Abstract
Reactivation of genes normally expressed during organogenesis is a characteristic of kidney regeneration. Enhancing this reactivation could potentially be a therapeutic target to augment kidney regeneration. The inductive events that drive kidney organogenesis in zebrafish are similar to the initial steps in mammalian kidney organogenesis. Therefore, quantifying embryonic signals that drive zebrafish kidney development is an attractive strategy for the discovery of potential novel therapeutic modalities that accelerate kidney regeneration. The Lim1 homeobox protein, Lhx1, is a marker of kidney development that is also expressed in the regenerating kidneys after injury. Using a fluorescent Lhx1a-EGFP transgene whose phenotype faithfully recapitulates that of the endogenous protein, we developed a high-content assay for Lhx1a-EGFP expression in transgenic zebrafish embryos employing an artificial intelligence–based image analysis method termed cognition network technology (CNT). Implementation of the CNT assay on high-content readers enabled automated real-time in vivo time-course, dose-response, and variability studies in the developing embryo. The Lhx1a assay was complemented with a kidney-specific secondary CNT assay that enables direct measurements of the embryonic renal tubule cell population. The integration of fluorescent transgenic zebrafish embryos with automated imaging and artificial intelligence–based image analysis provides an in vivo analysis system for structure-activity relationship studies and de novo discovery of novel agents that augment innate regenerative processes.
Introduction
High-throughput, in vitro screening has become the mainstay of contemporary drug discovery, but its ubiquitous use has not improved the efficacy with which new drugs are discovered. It is increasingly being argued that phenotypic assays should assume a greater role in drug discovery. 1 This view is supported by a retrospective analysis of Food and Drug Administration–approved drugs that showed phenotypic assays accounted for the majority of first-in-class new molecular entities. 2 It has long been proposed that better models are needed to improve discovery productivity.3,4 Whole multicellular organisms could provide such models because they provide relevant biological environments for drug testing, but to date, few in vivo systems have been found to be compatible with contemporary drug discovery paradigms encompassing systematic interrogation of chemical libraries. Of the multicellular organisms currently used for drug discovery, zebrafish have become a favorite because they are vertebrates, small in size, highly fecund, and genetically tractable. 5 In addition, optical transparency during development makes zebrafish embryos uniquely suited for image-based analysis and high-content screening. Over the past 5 years, our group has developed a phenotype- and platform-independent methodology for automated imaging of transgenic zebrafish embryos 6 that has proven useful in the discovery and characterization of allosteric dual specific phosphatase inhibitors,7,8 fibroblast growth factor pathway activators, 9 and antiangiogenic agents.10.11 In this report, we present the development of high-content imaging assays for zebrafish kidney progenitor cell expansion.
The mammalian kidney has an innate ability to regenerate after injury. Kidney regeneration is characterized by dedifferentiation of surviving renal tubular epithelial cells into progenitor cells, associated with reactivation of genes normally required during organogenesis.12,13 Agents that enhance this reactivation and increase progenitor cells could represent novel therapeutic modalities to accelerate the rate of recovery or reduce postinjury fibrosis. One of the genes reactivated during regeneration is the Lim homeobox protein, Lhx1 (formerly Lim1), which is essential during embryonic development for establishing the kidney field. 14 Using a fluorescent Lhx1a transgene, 15 we developed a quantitative high-content screening assay for agents that hyperactivate Lhx1a expression during development. Detection and quantitation of the fluorescent transgene was achieved through the use of cognition network technology (CNT), an object-based analysis method that is superior to traditional image analysis methods in heterogeneous samples that contain spectrally similar features across multiple scales, such as tissue sections or whole multicellular organisms. 6 CNT emulates human cognitive processes in a computer environment, 16 allowing an expert user to translate visual observations into object features and relations. It is this assembly of a hierarchical network of objects and their relationships that makes CNT uniquely suited for zebrafish analysis, because it permits the detection of structures of interest within the embryo without interference from surrounding areas. We thus developed a CNT rule set that correctly detected and quantified the Lhx1a-EGFP transgene at baseline and after activating stimuli. Implementation of the assay on a confocal high-content reader enabled real-time kinetics, dose-response, and in vivo structure-activity relationship (SAR) studies. Variability studies suggested the assay can be adapted to mass matings and chemical library screening. A secondary assay paradigm was developed that encompassed in situ hybridization to confirm expansion of endogenous Lhx1a and CNT analysis of cdh17-GFP, a kidney-specific transgene that labels the entire embryonic tubular segments at later points of development. The integration of transgenic zebrafish with automated imaging created an enabling tool for the discovery and characterization of novel agents that may have the potential to augment kidney regeneration after injury.
Materials and Methods
Reagents and Chemicals
Methyl-4-(phenylthio)butanoate (UPHD25), methyl 6-(phenylsulfanyl)hexanoate (UPHD-00047), and N-hydroxy-4-(phenylsulfanyl)butanamide (UPHD-00023) have been reported in the literature17,18 and were synthesized and characterized as described in the Supplemental Methods section. 4-(Phenylsulfonyl)butyric acid (PSOBA) was obtained from Matrix Scientific (Columbia, SC). SB-431542 (4-[4-(1,3 benzodioxol-5-yl)-5-(2-pyridinyl-1H-imidazol-2-yl]benzamide) (Sigma-Aldrich, St. Louis, MO) was stored as a 100 mM stock in DMSO at 20 °C.
Zebrafish Husbandry and Transgene Generation
All procedures involving zebrafish were reviewed and approved by the University of Pittsburgh Institutional Animal Care and Use Committee. The Tg(lhx1a:EGFP)pt303 line has been described.
15
The cdh17-EGFP construct was established from a 5066 bp genomic region (chromosome: Zv9:16:29019930:29024995:1) containing the promoter, and the 5′UTR sequence of the cdh17 locus was amplified from zebrafish AB genomic DNA by PCR using the following primers: 5′-gcgc
In Situ Hybridization
Lhx1a in situ hybridization was performed essentially as described.15,21 Embryos were arrayed into 12-well plates (maximum of 25/well) and treated between 256 and 512 cell stage in 2 mL of E3 (5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl2, 0.33 mM MgSO4) containing 0.8% DMSO or 800 nM UPHD25. Drug or vehicle was removed at the 10-somite stage by washing five times with E3 media, and embryos were fixed in 4% paraformaldehyde for whole-mount in situ hybridization with DIG antisense RNA probe for lhx1a. 15 A minimum of 20 embryos per condition were visually scored for increased Lhx1a expression. Data are presented as the percentage of embryos with expanded Lhx1a.
Automated Imaging and Analysis
Sample Preparation and Imaging
Heterozygous Tg(lhx1a:-EGFP)pt303 embryos were obtained by natural crossings and kept at 28.5 °C. At 256 cell stage (2.5 h postfertilization [hpf]), embryos were arrayed in groups of 10 to 15 embryos into 12-well plates and treated with vehicle or increasing concentrations of compounds in 2 mL E3 medium containing 0.5% DMSO. Single embryos (eight per condition) were placed in the wells of a 96-well round-bottom plate (TTP Labtech, Cambridge, MA) and kept at 28.5 °C until 30% epiboly. Plates were placed in an ImageXpress Ultra high-content reader (Molecular Devices, Sunnyvale, CA). Using a 4× objective at fully open aperture, images were acquired in the DAPI and GFP channel at excitation/emission wavelengths of 350/460 and 488/525 nm, respectively. For Lhx1a time course experiments, wells were imaged every 2 h for 22 h. To determine the window during which compounds could be administered without loss of assay performance, embryos were treated with vehicle or UPHD25 (800 nM) at different stages of development (256 cell, 512 cell, 1000 cell; high, sphere, and dome) and imaged every 2 h for 22 h as described above. For cdh17-EGFP measurements, embryos from a homozygous outcross of the Tg(cdh17:EGFP)pt305 line were allowed to develop for 48 h after a single treatment with test agents, dechorionated, arrayed in flat-bottom 96-well plates, and anesthetized with 40 µg/mL MS222 (tricaine methanesulfonate; Sigma) in E3 to restrict movement during imaging. Images were acquired at a single wavelength (GFP). Because the elongated morphology of the 48 hpf embryos necessitated capture of the entire well, a MetaXpress journal was generated that acquired two sites in each well, which were stitched together into a single montage.
Image Analysis and Transgene Quantitation
Archived fluorescence micrographs were uploaded into Developer (Definiens AG, Munich, Germany) using the Cellenger module. CNT rule sets were developed that quantified Lhx1a-EGFP and Cdh17-EGFP expression within the developing embryo.
Lhx1a-GFP
Because transgene expression was highly localized, the GFP channel image did not contain enough information to permit detection of the whole embryo, as previously described.6,8 We therefore used ultraviolet (UV) autofluorescence to detect the whole embryo and to eliminate fluorescent artifacts. Fluorescent regions with high variability in the DAPI channel were detected by a quadtree-based segmentation algorithm and merged. Resulting objects that touched the image border were eliminated, and from the remaining objects, the largest one that was closest to the center of the well was selected as a seed for embryo detection. The seed object was expanded into the background until its area slightly exceeded the size of the embryo. Candidate regions containing the Lhx1a-EGFP transgene were then detected as lower hierarchy subobjects of the autofluorescent embryo based on an interactively determined threshold. Transgene shape was refined by a closing morphology algorithm. Total GFP intensity of the transgene (=GFP intensity per pixel × area) was computed and used to generate graphs. For compound profiling, data were normalized to the magnitude of response seen with negative (DMSO) and positive (800 nM UPHD25) controls and expressed as percentage activation = ((total GFPsample – GFPDMSO)/(total GFPUPHD25 – GFPDMSO)) × 100, which enabled direct comparisons of compound EC50 and magnitude of response between experiments. EC50 curves were generated in Prism (Graph Pad software) using a four-parameter logistic equation with the bottom constrained to that of the negative control.
Cdh17-EGFP
The single montage image file was segmented and the outline of the embryo detected as for Lhx1a-EGFP, except that the GFP channel alone contained enough information to reliably identify the whole embryo. The embryo’s outline was refined until it accurately traced the interface between embryo and background. The interface then served as a reference point for identification of objects based on the proximity to the edge of the zebrafish. The fluorescent kidney was detected based on fluorescence intensity and refined by a closing morphology algorithm. Small objects that did not connect to the main tubular structures were eliminated. The kidney was subsegmented and the cloaca identified based on brightness and proximity to the zebrafish edge. In some images, morphological abnormalities prevented the appearance of a fully extended kidney. Therefore, a 250-pixel-long segment of the fluorescent kidney (extending from the cloaca) was isolated, enabling consistent measurements regardless of morphology. Total GFP intensity of the transgene (=GFP intensity per pixel × area) was computed and used to generate graphs. For EC50 curves, data were normalized to vehicle control and averaged over the course of multiple independent experiments.
Results
A Small-Molecule HDAC Inhibitor Increases Lhx1a Expression
We had previously shown that a histone deacetylase (HDAC) inhibitor (methyl-4-(phenylthio)butanoate, UPHD25 ( Fig. 1a ) increased kidney progenitor cells in a proliferation-dependent fashion. 21 Proof of principle that increasing pools of kidney progenitors by a small molecule can ameliorate recovery of the kidney after injury comes from a recent study in which UPHD25 accelerated recovery and reduced fibrosis in mouse and zebrafish models of acute kidney injury. 22 To quantify UPHD25-mediated Lhx1a expansion, we first performed a concentration-response curve for UPHD25 using our previously described Lhx1a in situ hybridization method15,21 and found that UPHD25 dose-dependently increased Lhx1a expression ( Fig. 1b ), with a concentration required to elicit a half-maximal response (EC50) of about 600 nM ( Fig. 1c ). This documented that small-molecule–mediated expansion of Lhx1a was graded and saturable and provided a reference point for transgene analysis.
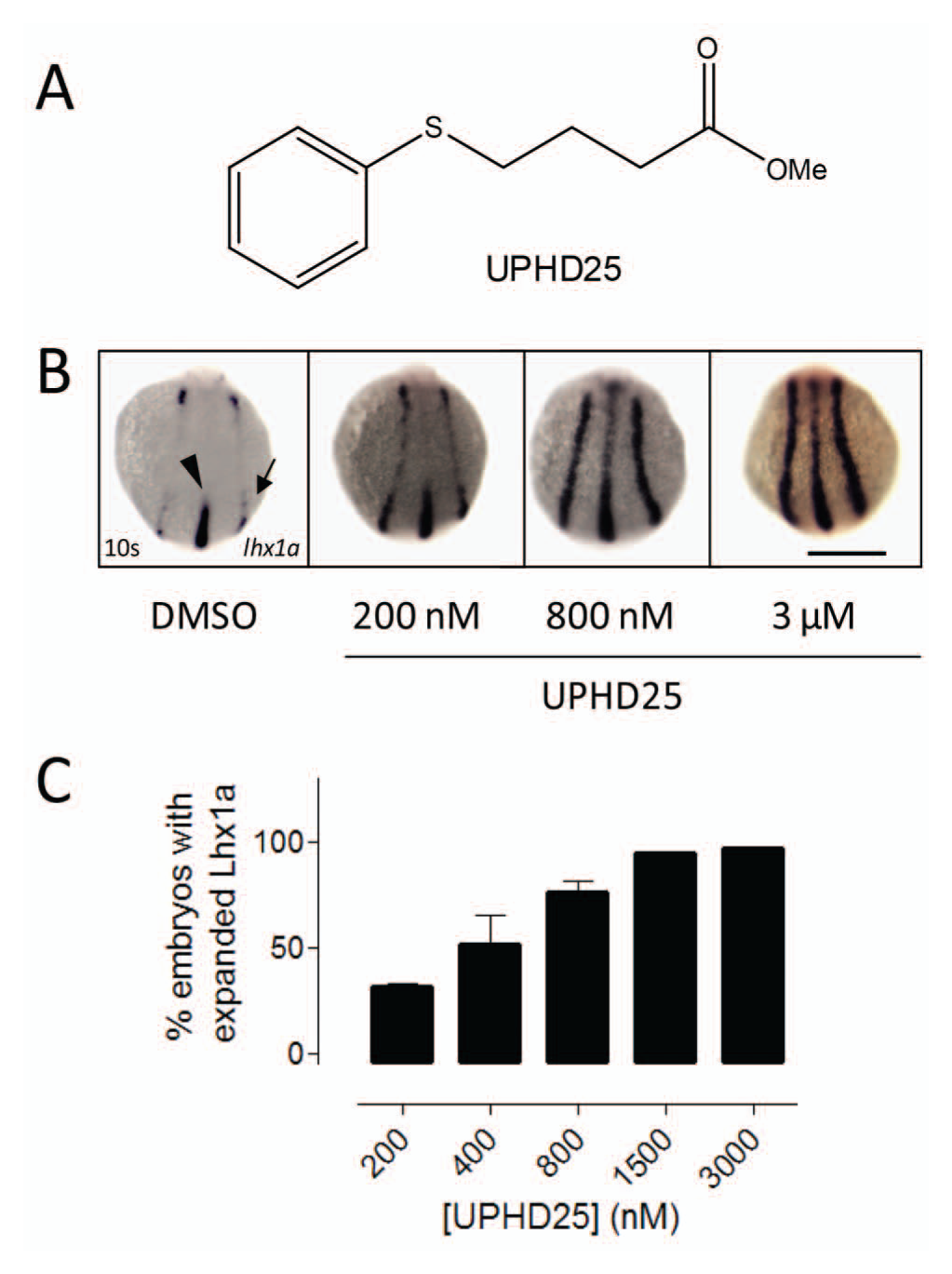
UPHD25 causes concentration-dependent increases in Lhx1a expression. (
Development of a Cognition Network Technology Rule Set to Quantify Lhx1a-EGFP in the Zebrafish Embryo
We developed an imaging algorithm to detect Lhx1a-EGFP expression. Tg(lhx1a:EGFP)pt303 embryos were grown to 256 cell stage, treated with vehicle (0.5% DMSO) or UPHD25 (800 nM), and imaged on a high-content confocal reader using a GFP-compatible filter set.
Figure 2a
shows that transgene expression was detectable and, as expected,
15
localized to the shield and germ ring. Because of the highly localized nature of the Lhx1a-EGFP transgene, the GFP channel image did not contain enough information to trace the whole embryo reliably. We therefore adopted a strategy in which the embryo body was detected by UV autofluorescence, followed by identification of the transgene as a lower hierarchy, GFP-positive subobject within the autofluorescent embryo (
Fig. 2b
,
c
). Further, locally specific segmentation and classification of the transgene enabled differential measurements of transgene expression in the shield (
Fig. 2c
,
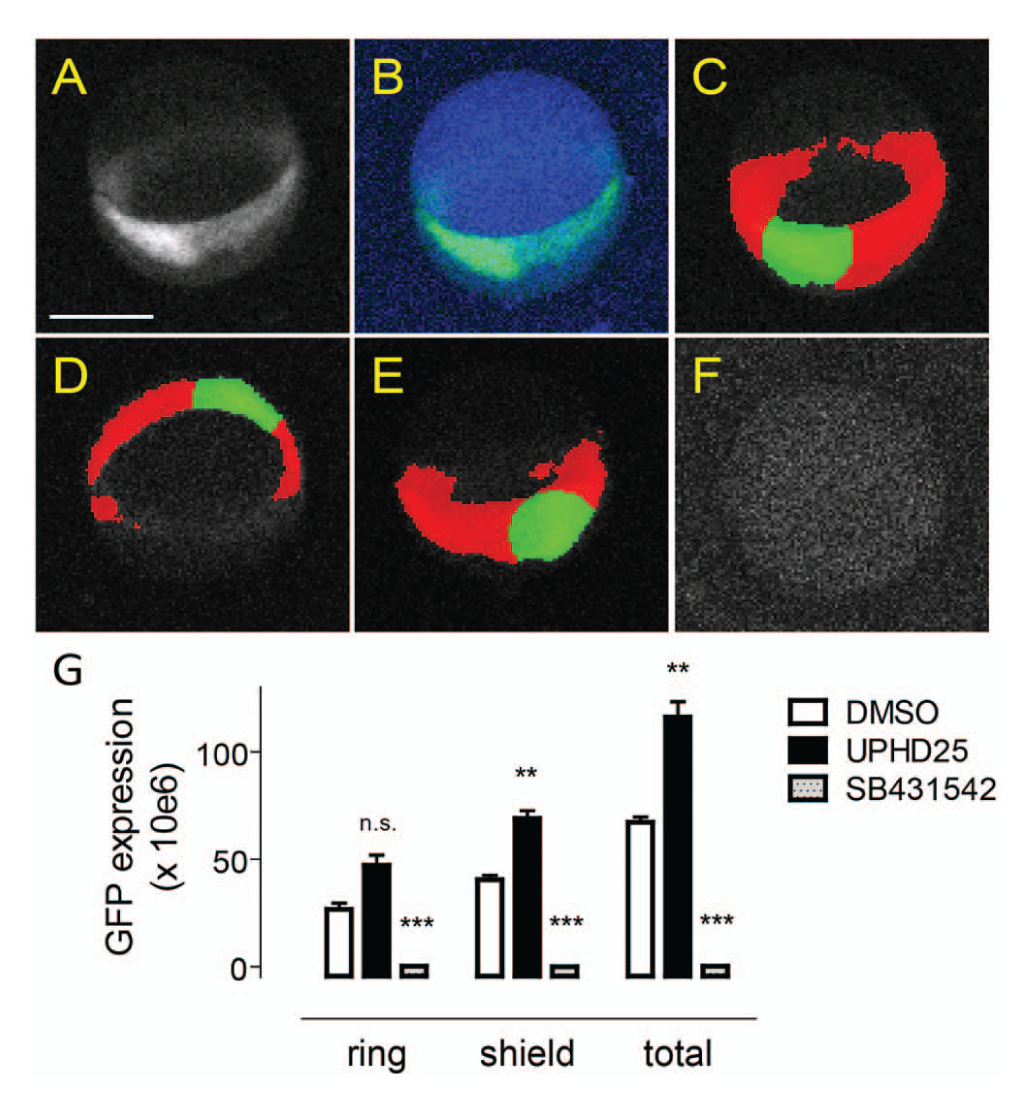
Image-based analysis of the Lhx1a:EGFP transgene at shield stage. (
Characterization of Lhx1a-EGFP Transgene Expression Kinetics
We examined the temporal characteristics of Lhx1a-EGFP transgene expression. Tg(lhx1a:EGFP)pt303 embryos (eight per condition) were treated at the 256 to 512 cell stage with vehicle or increasing concentrations of UPHD25. Single embryos were arrayed in the wells of a 96 well plate and imaged in the UV and GFP channel every 2 h for 22 h using identical image acquisition parameters. In the presence of vehicle alone ( Fig. 3a , upper panel), Lhx1a-EGFP became visible at 5 to 6 hpf, reached a maximum around 13 hpf, and remained elevated for several hours thereafter. UPHD25 increased levels of Lhx1a-EFGP but did not appear to prematurely activate Lhx1a expression ( Fig. 3a , lower panel). Quantitation by the CNT algorithm confirmed identical expression kinetics for vehicle- and UPHD25-treated embryos ( Fig. 3b ). GFP fluorescence reached a plateau at 14 hpf and remained elevated for several hours. The maximum assay window was a fivefold expansion with 3 µM UPHD25, which was stable over a period of 4 h (12–16 h posttreatment; Fig. 3b ). To determine optimal timing of compound addition, we treated embryos at various stages of development and found that expression kinetics and magnitude were similar over a 2-h treatment window (256 cell stage to dome stage; Fig. 3c ). The assay delivered graded responses; the EC50 of UPHD25 was about 900 nM and did not change over a period of 6 h ( Fig. 3d ).
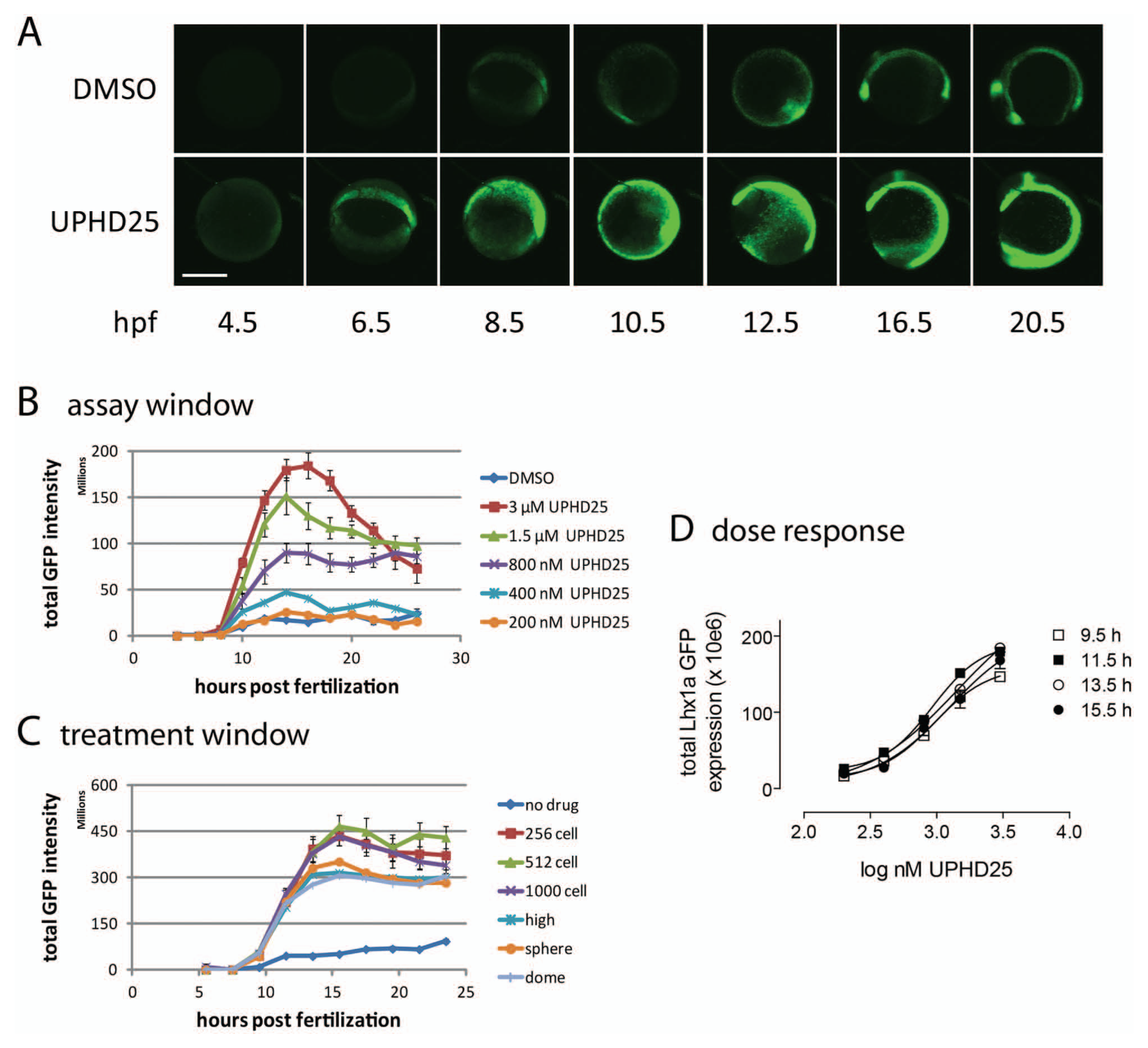
Assay optimization. (
Compound Activity Profiling
We next tested the ability of the assay to profile UPHD25 derivatives for in vivo expansion of Lhx1a-EGFP ( Fig. 4a ). UPHD25 is thought to exert its biological effects though the inhibition of Zn-dependent HDACs. 21 Invariably, the active site of all classic HDACs (HDAC1-11) consists of an approximately 11 Å long tubular channel containing a Zn2+ ion at its bottom. For HDACs of class I (a subset of the HDAC family that consists of HDACs 1, 2, 3, and 8), this 11 Å channel is also flanked by a foot pocket. 23 Changes on the UPHD25 core structure (i.e., length of the alkyl chain, the presence/absence of a strong zinc-chelating moiety, and the oxidation state of the sulfur atom) should influence activity toward HDACs and, in that regard, the in vivo expansion of Lhx1a-EGFP. We tested the effects of (1) an extended carbon chain (compound UPHD47), (2) a hydroxamic acid zinc-chelating group (compound UPHD23) as found in the HDAC inhibitors trichostatin and suberoylanilide hydroxamic acid, and (3) a change in the sulfur oxidation state (4-[phenylsulfonyl] butyric acid; PSOBA) on in vivo expansion of Lhx1a-EGFP. To enable direct comparison of compound activity across different experiments, a percentage activation value was calculated that considered the magnitude of response for both vehicle and the positive control (800 nM UPHD25) on each day of experiments (see the Materials and Methods section for details). Figure 4b shows that the assay profiled compounds by EC50 and magnitude of response. Oxidation of the sulfur moiety to a sulfone completely abrogated activity (PSOBA, closed squares). Lengthening the alkyl chain by two carbons (UPHD47, open circles) reduced the magnitude of response and EC50 compared with the UPHD25 benchmark (closed circles). Replacing the methyl ester with a hydroxamic acid resulted in a further reduction in EC50 and a similarly reduced magnitude of Lhx1a expression (UPHD23, closed triangles). In situ hybridization data qualitatively confirmed the results from the transgene studies ( Fig. 4c ). However, because of the quantal nature of in situ evaluations, which classify embryos as positive or negative for Lhx1a expansion, the assay did not capture differences in expansion levels. Collectively, the data not only indicate that the Lhx1a-EGFP assay is logistically simpler and higher throughput than manual scoring but also provide additional information that could be useful for in vivo SAR studies.
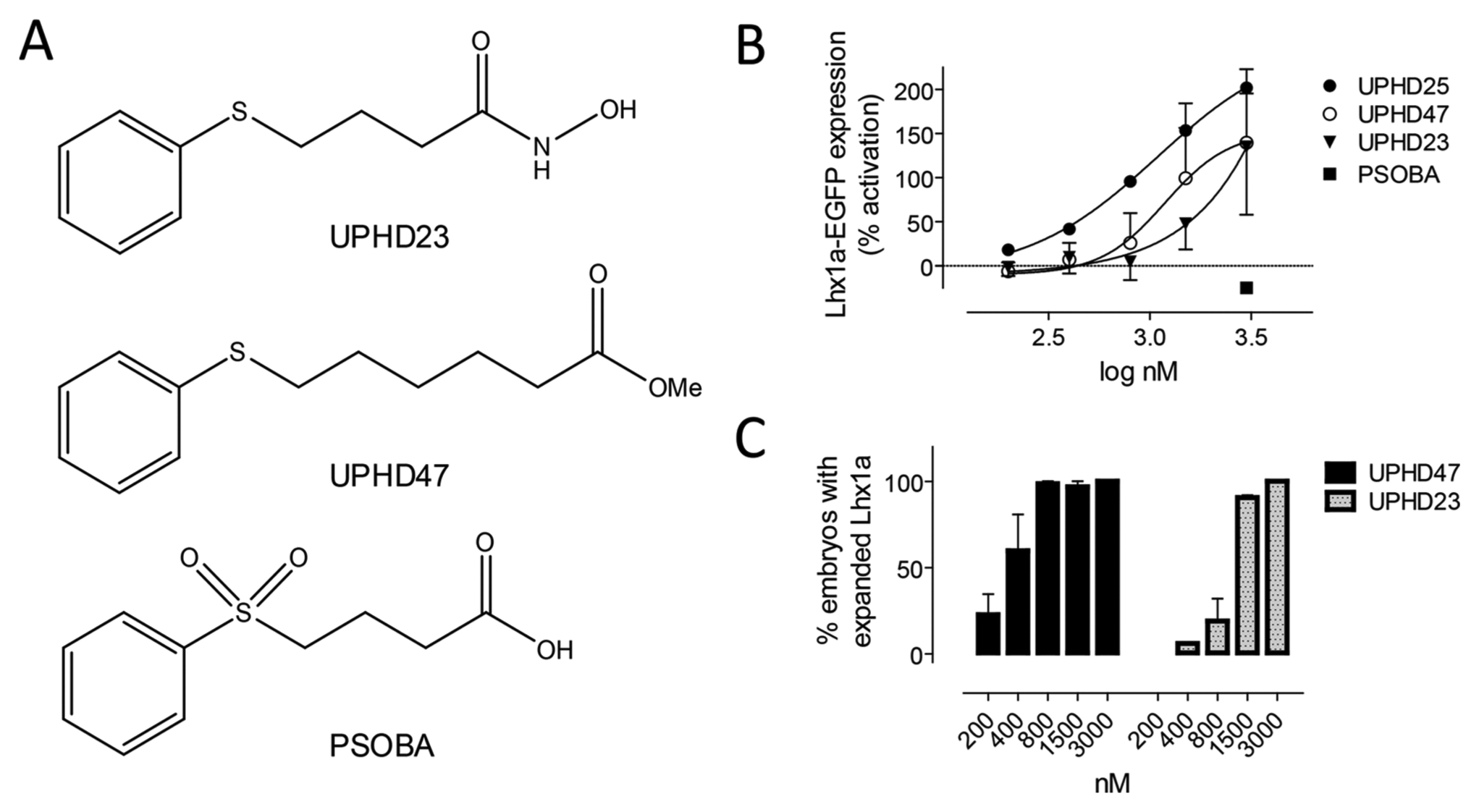
Analog profiling (A) Structures of UPHD25 analogs. (B) EC50 determinations. The assay profiled structurally related agents by EC50 and magnitude of response. The y axis shows the percentage activation relative to positive (800 nM UPHD25) and negative (DMSO) controls. Data are the averages of at least two independent experiments demonstrating week-to-week repeatability. (
Secondary Assay Development
Although Lhx1a is essential for kidney development 14 and is expressed in the kidney during recovery after ischemic injury, 13 it is also expressed in regions of the brain and in the notochord (see Swanhart et al. 15 ; Fig. 2a ). To ensure that compounds acted in a kidney-specific manner and increased pools of progenitor cells, we developed a secondary assay that measures expansion of the zebrafish embryonic kidney. We generated a fluorescently labeled transgene ( Fig. 5a ) that expresses EGFP under the control of a kidney-specific cdh17 promoter. The resulting Tg(cdh17:EGFP)pt305 line expressed EGFP in kidney tubule cells, enabling real-time observation of organ development. At 48 hpf, the kidney appears as a pair of tubular structures that converge at the cloaca ( Fig. 5b , c ). We developed a CNT rule set that detected the fluorescent zebrafish kidney (minus the glomeruli) and tubular subdomains therein. The cloaca was identified based on brightness and proximity to the zebrafish embryo domain edge, and EGFP intensity was measured in a 250 pixel long segment extending from the cloaca ( Fig. 5d – i ). While expansion of the embryonic kidney after compound treatment was visible but difficult to discern ( Fig. 5j – l ), quantitation of total transgene expression by the CNT rule set revealed that UPHD25 dose dependently increased the number of fluorescent cells within the kidney tubule. Its EC50 of 620 nM ( Fig. 5m , circles) was consistent with Lhx1a measurements ( Figs. 1c and 4d ). UPHD23 ( Fig. 5m , triangles) showed a lower magnitude of response and higher EC50 than UPHD25. These data documented that the increase in Lhx1a expression seen with both compounds translated into expansion of kidney progenitor cells and into increased organ size at later stages of development.
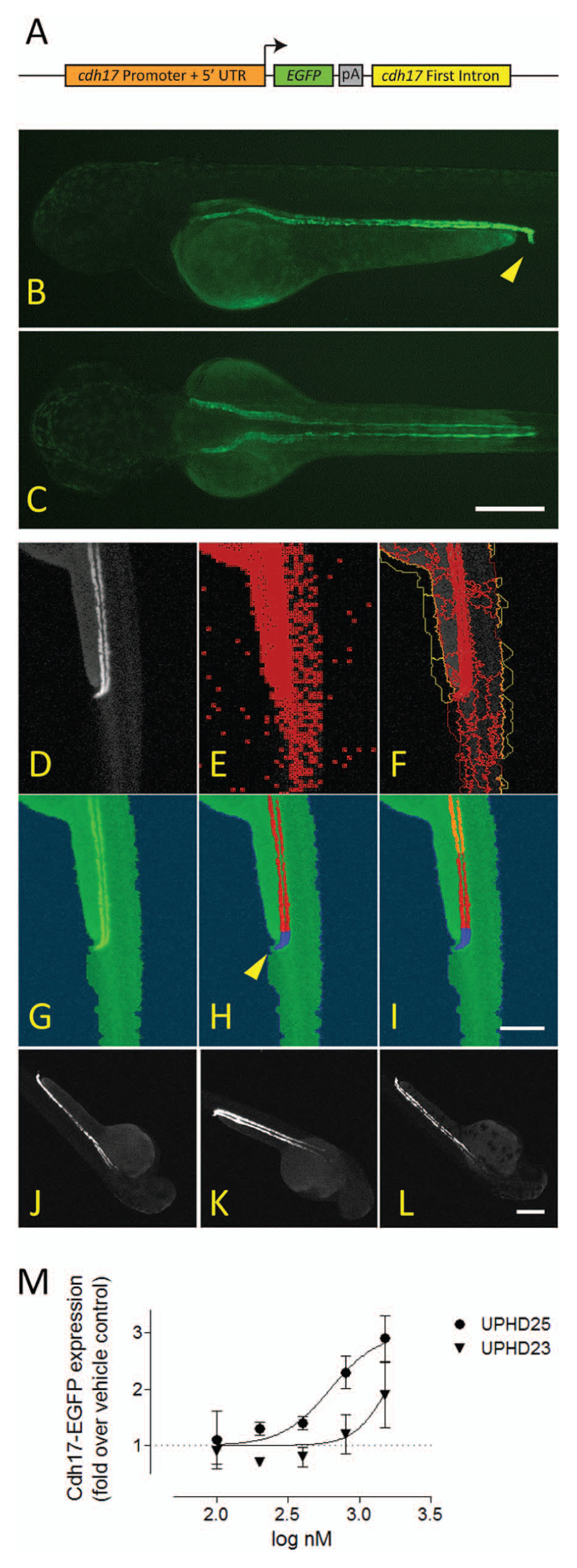
Secondary assay development. (
Heterogeneity Assessments
The availability of a quantitative in vivo assay on high-content readers opened the possibility for large-scale discovery efforts. We therefore investigated avenues to increase the number of embryos for experimentation. Because the Tg(lhx1a:EGFP)pt303 line was established from a single founder fish, we reasoned that it would be possible to combine embryos from multiple mating pairs if their baseline Lhx1a-EGFP expression and their responsiveness to UPHD25 were identical. Figure 6 shows the averages and standard errors from three different mating pairs. One-way analysis of variance documented no significant differences between the three clutches, neither at baseline nor after treatment with UPHD25 (p < 0.001). Importantly, neither egg clutch alone was statistically different from the average of all embryos. These data indicate that eggs from different mating pairs can be pooled without loss of assay performance and suggests the assay can be adapted to mass breeding.
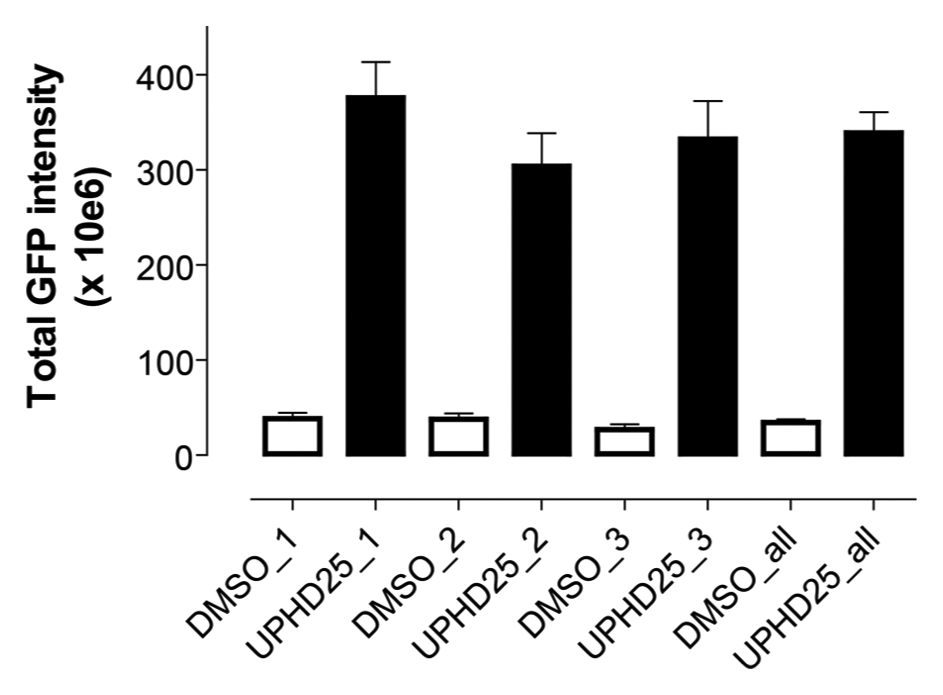
Mating pair heterogeneity. Tg(lhx1a:EGFP)pt303 (eight embryos per condition) from different founder fish were treated at the 256 to 512 cell stage with vehicle (open bars) or 800 nM UPHD25 (closed bars). Images were acquired and analyzed with the CNT rule set. One-way analysis of variance with Tukey’s post test revealed no statistical differences between mating pairs at baseline or after treatment with UPHD25 (p < 0.001). The fourth data set (DMSO_all and UPHD25_all) represents the aggregate of all three mating pairs.
Discussion
The availability of in vivo assays for systematic interrogation of small-molecule chemical libraries could substantially improve phenotypic drug discovery efforts. The impact of whole-organism models and assays will be most profound in diseases or biological processes that are not dependent on the expression or activity of a single target, are mechanistically poorly defined, or involve complex in vivo micro- and macroenvironments. One of those scenarios is the recovery after injury of organs with innate regenerative capacity, such as the kidney. During regeneration, genes normally involved in organogenesis are reactivated and contribute to the regenerative process. Because the zebrafish embryonic kidney developmentally closely resembles the mammalian embryonic kidney specification, 24 quantifying embryonic markers of zebrafish kidney development is a promising strategy for the discovery of agents that ameliorate organ regeneration after injury.
In this study, we have developed a quantitative, in vivo profiling assay for expansion of lhx1a expression in transgenic zebrafish. The assay is based on the measurement of an EGFP-tagged marker of organogenesis that is critical for development of the zebrafish and human embryonic kidney. Using an artificial intelligence–based image analysis method, we were able to rapidly perform quantitative measurements of transgene expression that confirmed earlier studies using in situ hybridization and visual scoring. The Lhx1a-EGFP transgene responded to activating and inhibitory stimuli, including a previously reported small molecule (UPHD25) that ameliorates recovery from kidney injury in zebrafish and mice. 22 The assay’s ability to perform real-time kinetics studies documented that UPHD25 expanded Lhx1a only at time points when the endogenous gene was expressed, indicating UPHD25 potentiated existing developmental processes rather than activating transcription by itself. This is an important feature from a therapeutic standpoint, as inappropriate activation of organogenesis genes could lead to tissue hyperproliferation.
The assay delivered graded responses, enabling determination of EC50 values for UPHD25 that were consistent with those obtained by manual scoring of Lhx1a-stained embryos. Assay development studies defined a treatment window of 2 h for compound addition and a stable analysis window of at least 4 h starting at 10 h posttreatment. The assay profiled structurally related compounds by both EC50 and magnitude of response. Therefore, automated imaging and analysis not only improved throughput compared with in situ hybridization but also provided higher information content. The assay’s ability to quantify EC50 and magnitude of Lhx1a expansion enables, for the first time, SAR studies to dissect the relative contributions of EC50s and the extent of Lhx1a expansion to compounds’ biological activity.
Zebrafish have become a favorite among multicellular organisms for systematic discovery of novel therapeutics. Although the zebrafish community has made great strides toward high-throughput screening, the throughput of zebrafish assays is still much smaller than that of traditional biochemical or even cell-based screens. Having eliminated image acquisition and analysis as limiting factors, a newly emerging bottleneck is the ability to generate large numbers of eggs for experimentation. A single zebrafish mating pair can generate egg clutches in the hundreds, but for large-scale screening, much greater numbers of eggs are needed, and this can be achieved only by mixed breeding. Variability assessments of the Tg(lhx1a:EGFP)pt303 line indicated that Lhx1a responses in embryos from different mating pairs were not statistically different from each other or from the pooled population from all pairs, indicating the line is suited for mass breeding.
It should be noted that although the Lhx1a assay is logistically simple and robust, expression of Lhx1a during development is not limited to the kidney but also occurs in areas of the brain and in the notochord. Therefore, discovery efforts using the automated Lhx1a assay will have to include functional follow-up assays that document kidney-specific biological activity. To that end, we developed an automated, quantitative organogenesis screen based on a fluorescently labeled cadherin-17 transgene (cdh17:EGFP) that is expressed in kidney tubular cells. The cdh17:EGFP transgene, which allows real-time observation of organ development in the living organism, was readily quantified by CNT. Because cdh17-EGFP fluorescence is a direct measure of kidney field expansion, 20 the assay can eliminate compounds for which increased expression of organogenesis markers does not translate into expansion of kidney progenitor cell pools. Therefore, the cdh17-EGFP assay is a kidney-specific, functional complement to the primary Lhx1a screen. Although the cdh17-GFP assay is quantitative and automated, its dynamic range and higher variability make it more suited for secondary screening. In summary, the Lhx1a:EGFP transgene, when combined with a functional organogenesis assay, is a powerful, enabling tool for the phenotypic discovery and characterization of novel agents that increase kidney progenitor cells.
Footnotes
Acknowledgements
The authors thank Eric de Groh, Cuong Diep and Alan Davidson for early discussions in the design of the cdh17 promoter.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by grants 1RC4 DK090770, 2R01 DK069403, and 2R01 HD053287 from the National Institutes of Health. This project used UPCI Shared Resources supported in part by award NIH P30CA047904 and O’Brien Kidney Center resources supported by 1P30 DK079307.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
