Abstract
Modern flow cytometers can make optical measurements of 10 or more parameters per cell at tens of thousands of cells per second and more than five orders of magnitude dynamic range. Although flow cytometry is used in most drug discovery stages, “sip-and-spit” sampling technology has restricted it to low-sample-throughput applications. The advent of HyperCyt sampling technology has recently made possible primary screening applications in which tens of thousands of compounds are analyzed per day. Target-multiplexing methodologies in combination with extended multiparameter analyses enable profiling of lead candidates early in the discovery process, when the greatest numbers of candidates are available for evaluation. The ability to sample small volumes with negligible waste reduces reagent costs, compound usage, and consumption of cells. Improved compound library formatting strategies can further extend primary screening opportunities when samples are scarce. Dozens of targets have been screened in 384- and 1536-well assay formats, predominantly in academic screening lab settings. In concert with commercial platform evolution and trending drug discovery strategies, HyperCyt-based systems are now finding their way into mainstream screening labs. Recent advances in flow-based imaging, mass spectrometry, and parallel sample processing promise dramatically expanded single-cell profiling capabilities to bolster systems-level approaches to drug discovery.
Keywords
Introduction
At the time of its development in the latter part of the past century, flow cytometry represented an unparalleled advance in the speed with which it was possible to make quantitative, multiparameter measurements on single cells.1–3 Large populations (thousands) of cells could be routinely evaluated in seconds as compared with the hundreds in minutes feasible with conventional microscopy of the time. Today, even the most modestly priced modern flow cytometers can distinguish and make quantitatively precise measurements from cells in a single sample that may differ by as many as five orders of magnitude in fluorescence intensity, ranging from as few as 50 fluorophores per cell to 5 million or more.4,5 Many flow cytometers use hydrodynamic focusing, a technique that encloses the sample stream within a more rapidly flowing sheath stream, to narrow the sample stream diameter to approximate that of a typical cell. A cell thus displaces most of the sample stream liquid in its immediate vicinity so that when it passes through the spot of the laser beam (also of diameter similar to the cell), it is predominantly cell-associated fluorophores that are excited and detected. This configuration geometry dramatically diminishes the contribution of unbound fluorophores to the fluorescence signal and enables the flow cytometer to make biologically significant optical measurements without wash steps.6–8 To a first approximation, the data acquisition capabilities of modern flow cytometers are relatively insensitive to the number of optical parameters that are detected. The ability to accurately measure each parameter is primarily limited by the properties of the fluorescent probes (e.g., quantum yield, spectral overlap) and features of the targeted particles (e.g., autofluorescence and number of determinants available for probe binding). Currently capable of detecting 20 or more discrete optical parameters per cell at tens of thousands of cells per second, 1 flow cytometry remains unequaled in sensitivity, dynamic range, and information content for applications requiring high-speed analysis of single cells or particles in suspensions.
The flow cytometer has been used both in pharma and biotech companies at every stage of the drug discovery cycle, including target identification and validation, hit identification, lead and candidate selection, and safety studies. 9 However, its use has typically been restricted to applications tolerant of relatively low rates of sample throughput. This has reflected limitations of the conventional flow cytometer “sip-and-spit” sample-processing approach in which each sample aspiration cycle is followed by a back-flush cycle to clean the sample uptake line. The back-flush cycle significantly lengthens sample-processing time. It also results in loss of the unanalyzed volume of sample that remains in the uptake line, often a significant fraction of the total aspirated sample, sending it to waste. Slowing the sampling process further is the need to save data acquired from each sample to a data file before beginning data acquisition for the next sample. Automated systems using this sampling approach have typically required up to an hour or more to read a 96-well plate. The sample-processing speed of this approach has been boosted several-fold with the commercialization of the High Throughput Sampler system by Becton Dickinson as an accessory for some of their flow cytometers. Although this has better enabled screening of small, focused libraries containing hundreds to thousands of compounds, 10 it has still been inadequate to accommodate the scale and size of assays run in many drug-screening laboratories.
Exploiting Time and Air
The concept of collecting data from multiple samples into a single temporally resolved data file was first advanced in 1991 as a solution for speeding multisample analysis by eliminating the time demands imposed by saving data from each sample as a separate file. 11 During a single extended round of data acquisition, sample tubes were introduced one at a time into the sample uptake pathway. Between samples, a coded voltage pulse was manually applied to one of the data acquisition channels. The pulse served as a time stamp by which the time domain boundaries for each sample’s data could be distinguished in subsequent analysis of the extended data file, a process designated as time interval gating.11,12
A faster and more automated approach, called plug flow cytometry, was subsequently developed that used a reciprocating, multiport flow injection valve with two 5 µL sample loops to serially process samples. 13 A sample would be aspirated into loop 1 while a second sample in loop 2 was being simultaneously delivered to the flow cytometer for analysis. A switch of the valve to its alternate position allowed filling of loop 2 with a third sample while the loop 1 sample was analyzed. Continuous reciprocation of the valve and acquisition of data in a single, time-resolved data file enabled processing of up to 10 endpoint assay samples per minute, 4 online sample-mixing experiments per minute, and a 15-point concentration gradient dose-response experiment in less than 2 min.14,15
Introducing air bubbles into the sample stream of a flow cytometer was (and still is) considered to be undesirable for many types of analysis because of disruption resulting from passage of a bubble through the laser beam. However, in time-resolved data acquisition, bubble passage events could easily be distinguished and used as an efficient means for segregating a data stream into discrete, unperturbed segments. 16 This finding was exploited to implement the more rapid and flexible sample-processing approach called HyperCyt; samples and air bubbles were alternately aspirated into a sample line and delivered to the flow cytometer as a segmented sample stream by a continuously rotating peristaltic pump, perhaps more akin to a “sip-and-whiff” approach17,18 ( Fig. 1 ). For highest throughput, the aspirated samples are typically 1 to 2 µL in volume to enable, for example, processing of a 384-well plate in 10 min or less. In contrast to the sip-and-spit approach, total assay volumes in a well can be as small as 4 to 5 µL, and each aspirated sample is analyzed in its entirety, with little measurable dead volume. Larger sample volumes are also manageable, limited only by the total volume available in the well. Because of the unique use of air bubbles to segment a continuous stream of samples, it has been considered important to characterize performance features of the system in some detail, including determinants for control of fluid and particle carryover between samples, 19 effective particle concentration ranges, 18 inline mixing of convergent sample streams,20,21 fluid carryover in association with inline receptor stimulation for delivery to a cell sorter, 22 immunophenotyping, 18 and compound screening. 23
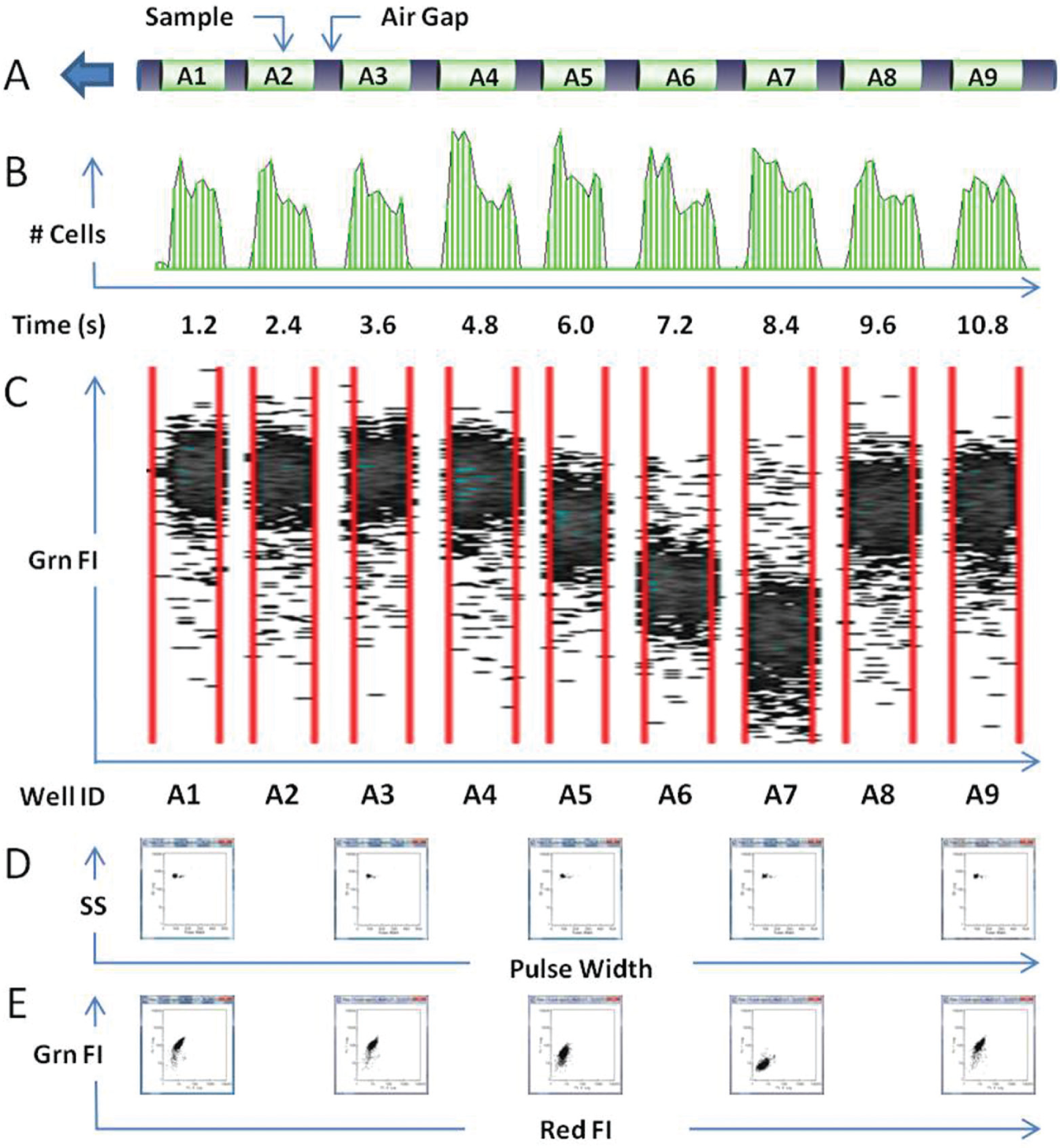
HyperCyt-based high-throughput flow cytometry. (
Since 2005, when the University of New Mexico Center for Molecular Discovery (UNMCMD; originally the New Mexico Molecular Libraries Screening Center) first joined the USA National Molecular Libraries Screening Center Network, the platform has been used to screen, predominantly in 384-well format, more than 60 biological targets to produce more than 13 million data values that are publically accessible on the PubChem Web site. 24 IntelliCyt Corporation (Albuquerque, NM) was founded in 2006 to commercialize the HyperCyt platform. Compatibility of the HyperCyt platform with the high-density, 1536-well screening format was first introduced at the UNMCMD in 2012 25 and is now in routine use.26–30 Two fully integrated HyperCyt-based systems enabled for high-density, 1536-well screening were subsequently introduced in 2014, one a commercial platform from IntelliCyt (iQue Screener HD, http://www.intellicyt.com/ique-screener.html) and another by a group at the Genomics Institute of the Novartis Research Foundation. 31 Thus, there is now a wealth of data to support the validity of this approach for compound screening as well as commercial systems capable of high-throughput screening (HTS) in low- to high-density plate formats.
The intent of this review is to (1) highlight selected screening projects at the UNMCMD and elsewhere that exemplify platform features of importance in HTS, (2) discuss evolution of the platform and trends that have influenced its adoption, and (3) review recent technological advances and industry trends likely to affect flow cytometry applications in drug discovery now and in coming years.
Target-Based Screening
A frequent screening objective is to identify compounds that disrupt or augment binding interactions between specified molecules. This is commonly referred to as target-based screening as the identities of the molecules with which compounds may interact are known and directly targeted. Typical examples are receptor-ligand interactions such as proteins binding to other proteins, peptides, or polynucleotides.
Target Multiplexing with Beads
A particularly attractive feature of flow cytometry is the high dimensionality of the data that can be obtained. Not only is it easily possible to discriminate from 6 to as many as 20 optical parameters in parallel, but the broad intensity response range of modern instruments (five orders of magnitude or more) can be exploited for intensity encoding of cells and beads within one or more optical parameter detection channels to produce even higher levels of dimensionality. Although such encoding occurs naturally (for example, large and small particles or cells discriminated on the basis of forward light scatter intensity), it can be artificially implemented as a means for discriminating multiple distinct biological targets in a single sample.
An extreme example is illustrated by the bead-based xMAP system from Luminex Corporation (Austin, TX) in which intensity encoding with 10 concentrations each of red and orange fluorescent dyes produces 100 sets of beads that can be distinguished in a correlated plot of two optical parameter channels 32 ( Fig. 2A ). Each bead set can be engineered to express or capture a different target analyte (protein, DNA, etc.), and a green fluorescent reporter molecule, detected in a third optical parameter channel, is used to quantify the analyte. In this fashion, only three fluorescence channels are required for quantifying 100 analyte targets in a single well.
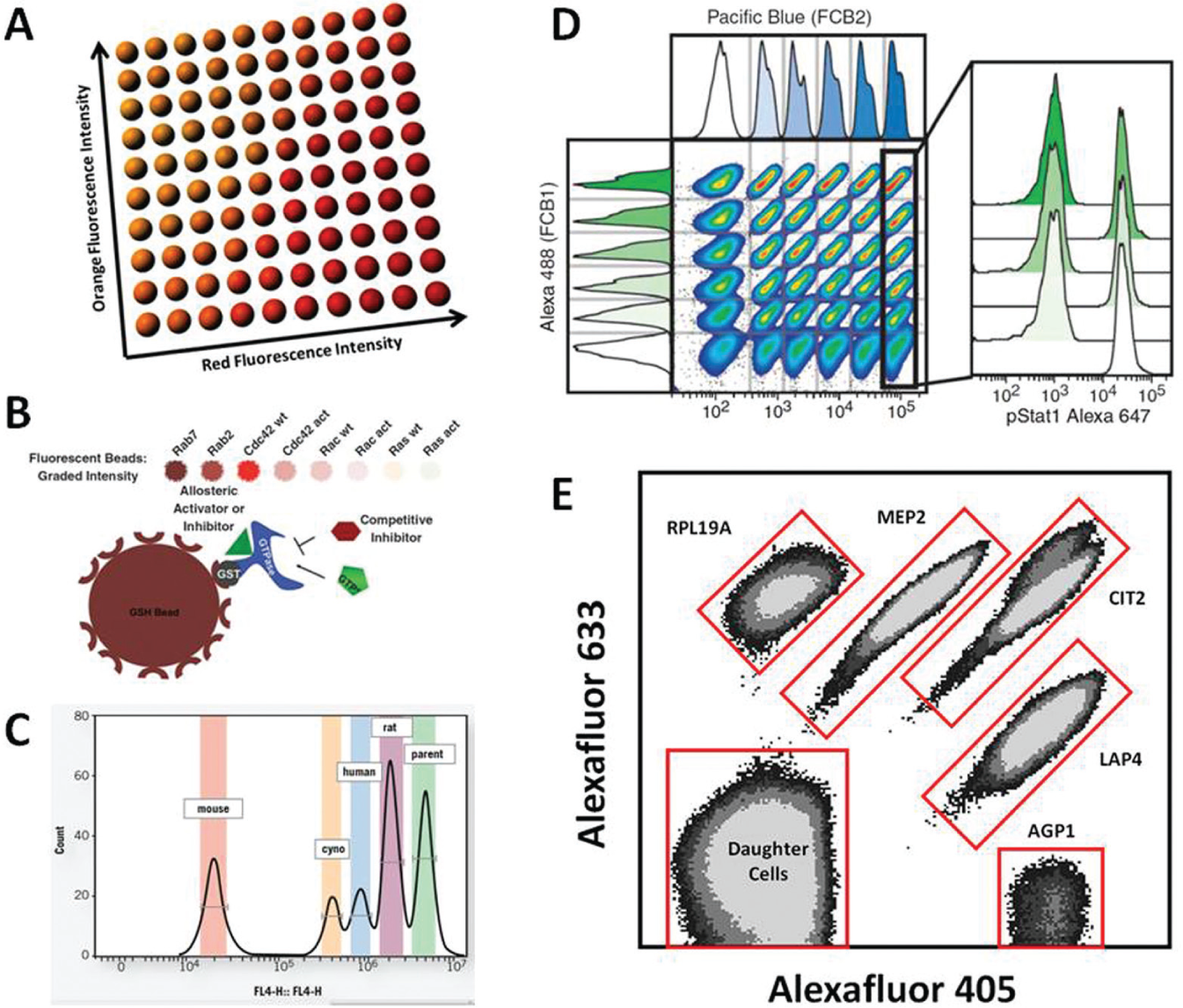
Configurations for multiplexed flow cytometry analysis in target-based and phenotypic screening. (
Bead-based platforms have been extensively used for high-throughput, multiplexed target-based screening flow cytometry. For practical purposes related to complexities of assay design and reagent costs, these assays have typically been limited to multiplexes of less than 10 targets (see the “Balancing Throughput and Content” section below). In a recent, representative example, such an approach was used to identify small-molecule inhibitors and activators of small GTPases, intracellular signaling proteins of 20 to 40 kDa that control diverse cellular functions. Rho family GTPases that were evaluated included wild-type versions of Rab7, Rab2, Ras, Rac, and Cdc42, plus mutant, constitutively activated forms of Ras, Rac, and Cdc42 ( Fig. 2B ). All were glutathione-S-transferase tagged, attached to red fluorescence intensity–encoded glutathione bead sets, and assessed for binding of green fluorescent GTP in the presence and absence of ~200,000 compounds from the Molecular Libraries Small Molecule Repository (MLSMR). 33 This screening approach has identified a pan inhibitor of Ras superfamily GTPases, 34 inhibitors of Rho family GTPases,35,36 a selective inhibitor of Cdc42 GTPase,37,38 and several activators that selectively promote increased binding of GTP to Rho family GTPases. 39 An attractive feature of this approach was that it enabled assessment of compound selectivity at the primary screening stage, streamlining compound prioritization for later stages in the pipeline. Immobilizing GTPases on the beads (2000 beads/well, 10 6 binding sites/bead) reduced total enzyme requirements as compared with solution-based assays compatible with other screening platforms. This highlights a general rule of thumb for assembling flow cytometry binding interaction assays: it is most efficient and cost-effective to attach the most expensive or quantity-limited binding partners to the beads. Additional examples of molecular binding interactions screened with bead-based, high-throughput flow cytometry applications include the binding of five regulators of G-protein signaling proteins to the G-protein Gαo,40–42 binding of six antiapoptotic members of the human Bcl2 family to a common proapoptotic family member (Bim),43–46 binding of an RNA aptamer to GRK2 Ser/Thr protein kinase outside of its active site,25,47 binding to three MEK kinase targets (MEKK2, MEKK2 mutant, and MEKK3) of the PB1 domain of MEK5, 48 binding of pathogenic antibodies to a pemphigus-associated desmoglein epitope, 49 binding interactions governing the catalytic cycle of the proteasome, 50 and multiplexed analysis of fluorescent substrate cleavage by proteases, including lethal factor, factor Xa, and botulinum neurotoxin light chains A and F.26–28,51–53
Target Multiplexing with Cells
Target-based assays in cells are most practical when it is important that the target be displayed in a physiologically relevant context. Flow cytometric analysis of receptor-ligand interactions in intact cells may minimize ligand depletion issues associated with solution-based assays that use cell lysates. 54 A more important advantage of cell-based flow cytometry assays is the ability to evaluate multiple targets in parallel by assay multiplexing.
Fluorescence intensity-based multiplexing methods in cells, popularized in nomenclature as cell fluorescent barcoding, 55 have been used for discriminating collections of cells with different innate biological characteristics (e.g., clonal origin) or environmental history (e.g., previous exposure to stimulants). Cell-barcoding methods have incorporated the use of varying concentrations of fluorescent dyes 56 ( Fig. 2D ), antibodies precoupled with varying ratios of fluorescence-amplifying reagents, 57 and varying cellular expression levels of fluorescent proteins. 58 To a large extent, flow cytometry screening applications using cell-barcoding schemes have been for unknown target molecules for which the screening assay endpoint is a cell-associated phenotype (phenotypic screening, discussed below). One target-based screen in cells, targeting a pair of G-protein–coupled receptors (GPCRs) belonging to the formylpeptide receptor family, is referenced in a separate section below. Others, highlighted here, reflect the utility recognized for target multiplexing in hybridoma screening.
Screening for biologics, in particular antibody drugs, has evolved significantly over the past 20 years. 59 Screening hybridoma supernatants for specific antibodies that bind cell-based antigen is a critical component of monoclonal antibody generation and often a bottleneck in the process. Also, even when an antibody is identified that binds a human target antigen with high specificity, it may have limited ability to bind the orthologous antigen in other species, an impediment to preclinical testing. A recent, illustrative solution to address this issue is a flow cytometry–based hybridoma antibody screening application used at GlaxoSmithKline. 9 The objective was to assess each of thousands of hybridoma supernatants for antibodies binding to (1) cells expressing a human protein of interest, (2) negative control cells expressing a related but different protein, and (3) cells expressing the orthologous protein from two different animal species of interest (cynomolgus monkey and mouse). Rather than testing supernatants against each cell line separately, the four different cell populations were barcoded by incubation with four different concentrations of a fluorescent dye (calcein-AM), then combined in wells together with hybridoma supernatants and reporter antibodies that had a fluorescence signature distinct from the barcoding dye. In subsequent analysis in a HyperCyt flow cytometry–based platform, each cell population was distinguished on the basis of barcode fluorescence, and the binding profile of antibodies in each hybridoma supernatant was quantified for all four simultaneously. A similar hybridoma screening approach has been used at XOMA Corporation, an antibody therapeutics biotech in Berkeley, California, to streamline the determination of hybridoma antibody binding profiles to orthologous antigens from four different species 60 ( Fig. 2C ). Such target-multiplexing methodologies enable identification of lead candidates with cross-reactivity against multiple species early in the discovery process, when the greatest numbers of candidates are available for evaluation.
Phenotypic Screening
There is an ongoing paradigm shift in drug discovery from a target-centric focus to one in which pathway-driven functional responses are the object of equal if not paramount interest. 61 Reflecting this shift is an increasing emphasis on cell-based assays in which the objective is to identify compounds that disrupt, augment, or restore a cellular phenotype such as output from a targeted signal transduction pathway or expression of a physiological profile. Cell-based phenotypic screening is attractive because it allows interrogation of entire signaling pathway networks. Even though the molecular basis of compound activity may be poorly understood, the expression of such activity in a cellular context is evidence of physiological relevance not available in a cell-free biochemical assay. Cell-based assays are also more likely to detect undesirable or toxic side effects of compounds, another impetus for their inclusion at early stages of the drug discovery pipeline. A complexity of cell-based phenotypic screening is that one or more molecules may be targeted by active compounds, so that follow-up experiments are required to elucidate molecular mechanisms of action. On the other hand, because the molecular targets are unknown, this screening approach avoids the potential pitfall of poor target validation that has been advanced as a contributing cause of late-stage clinical failures of drugs developed using target-centric screening approaches.62–65 Limited assumptions about target identity decrease the probability of committing resources to an invalid target.
There has been much recent debate about the relative efficacy of target-based and phenotypic drug discovery approaches in discovery of first-in-class small-molecule drugs approved by the Food and Drug Administration. Although evidence for superior success rates of phenotypic screening approaches has been documented for the period of 1999 to 2008,66,67 a more restricted definition of phenotypic screening as entirely target agnostic together with data from a more extended time period (1999–2013) have been used and interpreted to suggest otherwise. 68 Because a case can be made that many phenotypically driven screening projects are not strictly target agnostic, perhaps a more useful concept that has recently been advanced is that of “mechanism-informed phenotypic drug discovery.” 69 In this approach, phenotypic assays incorporate knowledge about molecular pathways or processes that are implicated in clinically relevant responses irrespective of which molecular component(s) may be targeted. The guiding design objective of successful in vitro phenotypic assays is to capture key aspects of the relevant in vivo biology.
One approach for achieving high-throughput phenotypic screening uses conventional flow cytometry in combination with complex fluorescent cell-barcoding schemes55,70 ( Fig. 2D ). For example, by using three fluorescent dyes (four different concentrations of two dyes and six of the third), a unique fluorescence signature can be conferred upon cells in each well of a 96-well plate. At the end of an assay, cells from all wells can be pooled together and analyzed in 5 to 7 min as a single sample using three fluorescence channels to decode source well identity and additional fluorescence channels for purposes such as phosphoprotein pathway analysis (Phosflow) or cell type identification. 55 Pathway profiling analyses of very high content have been achieved with variations of this approach, with as many as 27 different cell type–pathway combinations assessed per test compound in one example. 70 In these types of experiments, fluorescent barcoding is performed after the assay endpoint, with barcoding dyes applied in the wells to fixed and permeabilized cells. In general, this approach trades the number of compounds tested with the number of parameters assessed.
Simpler multiplexing approaches that minimize requirements for cell numbers, reagents, sample volumes, and time-consuming assay preparation steps are preferred when working with larger compound libraries of tens to hundreds of thousands of compounds (see the “Balancing Throughput and Content” section below). They can have a significant cost benefit impact irrespective of library size (for example, by minimizing the amounts of compound consumed). Exemplifying such an approach was a screening project designed to discover small molecules targeting target of rapamycin (TOR) proteins,71,72 Ser/Thr protein kinases phylogenetically conserved from yeast to human, which are fundamental controllers of cell growth.73–75 The impetus for this project was a need for new TOR inhibitors to improve on the moderate clinical benefit of rapamycin in mTOR-based therapy of many cancers. Five green fluorescent protein (GFP)–tagged yeast clones representing the readouts of four branches of the TORC1 signaling pathway were first selected by screening the Yeast GFP Clone Collection 76 (Life Technologies) for clones with high responsiveness to rapamycin. The five clones were barcoded with two dyes that had fluorescence emission spectra distinct from GFP. Importantly, barcoding was performed on separate bulk preparations of live cells, which were then combined and distributed into wells of 384-well plates so that all five clones were present in each well during exposure to compounds for induction of GFP response and subsequent analysis ( Fig. 2E ). In a primary screen of ~320,000 compounds from the MLSMR, multiplexed analysis of the five clones allowed evaluation of compound activity on the four pathway branches simultaneously.71,72 This fostered rapid prioritization of molecules that functionally mimicked rapamycin as well as molecules selective for individual branches that could target effectors in the TORC1 pathway or interfere with other non-TOR, cross-talk signaling mechanisms. It is noteworthy that both the Phosflow and TOR pathway screening approaches illustrated above represent examples of mechanism-informed phenotypic screening.
Numerous cell-based phenotypic screening projects have been performed in recent years, most using HyperCyt platform technology ( Table 1 ). Reporters used in such assays have included endogenously expressed fluorescent proteins in bacteria,77,78 yeast,71,72,79,80 and acute myeloid leukemia cells 81 ; fluorescent antibodies (homogeneous, no-wash format) to detect surface proteins in primary murine T cells10,82 and human cytotoxic T cell29,30,83 and myeloid81,84 cell lines; and fluorescent substrates used to monitor activity of cell membrane efflux pump transporters in fungal cells 85 as well as human cell lines.86–88 A group at the National Center for Advancing Translational Sciences recently validated the HyperCyt platform for use in their dose-response–based method of screening, called quantitative high-throughput screening (qHTS), using a multiplexed apoptosis assay with a human lymphoma cell line. 89 Also, a new class of biosensor has been recently employed that enables flow cytometry for detection of receptor transit from cell surface to cell interior. 90 This reporter system is composed of a fluorogen-activating protein (FAP) and a small molecule (fluorogen) whose fluorescence increases dramatically when noncovalently bound by the FAP. 91 Cells that expressed an external N-terminal FAP fused to the β2-adrenergic receptor (β2AR) were screened against ~350,000 compounds from the MLSMR to detect novel agonists. 92 The assay used a membrane-impermeant fluorogen to detect loss of signal resulting from agonist-induced β2AR internalization. A group at Pfizer has recently implemented FAP technology for a GPCR project using integrated HyperCyt flow cytometry and imaging platforms. 93 It is anticipated that this reporter system will be a useful tool for identifying ligands for orphan GPCRs.
Examples of Cell-Based Phenotypic Screening of Small-Molecule Libraries with Flow Cytometry.
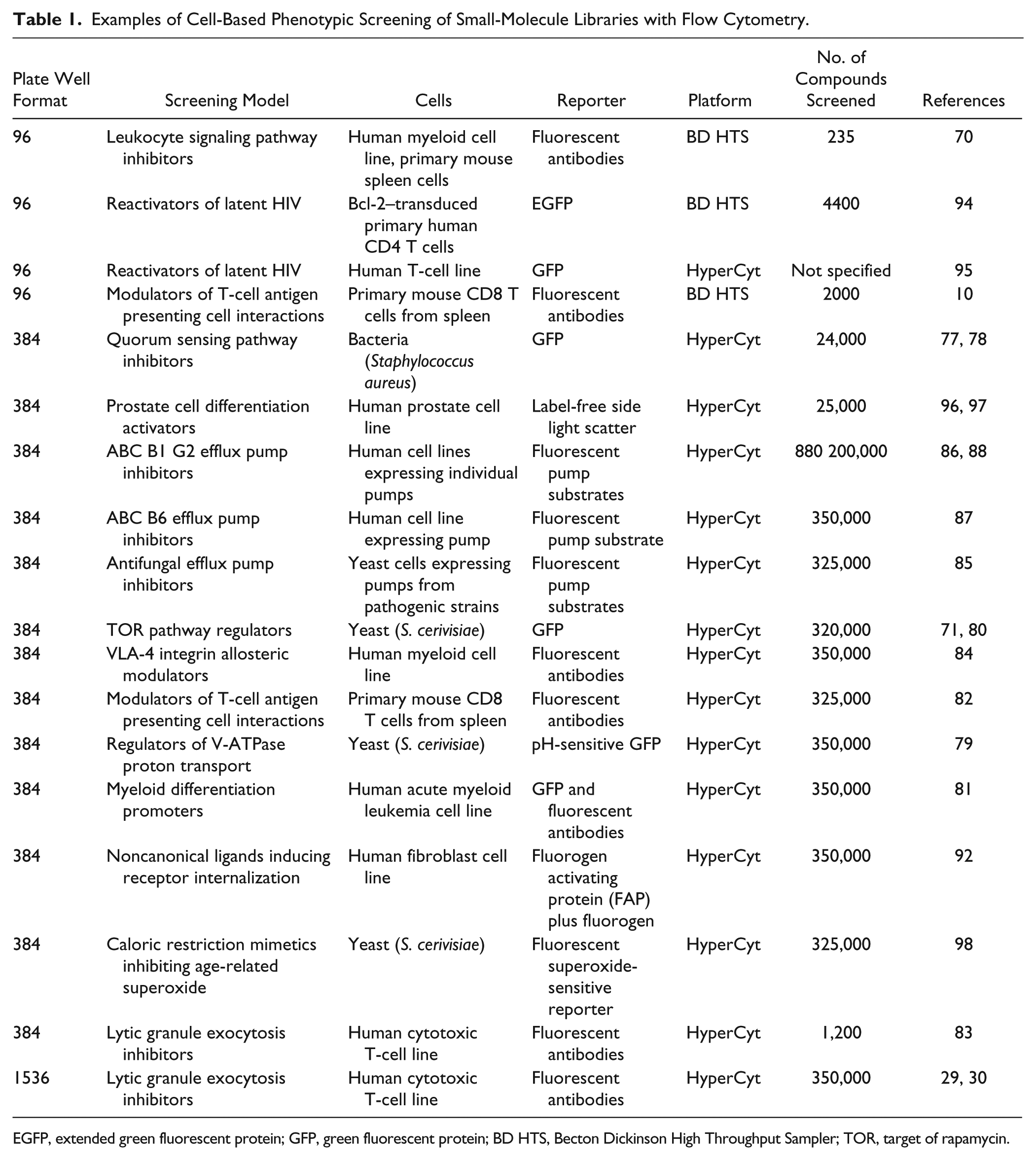
EGFP, extended green fluorescent protein; GFP, green fluorescent protein; BD HTS, Becton Dickinson High Throughput Sampler; TOR, target of rapamycin.
Unique Value Added and Impact on Early Drug Discovery Progress by High-Throughput Flow Cytometry
Almost all the examples of phenotypic screening listed above involved analysis of suspension adapted cells, predominantly mammalian leukocytes but also including a diversity of single-cell organisms used as experimental models. Automated high-content imaging (HCI) is an established, alternative single-cell analysis technology common in many screening laboratories, where it has been the workhorse technology for phenotypic screening for many years. HCI is a planar technology that is well suited for analysis of cells with adherent morphology but requires potentially assay-compromising cell immobilization procedures to accommodate analysis of cells that do not naturally attach to surfaces. This has led to an apparent bias in the types of cells subject to HCI analysis, a trend highlighted in a recent compendium of research articles and reviews focused on phenotypic screening (December 2013 and January 2014 issues of Journal of Biomolecular Screening). In 18 articles that featured HCI screening approaches,99–116 suspension adapted cells were notably absent. Surface adherent cells were the focus of 16 of the articles. The other two described HCI analysis of multicellular zebrafish embryos 101 and 3D colony formation by carcinoma cells immobilized in soft agar. 106 Thus, high-throughput flow cytometry complements HCI by offering a facile approach for early phase, single-cell phenotypic screening of otherwise underrepresented cellular phenotypes. Other HTS technologies that rely on bulk fluorescence measurements of cells in suspension are incapable of the high degree of multiplexing used in many of the flow cytometry screening applications referenced above. Such multiplexing can affect early drug discovery by enabling identification of lead candidates that simultaneously satisfy multiple important phenotypic criteria at the stage when the greatest number of candidates are being evaluated. Just as in target-based screening, this minimizes the risk that an inappropriately prioritized single criterion might lead to early rejection of promising candidates. The benefits of such multiplexing were highlighted above for target-based projects to identify antibodies with cross-species reactivity ( Fig. 2C ) and a phenotypic screening project to identify “rapalogs,” compounds that mimic rapamycin with respect to their parallel impact on four different mTOR-associated pathways ( Fig. 2E ). Thus, high-throughput flow cytometry offers a capability for highly multiplexed, single-cell analysis of suspension adapted cells, an important approach to cell-based screening that appears to be inadequately addressed by other screening technologies in current use for early-phase drug discovery.
Multiplexing Squared with Combinatorial, Mixture-Based Libraries
Phosflow flow cytometry, with the ability to characterize elements of multiple signaling pathways in parallel, represents a powerful and potentially lucrative approach for pathway-focused phenotypic screening. 70 As currently implemented, the assay protocol requires multiple wash steps, a bottleneck for automation processes compatible with screening of sizable compound libraries. Other types of useful phenotypic screens may, for example, involve parallel analysis of complex collections of cell subsets. To obtain statistically meaningful numbers of cells from each subset, especially when one or more is low in frequency, the sample acquisition time may need to be significantly extended to enable analysis of an appropriately large number of cells. Assay formats such as these and others may be practically best suited for screening of smaller, focused libraries in the range of 10,000 compounds or fewer. 70 An alternative approach would be to use mixture-based libraries in which cells are screened against a series of mixtures, each containing multiple compounds, rather than one compound at a time.
The utility of such an approach was recently demonstrated by a screening project 117 to identify small-molecule ligands that selectively bound two members of the formylpeptide receptor family, FPR1 and FPR2, GPCRs associated with host defense and inflammation. 118 The screen used a target-based assay in cells in which compounds were assessed for the ability to displace a green fluorescent ligand from FPR1 or FPR2, each expressed on a separate set of red fluorescently barcoded cells. The compound collection was constructed at the Torrey Pines Institute for Molecular Studies by synthetic, combinatorial methods119,120 and contained more than 5 million small molecules representing 37 unique chemical scaffolds. These were arrayed as mixtures in 5261 wells with 48 to 216,000 compounds in each mixture. Figure 3 illustrates an example of one of the chemical scaffolds ( Fig. 3A ), the mixture-based library generated from it ( Fig. 3B ), and results of screening it for activity against FPR1 ( Fig. 3C ). A first round of screening that identified several active scaffolds was followed by three rounds of confirmatory screening to confirm and rank activity in several hundred wells representing two of the most active scaffolds. At that point, positional scanning deconvolution of the activity profile identified 106 candidate molecules from one scaffold library and 8 from the other. When purified and tested, 15 and 12 of these 114 compounds had inhibitory constants (Ki) of <100 nM for FPR1 and FPR2, respectively, all with >100-fold selectivity for the targeted receptor. This result, obtained by screening of fewer than 10,000 total wells, was a remarkable improvement over two previous projects using the same assay in which screening of ~5000 small molecules from two focused libraries54,121 and of ~24,000 from a diversity library 122 yielded only one compound of similar potency and selectivity. Selective antagonists and agonists for both receptors were represented, two (an FPR1 agonist and FPR2 antagonist) with unprecedented potencies. 117
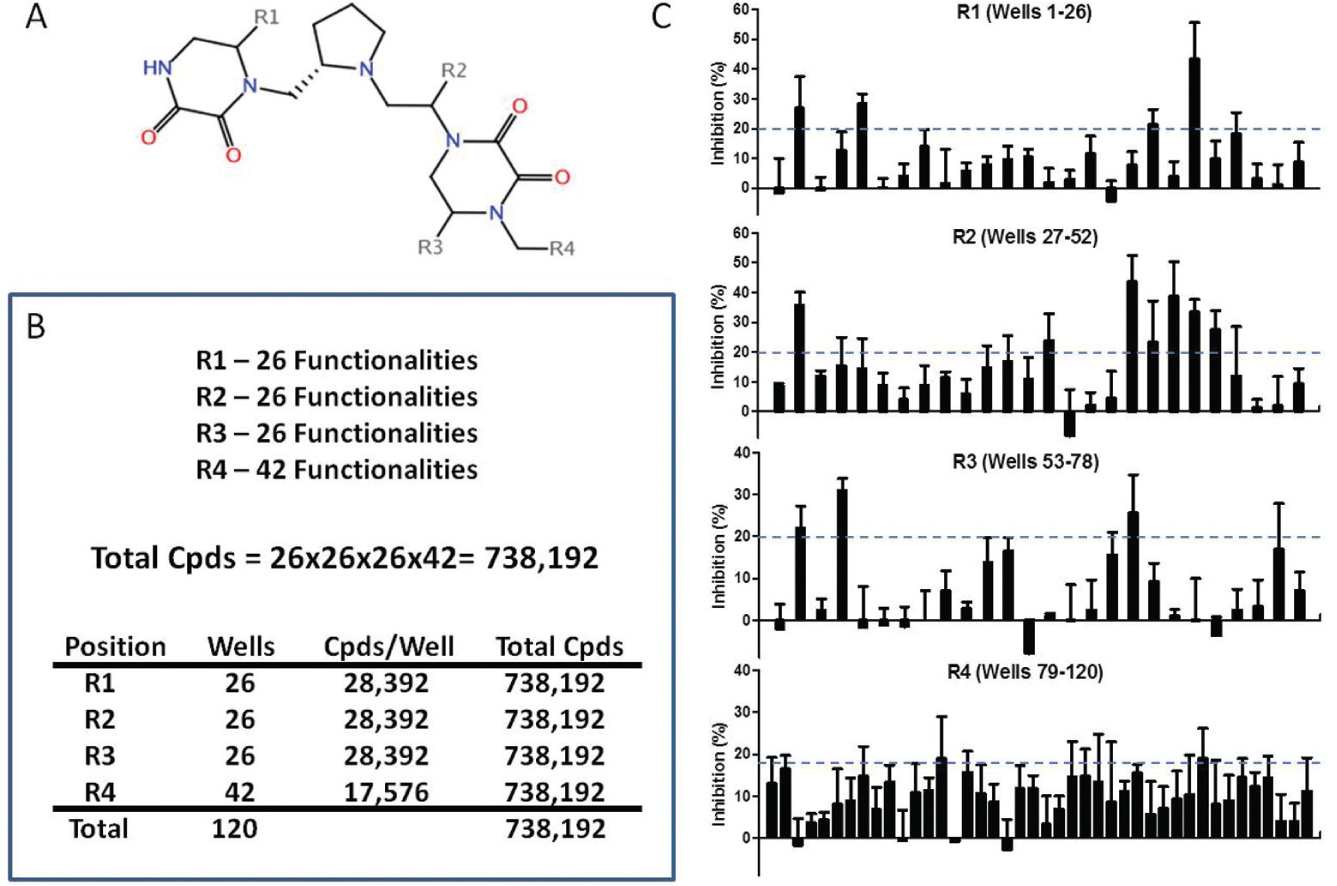
Screening with combinatorial, mixture-based small-molecule libraries. (
Evolution of HyperCyt Platform
HyperCyt sampling technology has been available as a commercial product since 2006. Early adopters of the platform were companies that had already targeted flow cytometry as a preferred approach for accomplishing goals such as small interfering RNA–based functional genomics screens of plasmacytoma cells, hybridoma screens for therapeutic antibodies, and screens of immune cells from primary tissues. Attractive to these companies was not only the faster sample processing speed but also the ability to process smaller sample volumes and to avoid loss of cells as dead volume in sample back-flushing operations. For example, a group at GSK had for many years been using an automated system with conventional flow cytometry sampling for drug concentration profiling of target-based assays in 96-well plates, a task requiring an hour or more for each plate depending on the assay. Implementation of HyperCyt technology enabled screening of 10 µL samples of whole blood arrayed in a 96-well plate in less than 10 min. 9 This would have been impractical if not impossible with the original system. More than 200 HyperCyt platforms are currently in laboratories worldwide, but only recently are they being placed with increasing frequency in mainstream screening labs. This is likely attributable to capabilities associated with evolution of the platform as well as recent trends in strategies for drug discovery.
Hardware
Initially, the platform was available as an autosampler that could be used only as a front end for a variety of commercial flow cytometers. Although this was (and still is) an attractive approach for expanding capabilities of an existing flow cytometer, most flow cytometer manufacturers were reluctant to provide a nonproprietary automation interface to coordinate operation with external devices from other instrument vendors. System installation often required that holes be drilled in the flow cytometer casing or that built-in mechanical and electronic control systems be bypassed to enable a working physical interface. Such installation issues raised concerns about invalidating instrument warranties. Also, two computers were required to operate in parallel, one to run the flow cytometer and the other to run the autosampler, and the handshake between the two was sometimes awkward in its implementation. These concerns were first addressed by creation of the HTFC platform option with its dedicated flow cytometer and fully integrated operation of flow cytometer and autosampler. More popular has been the recently introduced, next-generation iQue Screener System, a fully integrated solution in which sampling and reader functions are combined in a single benchtop unit, controlled by a single operating system, and provided with an automation interface to facilitate integration into screening laboratory work flows. An advanced HD option enables the platform for analysis of assays in 1536-well format. As with the HTFC platform, it is a turnkey system that requires little user intervention or esoteric knowledge to operate.
Software
Early versions of software for analyzing flow cytometry data were adequate for basic needs of plate-based analysis but lacked some of the more specialized features to which practitioners of flow cytometry had become accustomed. Users would feel compelled to export the data from each well to a separate file for analysis with their favorite third-party analysis package, a procedure that defeated advantages of data organization, manipulation, and storage efficiency associated with one-plate, one-file methodology. The current ForeCyt control and analysis software represents a significant advance that provides most of the capabilities expected of modern flow cytometry analysis packages. It also enables a range of sophisticated multiwell data operations such as custom intraplate data normalization capabilities and dose-response plots with curve-fit statistics. Particularly useful is a PlateView function that displays histograms, 2D plots, or overlay plots for each well in a consolidated view of all plate wells or selected well groups. Together with heat map display functions that profile assay response data over multiwell regions and well-scanning features for selectively viewing data in individual wells, there is remarkable built-in versatility to streamline hit detection as well as identify anomalous response distribution patterns (e.g., edge effects) that can skew screening data. Further bolstering the utility of flow cytometry HTS is software developed by third parties to accommodate analysis of plate-centric flow cytometry data files.123,124
Drug Discovery Trends
Two recent trends in drug discovery are also likely to have benefitted high-throughput flow cytometry platforms. One is the ongoing shift in emphasis from target-centric screening to target-agnostic or mechanism-informed phenotypic screening approaches discussed above.61,66–69 The other is the increasing interest and investment of major pharma companies in developing biologic drugs, especially novel antibodies. 125 As discussed above, automated HCI has been the predominant workhorse technology for phenotypic screening for many years, but its use has been heavily biased toward analysis of cells with adherent morphology. With the advent of high-throughput capabilities, flow cytometry is achieving recognition as a complementary technology ideally adapted for high-content phenotypic analysis of suspension-adapted cells such as leukemias and lymphomas as well as leukocytes responsible for immune system regulation and inflammation-associated disease processes. Also, in contrast to imaging-based systems, the amount of time required for flow cytometry data acquisition is largely insensitive to the number of optical parameters that are detected. The advantages of high-throughput flow cytometry for multiplexed screening of antibody-producing cells have been outlined above in the section on target-based screening. An alternative technology for this is Mirrorball, a laser scanning instrument that has been recently introduced to replace the aging fluorimetric microvolume assay technology (FMAT) platform, for many years an established platform for antibody screening with whole cells.59,126 The Mirrorball platform can detect up to five colors of fluorescence simultaneously, along with a single light scatter parameter from beads but not from cells (http://ttplabtech.com/cell-imaging/mirrorball/). It is more versatile than FMAT for analysis of intact cells but relies like HCI platforms on cells immobilized and displayed in a planar format at the bottom of wells. Thus, flow cytometry offers unique advantages for addressing both emerging drug discovery trends, on one hand by enabling multiplexed, phenotype-driven primary screening of suspension cells and on the other by providing more efficient and versatile approaches for screening cellular sources of biological drugs such as antibodies.
Impact on Drug Discovery
Recent advances in flow cytometry are likely to affect drug discovery now and in coming years in several ways. One area of impact is the emerging appreciation of how sample-sparing approaches such as HyperCyt may make HTS of primary tissues more accessible. Advances in single-cell throughput are also on the near horizon that promise to significantly reduce sample analysis times when massive numbers of cells must be scrutinized for access to rare cell subsets. The number and types of parameters that can be measured in parallel on single cells are also on the increase to bolster systems approaches to drug discovery.
Screening of Primary Cells
Primary cells have been widely used in screening laboratories, predominantly in throughput-limited secondary assessments such as concentration-response, metabolism, and toxicity profiling of compounds.9,127 The attraction of primary cells is the premise that they more closely resemble cells in vivo than immortalized cell lines with respect to endogenous protein levels and the physiology of signaling pathways and receptors. Also, because many types of primary cells exhibit minimal proliferation unless stimulated to do so, off-target effects of compounds on cell division are less likely than with cell lines to obfuscate screening results. 127 Because most high-throughput technologies require large numbers of cells, a major feature limiting primary cell use in high-throughput compound screening is low abundance. Another confounding factor is the innate heterogeneity of primary cell cultures. This provides an element of uncertainty as to which subset of cells within a culture might be the basis of observed compound activity. The multiparameter analysis capabilities of flow cytometry are well suited for discriminating cell subsets in a sample to resolve differential targeting by compounds. 55 Sample-sparing characteristics of HyperCyt sampling technology allow efficient quantitative analysis of as few as hundreds to thousands of cells in a well with minimal loss to waste. Thus, high-throughput flow cytometry may be of immediate use for intermediate- to large-scale primary cell screening applications using suspension cells from animal model systems as well as humans. Such considerations are the basis of a precision medicine concept currently in development at UNMCMD. 24 Primary tissues from leukemia patients are to be screened for responsiveness in vitro to both old and new oncology drugs and their combinations. Single-cell analysis capabilities of flow cytometry are ideal for assessing the degree to which heterogeneity of the original tumor is retained in such an in vitro therapy model and how different subpopulations of cancer cells are affected by the treatment regimen. This effort, an expansion on an approach reported by a group in Finland, 128 is expected to lead ultimately to an understanding of the basis of durable responses and translation to clinical therapy. A recently introduced alternative to using primary cells is a “disease-in-a-dish” approach, in which a patient’s somatic cells can be reprogrammed into an embryonic pluripotent state by the forced expression of a defined set of transcription factors. The resulting induced pluripotent stem cells recapitulate the disease phenotype and can be expanded for in vitro disease modeling and drug discovery.129,130
Increasing Volumetric and Single-Cell Throughput to Assess Low-Frequency Subpopulations
There is great interest in improving methods for screening of rare cells such as circulating tumor cells (CTCs), cancer stem cells implicated in metastases and acquisition of resistance to therapy.130–134 Current procedures for detection and analysis of CTCs, at estimated levels of 100 or fewer per milliliter of blood, 135 involve capture or enrichment procedures that are lengthy and conducive to cell loss.136,137 How much better it would be if all cells in a blood sample could be efficiently examined to detect and potentially isolate rare cells such as CTCs without such preliminary manipulations. A similar argument can be made for other rare cell populations such as fetal cells in maternal blood or endothelial progenitor cells. At current maximal analysis rates of modern flow cytometers, it would require hours to analyze all cells in 1 mL of whole blood. A cluster of flow cytometers might be used in parallel to divide the workload and speed the process, as has been previously done with four flow cytometers to enhance processing of 1536-well plates. 25 However, the number of instruments required would greatly exceed anything remotely practical with respect to cost, space requirements, and complexity of operation. An alternative approach has been to implement chip-based systems whereby tens to potentially as many as hundreds of tightly focused, parallel streams of cells can be generated in the presence of an appropriately modulated acoustic field138,139 ( Fig. 4A ). Acoustic focusing techniques such as this can position biological cells and other particles over a wide range of sizes and support high volumetric flow rates for flow cytometry.140,141 Creation of simple methods to analyze the multiple flow streams will be critical and are currently in early stages of development. With the potential to analyze millions of cells per second, such methods, in combination with high-throughput sampling systems, promise to bring efficient screening of low-frequency cells in heterogeneous cell populations closer to reality.
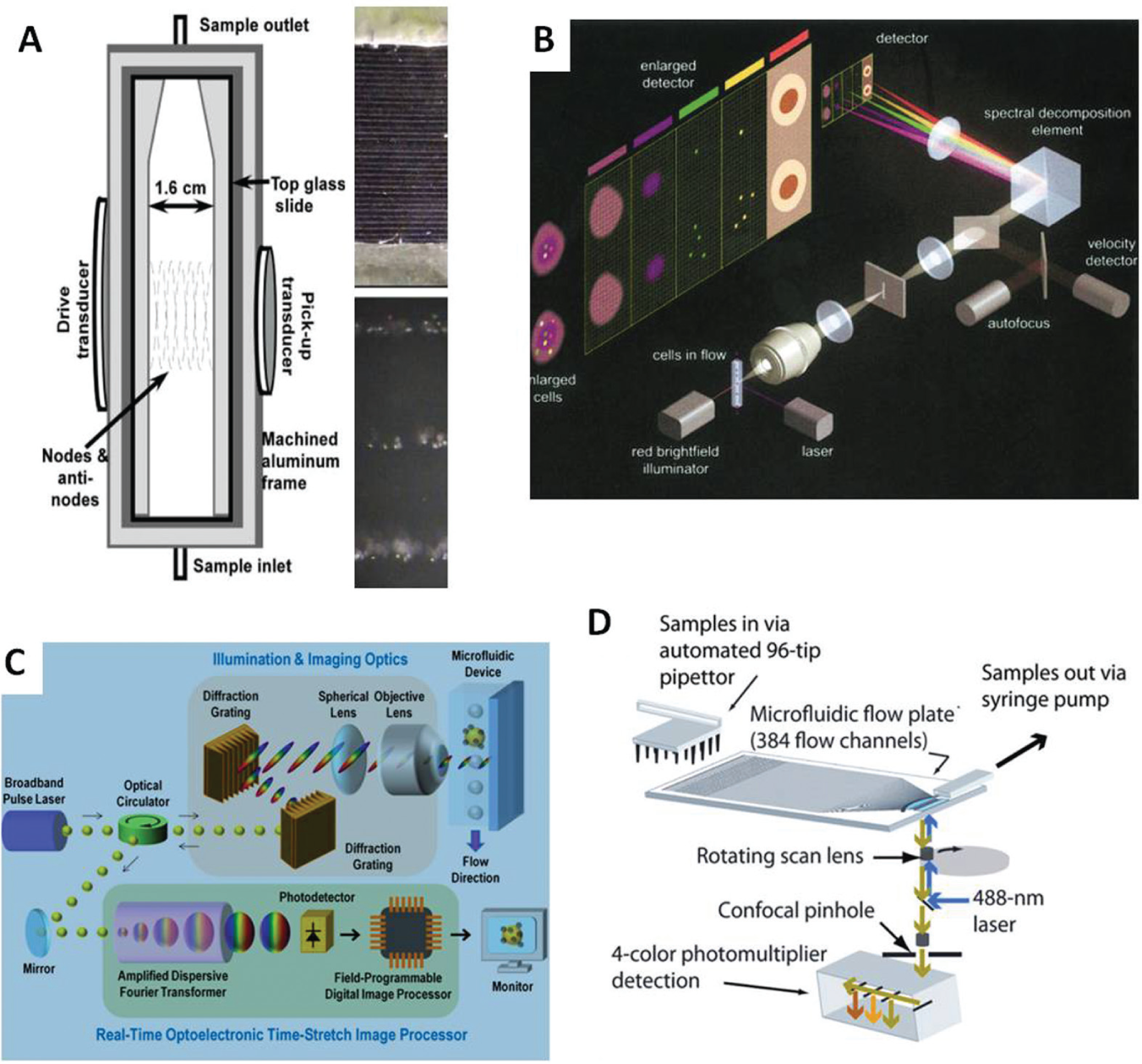
Technologies for rare cell detection and imaging in flow-based systems. (
Expanding the Content of Suspension Cell and Particle Analysis
Flow-based imaging systems are commercially available that can record images of hydrodynamically focused cells in suspension at rates as high as 5000 cells/s. Simultaneous image modes include brightfield (transmitted light), darkfield (side scatter), and four channels of fluorescence.142,143 These systems offer the advantage of spatial resolution in addition to optical parameters available in conventional flow cytometers ( Fig. 4B ). Recently documented has been an even faster imaging flow cytometry platform capable of imaging rates as high as 100,000 cells/s. 144 This platform uses an ultrafast optical imaging modality known as serial time-encoded amplified microscopy 145 for blur-free imaging of particles in high-speed flow ( Fig. 4C ). It also uses hybrid optoelectronic image-processing circuitry to enable fully automated real-time image recording and classification of a large number of cells or particles through their morphological and biochemical features. A different flow-based imaging approach, embodied in an instrument called the parallel microfluidic cytometer (PMC), uses a high-speed scanning photomultiplier-based detector to combine low-pixel-count 1D imaging with flow cytometry 146 ( Fig. 4D ). A confocal scanner records fluorescence values of cells at 1 µm intervals as they cross the detection window, and up to 384 separate streams of cells can be measured in parallel. At up several thousand cells per second, six-pixel 1D images captured by the PMC were adequate to quantify protein localization in a yeast model and nuclear translocation in Chinese hamster ovary cells. 146 A wide range of microfluidics-based systems have been integrated with flow cytometry technology, as recently reviewed in some detail. 139
New instrument and reporter technologies are also increasing the numbers of cell-associated parameters that can be measured in parallel. One approach uses surface-enhanced Raman scattering (SERS) from metal nanoparticles to produce signal intensities that rival fluorescence but with narrower spectral features that allow a greater degree of multiplexing. 147 Raman resonant compounds are adsorbed on the nanoparticles to confer a unique spectral fingerprint on each SERS tag. Tags are then encapsulated in a polymer coating for conjugation to antibodies or other targeting molecules. SERS spectral measurements are made with a high-resolution spectral flow cytometer that is also capable of making conventional flow cytometry measurements on thousands of individual cells per minute. The narrow spectral features of the SERS signal enable more distinct probes to be measured in a smaller region of the optical spectrum with a single laser and detector, allowing for higher levels of multiplexing and multiparameter analysis.
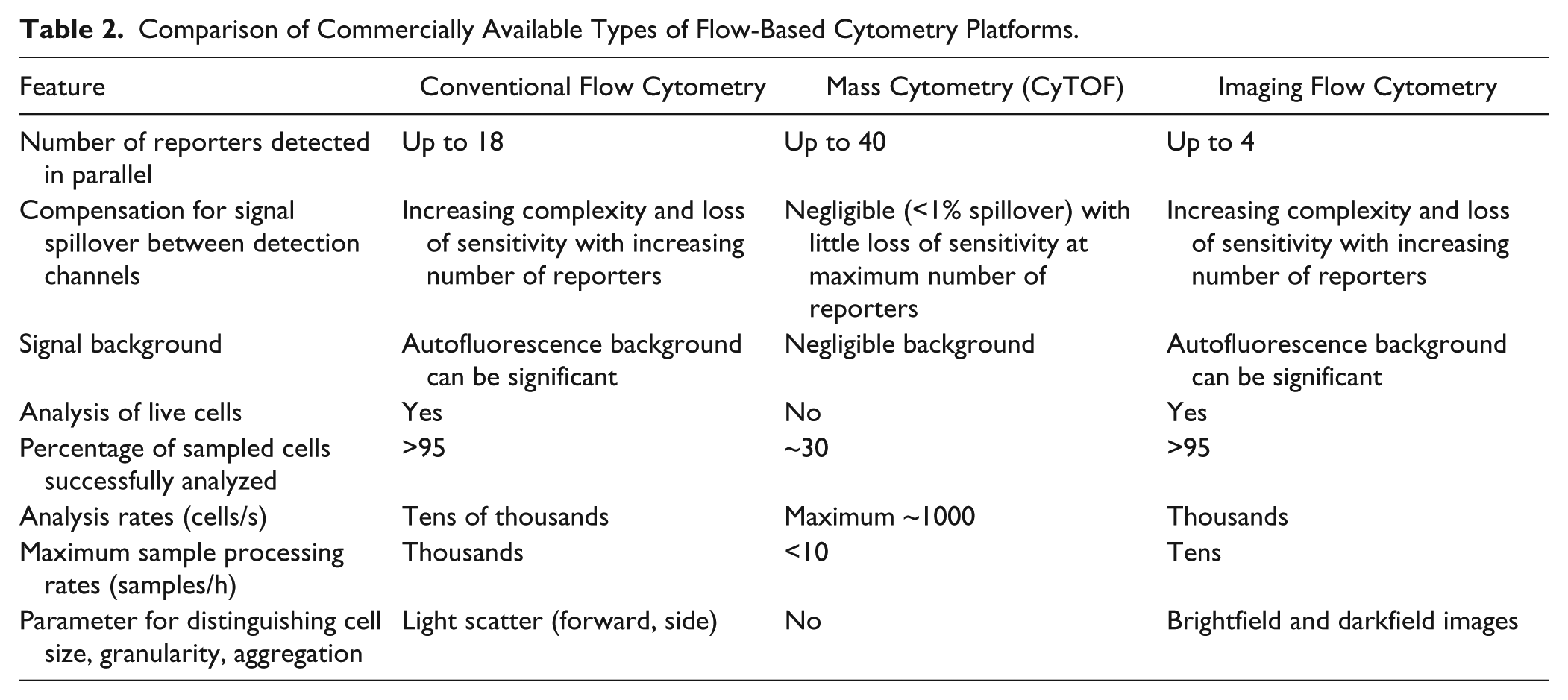
Another rapidly evolving high-content technology is mass cytometry, a mating of flow cytometry and mass spectrometry that, even its current state of infancy, is revolutionizing single-cell profiling capabilities. Using stable isotopes of rare earth metals as reporters rather than fluorophores, mass cytometry is currently capable of multiplexing up to 40 cellular measurements. Importantly, it does so with little measurable background signal and negligible (~0.1%) signal spillover between measurement channels, eliminating the need for interchannel signal compensation that can add significant complexity to highly multiplexed fluorescence measurements. 148 Just as in fluorescence-based flow cytometry, cell barcoding can be performed to enable assessment of the signaling state of cells at one or more time points or pooling of cells from multiple wells to speed analysis of multiple samples. At present, mass cytometry is limited to analyzing only dead (fixed) cells at analysis rates of ~1000 cells/s, and only ~30% of the cells in a sample are analyzed because of sampling inefficiencies.148,149 Nonetheless, recent profiling studies of hematopoietic cells, 150 virus-specific T cells, 151 and NK cells, 152 each involving simultaneous measurements of up to 37 features, have led to unprecedented understanding of these cellular biological systems. Mass cytometry was developed by a group in Toronto in 2009, 153 and the commercial product, called CyTOF, for cytometry by time of flight (www.dvssciences.com), is likely to improve over time with respect to sensitivity, cell analysis rates, sampling efficiencies, and even greater numbers of simultaneously measurable parameters. Its primary role in drug discovery is anticipated to be in deep profiling studies to elucidate compound mechanism of action and off-target effects. Complementing this can be the role of fluorescence-based flow cytometry in studies of living cells and targeted screening of compound libraries. 149 Table 2 summarizes a number of the salient features of the CyTOF platform as compared with conventional and imaging flow cytometry platforms that are currently available from commercial sources.
Comparison of Commercially Available Types of Flow-Based Cytometry Platforms.
Improving High-Throughput Flow Cytometry
Although some of the above technologies are unlikely to be applicable to high-throughput flow cytometry platforms such as HyperCyt in the near future (e.g., mass cytometry), there are others that may be of more imminent promise for improvement. Key areas of the most immediate potential impact on early phase drug discovery include (1) increased single-cell and sample throughput, (2) expanded range of particle sizes amenable to analysis, (3) ability to analyze cells with unperturbed adherent morphology, and (4) addition of high-throughput imaging capability. Many of these areas of improvement have been demonstrated or are in early proof-of-principle stages in conjunction with chip-based systems described above that are in current development.138,139,146 For example, acoustic focusing methods represent one approach by which multiple, tightly focused parallel streams of cells can be generated from one or more input sources (e.g., microplate wells) to dramatically increase throughput of cells, samples, or both depending on platform configuration. Acoustic focusing methods are also applicable to particles of larger size than conventional hydrodynamic focusing methods. For example, multicell spheroids of 100 µm or more in diameter can be aligned in a focal plane for optical analysis without the danger of clogging encountered in conventional flow cytometers (unpublished results). The ability to analyze spheroids or cells attached to large-diameter microspheres at hundreds to thousands per second would represent a novel capability for facile flow cytometry analysis of cells with unperturbed adherent morphology. Cells growing in a 3D spheroid configuration would be of particular interest because of their better representation of in vivo cell physiology as compared with cells grown in monolayers.154,155 Systems by which to achieve simultaneous optical analysis of multiple parallel streams have been demonstrated (see ref. 146, and manuscript in preparation). High-speed camera technologies and imaging methods are available 146 and rapidly advancing to allow ever improved ability to capture images of cells in parallel streams.
Balancing Throughput and Content
As discussed above, the opportunities for cytometric analysis range from high-throughput to multiparameter or high-content analysis and/or rare event analysis. In general, there is a tradeoff in sampling rates and data complexity that can involve not only sample preparation or analysis times but also the costs per well. Although there is not a single answer to optimizing workflow in a discovery campaign, one should take into account the number of samples to be tested, the number of parameters to be evaluated, and the number of events in the target population. Typically, a primary screen for an HTS campaign would involve a cost-effective number of parameters with a large number of compounds (upwards of tens of thousands). Although dose response is possible at the primary screening phase (qHTS), the secondary screening campaigns for active compounds would allow increasingly detailed analysis. Here, time and costs would dictate the preference for extremes of multiparametric analysis as compared with rapidly executed sequential analyses of lesser complexity.
In concert with throughput considerations is a misperception that persists even among many flow cytometry practitioners that an excessively large number of cells need be evaluated by a flow cytometer to obtain statistically valid results (10,000 is the number most frequently espoused). In reality, the relationship between cell-counting statistics and data quality is no different for flow cytometry than for any other single-cell analysis technology. High-quality flow cytometry screening results have frequently been obtained with as few as 100 to 200 cells analyzed per well, although higher numbers are preferred when practical. For HTS, the flow cytometer typically samples only a fraction of the cells in a well, with the objective being to achieve maximum processing speed by sampling the minimum volume required for evaluating a statistically adequate number of cells. With HyperCyt, it is also possible to aspirate and analyze the entire volume contained in a well. In this way, a few hundreds of cells in a well could all be analyzed with a sampling time that is a function of total volume (i.e., cell concentration). Thus, tradeoffs between sampling time and cell concentration apply in a similar way to flow cytometry as to other screening technologies such as HCI.
The advent of HyperCyt sampling technology has moved flow cytometry from a platform restricted to secondary, low-throughput drug-profiling applications to one suitable for primary screening applications in which tens of thousands of compounds are analyzed per day. The high multiplexing capabilities of flow cytometry make it possible to analyze multiple targets in parallel without appreciably affecting sample analysis time. Together, these capabilities facilitate high-content compound profiling at the earliest screening stages, streamlining compound prioritization with respect to features such as target specificity and off-target effects. The ability to sample small volumes with negligible waste reduces reagent costs and compound usage and makes quantity-limited sources of cells more accessible to testing against a broad array of compounds. Improved combinatorial approaches to formatting of compound libraries promise to further extend opportunities for flow cytometry–based primary screening when samples are scarce or when sample throughput is limited. Recent advances in flow-based imaging and mass spectrometry have dramatically expanded the numbers and types of measurements that can be made simultaneously on single cells. Such technological innovations offer opportunities for pathway-based compound-profiling applications of unprecedented depth and complexity to help usher in systems-level approaches to drug discovery. Novel microfluidics-based approaches for parallel sample processing and analysis promise to boost single-cell analysis rates by orders of magnitude. A resulting improved ability to efficiently detect and characterize low-frequency cells such as CTCs in heterogeneous cell mixtures will likely be of most immediate utility for drug efficacy evaluation in animal models and clinical trials as well as precision medicine applications. Flow cytometry will inevitably continue to play an important role in drug discovery. Although recent trends are promising, it remains to be seen how successfully new high-throughput flow cytometry platforms will contend with established and emerging instrument technologies for space on the HTS lab bench.
Footnotes
Acknowledgements
Supporting data for this article were generated in the University of New Mexico Center for Molecular Discovery and the UNM Cancer Center Flow Cytometry and High-Throughput Screening Shared Resource.
Declaration of Conflicting Interests
The authors hold patents for HyperCyt technology and are co-founders of IntelliCyt, a company that manufactures and sells instruments incorporating HyperCyt technology. No financial support was received from any commercial entity for production of this work.
Funding
This work was suported in part by NIH grants R01HG005066 (BSE), U54MH084690 (LAS), U54 MH074425 (LAS) and CCSG P30CA118100.
