Abstract
Triclosan is an effective inhibitor for enoyl acyl carrier protein reductase (ENR) in fatty acid biosynthesis. Triclosan-resistant mutants of ENR have emerged. Thus, it is important to detect these triclosan-resistant mutations in ENR. Generally, enzyme activity assays on the mutants are used to determine the effect of triclosan on ENR activity. Since the substrates are linked to acyl carrier protein (ACP), the assays are challenging due to the need to prepare the ACP and link it to the substrates. Non-ACP-linked (coenzyme A [CoA]-linked) substrates can be used in some ENR, but not in all. Consequently, screening for triclosan-resistant mutants is also challenging. We have developed a simple thermal shift assay, which does not use ACP-linked substrates, to determine the binding ability of triclosan to the ENR active site, and thus it can be used for screening for triclosan-resistant mutants. Staphylococcus aureus FabI enzyme and its mutants were used to demonstrate the binding ability of triclosan with NADP+ to FabI. The direct correlation between the binding ability and enzyme activity was demonstrated with Francisella tularensis FabI. This method may also be applied to select effective triclosan analogues that inhibit ENR activity.
Keywords
Introduction
Triclosan, 5-chloro-2-(2,4-dichlorophenoxy)phenol or 2,4,4′-trichloro-2′-hydroxydiphenyl ether, has activity against many, but not all, types of Gram-positive and Gram-negative bacteria,1–3 including methicillin-resistant Gram-positive Staphylococcus aureus strains, 4 Mycobacterium tuberculosis,5,6 and Plasmodium falciparum. 7 It has also been reported that triclosan is cytotoxic to breast cancer cells. 8 The mode of action of triclosan on bacterial cells is well studied3,9–12 and it targets enoyl-acyl carrier protein (ACP) reductase (ENR), which is encoded by the gene FabI 11 and catalyzes the final step of the type II (bacterial) fatty acid biosynthesis pathway to reduce the double bond of enoyl-ACP to single-bond acyl-ACP with cofactor NAD(P)H. 13 Triclosan inhibits the enzyme action by forming a stable ternary complex with the oxidized cofactor NAD(P)+ and the side chains of FabI active site residues. 13 The triclosan effect on the human fatty acid synthase study suggests that inhibitors like triclosan may have chemotherapeutic potential. 8
Due to its usefulness toward inhibiting bacterial growth, triclosan has been widely used, 14 and species resistant to triclosan can arise from mutations in ENR/FabI. For example, S. aureus FabI (SaFabI) mutations such as A95V, I193S, and F204S have been identified in selection experiments, 15 suggesting that other mutations in S. aureus may also survive triclosan treatment and need to be identified. To determine whether triclosan inhibits certain ENR/FabI mutants, enzymatic activity in the presence of triclosan is commonly measured to determine triclosan inhibition activity. For ENR/FabI enzymes of many species such as Escherichia coli and Francisella tularensis, it is possible to substitute the substrate enoyl-ACP with coenzyme A (CoA) esters, such as crotonoyl-CoA, as experimental substrates,16,17 but with much reduced activities. 17 For the ENR/FabI of species where the substrate cannot be substituted with crotonoyl-CoA, such as that of S. aureus, substrate enoyl-ACP needs to be used to identify triclosan-resistant mutations or to select effective triclosan analogues as useful antibiotic leads. Obtaining ACP-linked substrates involves the purification of apo-ACP from cellular extract followed by either enzymatic-driven15,18 or chemically driven 19 linkage of the enoyl chain to the ACP. Consequently, studies that require enoyl-ACP for substrate are laborious. It is speculated that SaFabI diverges from the classical FabI structure and behavior owing to the wealth of branched-chain fatty acids occupying the membranes of S. aureus. 20 If so, the traits of SaFabI may be repeated in other organisms with similar membrane compositions, such as the genus Bacillus, in which all of the species are with branched fatty acid chains in their membranes.20,21
We have developed a thermal shift method to determine whether triclosan binds specifically to the active site of FabI without the use of enoyl-ACP. This thermal shift assay is a sensitive fluorescence-based assay, 22 making it amenable to high-throughput screening. We show that triclosan, together with NADP+, binds to SaFabI wild-type (WT) and triclosan-sensitive mutant M99T, but not to triclosan-resistant mutants A95V, I193S, and F204S. We used the F. tularensis FabI (FtFabI) system, WT and a triclosan-resistant mutant, to demonstrate the direct correlation between their thermal shifts and enzyme activities. We also show that the thermal shift is not due to the fluorescent probe used in the assay. We demonstrate that this simple and economical method, without using enoyl-ACP substrate, can be used to screen for triclosan-resistant mutants of ENR/FabI.
Method and Materials
Recombinant Plasmids
The SaFabI gene (NWMN_0881) was amplified from S. aureus gDNA and inserted in a pET-15b vector to express recombinant His-tagged SaFabI WT. An AFN-1252-resistant, but triclosan-sensitive M99T mutation 23 and triclosan-resistant mutations A95V, I193S, and F204S 15 were constructed with a site-directed primer mutagenesis method. 24 The FtFabI gene (FTT_0782) was amplified from F. tularensis gDNA and inserted in a pET-15b vector to express recombinant His-tagged FtFabI WT. The A92V mutation, a homologous mutation to the E. coli FabI (EcFabI) G93V triclosan-resistant mutation, 10 was also constructed using the same method. All resulting gene constructs were sequenced at the University of Illinois at Chicago Research Resource Center (UIC RRC).
Protein Expression and Purification
E. coli competent cells (BL21 CodonPlus(DE3)-RIL; Zymo, Irvine, CA) were used to express both WT and mutant proteins of both S. aureus and F. tularensis. An overnight culture (100 mL) was added to fresh lysogeny broth medium (1 L in a 2.8 L culture flask). Cells were grown to an optical density at 600 nm of 0.5–0.8 at 37 °C with shaking. Isopropyl β-D-1-thiogalacto-pyranoside was then added to a final concentration of 0.5–1.0 mM to induce protein expression. Cells were grown for an additional 3 h after induction and harvested at 4 °C.
The cells in 50 mM sodium phosphate buffer at pH 8 with 300 mM NaCl (phosphate buffer) plus 10 mM imidazole and 1% Triton X-100 were sonicated for 20 min on ice slurry. The mixture was centrifuged at 35,000 g for 30 min to give cell lysate. The lysate was loaded to a nickel affinity column (Qiagen) preequilibrated with the phosphate buffer plus 10 mM imidazole. The His-tagged protein was released from the column with the phosphate buffer plus 300 mM imidazole and dialyzed in 50 mM Tris buffer at pH 8 with 100 mM NaCl (Tris buffer) before concentrating to ~100–300 µM. The protein samples were frozen dropwise (20 µL) in liquid nitrogen and stored at −80 °C until needed. The protein purity was monitored with 16% polyacrylamide gel electrophoresis, and the molecular mass of the proteins was checked with high-resolution mass spectrometry at UIC RRC.
The concentration of each protein was determined from the absorbance at 280 nm and the extinction coefficient derived from its sequence, with 13,410 M−1 cm−1 for SaFabI WT, A95V, M99T, I193S, and F204S, and 17,420 M−1 cm−1 for FtFabI WT and A92V.
Enzymatic Assay for FtFabI
The published method for the enzymatic activity of EcFabI WT using crotonyl-CoA as substrate 12 was adapted for that of FtFabI. Briefly, FtFabI WT (450 nM) or A92V (450 nM and 1 µM) in Tris buffer was incubated with 250 µM NADH and 200 µM NAD+, without and with triclosan (2.5 molar ratio to FabI; triclosan from Sigma, St. Louis, MO, 72779-5G-F), for 5 h at 4 °C. 12 Since triclosan was in DMSO, both samples also consisted of 1% (v/v) DMSO. The relatively long incubation time (5 h) was consistent with the condition used for SaFabI in the thermal shift assay to ensure the complex formation among SaFabI, NADP+, and triclosan. It is known that triclosan exhibits its inhibition activity by forming a tight complex with FabI and NAD(P)+, and it is also known that NAD(P)+ itself exhibits relatively low affinity toward FabI and is in competition for the binding site.12,20 At the end of the incubation, the mixtures were warmed to 25 °C (in about 10 min), followed by the addition of an experimental substrate, crotonyl-CoA (200 µM final concentration; Sigma, C6146-5MG), to start the reaction. The decrease in NADH concentration, upon oxidation to NAD+ during the catalytic reaction, was monitored by UV absorption at 340 nm (A340) for ~2.5 min with a plate reader (Victor 3 V; PerkinElmer, Waltham, MA). The slope of a linear fit of A340 versus time plot (ΔA340/Δmin) was converted to a rate of NADH consumed, using the NADH extinction coefficient of 6220 M−1 cm−1. Specific activity for FtFabI was calculated as the rate of NADH consumed (µmol/min) per milligram of FtFabI.
Thermal Shift Assay for Binding Triclosan
Samples consisting of SaFabI (7 µM in Tris buffer) and 5× Sypro Orange (Invitrogen, Grand Island, NY) alone (control), or with only triclosan (100 µM) (sample-T), with only NADP+ (200 µM) (sample-N), or with both triclosan and NADP+ (sample-TN), were prepared side by side, with the same batch of protein, triclosan, and Sypro Orange to reduce sample variability. The control and three samples (sample-T, sample-N, and sample-TN) also included 1% (v/v) DMSO since both Sypro Orange and triclosan stock solutions were in DMSO. Sypro Orange was supplied in DMSO as 5000× stock solution. The mixtures were incubated for 5 h at 4 °C, as in the activity studies. Some samples were incubated for only 30 min. In the FtFabI samples, instead of NADP+, NAD+ was used.
The fluorescence intensity of Sypro Orange at 570 nm with excitation at 472 nm was monitored with a FP-6200 Jasco (Easton, MD) Spectrofluorometer as a function of temperature in the range of 25–75 °C. The temperature ramp rate was 1.0 °C/min. The intensity values were converted to fractions unfolded.
A Boltzmann sigmoidal fit, modeling a two-state unfolding process, was applied to the data to give transition temperatures measured by Sypro Orange (TmSO). A difference in the transition temperatures (thermal shift, ΔTmSO) between control and samples was obtained, and the ΔTmSO between control and sample-TN was used to indicate a degree of stabilization of FabI upon binding triclosan and NAD(P)+ to the active site of FabI.
Effects of Sypro Orange on Thermal Unfolding
The circular dichroism (CD) ellipticity (θ) signals at 222 nm (with a Jasco J-810 Spectropolarimeter) of FtFabI and SaFabI (15 µM) in the Tris buffer were measured from 25 to 75 °C with a ramp rate of 1.0 °C/min. θ values were converted to fractions of unfolding, with θ at 25 °C as 0 and θ at 75 °C as 1. Again, a Boltzmann fit was applied to the data to give transition temperatures measured by CD (TmCD). Also measured were FabI samples with 5× Sypro Orange and/or 0.1% (v/v) DMSO. We were not able to do CD measurements in the presence of 1% DMSO (as in the activity assay and thermal shift assay) due to the strong CD signal from 1% DMSO. For the same reason, the CD method was not used to monitor thermal shifts of samples with and without triclosan and NAD(P)+, since the samples included 1% DMSO.
Results
Enzyme Solution Properties
The polyacrylamide gel showed a band around 30 kDa for each of the seven overexpressed proteins, with 80–90% purity. High-resolution mass spectrometry results indicated that the initial methionine in the His-tag was missing in all our proteins, and the masses were within 1 Da of the expected values derived from sequences. For example, for SaFabI WT, the difference between the sequence-derived mass (30,397.4 Da) and experimental mass (30,396.7 Da) was −0.7 Da. For SaFabI M99T, A95V, I193S, and F204S, the sequence-derived masses were 30,367.3, 30,425.3, 30,371.3, and 30,337.3 Da, respectively, and the experimental masses were 30,366.6, 30,424.8, 30,370.6, and 30,336.6 Da, respectively.
The FtFabI WT enzyme exhibited a specific activity of 0.66 µmol min−1 mg−1 ( Fig. 1 ). In the presence of triclosan, the specific activity decreased to 0.06 µmol min−1 mg−1, with only 9% activity remaining, clearly demonstrating the inhibitory effect of triclosan on FtFabI. For the A92V mutant at the concentration used for WT measurements (450 nM), the decrease of A340 was small, making it difficult to obtain an accurate value for inhibition ( Fig. 1 ). However, with the enzyme concentration increased to 1 µM, the decrease in A340 as a function of time was more pronounced, and the normalized specific activity value for A92V was 0.09 µmol min−1 mg−1. In the presence of triclosan, the specific activity was measured as 0.10 µmol min−1 mg−1. Thus, A92V, with much reduced activity compared to that of the WT, was not inhibited by triclosan.
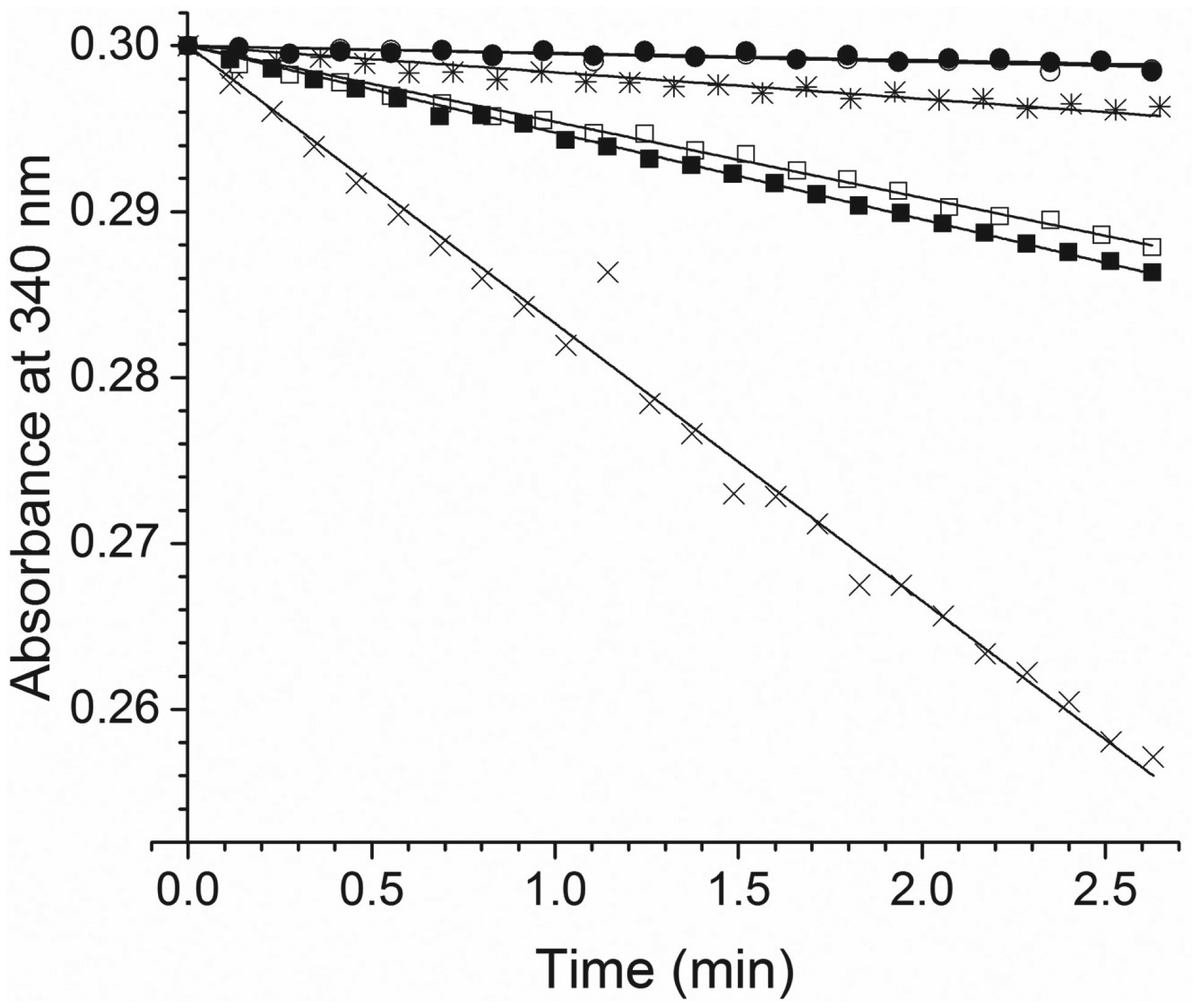
The decrease of 250 µM NADH absorbance upon the addition of 200 µM crotonyl-CoA by 450 nM FtFabI WT with no triclosan (cross) and with 1.125 µM triclosan (asterisk). Also shown are those of 450 nM FtFabI A92V with no triclosan (empty circle) and with 1.125 µM triclosan (filled circle), and an increased concentration of FtFabI A92V (1 µM) with no triclosan (empty square) and with 5 µM triclosan (filled square). The slope of the linear fit of the data points (ΔA340/Δmin) was converted to a rate of NADH consumed, using the NADH extinction coefficient of 6220 M−1 cm−1. Specific activities for FtFabI (WT and A92V) were calculated as the rate of NADH consumed (µmol/min) per milligram of FtFabI. For FtFabI WT, the activity was 0.66 µmol min−1 mg−1. See text for details.
The enzyme activity for SaFabI was not measured, as it requires enoyl-ACP as the substrate, as mentioned above.
Thermal Shifts for Triclosan Binding to Active Site
SaFabI
The TmSO average value from thermal unfolding profiles of SaFabI WT ( Fig. 2 ) was 40.5 ± 1.0 °C (n = 6). With the addition of just triclosan alone, the average value was 37.6 ± 0.9 °C (n = 2), and of just NADP+, the average value was 40.2 ± 1.2 °C (n = 2). Neither triclosan nor NADP+ increased the TmSO values. For SaFabI WT with both triclosan and NADP+ ( Fig. 2 ), the average value was 58.8 ± 1.6 °C (n = 2). A thermal shift of about 20 °C was observed for SaFabI WT upon addition of both triclosan and NADP+, but no shift upon addition of just triclosan or NADP+. Since the addition of triclosan or NADP+ alone did not stabilize SaFabI, and only when both were added was SaFabI stabilized, the results, together with the knowledge that triclosan and NADP+ bind as a complex to the active site of SaFabI 20 , suggest that the observed thermal shift reflects not only binding to SaFabI, but also specific binding to the active site of SaFabI.
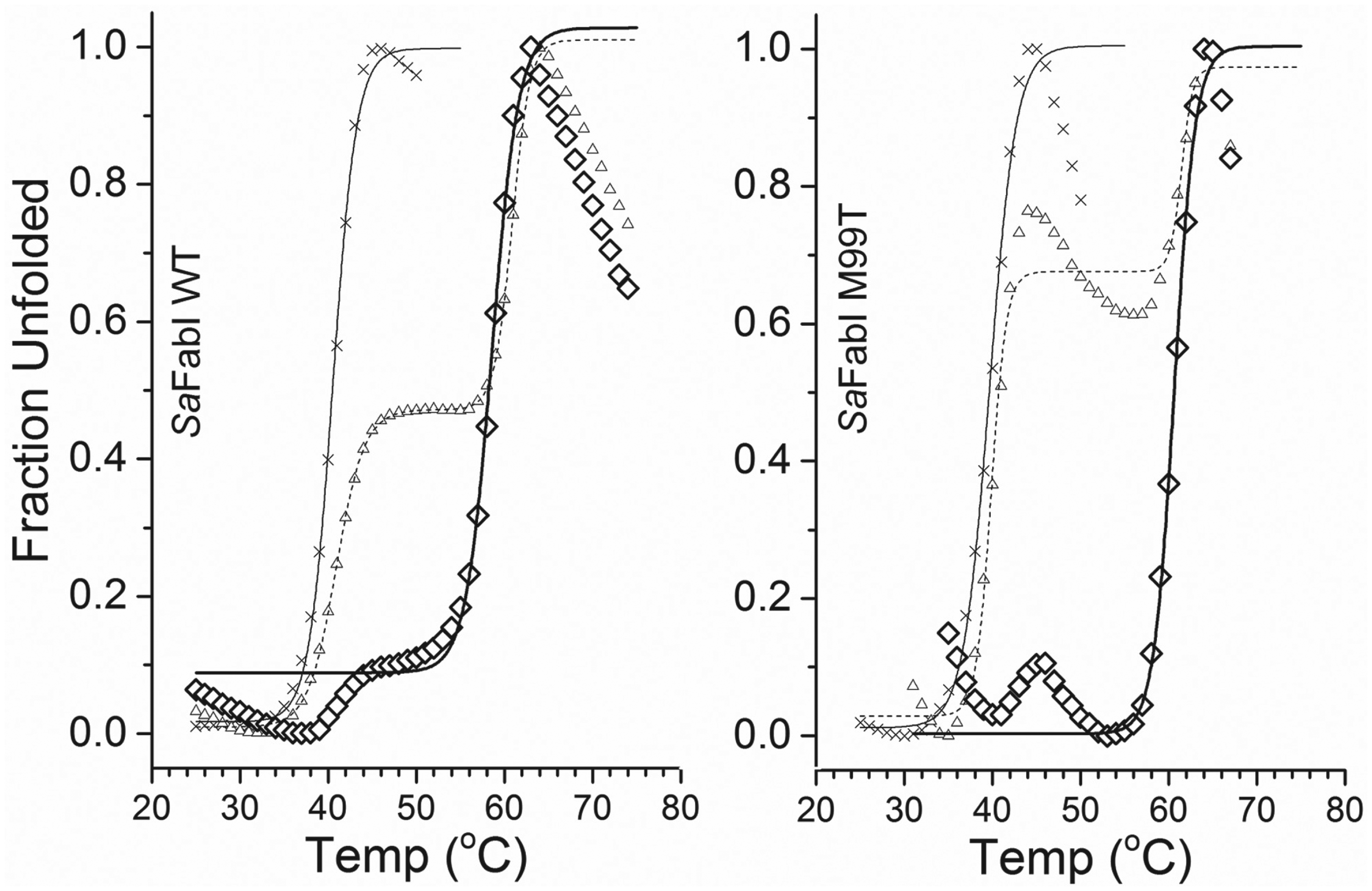
Representative thermal unfolding profiles of SaFabI WT and M99T samples, derived by monitoring the fluorescence emission intensity of Sypro Orange at 570 nm, with excitation at 472 nm. SaFabI WT (cross, left panel), with a TmSO of 40.5 °C, and SaFabI WT incubated with 100 µM triclosan and 200 µM NADP+ for 5 h (diamond), with a TmSO of 58.6 °C. We found that incubating triclosan and NADP+ for 30 min (triangle) was not sufficient for SaFabI, and the sample consisted of two populations of SaFabI, one without and one with triclosan/NADP+. The first transition temperature, 40.9 °C, was that of SaFabI without triclosan and NADP+, and the second transition temperature, 60.9 °C, was that of SaFabI with triclosan and NADP+. With 5 h incubation, the transition temperature shifted to 58.6 °C in this typical paired run. SaFabI M99T (cross, right panel), with a TmSO of 39.7 °C, and SaFabI M99T with 100 µM triclosan and 200 µM NADP+ after 5 h incubation (diamond), with a TmSO of 60.6 °C. The data from the sample with 30 min incubation (triangle) showed two transitions at 39.8 and 61.4 °C.
The TmSO average value of SaFabI M99T ( Fig. 2 ) was 40.2 ± 0.8 °C (n = 3). With the addition of just triclosan alone, the average value was 40.8 ± 0.1 °C (n = 2), and of just NADP+, the average value was 41.2 ± 0.1 °C (n = 2). For SaFabI M99T with triclosan and NAD+, the average value was 60.2 ± 1.1 °C (n = 2). An approximate 20 °C thermal shift was observed for SaFabI M99T upon binding triclosan and NADP+. It is interesting to note that 30 min incubation allowed some of the triclosan and NADP+ to bind to WT and M99T molecules, but not all, resulting in a thermal shift profile of two transitions ( Fig. 2 ). The SaFabI WT sample consisted of two populations, one without and one with triclosan and NADP+; the first transition was at 40.9 °C, representing the thermal transition of the WT without triclosan and NADP+, and the second transition was at 60.9 °C, representing the thermal transition of WT stabilized by triclosan and NADP+. Similarly, for M99T, the triclosan-sensitive mutant, we also found a transition at 39.8 °C and another transition at 61.4 °C ( Fig. 2 ).
The TmSO average value for A95V ( Fig. 3 ) without triclosan and NADP+ was 39.4 ± 0.2 °C (n = 3), quite similar to that of the WT ( Fig. 2 ), and with triclosan and NADP+, it was 40.3 ± 0.1 °C (n = 2). No thermal shift was observed for SaFabI A95V. Similarly, the average value for I193S ( Fig. 3 ) without triclosan and NADP+ was 40.3 ± 0.0 °C (n = 2), and with triclosan and NADP+, it was 40.2 ± 0.1 °C (n = 2). No thermal shift was observed for SaFabI I193S. The average value for F204S ( Fig. 3 ) without triclosan and NADP+ was 41.1 ± 0.1 °C (n = 2), and with triclosan and NADP+, it was 41.2 ± 0.1 °C (n = 2). No thermal shift was observed. Our results showed no binding of triclosan and NADP+ to SaFabI mutants A95V, I193S, and F204S.
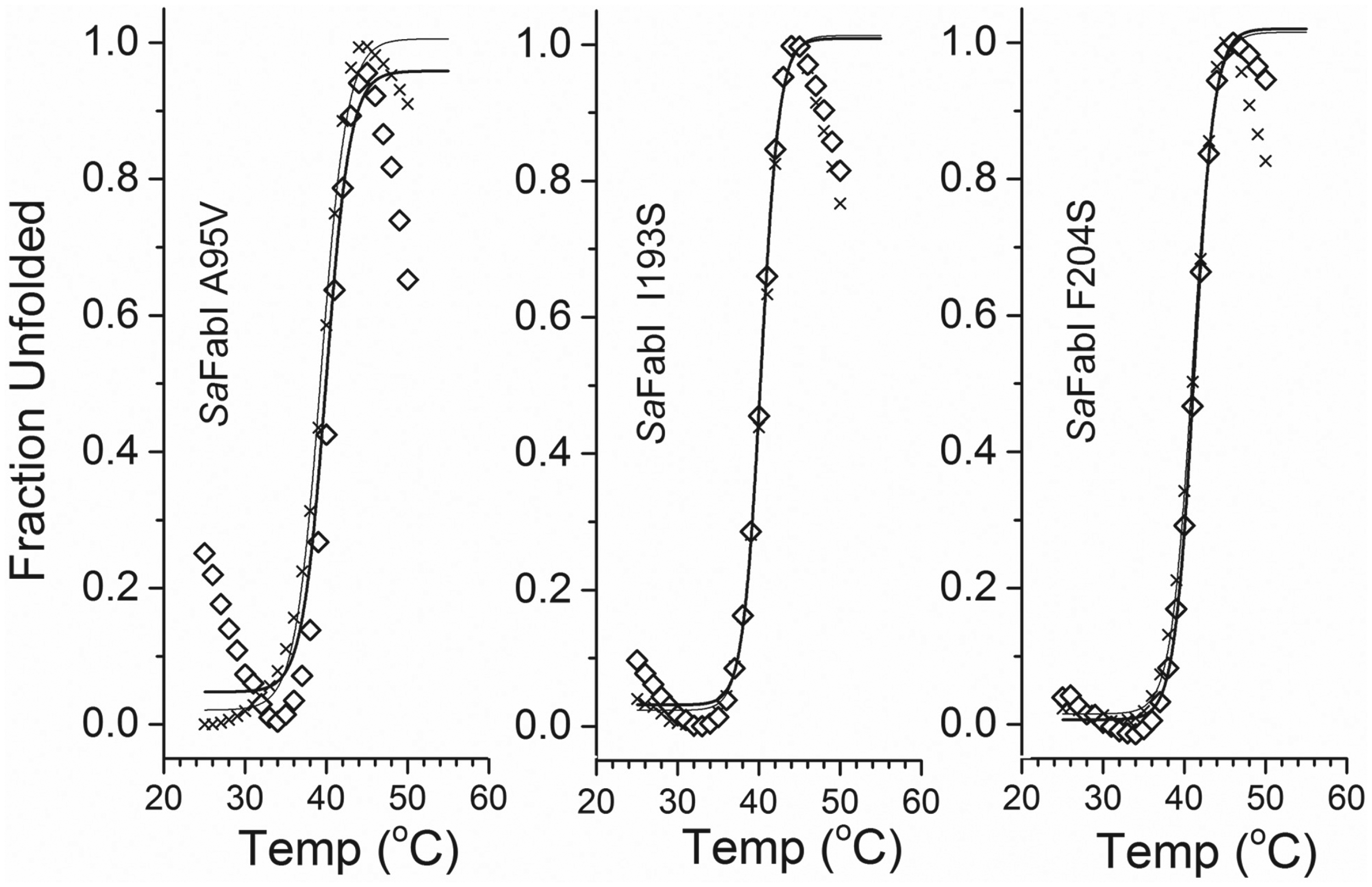
Representative thermal unfolding profiles of SaFabI A95V, I193S, and F204S samples, derived by monitoring the fluorescence emission intensity of Sypro Orange at 570 nm, with excitation at 472 nm. SaFabI A95V (cross, left panel), with a TmSO of 39.3 °C, and SaFabI A95V incubated for 5 h with 100 µM triclosan and 200 µM NADP+ (open diamond), with a TmSO of 40.1 °C. SaFabI I193S (cross, middle panel), with a TmSO of 40.3 °C, and SaFabI I193S incubated for 5 h with 100 µM triclosan and 200 µM NADP+ (open diamond), with a TmSO of 40.3 °C. SaFabI F204S (cross, right panel), with a TmSO of 41.2 °C, and SaFabI F204S incubated for 5 h with 100 µM triclosan and 200 µM NADP+ (open diamond), with a TmSO of 41.0 °C. No thermal shift was observed for SaFabI A95V, I193S, or F204S mutants.
FtFabI
The TmSO average value for FtFabI WT ( Fig. 4 ) was 53.2 ± 0.4 °C (n = 4). With the addition of just triclosan alone, the average TmSO was 52.3 ± 0.0 °C (n = 2), and of just NAD+, the average TmSO was 52.4 ± 0.9 °C (n = 2). For FtFabI WT with triclosan and NAD+, the average TmSO was 60.2 ± 3.4 °C (n = 2). About a 7 °C thermal shift was observed for FtFabI. A similar thermal shift was observed (60.8 ± 0.5 °C; n = 3) for samples incubated for 30 min. The average value for A92V without triclosan and NAD+ was 51.7 ± 0.5 °C (n = 3), and with triclosan and NAD+, it was 52.1 ± 0.1 °C (n = 2). No thermal shift was observed for FtFabI A92V. Together with the enzyme activity values ( Fig. 1 ), we showed that, using the FtFabI system, the thermal shifts and the activity decreases upon addition of triclosan and NAD+ are well correlated. Triclosan and NAD+ together stabilized FtFabI WT to increase the thermal transition temperature by 7 °C, and inhibited the enzymatic activity by 91%. For FtFabI A92V, no thermal shift was observed, and there was no change in the residual enzymatic activity in the presence of triclosan and NAD+. It was interesting to note that, unlike in the SaFabI system, 30 min incubation of triclosan and NAD+ with FtFabI was sufficient to fully stabilize FtFabI WT ( Fig. 4 ).
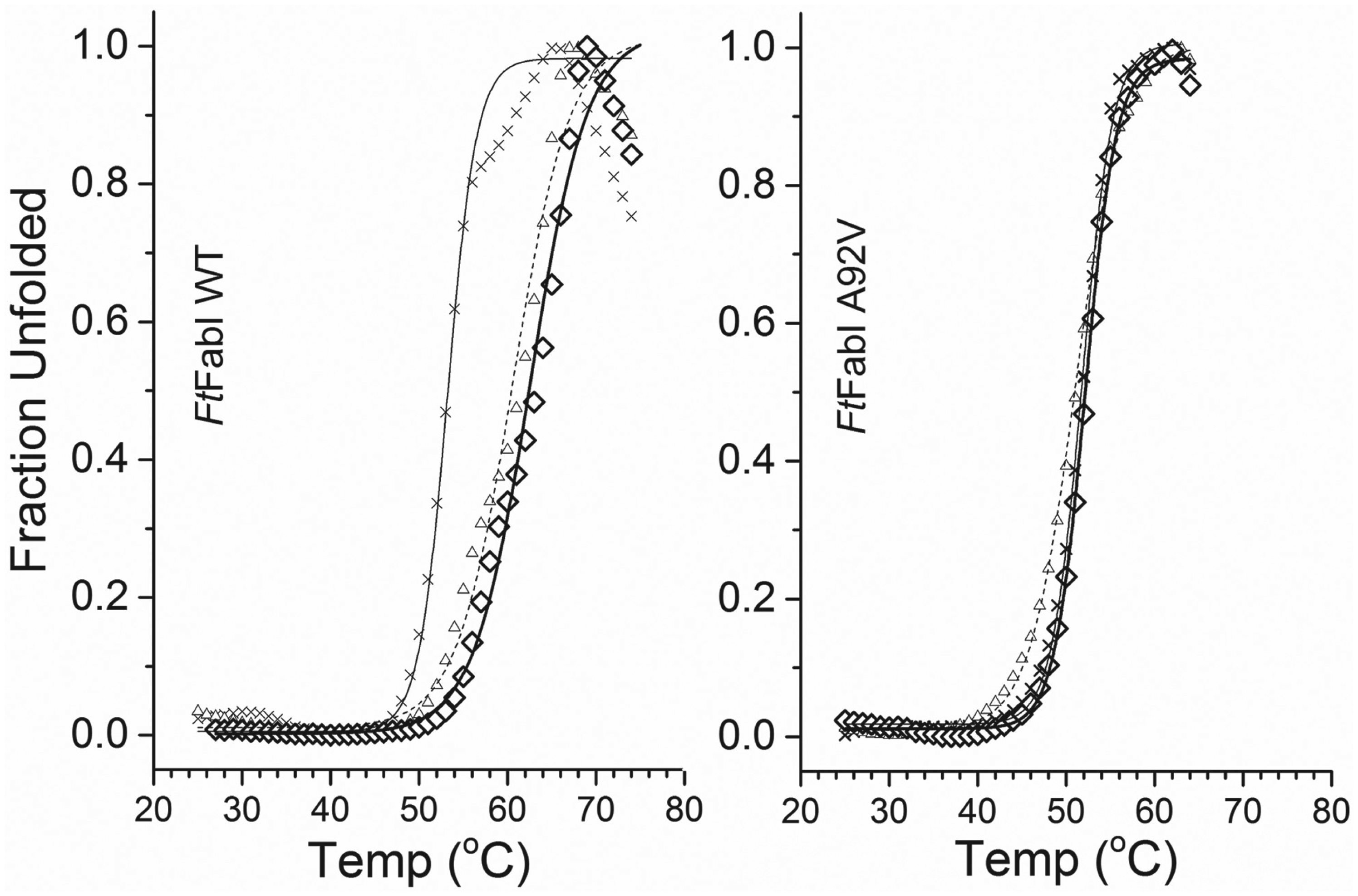
Representative thermal unfolding profiles of FtFabI WT and A92V samples, derived by monitoring the fluorescence emission intensity of Sypro Orange at 570 nm, with excitation at 472 nm. The TmSO of FtFabI WT (cross, left panel) was 53.2 °C, and of FtFabI WT with 100 µM triclosan and 200 µM NAD+ after 30 min incubation (triangle), it was 60.3 °C, and after 5 h (diamond), 62.6 °C. For A92V FtFabI (cross, right panel), the TmSO was 51.8 °C, and for A92V FtFabI with 100 µM triclosan and 200 µM NAD+ after 30 min (triangle), it was 51.0 °C, and after 5 h (diamond), 52.1 °C.
Sypro Orange Effect on FabI Stability
Since we used Sypro Orange to report the thermal unfolding, we investigated whether Sypro Orange in DMSO affected the thermal unfolding by using CD to monitor the unfolding of FabI with and without 5× Sypro Orange in DMSO (0.1%, v/v). Due to a large CD signal for DMSO, we used only 0.1% DMSO, rather than 1%, as used in the TmSO studies. Enzyme activities with and without 1% DMSO were about the same. FtFabI WT without DMSO, with 0.1% DMSO, or with 0.1% DMSO and 5× Sypro Orange revealed similar thermal unfolding profiles of secondary structures ( Fig. 5 ). The TmCD average values for samples without (n = 3) and with (n = 2) DMSO (0.1%) were both 57.7 ± 0.1 °C, and for samples with 0.1% DMSO and 5× Sypro Orange, the value was 56.4 ± 0.2 (n = 2). Similar profiles, except shifted to lower temperatures, were observed for SaFabI WT, with a TmCD of 43.7 ± 0.0 °C (n = 2). For SaFabI WT in 0.1% DMSO, the TmCD was 43.6 ± 0.0 (n = 2), and in 0.1% DMSO and 5× Sypro Orange, the TmCD was 42.7 ± 0.0 (n = 2). Little DMSO or 5× Sypro Orange effect was observed.
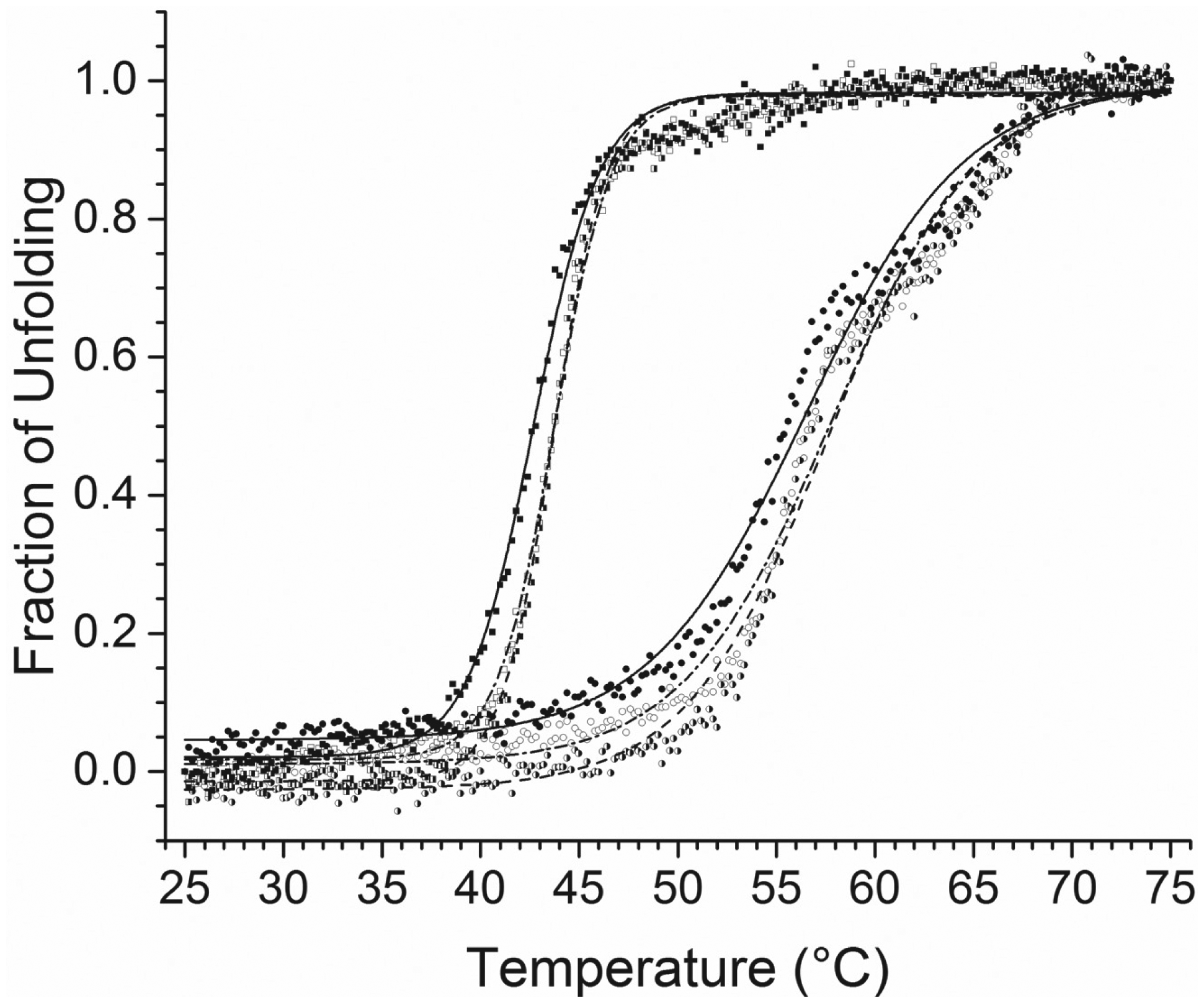
Representative thermal unfolding profiles and Tm values derived by monitoring the CD signal at 222 nm from 25 to 75 °C of 15 µM FtFabI WT with no DMSO (empty circle), Tm = 57.7 ± 0.1 °C; with 0.1% DMSO (half-filled circle), Tm = 57.7 ± 0.1 °C; and with 0.1% DMSO and Sypro Orange (filled circle), Tm = 56.4 ± 0.2 °C; and 15 µM SaFabI WT with no DMSO (empty square), Tm = 43.7 ± 0.0 °C; with 0.1% DMSO (half-filled square), Tm = 43.6 ± 0.0 °C; and with 0.1% DMSO and Sypro Orange (filled square), Tm = 42.7 ± 0.0 °C. These results show little DMSO (0.1%) or Sypro Orange effect on the thermal unfolding of either FtFabI or SaFabI.
The TmCD and TmSO values of the same sample differed slightly since the two methods measured slightly different unfolding events, with the CD reporting the unfolding of the secondary structures and the SO reporting the exposed hydrophobic moieties during unfolding.
Discussion
When Sypro Orange is added to intact protein in solution, which has little hydrophobic moieties, it exhibits little fluorescence intensity. However, when protein is thermally unfolded to expose hydrophobic moieties, Sypro Orange molecules around these moieties exhibit fluorescence intensity. 25 By following Sypro Orange fluorescence intensity in a protein sample as a function of temperature, protein unfolding profiles with characteristic transition temperatures are obtained. Since compounds binding to a protein molecule lower the free energy of the system, 26 Sypro Orange has been widely used to track the stabilization of a protein molecule due to compound binding.22,27,28
We showed that we may use this principle to determine whether triclosan and NAD/P+ bind to FabI mutants. The characteristic transition temperature in thermal unfolding of SaFabI WT in Tris buffer was about 40 °C. The temperature did not change upon addition of just triclosan or just NADP+. However, SaFabI WT and M99T were stabilized by the triclosan and NADP+ with the transition temperature shifted about 20 °C. For known triclosan-resistant mutants, A95V, I193S, and F204S, no thermal shift was observed upon addition of triclosan and NADP+. It should also be noted that all SaFabI samples, WT and the mutants, exhibited similar transition temperatures (~40 °C) in the absence of triclosan and NADP+. Since the thermal shifts were observed only when both triclosan and NADP+ were present, but not when only one was present, we suggest that the binding we observed was the active site binding observed previously. 20
To demonstrate that the active site bindings indicated by thermal shifts are related to enzyme activities, we used the FtFabI system. The transition temperature for FtFabI in Tris buffer was about 53 °C, but shifted to 60 °C upon addition of triclosan and NAD+. We also showed that the specific activity for FtFabI was inhibited by triclosan and NAD+. We further showed that a FtFabI triclosan-resistant mutant, A92V, with much reduced specific activity compared with that of the WT, was not further reduced upon addition of triclosan and NAD+. This mutant exhibited little thermal shift upon addition of triclosan/NAD+ in its thermal unfolding profile.
It was interesting to note that a 30 min incubation period was sufficient to fully stabilize FtFabI WT with triclosan and NAD+. However, the unfolding profiles of SaFabI WT and M99T samples with 30 min incubation clearly showed incomplete stabilization compared with those of 5 h incubation. These results suggest different binding kinetics, implying that this method may also provide kinetic information on active site binding of triclosan and NAD(P)+.
In this study, we used Sypro Orange thermal shift on purified protein. With the recent development of cellular thermal shift assay,29,30 it is certainly possible, using the principles discussed in this work, to apply cellular thermal shift assay on cells to screen for triclosan-resistant ENR/FabI mutants of species requiring enoyl-ACP-linked substrate. This method may also be used to select effective triclosan analogues that, together with NAD(P)+, bind to the active site of ENR/FabI to inhibit the enzymatic activity, and thus inhibit cell growth.
In summary, we have shown that the Sypro Orange–based thermal shift binding assay, with fluorescence sensitivity, can be used easily and reliably to show the binding of the antibiotic triclosan in the presence of NAD(P)+ to the active site of ENR/FabI enzymes, but not to a triclosan-resistant mutant. Thus, the method can be applied to FabI systems that require ACP-linked substrates without the need for the substrate to assay for activities. We have shown the lack of a thermal shift in a triclosan-resistant mutant. This approach offers a quick and convenient method to screen for triclosan-resistant mutations in ENR/FabI. This approach may also be applied to other antibiotics. In addition, it may also provide mechanistic insights into inhibition by studying the kinetics of the binding of triclosan to ENR/FabI.
Footnotes
Acknowledgements
We would like to thank Shahila Mehboob and Michael Johnson at the UIC Center of Pharmaceutical Biotechnology for the FtFabI and SaFabI WT plasmids and Marta Witek for the FtFabI A92V plasmid.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Proteomics and informatics services were provided by the Chicago Biomedical Consortium (CBC)—UIC Research Resources Center Mass Spectrometry, Metabolomics and Proteomics Facility, which was established in part by a grant from the Searle Funds at the Chicago Community Trust to the CBC. R.D.D. is an awardee of the Bridge to the Doctorate Fellowship at UIC. The work was supported by a grant from the National Institutes of Health (U01 AI-077949).
