplexing strategy, we need to be able to bar-code cells to allow us to identify the sensor each cell expresses. To exceed the level of multiplexing currently possible in microscopy, we wanted to be able to measure FRET signals from at least four sensors. Obviously, dyes used for bar coding should not overlap with CFP or FRET signals, as that could affect FRET measurements. Furthermore, bright YFP signals from the sensors render the use of green labels problematic. Because the instrument we used for these experiments lacked a UV laser, we decided to label cells with combinations of orange and red labels. We settled on calcein red-orange and Far Red DDAO-SE as bar-coding dyes. Neither of these dyes contribute signal in the CFP or FRET channels (see
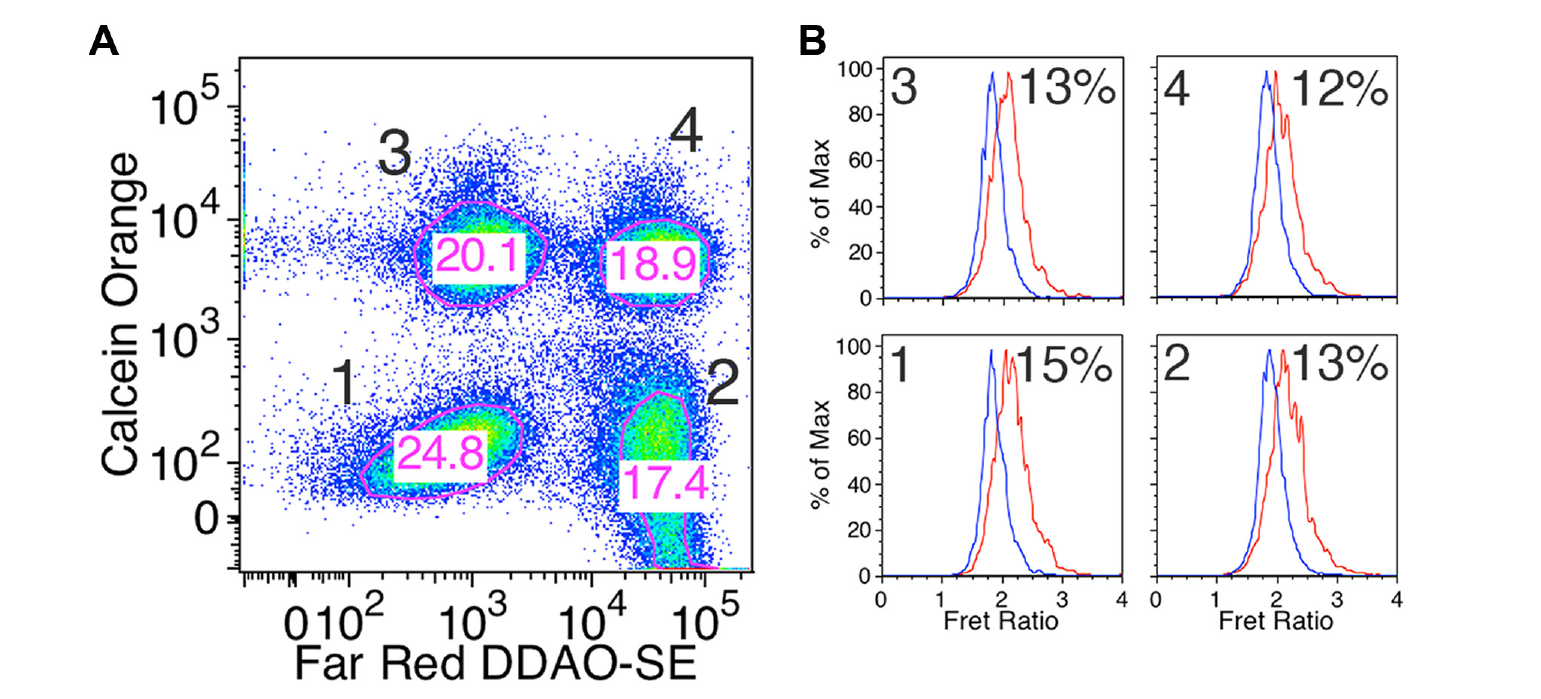
Multiplexed measurements of EKAR responses. (
Multiplexed Measurements with Kinase and Calcium Sensors
As an additional means of testing our multiplexing strategy, we examined the behavior of K562 long-term transfectants expressing D3CPV, EKAR, CKAR, and JNKAR, stimulating cells with agents intended to change the activity of different subsets of sensors ( Fig. 4 ). We first tested each by expressing each sensor individually; stimulating each with PMA, anisomycin, or thapsigargin; and measuring the FRET ratio as in Figure 2 . Histograms of the FRET ratio are shown in Figure 4A (thin lines denote the data from individual measurements). For each treatment, we also calculated the percent change in the geometric mean of the FRET ratio relative to unstimulated cells expressing the same sensor ( Fig. 4B , left). PMA activated CKAR and EKAR, as expected. It also activated, to a lesser extent, JNKAR. JNK activation by PMA has been reported previously in a lung cancer cell line. 29 Anisomycin activated JNKAR as expected, but also activated EKAR. ERK activation by anisomycin has been reported previously in PC-12 cells. 30 Anisomycin was without effect on CKAR or D3CPV. TG activated D3CPV, but was without significant effect on the other sensors.
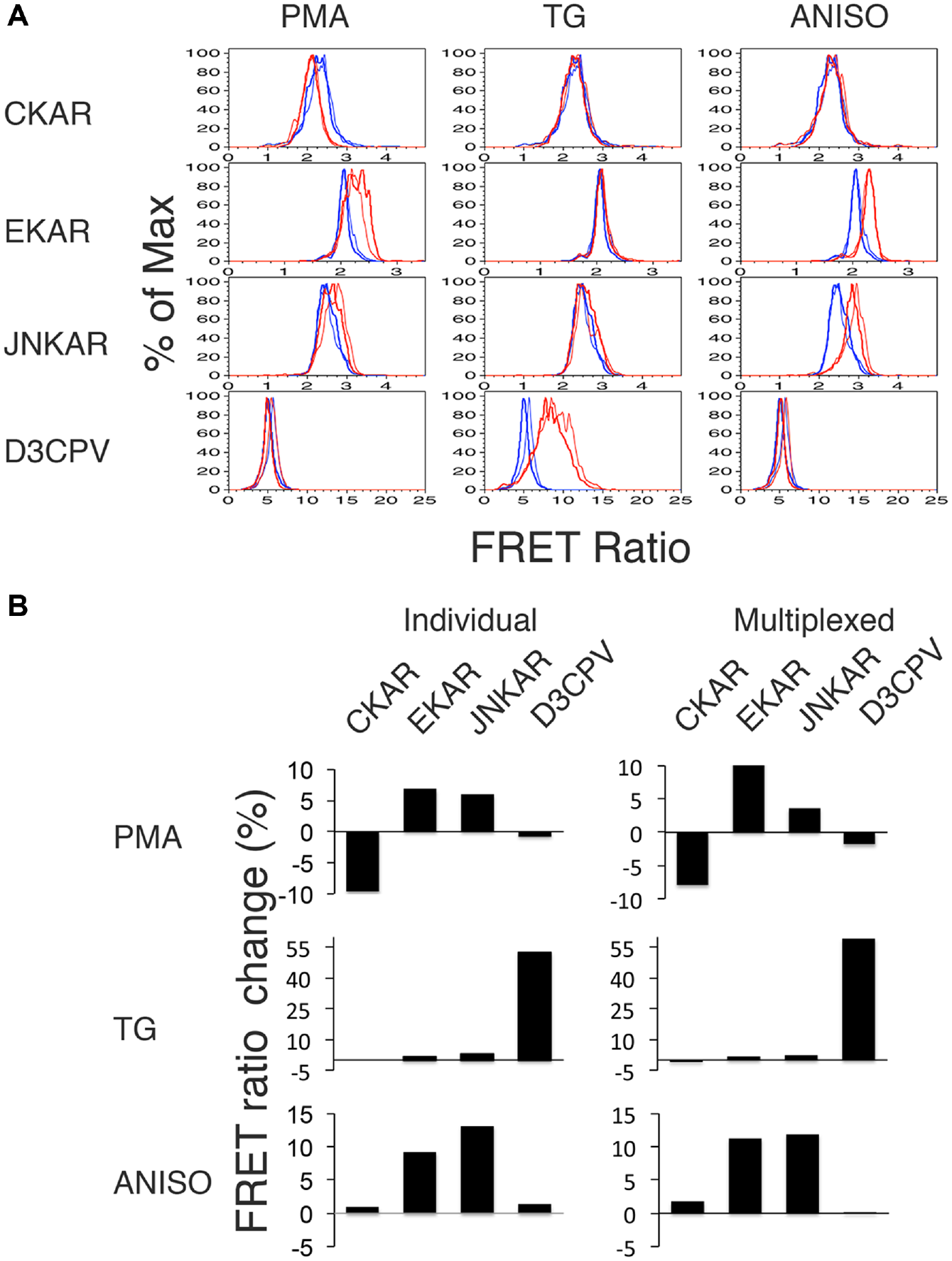
Comparison of signals from sensors for kinase activity and intracellular calcium individually and in multiplexed format. (
We next mixed bar-coded long-term transfectants together and stimulated them with the same agents, comparing responses to unstimulated bar-coded cells. The histograms shown in Figure 4A (thick lines denote the data obtained from the multiplexed sensors) and the percentage change in FRET ratio we calculated ( Fig. 4B , right) indicate that the pattern of FRET ratio changes we measured was generally similar to that observed when the sensor-expressing cells were examined separately. While there were some quantitative differences, as in the amplitude of the EKAR and JNKAR FRET ratio changes in response to PMA stimulation, these can likely be explained by the fact that different amounts of time elapsed between stimulating cells and measuring responses. We believe the important point is that the overall patterns of response are similar when cells are measured singly or in multiplex.
Multiplexed Measurements of PMA-Stimulated C1 Domain–Mediated Signaling
As a final means of testing our multiplexing strategy, we examined the behavior of four sensors that probe signaling events stimulated physiologically by diacylglycerol (DAG), a pathway critical for many biological responses triggered by G protein–coupled receptors and tyrosine kinase–linked receptors (see, e.g., Yang and Kazanietz
31
). We bar-coded and mixed cells expressing DAGR, CKAR, DKAR, and EKAR, and analyzed FRET signals at different times after stimulating the samples with 50 nM PMA (
Fig. 5A
). Treating cells with PMA caused time-dependent changes in FRET ratio for all sensors. The expected direction of FRET ratio change was captured (increases in FRET ratio for DAGR and EKAR cells, decreases in CKAR and DKAR cells). The magnitude of responses measured from the multiplexed samples at 30 min agreed well with responses measured from the long-term transfectants individually (compare to data in
Fig. 2
and
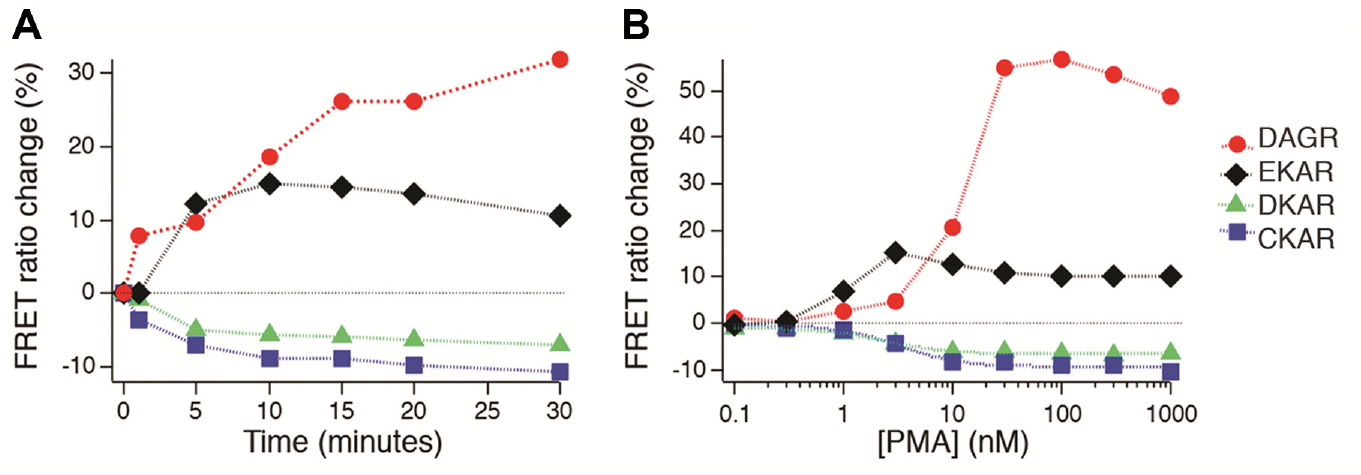
Multiplexed measurements of PMA-stimulated C1 domain signaling. (
We next performed dose–response measurements with PMA, measuring responses after 30 min ( Fig. 5B ). Clear dose-dependent responses were measured. Together with the time course data, this demonstrates that the method can resolve submaximal levels of activity. EKAR signals reached a maximum value at much lower concentrations of PMA than either CKAR or DKAR did, which may reflect the fact that activation of ERK by PMA is thought to depend on two C1 domain proteins, PKC and RAS-GRP, 32 whereas activation of PKD is PKC dependent, and thus would be expected to display a tight correlation to PKC activity.
Multiplexed Measurements in 96-Well Plate Format
As noted previously, FRET ratio changes from FP FRET sensors are quite small. However, Allen et al. showed that signals from a protein kinase A reporting FRET construct, which gives similarly small FRET ratio changes, could be measured with a plate reader to form the basis of a high-throughput screen. 33 Additionally, Kelly et al. have shown that fluorescence lifetime imaging microscopy can be performed in plates. 34 To determine whether multiplexed FRET measurements performed with flow cytometry are robust enough to serve as the basis for high-throughput screening (HTS), we developed a three-sensor 96-well plate assay using bar-coded DAGR-, CKAR-, and DKAR-expressing cells ( Fig. 6A,B ). For these experiments, we decided to omit EKAR, as the time and dose dependence of its responses to PMA (see Fig. 5 ) proved in preliminary experiments to pose problems, particularly when coupled with the relatively slow HTS sampler (BD HTS sampler attached to a BD Fortessa X20 cytometer) we had available, which required ~45 min to acquire a plate.
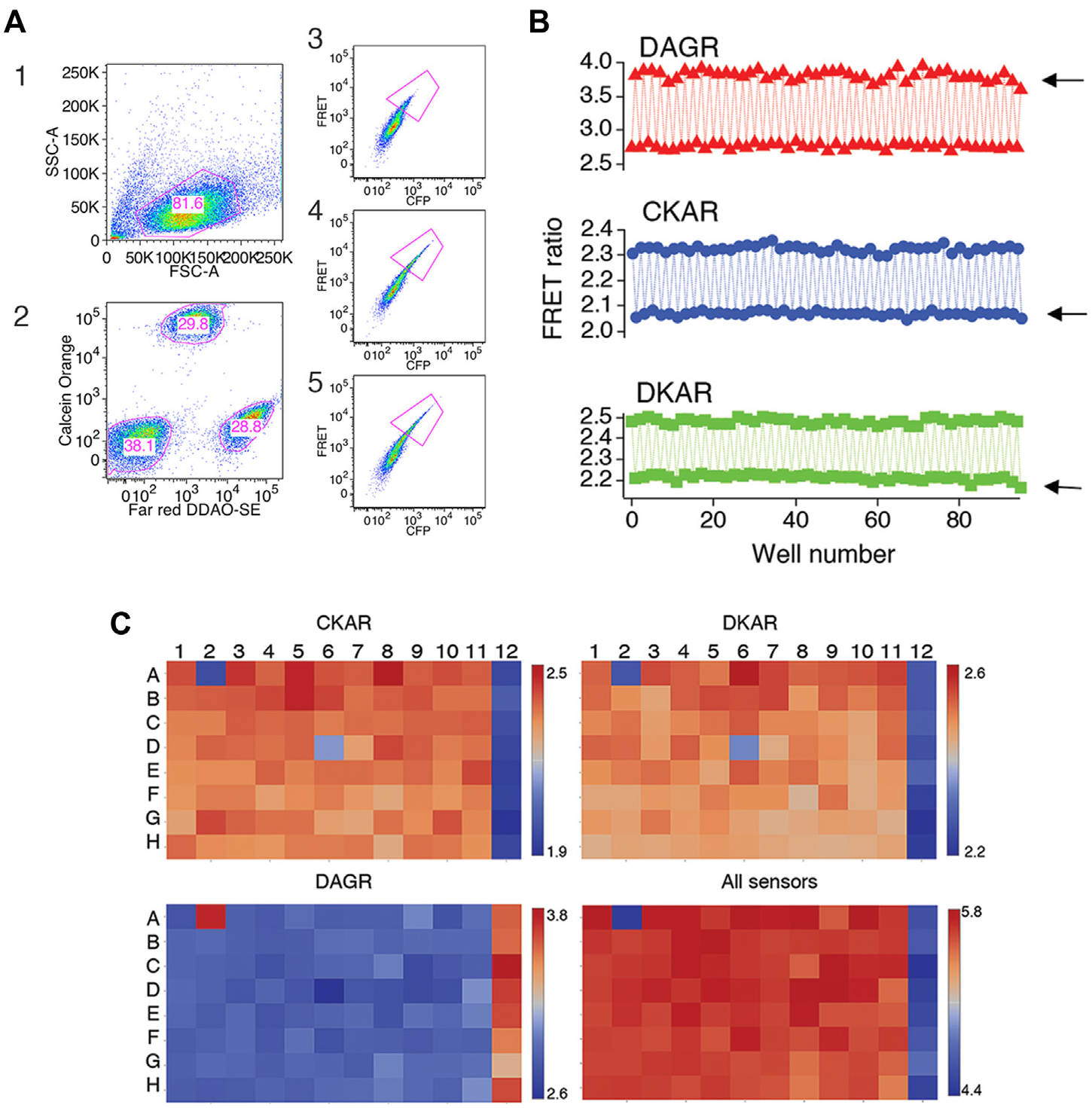
Measuring multiplexed FP FRET in plate format. (
We laid out plates with alternating columns of unstimulated cells and cells stimulated with 1 µM PMA. Fifty microliters were sampled from each well, and data were gated on forward and side scatter (the steps in gating samples are outlined in Fig. 6A ). In the experiment shown, we acquired a total of ~29,000 live cells from each well. Applying gates to plots of calcein red-orange versus Far Red DDAO-SE resulted in selection of ~7300 CKAR cells, 8800 DAGR cells, and 7400 DKAR cells per well. The sum of the selected cells is less than the total of number of live cells because gates were drawn fairly stringently to exclude cells in whose identity we were not confident. Applying gates to plots of FRET versus CFP resulted in the selection of ~1100 highly positive CKAR cells, ~600 highly positive DAGR cells, and ~1000 highly positive DKAR cells per well. FRET ratios were calculated for these cells. FRET ratio changes measured from these samples were stable over time, and their amplitude agreed well with our other measurements ( Fig. 6B ).
To assess whether signals we measured, which are quite small, are robust enough for HTS screening purposes, we calculated Z′, a measure of assay robustness in which values of >0.5 are taken as indicating a suitably robust assay, 35 for each of the sensors. Z′ values were calculated according to eq 3:
where m denotes the mean, σ denotes the standard deviation, and C+ and C– indicate PMA-stimulated and unstimulated wells, respectively. For the experiment shown, Z′ values calculated for all of the wells in the plate were 0.75, 0.68, and 0.7 for CKAR, DAGR, and DKAR, respectively. We found that Z′ values depended on both the number of cells acquired and the amplitude of the FRET ratio change. We estimated the minimum number of cells we would need to acquire to obtain Z′ values of >0.5 by calculating Z′ values for time subsets of the entire sample (data not shown). For DAGR, only ~100 highly positive cells would be sufficient to give Z′ values of >0.5. For CKAR and DKAR, which give smaller FRET ratio changes, we estimate that ~500 cells would be needed.
As an additional means of assessing the suitability of multiplexed flow FRET measurements for screening, we measured FRET signals from cells preincubated in a plate that contained 80 different extracts prepared from bacteria that were cultured from tunicates ( Fig. 6C ). We had speculated that these bacteria might produce compounds that modulate immune function in the host animal (the tunicate). Using an assay based on cytotoxic T-lymphocyte granule exocytosis, we identified well A2 as an enhancer of immune function and determined that it contains a PKC-activating natural product, which we subsequently identified as teleocidin A1 (Zhao et al., submitted manuscript; see Supplemental Methods), a known activator of PKC 23 that interacts with C1 domains. 28 When we screened this plate, cells in well A2 displayed decreased FRET ratios for CKAR and DKAR and an increased FRET ratio for DAGR, as expected for activation of C1 domain–dependent signaling. Compared to DMSO-treated control wells, PMA decreased the FRET ratio for CKAR from 2.7 ± 0.02 to 2.4 ± 0.02, increased the FRET ratio for DAGR from 3.5 ± 0.05 to 4.5 ± 0.1, and decreased the FRET ratio for DKAR from 3.1 ± 0.02 to 2.8 ± 0.03. The summed FRET ratio, calculated according to eq 2, decreased from 7.3 ± 0.1 to 5.7 ± 0.11. Well A2 had values essentially equal to those of PMA-stimulated cells: the CKAR FRET ratio was 2.4, the DAGR FRET ratio was 4.7, the DKAR FRET ratio was 2.8, and the summed FRET ratio was 5.5. All other wells except well D6 gave FRET signals essentially identical to unstimulated controls. Cells in well D6 showed decreased FRET ratios in CKAR and DKAR cells, but also decreased FRET ratios for DAGR cells, which was not expected for a ligand that binds to and activates C1 domains. This well was not active using the assay that originally detected the activity of well A2, which it should have been if it contained a PKC-activating compound. The extract was not notably fluorescent, which could have generated such results, and we are at present unable to account for its effects.
Discussion
Flow cytometry offers an extremely convenient and powerful way to measure intramolecular FP FRET signals. When we directly compared results with the transiently transfected EKAR in TALL-104 cells ( Fig. 1 ), microscopy and flow cytometry measured essentially identical changes in FRET ratio. Performing the measurement using flow cytometry was notably easier and faster than with imaging. Flow cytometry also makes it easy to examine the fluorescence profile of large numbers of cells, allowing gates to be drawn (e.g., Fig. 2 ) that can be used to exclude outliers that could potentially skew results if they were to be included. Our analysis of the dependence of FRET ratio on expression level ( Fig. 1B3 ) indicates that accurate FRET ratio measurements can only be made for high-expressing cells, as FRET ratios are poorly defined when cells express low levels of the sensor, most likely due to noise resulting from the subtraction of cellular autofluorescence. For this reason, we feel that determining the range of expression levels for which accurate estimates of FRET can be made is critical. However, it is not clear that the limitation to high-expressing cells is unique to flow cytometry. We are unaware of any studies that have examined the dependence of microscopy-based FRET measurements on expression level, but we would expect noise to be a problem in those measurements as well. In our imaging experiments, we selected relatively bright cells for analysis, but made no attempt to determine how FRET ratio varies with expression level. The requirement that cells express high levels of the reporter is not a problem using flow cytometry, since high-expressing cells can easily be selected for analysis via gating, and the method is sufficiently high throughput that obtaining sufficient numbers of bright cells is feasible. Given the ease of measuring intramolecular FP FRET ratio changes with cytometry compared to microscopy, we find it surprising that the method has not been more widely used.
Our primary interest in exploring the use of flow cytometry to measure FRET signals was to determine whether it would make multiplexed measurements possible, and whether those measurements would be suitable for screening. The ability to enable multiplexed measurements is one of cytometry’s key advantages over other analysis platforms (see discussion in Edwards and Sklar 36 ). The cytometer we used for most experiments was equipped with 405, 488, and 640 nm lasers, and we were readily able to acquire signals from four sensor-expressing long-term transfectants using orange and red dyes for bar coding. However, we found it difficult to accurately resolve more than four populations using these dyes, in part because of spread in apparent fluorescence introduced by compensation and in part because of a tendency for the red dye to be transferred between cells after mixing. We suspect that it will be possible to multiplex signals further using a cytometer equipped with a UV laser, which would enable the use of an additional dye, such as calcein blue, as a label and would in theory allow at least eight FP-expressing samples to be multiplexed. Note that we did not systematically examine whether different bar-coding dyes worked better for some sensors than others. However, we did notice that responses of DAGR decreased in a concentration-dependent manner when cells were labeled with calcein orange-red. This sensor does not undergo a conformational change to generate a change in FRET ratio, but rather, its local concentration increases when it translocates from cytosol to membrane, resulting in intermolecular FRET.2,26 We speculate that this may create a situation in which dye molecules can quench energy transfer. However, we did not examine whether similar effects are seen with Far Red DDAO-SE.
Despite the relative ease of measuring multiplexed signals with cytometry and bar coding, it is important to stress that signals are not being acquired from single cells, as they can be when imaging is used (see Welch et al. 37 for a discussion of the many advantages of measuring multiple signals from single cells). This means that the method will be suited to different uses than multiplexed single-cell FP FRET imaging—it can only provide information about population average behavior, not the relationship between parameters in a single cell. We envision that multiplexed cytometric analysis of intramolecular FP FRET signals could be used as a convenient means of investigating signal transduction cascades, ion fluxes, and metabolism, as FP FRET sensors have been created for these parameters. This may be useful to the screening community for studies to investigate the molecular mechanism of action (MMOA) of compounds identified in phenotypic HTS.
However, we believe that the main significance for screening of multiplexed cytometric FP FRET measurements is that it will allow the development of powerful new kinds of cell-based HTS. There are FRET sensors available that can enable screens for a variety of important targets in their native cellular environment—screens that cannot be conducted in any other way than with FP FRET sensors. An obvious example is provided by protein kinases, which constitute a very important class of drug target. 38 A variety of platforms, including time-resolved FRET assays based on energy transfer from Tb- or Eu-labeled phosphosite-specific antibodies to fluorescein- or GFP-labeled proteins or peptides, 39 are available to measure kinase activity in vitro. Previous work that used a fluorescence plate reader to detect signals from a PKA reporter construct 33 demonstrated the suitability of FP FRET sensors for screening compound libraries in cellulo, although the method does not seem to have been widely adopted. Multiplexing FP FRET sensors via bar coding, which cannot be accomplished using a conventional plate reader, will likely allow the development of even more powerful cell-based screens.
We envision three basic ways the method could be used. First, multiplexing could be used to increase assay robustness and enhance rejection of false positives, creating screens more likely to identify genuinely active compounds. An example of this is provided by the results shown in Figure 6 , where we used FRET sensors that change ratio in opposite directions to screen for C1 domain ligands. This enabled us to identify a well that contained a known C1 domain ligand, and also enabled us to reject as likely spurious a well that gave expected signals for only two of the three sensors. We note that while it seems intuitively obvious that multiplexing assays should result in more power, this issue has not received much attention (see Kümmel et al. 40 for a suggested implementation of a multiplexed Z′ factor for screening). We attempted to assess overall assay power by creating a summed signal from the individual sensors ( Fig. 6B ). Doing so gave a Z′ that was higher than for any of the individual sensors (0.8 vs. 0.7 for the experiment shown). However, as we also found that Z′ for the individual sensors increased as the number of cells analyzed increased, it is not clear whether Z′ for the summed signals was significantly higher than would be expected simply from collecting three times more events for any one of the individual sensors. As multiplex assays become more common, the issue of their power will need to be addressed systematically.
The second way we envision multiplexed FP FRET being used to increase assay power is by enabling assays that assess compound selectivity at the earliest stages of screening. Again using reporters for kinases as an example, it should be possible to mix cells expressing a panel of kinase FP FRET reporters, and then screen for activators and/or inhibitors that are selective for one or more of the kinases tested. Employing this strategy—which could be combined with qHTS 41 —would allow investigators to discard nonselective agents as early as possible. A similar strategy for assessing selectivity for two kinases simultaneously has been explored using TR FRET. 42
A final way that multiplexed FP FRET assays could increase assay power is by allowing multiple nodes in a complex process to be examined in a single assay step, making it possible to directly screen for functions that would otherwise require multiple follow-up experiments after screening. For example, using DAGR, CKAR, and DKAR, screens could be conducted to look for direct activators or inhibitors of PKD. PKD is activated by a combination of ligand binding to its C1 domain and phosphorylation by PKC, which in turn requires ligand binding to its C1 domain for activation. 43 Analysis of patterns of responses of DAGR, CKAR, and DKAR would make it possible to determine where in the network effects were occurring, eliminate compounds that act upstream, and identify those that act at the desired point.
Footnotes
Acknowledgements
We thank Dr. Carol Norris and the UConn Flow Cytometry Facility. Funding was provided by the University of Connecticut.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.