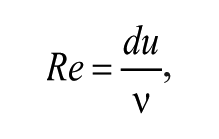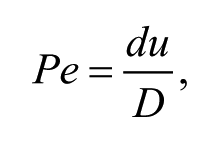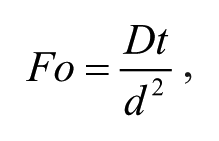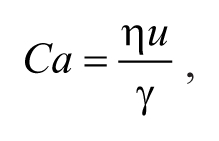Abstract
Over the past two decades, the application of microengineered systems in the chemical and biological sciences has transformed the way in which high-throughput experimentation is performed. The ability to fabricate complex microfluidic architectures has allowed scientists to create new experimental formats for processing ultra-small analytical volumes in short periods and with high efficiency. The development of such microfluidic systems has been driven by a range of fundamental features that accompany miniaturization. These include the ability to handle small sample volumes, ultra-low fabrication costs, reduced analysis times, enhanced operational flexibility, facile automation, and the ability to integrate functional components within complex analytical schemes. Herein we discuss the impact of microfluidics in the area of high-throughput screening and drug discovery and highlight some of the most pertinent studies in the recent literature.
Keywords
What Is Microfluidics?
Most processes in the chemical and biological sciences are performed on samples dissolved or dispersed in liquid media. Significantly, recent years have seen enormous improvements in the sensitivity and applicability of analytical detection techniques, which in turn have enabled the routine analysis of picoliter to nanoliter volumes.1,2 The benefits in terms of improved cost-effectiveness are obvious; however, for manual or even robotic processing, a decrease in sample size does not automatically result in higher analytical throughput, since simple scaling laws predict, for example, limited gains in pipetting times as sample volumes are reduced. Accordingly, completely new methods for handling ultra-small sample volumes are required.
Microfluidic technologies provide ideal environments for precise, controlled, and rapid processing of liquids on the sub-microliter scale when confined in channels with cross-sectional dimensions below a few hundred microns. 3 In principle, such microfluidic tools enable experimentalists to exploit many of the advantages that the microscale has to offer. 4 The benefits of microfluidics are not simply confined to throughput but also extend to improvements in analytical efficiency, instrumental footprint, process control, process automation (through integration of functional components), and reagent usage. 5 An excellent example in this respect is the development of handheld, low-cost, disposable microfluidic devices for use in point-of-care diagnostics.6–9 It is expected that such technologies will have a significant impact on health care in developing countries, 10 as well as in the monitoring of chronic conditions in the developed world. 11
The fundamental behavior of fluids at the micron scale is both complex and distinct from our everyday experience. This difference is often best described through the use of dimensionless numbers. 12 The most discussed, in relation to microfluidics, is the Reynolds number 13 (Re), which quantifies the relative importance of inertial and viscous forces. In simple terms, the Reynolds number is given by
where u and d are the characteristic velocity and length of the system, respectively, and ν is the kinematic viscosity of the fluid. Generally speaking, Reynolds numbers below 2000 indicate laminar flow, where streamlines are parallel and time reversible. Conversely, Reynolds numbers above 2000 characterize turbulent flow environments, where streamlines are chaotic and time irreversible. For microfluidic systems, Re is almost always smaller than 1, rendering the flow regime exclusively laminar. 14 The deterministic nature of such microflows engenders numerous applications that are impossible on the large scale, such as the controllable generation of monodisperse microdroplets. 15 In addition, within laminar flow environments, fluidic mixing occurs solely via diffusion. This provides for exquisite control over the distribution and concentration of molecules in the system but does affect mixing timescales. 16
The mixing behavior of a fluidic system can be described by the Péclet (Pe) and Fourier (Fo) numbers. Pe measures the relative rate of convective transport to diffusive transport and is defined for mass transport as
where D is the molecular diffusion constant. In the simplest case, the magnitude of Pe is directly proportional to the length of a channel or conduit that two fluids must flow through to mix completely. This allows controlled delivery of reagents through variation of Pe. 17 Similarly, the Fourier number is given by
where t is time. This provides a direct assessment of diffusive mixing efficiencies in a microfluidic channel. As a rule of thumb, a Fourier number of 1 indicates “complete” mixing, a value of 0.5 describes “adequate” mixing, and a value of 0.1 indicates “poor” mixing. 18
A final dimensionless number of utility in the current discussion is the capillary number (Ca). In flows that consist of separate, immiscible fluid streams, the capillary number is given by
where
The preceding discussion of fluid flow on the micron scale is purposely brief and solely intended to highlight some key features of fluid flow on the small scale. The interested reader is directed to more complete considerations provided elsewhere. 12
Microfluidic Platforms
A variety of different microfluidic tools, techniques, and platforms have been developed to address different biological problems. Although many microfluidic devices are specifically designed for a single task, most can be assigned to one of three main categories that we term continuous-flow, batch (or semi-batch), and droplet-based microfluidics.
Continuous-flow microfluidic systems are characterized by the continuous manipulation and flow of a uniphasic fluid through enclosed microchannels or conduits. Fluids are typically driven by hydrodynamic pumps through micron-sized channels made from glass, 19 silicon, metals, ceramics, or plastics such as polydimethylsiloxane 20 (PDMS) and Teflon. 21 This format has found wide application in the biological sciences but is limited by the fact that parabolic flow profiles (which generate residence time distributions) are inherent and the fact that analyte molecules are in intimate contact with channel walls, thus causing precipitation, fouling, or contamination. 22 Nevertheless, it should be noted that there are many reports of continuous-flow microfluidic systems being used successfully in drug discovery applications. 23
Batch or semi-batch methods describe techniques where the liquid phase is uniphasic, but discrete volumes may be separated from the stream to undergo further processing. A good example of such technology was reported by Hansen et al. 24 in the investigation of protein phase behavior. In this device, a mixing ring is interfaced with a series of continuous-flow channels that contain different stock reagents. The ring is filled with various amounts of the reagents, separated using valves, mixed, and the resulting solution assayed.
Finally, droplet-based microfluidic systems operate on discrete, physically separated fluid volumes or droplets. Within this broad category, two types can be distinguished: digital and segmented-flow microfluidic systems. In a segmented-flow setup, the sample or reagent phase is interrupted by an immiscible carrier phase, forcing it to split into discrete droplets. This, on first sight, mundane difference from continuous-flow microfluidic systems has profound implications on the physical behavior of the liquid. Such droplet flows do not suffer from many of the disadvantages found in continuous flows such as residence time distributions or fouling. This is because droplets are separated from each other and the channel walls by the immiscible carrier phase. Furthermore, the experimentalist can choose from a large number of readily available methods for droplet manipulation such as mixing, 25 dilution, 26 splitting, 27 merging, 28 and incubation. 29 Conversely, digital microfluidics30,31 describes the manipulation of discrete nL to µL droplets on essentially flat surfaces. Various unit operations such as transport, mixing, or dispensing can be combined to build a chip-based system able to perform a range of analytical operations. 32 A variety of techniques, based on electric and acoustic actuation, for the manipulation of droplets have been developed. 33 These actuation methods can be used to independently move droplets on a flat surface, thus eliminating the need for channels or pumps. However, significant problems such as surface fouling, droplet volume limitations, and difficulties in assembling the complex control electronics arise quickly. Some applications of digital microfluidic technology in drug discovery have been reported, due to the potential ease of integration with conventional high-throughput screening setups, but these represent a relatively niche activity. 34 Because of their ability to process large numbers of biologically distinct droplets, segmented-flow microfluidic systems have proved far more applicable to high-throughput experimentation and will form the basis of most of the proceeding discussion.
Droplet Generation
Many methods for droplet generation within chip-based formats have been reported. Active methods use time-dependent perturbations of the flow, for example, via electric fields, 35 pneumatic pressure, 36 optical fields, 37 or thermal control. 38 However, the most widely used methods for making droplets are passive in nature and rely on fluid physics. Briefly, droplets are most commonly created from two immiscible phases using either flow-focusing 39 or T-junction 40 geometries. In both geometries, the liquid with the highest affinity to the channel walls will form the continuous phase, whereas the one with lower affinity splits up into droplets. 41 In a T-junction, the carrier phase flows through a straight channel with an immiscible phase being injected from a perpendicular channel and broken into discrete droplets via shear forces. In a flow-focusing geometry, the discrete fluid is pumped through an orifice and is flanked or surrounded by two streams of carrier phase. Surface tension causes the central stream to extend and ultimately form discrete droplets. The deterministic nature of flow at the microscale allows droplet formation at kHz rates while maintaining exceptional control over droplet size. 42
Droplet Library Creation
Droplets provide many advantages over traditional storage formats for chemical or biological libraries such as multiwell plates. Picoliter-volume droplets can be stored at extremely high probe densities 43 (in excess of 2000 per mm2) and can be controllably interrogated and manipulated at extremely high frequencies (in excess of 1 kHz). 44 Importantly, long-term storage of droplet libraries has been demonstrated both on-chip (in traps and reservoirs) and off-chip (in vials, tubing, and syringes).45,46 Significantly, it should be noted that addition of appropriate surfactants to the continuous phase is hugely effective in improving droplet stability. 47 Surfactants are key to the application of droplet-based microfluidic technology and act to stabilize droplet interfaces. Traditional surfactants, such as sodium dodecyl sulfate, are generally toxic to most organisms and thus a range of bespoke biocompatible surfactants have been developed. 48
While droplets can be formed at very high frequencies, it remains challenging to form droplets of distinct chemical composition on demand.27,49 Droplet populations with nonuniform chemical contents can be created in a facile manner using the concept of dilution (see Fig. 1 ). Indeed, Niu et al. 26 recently reported a passive dilution device, which allowed the creation of a linear dilution series of droplets (on a millisecond scale) by continuously merging, mixing, and resplitting smaller dilutor droplets with a large, “mother” droplet localized within a trapping chamber. This approach was successful in generating chemically distinct droplets that could be used to assay DNA binding reactions. Droplet dilution series have also been generated by exploiting the inherent residence time distribution of continuous fluids flowing through a microfluidic channel, where contained molecules exhibit a parabolic elution profile. 50 In addition, Sun and Vanapalli 51 have used an array of static droplets, containing solution to be introduced into a dilutor plug. The dilutor plug could be moved along this array, allowing contact with each static droplet for a given period. This process resulted in the formation of a chemical gradient over the length of the dilutor plug, which spanned a concentration gradient range of three orders of magnitude.
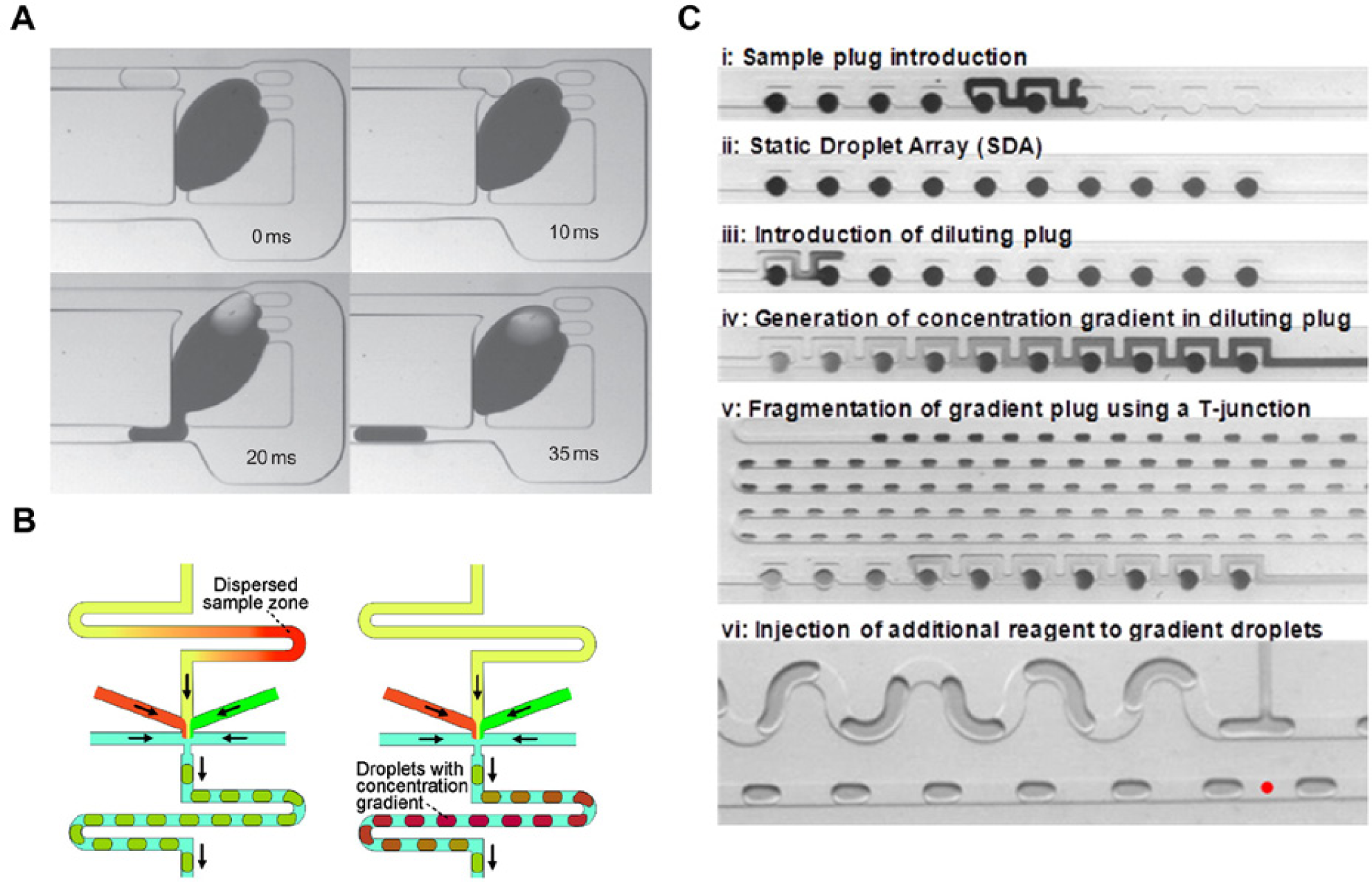
Microfluidic dilution structures. (
Recent progress in interfacing multiwell plates with microfluidic devices has also been shown to aid droplet library creation.52–54 For example, Brouzes and coworkers 52 created a mixed population of droplets from different solutions in a microwell plate. Droplets from this population could then be merged with droplets containing cells or virions, incubated and finally screened for cytotoxicity. In addition, a library of droplets with combinatorial chemical payloads was created by coalescing droplets from a preformed library with droplets formed from larger sample plugs that varied over time. This approach was used to generate 106 droplets of each of the 21 members of a combinatorial library. 55 On-chip dosing was previously used by Zec et al. 54 to selectively add contents to a preformed droplet population. In this study, the authors digitized large, serially stored sample plugs into smaller daughter droplets and used on-chip valves to selectively add reagents to each daughter droplet. Using a different approach, Kaminski and colleagues 56 created a library of nL-volume droplets from µL-volume plugs by serially separating monodisperse droplet populations with a third immiscible spacer fluid. In this way, daughter droplets could be formed at rates of up to 1 kHz.
In theory, “barcoding” individual droplets allows for direct identification of droplets over extended periods without the need to preserve the spatial position or order of droplets within a microfluidic system. This process significantly facilitates droplet incubation since droplets can be stored in large chambers or external capillary lines, thus reducing back-pressure constraints set by the fluidic system. In this respect, continuous-flow lithography has been successfully used to create particles with unique signatures or codes. 57 For example, particles can be formed by co-polymerization of two co-flowing PEG monomer solutions in a microfluidic channel that were loaded with fluorophore and biomolecules, respectively. Barcodes could be introduced using a dot-coding scheme during the polymerization process, which enables in theory the creation of more than a million distinct barcodes. 57 Barcoding based on nucleotide sequences has also been suggested as a high-efficiency route to droplet identification but would require some form of sequencing to read out the barcode. 58 Recent attempts at barcoding have typically used mixtures of fluorophores to provide droplets with unique fluorescence signatures. While dilution structures can produce droplets containing varying concentrations of a dye, they are typically limited to generating dilution series of only one dye at a time. 59
Quantum dots or compound semiconductor nanoparticles have also been used in barcoding applications due to their narrow fluorescence excitation and emission bands. This means that they can be engineered or tuned so as not to interfere with other fluorophore emission bands in complex mixtures. To this end, barcoding droplets with quantum dots has been demonstrated by incorporating quantum dots of varying concentrations into the core of pL-volume double-emulsion droplets. 60 In addition, barcoded alginate particles have been formed while concurrently mixing two quantum dot–containing solutions at varying ratios during gelation. 61 Unfortunately, a system able to generate large numbers of barcoded droplets based on fluorophores has remained elusive to date. This is most likely closely linked to the difficulties encountered when creating a chemically diverse droplet population.
Probing Droplet Contents
Fluorescence detection is most commonly used to probe the small volumes associated with microfluidic systems, because of its high sensitivity and low detection limits. 2 In addition, many materials from which microfluidic devices are made are transparent in the visible, near-infrared (IR) and UV regions of the electromagnetic spectrum.20,62 Furthermore, fluorescence signals can be acquired and processed on extremely short timescales, which facilitates in-line processing and aids the realization of real-time feedback loops. 63
A nice example of the utility of fluorescence detection in a high-throughput, continuous-flow microfluidic experimentation was presented by Taniguchi and coworkers 64 for the screening of an Escherichia coli YFP-fusion library consisting of 1018 strains. To increase experimental throughput, the authors used a microfluidic-processing platform, which allowed them to monitor 96 bacterial strains concurrently. This, in turn, enabled the determination of both variations in protein and mRNA levels of all strains in the library based on single-cell measurements.
There are many reports of using fluorescence to probe droplet-based microfluidic systems in high throughput. As an example, Brouzes and coworkers 52 have screened a library of chemical compounds stored within droplets by merging them with cell-containing droplets, incubating the merged population, and assessing the survival rate of the contained cells via fluorescence.
Fluorescence-activated droplet sorting has also been demonstrated by a number of researchers, where the fluorescence signal originating from an individual droplet is assessed in line prior to physical sorting. For sorting, droplets are typically deflected into a target channel using various actuation methods, which include alternating current, 65 high-frequency ultrasound, 66 or on-chip valves. 67 Such an approach has been shown to be successful in sorting droplets containing E. coli cells based on their fluorescence signature, at kHz rates and with extremely low false-positive rates. 65
A variety of image-based assays have also been implemented in microfluidic devices. Hofmann and colleagues 68 reported the label-free detection of reactant consumption by exploiting differences in osmotic pressure between droplets. The authors were able to observe that droplets containing cells consumed certain metabolites present in the droplet. In addition, water diffusing from cell-containing droplets through the continuous phase into non–cell-containing droplets resulted in variations in droplet size. This allowed the authors to gauge metabolite consumption based on the variation of droplet size. It should also be noted that structures that exploit “mechanical biomarkers” of cell status such as size, shape, or deformability have been used to achieve passive sorting and culture enrichment. 69 Microfluidic structures are especially suited to the assessment of mechanical biomarkers because typical feature sizes are often similar to the characteristic length scales of cells. Using a continuous-flow microfluidic device, Bow et al. 70 were able to sort low-abundance, malaria-infected, red blood cells from uninfected cells due to the decreased cell deformability upon infection.
Ligand-Binding Assays
Many biochemical assays used in drug discovery depend on ligand binding of biomolecules or whole cells. Several such assays have been implemented in microfluidic formats. Because purified ligands are precious and expensive, such assays mostly profit from the inherent low-volume environment used in continuous-flow and droplet-based microfluidic devices, which typically leads to reduced reagent consumption.3,9,71
Several groups have used surface treatment of microfluidic channels to coat channel walls with biomolecules.71–73 For example, Stott et al. 74 coated the walls of a PDMS microdevice with antibodies and showed that patterning the bottom of the microfluidic channel with herringbone structures introduced micro-vortices into the flow, greatly increasing the interaction of biomolecules with the coated walls. Numerous variants of phage-display on chip have also been reported in the literature. For example, Wang et al. 75 realized phage-display within a microfluidic chamber, enabling the discovery of peptides with higher affinity and specificity against surface markers from live mammalian cells through increased control and reproducibility of the chemical environment. Since phage-display is inherently a high-throughput experimental technique, it lends itself to microfluidic implementation and optimization. Microfluidic environments are also enabling when performing multiple phage displays in parallel and without the need for bacterial infection. This resulted in a 15-fold reduction in the experimental time needed for the analysis of four targets compared with traditional phage-display formats. 76 Other recent efforts in this area include the functional screening of hybridoma cells for the release of antibodies inhibiting a drug target at a screening rate of 300,000 cell clones per day 77 and chromatin immunoprecipitation on a chip to select for antibodies against specific proteins. 78
In 2007, Maerkl and Quake 79 introduced the MITOMI (Mechanically Induced Trapping of Molecular Interactions) platform (see Fig. 2 ). Although continuous flow rather than droplet based, the platform has proved to be highly versatile for high-throughput ligand-binding experiments. Examples of its utility include assessment of RNA binding of membrane proteins after in vitro protein synthesis 80 and identification of preferred binding sequences by assessing binding of transcription factors to all possible eight base-pair DNA sequences. 81 Recently, MITOMI has also been used to perform co-immunoprecipitation, allowing the inference of protein functions for uncharacterized proteins of Streptococcus pneumoniae through analysis of protein-protein interactions. 82
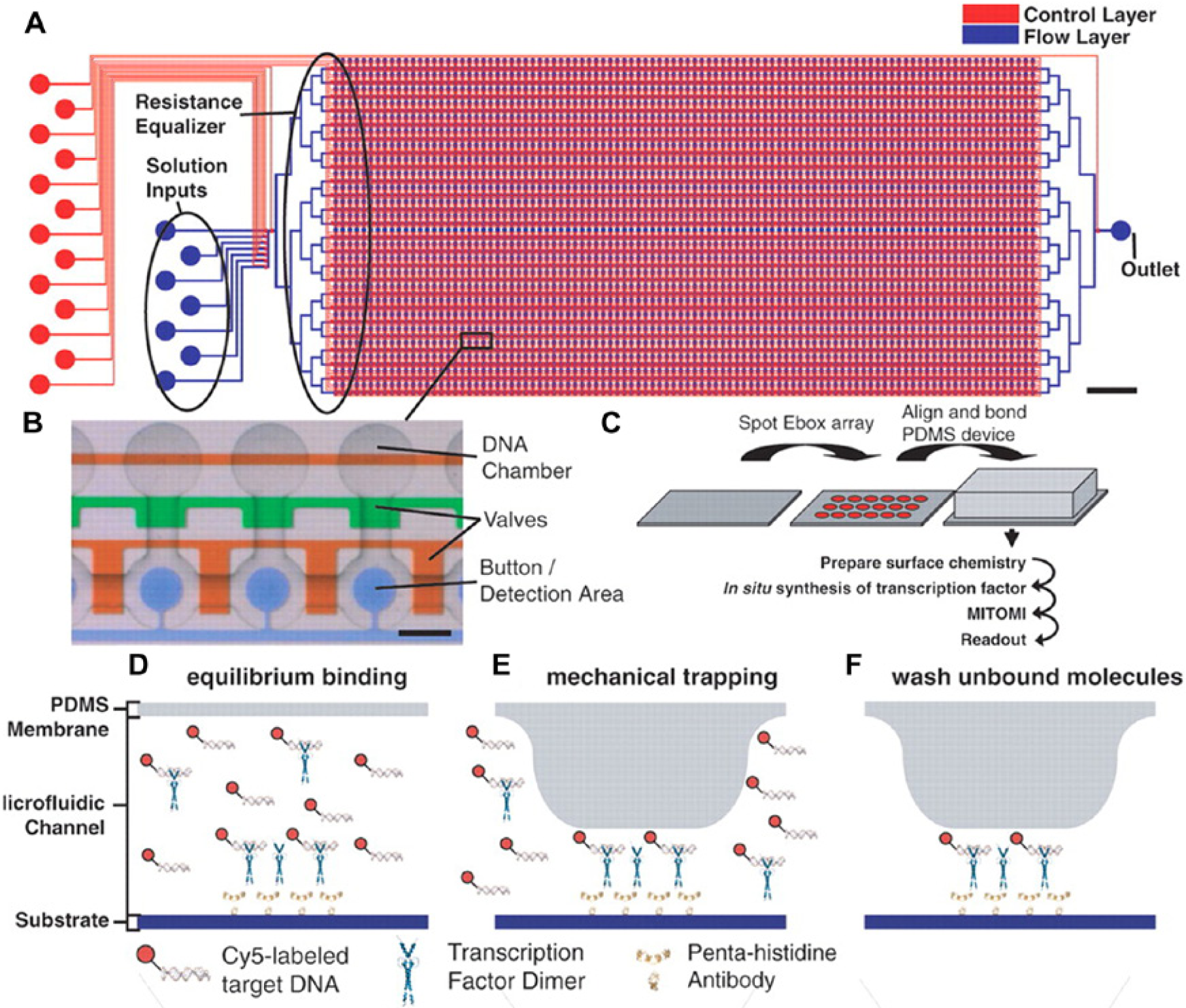
MITOMI (Mechanically Induced Trapping of Molecular Interactions) platform. MITOMI can be used to determine DNA transcription factor binding.
79
Large-scale integration (
PCR-Based Assays
PCR is one of the most indispensable reactions in molecular biology and used to determine the presence of a specific nucleotide sequence in a biological sample. 83 It is based on repeatedly cycling a reaction mixture between two or three different temperatures. Since heat transfer is increased as reaction volumes decrease, overall reaction times for PCR can be significantly reduced when performed on chip.82,84 Several methods have been used to achieve temperature control of the reaction mixture in a microfluidic chip. These have included microfabricated heater elements,85,86 Joule heating, 87 noncontact heating, 88 and Peltier elements, 89 among others.
Continuous-flow PCR presents an exciting alternative to traditional (bulk) PCR. Here a reaction mixture contained within a fixed vessel is thermally cycled between different temperatures to achieve strand separation, primer annealing, and polymerase-induced strand extension. During continuous-flow PCR, the reaction mixture is physically moved through different, spatially localized temperature zones. The first example of continuous-flow PCR in a microfluidic system was reported in 1998 and shown to reduce the total time needed for 20 PCR cycles down to 90 s. 84 Subsequently, RT-PCR was achieved in continuous flow, performing reverse transcription in a first step and PCR in a second step on a single monolithic chip. 90 Quantitative PCR in continuous flow can also be performed by optically monitoring the reaction progress at different points along the length of the channel. 91
Parallelization of PCR in microfluidic formats can be realized by operating many chambers concurrently. For example, Marcus et al. 92 reported that low-abundance RNA templates could be detected down to 34 copies by performing RNA RT-PCR in 72 parallel chambers. However, droplet PCR is a more promising approach for performing multiplexed PCR, since each droplet defines a separate reaction container91,93. (see Fig. 3 ) To this end, Schaerli et al. 89 reported a radial droplet PCR device incorporating a single Peltier element as a heat source and exploiting the radial temperature gradient created through heat dissipation from the element. The authors demonstrated successful PCR using a single-template DNA molecule per droplet. More recent work has demonstrated both DNA extraction and RT-PCR on a single chip, using an aluminum oxide membrane for DNA extraction. This allowed the authors to forego complicated architecture for sample transport. 94
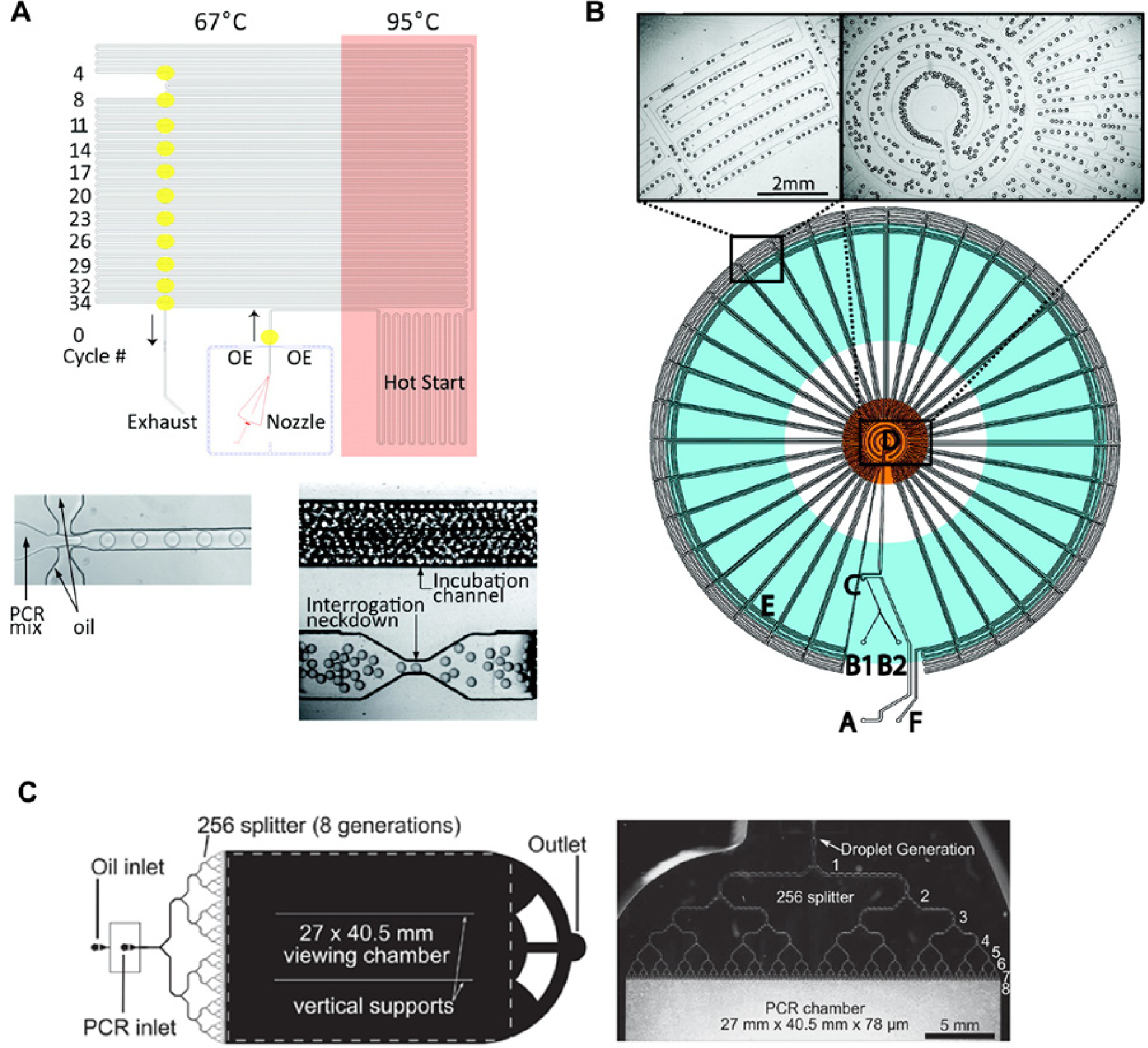
Digital PCR in Droplets. (
Quantifying very low-abundance nucleotide sequences such as rare mutants can be challenging using PCR because they are often masked by other high-abundance sequences. Digital PCR can be used to circumvent such problems. Here many separate PCR reactions are performed using a highly diluted sample, such that only a portion of the reactions contains a single template sequence. The ratio of reactions showing amplification versus nonamplifying reactions can be used to directly infer the concentration of the sequence in the initial mixture. Digital PCR in microfluidic droplets has been shown in multiple studies.91,95 Shen and coworkers 96 reported digital PCR using a SlipChip-format, which allowed them to perform 1280 PCR reactions in parallel. Hatch et al. 97 have further reported a quantitative digital PCR system that forms one million droplets in a single microfluidic device. This was achieved by repeatedly splitting large sample plugs into smaller “daughter” droplets, all of which were subsequently stored in a large chamber. PCR was then performed on all droplets in parallel while they were monitored using wide-field fluorescence imaging.
Proteomics Assays
Multiple studies have used continuous-flow–based microfluidic devices to probe and analyze complex mixtures of proteins and peptides. Hughes et al. 98 showed a device capable of performing protein isoform analysis via isoelectric focusing. Notably, the authors forewent the use of valves and pumps by using electrokinetic fluid transport. In addition, there have been several attempts at interfacing microfluidic chips with mass spectrometers. This is an especially important development for proteomics because mass spectrometry–based methods have become the de facto standard in the field. 99 To this end, Mao et al. 100 reported a multinozzle emitter array able to perform liquid chromatography–mass spectrometry (LC-MS) using a microfluidic chip in line with a mass spectrometer. This format was shown to be highly useful for small-volume proteomics, with the authors showing parallel, on-chip, and online LC-MS analysis of hemoglobin and its tryptic digests directly from microliters of blood. In addition, Chao and Hansmeier 101 reviewed the possibility of implementing all the necessary preprocessing steps for shotgun proteomics (cultivation, cell lysis, purification, digestion, and separation) on a single microfluidic chip.
In recent years, droplet-based microfluidic systems have also been interfaced with mass spectrometers to afford high-throughput analysis of large numbers of isolated samples. For example, Küster and coworkers 102 reported a platform able to deposit up to 26,000 droplets MALDI-MS (matrix-assisted laser desorption/ionization–MS) target patterned with hydrophilic spots at a rate of 0.7 Hz. Upon deposition, droplets were located on the hydrophilic patches, and the continuous phase was removed by evaporation. Directly separating the dispersed phase from the continuous phase poses an additional experimental challenge but is often necessary, since the overabundant continuous phase can often swamp sought after (desired) peaks (fragments). A recent innovation by Pereira et al. 103 used a passive structure to remove the oil phase from a microfluidic segmented flow in a rapid manner. This allowed direct deposition of analytes onto a conventional (unmodified) MALDI-MS target and obviated the need to rely on evaporation of the continuous phase after deposition. By analyzing tryptic digests of bovine serum albumin and cytochrome c, the authors demonstrated that their microfluidic interface enhanced analytical performance by at least 50% compared with conventional interface technologies.
Cell-Free Protein Synthesis on Chip
Cell-free protein synthesis provides many advantages for high-throughput applications compared with cell culture–based protein synthesis, since it does not require the transformation or culturing of cells and thus greatly accelerates the screening of large protein libraries. Furthermore, cell-free protein synthesis allows for easy modification of reaction conditions, as well as minimizes required reaction volumes and process times. 104 Microfluidic technology can be used to further leverage these advantages and aid in high-throughput cell-free protein synthesis. In vitro protein synthesis in a droplet-based microfluidic system was recently demonstrated by Mazutis et al. 46 (see Fig. 4 ) by combining both on-chip and off-chip operations. This system was used for in vitro transcription and translation of Bacillus subtilis cotA laccase genes and kinetic analysis of the catalytic activity of the translated protein. Subsequently, a macroscale array for parallel cell-free protein synthesis was reported by Khnouf and coworkers. 105 Another recent study in this area used a microfluidic device to perform droplet-based PCR (for single-gene amplification) and then translation of the amplified genes into proteins within a cell-free system. The authors showed that it is possible to screen one million genes using only 150 µL of template solution. Since droplets can be sorted at rates in excess of 2000 s–1, the complete experiment could theoretically be performed in less than 1 h. 44
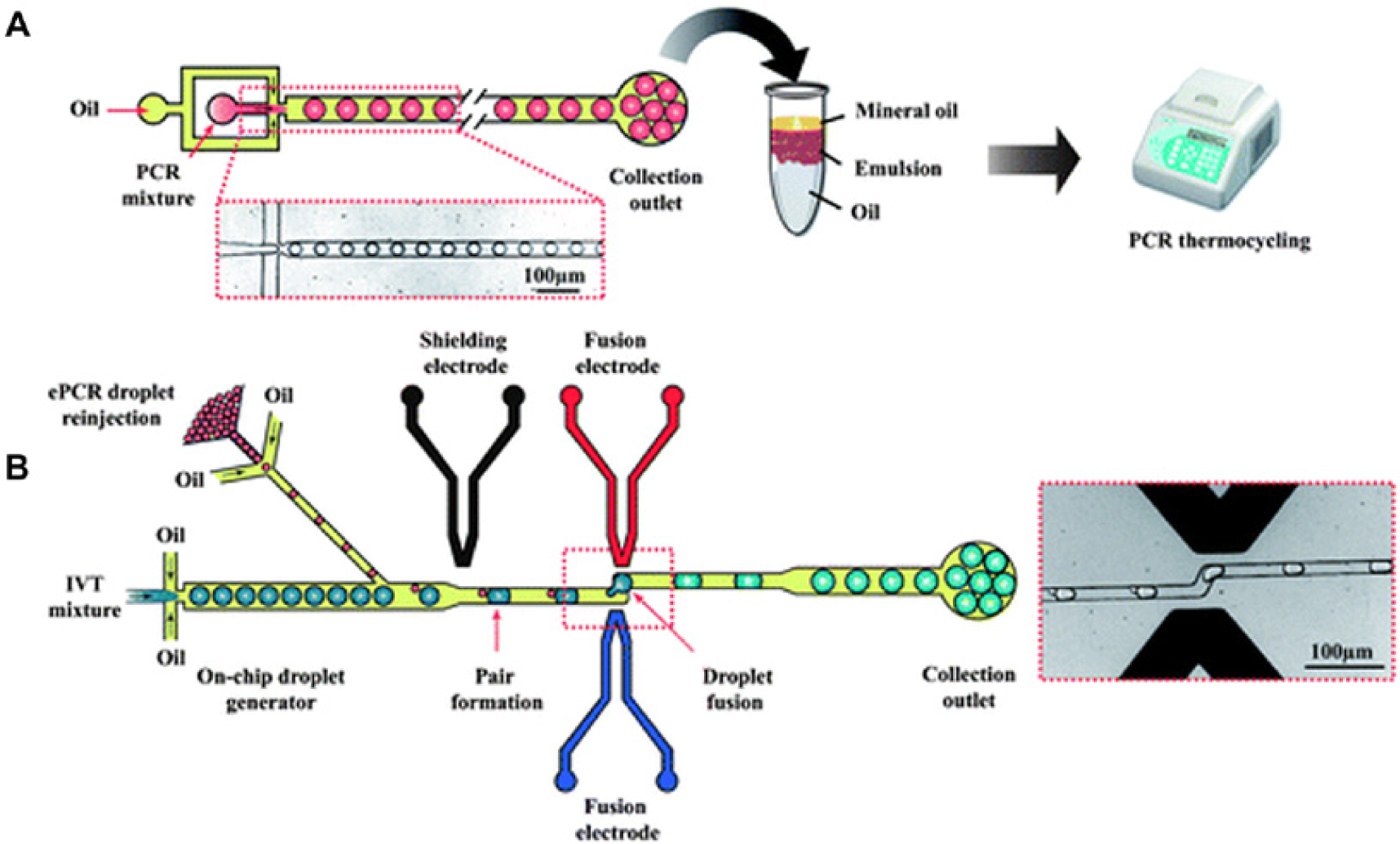
Cell-free protein synthesis. (
3D Cell Culture on Chip
Current cell-based disease models provide only limited diagnostic information because they generally analyze the drug response of isolated cells in an artificial environment. Animal testing, on the other hand, enables the analysis of systemic effects of a compound. Unfortunately, the obtained information is only transferable or applicable to humans within certain boundaries due to metabolic and physiological differences. In addition, animal testing is subject to various ethical constraints. Accordingly, microfluidic devices may serve to bypass or mimic animal testing by re-creating tissues using co-culture of multiple cell types. 106 These devices generally aim to achieve controlled growth of two or more cell types on a 2D or 3D scaffold. Typically, these devices aim to model key processes in human organs involving the interplay of multiple cell types and thereby providing insight into human physiology. 106
Several efforts have been made to re-create liver function on chip because of its central position in drug metabolism and the fact that liver toxicity is a common side effect of many drugs.107–111 Moreover, intestines-on-a-chip have proven useful for the study of drug uptake and toxicity.112–114 Here devices typically serve to study polarized transport activity using intestinal cell models. 106 Huh et al. 115 have re-created a “lung-on-a-chip” by mimicking the alveolar-capillary interface lithographically. In such a system, the authors showed the importance of including mechanical actuation of the interface into the model system. This was achieved by applying a fluctuating vacuum to the alveolar side of the membrane. This led to repeated stretching of the PDMS membrane and initiation of cell growth. This resulted in behavior more comparable with in vivo measurements. Among other uses, the authors applied their lung-on-a-chip to investigate the pulmonary inflammatory response and mechanosensitive response to nanoparticles. Other notable examples of organs-on-a-chip include the incorporation of a mouse arteric segment into a microfluidic chip 116 and the growth of a tumor-on-a-chip, which in turn allowed photodynamic therapy-based measurements. 117
Organisms-on-a-Chip
While drug discovery using cell-based assays is ubiquitous, such assays often do not allow the testing of systemic effects of the investigated chemical compounds. Accordingly, testing novel compounds on multicellular model organisms greatly increases the understanding of such systemic effects but is generally more challenging and time-consuming. To this end, a number of microfluidic platforms have been developed to facilitate culture, manipulation, and analysis of several multicellular biological model organisms.
The roundworm Caenorhabditis elegans is well suited to microfluidic culture and manipulation because of its size and short generation time of 4 days. C. elegans eggs can be packaged into droplets and nematodes grown and monitored for up to 9 days. 45 Such operations have allowed for the quantification of the effects of chemical compounds on C. elegans growth and development. Moreover, investigation of C. elegans in droplets immobilized in traps over 120 min was shown by Shi and coworkers. 118 In recent years, C. elegans has been increasingly used as a model organism to study the effects and mechanics of aging. Traditional experiments often include manual sorting of C. elegans based on physical traits such as size and motility. This is typically a time-consuming process and does not allow for a high number of experiments per unit time. To address this limitation, Chung et al. 119 introduced a microfluidic worm sorter, which allowed the sorting of C. elegans nematodes based on gene expression patterns. Automatic sorting of C. elegans based on subcellular traits, using a fluorescence readout, has also been shown, and resulted in a 100-fold increase of sorting speed compared with manual sorting. 119 Electrotaxis (the directional movement of motile cells in response to an electric field) has also previously been used to sort worms based on their speed of movement, which allowed sorting based on the age of the nematodes. 120 Interestingly, Casadevall i Solvas and colleagues 121 reported a passive microfluidic device capable of sorting worms according to their size and behavior within a continuous-flow microfluidic environment. The passive nature of the chip allowed for high-throughput age-based sorting of more than one million nematodes per day with minimal larval contamination of adult populations. Liu and colleagues 122 have also proposed a method for lens-less tracking of C. elegans movement in a microfluidic chip, where two microelectrode grids were used to locate a worm via conductance measurements. Other organisms cultured on a microfluidic chip included Drosophila melanogaster embryos 123 and Danio rerio larvae. 124
Market Penetration
Despite the myriad benefits afforded by microfluidic tools and platforms, it is sobering to note that over the past two decades, very few “killer apps” have been identified and a surprisingly small number of commercial products (based solely on microfluidic components) have established themselves as core tools in the chemical and biological sciences. Despite this lack of market penetration, it is clear, however, that success has typically been encountered in applications where reduced reaction volumes and high analytical throughput are demanded. In molecular biology, chip-based electrophoresis systems, using gel-filled channels in glass or plastic devices, have had some success in providing a robust format for efficient and fast separation of nucleic acids and proteins. 125 However, recent developments in next-generation sequencing and DNA amplification perhaps highlight a more significant commercial adoption of microfluidic tools. For example, RainDance Technologies (Billerica, MA) has developed commercial droplet microfluidic systems that allow for targeted DNA sequencing and digital PCR within millions of pL-sized droplets. 126 Through the use of segmented-flow microfluidics, such technologies provide access to unparalleled experimental throughput and have significantly opened up the utility and application of digital PCR as a basic research tool. In addition, it should be noted that many next-generation sequencing systems use microfluidic flow cells to process genomic libraries in parallel, which again improves analytical throughput while minimizing sample consumption. 127
Outlook
Over recent years, activities within the field of microfluidics have evolved from pure technology-driven, proof-of-concept experimentation toward application-driven research. In this review, we have highlighted a small number of experimental approaches where microfluidic technologies have directly or indirectly affected the process of drug discovery. Nevertheless, it should be stressed that commercial applications of microfluidic technology are still rare with experiments mostly being performed using custom-built apparatus in research laboratory environments. We expect that future developments and standardization of chip-to-world interfaces will further facilitate microfluidic adaptation, since there are few (if any) standardized sample vessels for microfluidic volumes, such as multiwell plates commonly encountered on the macroscale. We are also convinced that further efforts in commercialization will drive broader adaptation of microfluidic techniques outside the core research community and into commercial drug discovery environments.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported in parts by a National Research Foundation of Korea (NRF) grant funded by the Ministry of Science, ICT & Future Planning (MSIP) of Korea (Grant Numbers K20904000004-12A0500-00410).