Abstract
Natural products are considered an extremely valuable source for the discovery of new drugs against diverse pathologies. As yet, we have only explored a fraction of the diversity of bioactive compounds, and opportunities for discovering new natural products leading to new drugs are huge. In the present study, U2nesRELOC, a previously established cell-based imaging assay, was employed to screen a collection of extracts of microbial origin for nuclear export inhibition activity. The fluorescent signal of untreated U2nesRELOC cells localizes predominantly to the cytoplasm. Upon treatment with the nuclear export inhibitor leptomycin B, the fluorescent-tagged reporter proteins appear as speckles in the nucleus. A proprietary collection of extracts from fungi, actinomycetes, and unicellular bacteria that covers an uncommonly broad chemical space was used to interrogate this nuclear export assay system. A two-step image-based analysis allowed us to identify 12 extracts with biological activities that are not associated with previously known active metabolites. The fractionation and structural elucidation of active compounds revealed several chemical structures with nuclear export inhibition activity. Here we show that substrates of the nuclear export receptor CRM1, such as Rev, FOXO3a and NF-κB, accumulate in the nucleus in the presence of the fungal metabolite MDN-0105 with an IC50 value of 3.4 µM. Many important processes in tumor formation and progression, as well as in many viral infections, critically depend on the nucleocytoplasmic trafficking of proteins and RNA molecules. Therefore, the disruption of nuclear export is emerging as a novel therapeutic approach with enormous clinical potential. Our work highlights the potential of applying high-throughput phenotypic imaging on natural product extracts to identify novel nuclear export inhibitors.
Keywords
Introduction
Throughout recent history, metabolites of microbial origin have had an extraordinary impact on the welfare of humanity. Antibacterial and antifungal antibiotics, immunosuppressors, and antitumoral and hypolipidemic agents, derived from compounds of microbial origin, have been increasingly used in clinical practice over the past 60 years. 1 They have significantly contributed to the well-being and the remarkable enhancement of life expectancy observed globally. The outstanding diversity of chemical structures that microorganisms are able to produce, including polyketides, nonribosomal peptides, terpenoids, and their combinations, derive from a relatively limited number of basic biosynthetic pathways, and these pathways have diversified significantly during evolution. In view of such notable structural diversity, it should not be surprising that we keep finding microbial metabolites with biological activity of interest for a wide range of therapeutic fields.2,3 However, the validation and selection of primary screening assays are vital to guaranteeing a selection of extracts or molecules with relevant pharmacological action and need to be followed up. The assay must be rapid, simple, and easy to implement, and it should produce robust results, preferably at a low cost. Cellular imaging and high-content screening (HCS) technology are widely used to measure cellular function and behavior in cells. 4 HCS combines the efficiency of high-throughput techniques with the ability of cellular imaging to collect quantitative data and facilitates the integration of disease-relevant screens in the drug discovery process. 5 In this work, we applied and validated a fluorescent microtiter assay based on mammalian cells stably expressing green fluorescent protein (GFP)–labeled Rev protein, which contains a strong heterologous nuclear export signal (NES). 6 NESs can be defined as sequences that are necessary and sufficient for nucleocytoplasmic translocation through the nuclear pore.7,8 NES on a cargo protein binds to the nuclear exporting receptor CRM1, forms a tripartite complex with Ran-GTP in the nucleus, 9 and allows the release of the transport complex at the cytoplasmic side of the nuclear pore. 10 Aberrant localization of tumor suppressor proteins is one of the mechanisms used by cancer cells to evade their antineoplastic mechanism.11,12 In addition, the aberrant localization of oncoproteins can lead to their inadequate activation. 13 Extracts and molecules that interfere with the general export machinery and in turn accumulate NES-containing proteins in the cell nucleus present an exciting therapeutic profile for the treatment of cancers that display aberrant protein subcellular localization. Leptomycin B (LMB), a fungal metabolite that binds covalently to a cysteine residue in the active center of CRM1, preventing the efflux of NES-containing proteins,14,15 is the best-studied nuclear export inhibitor. Therefore, natural products of microbial origin appear to be a valuable source for the discovery of new nuclear export inhibitors. A high-throughput screening (HTS) of 14,000 microbial extracts was carried out in U2nesRELOC, and 1.6% of the extracts displayed nuclear export inhibitor activity (average Z′ higher than 0.76). To our knowledge, our group was the first to use this methodology for screening a collection of natural extracts to identify compounds with nuclear export inhibitor activity.
Materials and Methods
Plasmids
The U2nesRELOC assay uses the reporter construct pRevMAPKKnesGFP, which has been described earlier. 16 pRevMAPKKnesGFP carries the NES from mitogen-activated protein kinase kinase (MAPKK) (or MEK) cloned between the BamHI and AgeI sites of pRev(1.4)-GFP, sandwiched between the Rev and the GFP coding sequences. 16 The EGFP-FOXO3a fusion protein was kindly provided by T. Finkel (National Institutes of Health, Bethesda, MD).
Cell Culture and Stable Transfections
The human osteosarcoma cell line U2OS was obtained from the American Type Culture Collection (Manassas, VA) and cultured in Dulbecco’s modified Eagle’s medium, supplemented with 10% fetal bovine serum (Sigma, St. Louis, MO) and antibiotics-antimycotics. Cell cultures were maintained in a humidified incubator at 37 °C with 5% CO2 and passaged when confluent using trypsin/EDTA. The stable transfected U2nesRELOC and U2foxRELOC cells have been described previously. 6
Upscaling and Automation of the U2nesRELOC Assay
The U2nesRELOC-based assay was formatted in 96-well plates (Greiner Bio-One, Frickenhausen, Germany), and workflow was automated (
Image Acquisition and Processing
The BD Pathway 855 High Content Bioimager (BD Biosciences, San Jose, CA) was used for automated image acquisition. Acquired images were processed using AttoVision software (BD Biosciences). The Bioimager was equipped with a 488/10-nm enhanced GFP (EGFP) excitation filter, a 380/10-nm DAPI excitation filter, a 515-nm long-pass (LP) EGFP emission filter, and a 435-nm LP DAPI emission filter. Images were acquired in the DAPI and GFP channels of each well using a 10× dry objective. The plates were exposed 0.066 ms (gain 0) to acquire DAPI images and 0.85 ms (gain 30) for GFP images. Cells were stained with DAPI to facilitate autofocus of the microscope and to aid the image segmentation. An image algorithm was applied to allow the cell nucleus segmentation based on a local thresholding. Our segmentation strategy assumes that the cell’s cytoplasm surrounds the nucleus. Consequently, cytoplasmic fluorescence intensity is calculated from all the pixels within a circumferential ring surrounding the nuclear ring mask. The width of the ring was defined to be small enough to avoid ambiguities due to irregular cell shape. Based on the definition of cell compartments, the nuclear and cytoplasmic levels of GFP fluorescence were quantified.
Data Analysis
The BD Pathway Bioimager outputs its data in standard text files. Data were imported into the data analysis software BD Image Data Explorer. The nuclear/cytoplasmic ratios of fluorescence intensity were determined by dividing the fluorescence intensity of the nucleus by the cytoplasmic fluorescence intensity. A threshold ratio greater than 1.8 was employed to define nuclear accumulation of fluorescent signal for each cell. On the basis of this procedure, we calculated the percentage of cells per well displaying nuclear translocation or inhibition of nuclear export. Compounds that induced a nuclear accumulation of the fluorescent signal greater than 60% of that obtained from wells treated with 4 nM leptomycin B (LMB) (LC Laboratories, Woburn, MA) were considered hits. We also monitored nuclear shrinkage known to be associated with apoptosis by analyzing the number of nuclear vertices. The term nuclear vertices refers to the number of pixels that lie on the nuclear boundary. 6 To estimate the quality of the HCS assay, we calculated the Z′ factor with the following equation: Z′ = 1 – [(3 × SD of positive controls) + (3 × SD of negative controls)/(mean of positive controls) – (mean of negative controls)] as previously described by Zhang and colleagues. 17
Fundación MEDINA’s Natural Extracts Collection
Six thousand actinomycetes, 2000 marine actinomycetes, and 6000 fungal strains were cultivated in different production media from 7 to 28 days at temperatures ranging from 22 °C to 28 °C, according to the microbial strain requirements. 18 Secondary metabolites produced in the cultivation broths were collected and extracted with acetone. Extracts were prepared by mixing 10 mL of culture broth with 10 mL of pure acetone. After 1 h of shaking in an orbital shaker (Adolf Kühner AG, Birsfelden, Switzerland), the acetone/water extracts were transferred to 16-mm glass tubes and evaporated to half of their volume after centrifugation at 1500 g for 15 min. The final aqueous extract was approximately 2× whole-broth equivalent with 20% DMSO. After storage at −20 °C, an aliquot of 500 µL was thawed, stored at 4 °C, and mixed briefly before testing.
Extract Fractionation
The active extracts were fractionated on a Gilson Preparative HPLC system (Gilson, Middleton, WI) using a Zorbax RX-C8 column (9.4 × 250 mm, 5 µm), a gradient H2O/CH3CN from 5% to 100% CH3CN in 35 min + 5 min at 100% CH3CN, a flow rate of 3.6 mL/min, and UV detection at 210 and 280 nm. A total of 80 fractions were collected every 30 s from each chromatographic run, dried under vacuum, and tested for their U2nesRELOC inhibitory activity in the above-mentioned conditions. Active fractions from each extract were analyzed by liquid chromatography/mass spectrometry and nuclear magnetic resonance to establish the identity and/or novelty of the bioactive components.
Determination of IC50 Values in U2nesRELOC Assays
Determination of IC50 values was carried out in automated U2nesRELOC assays. Cells were cultured as described above and exposed for 1 h to equal volumes of test compounds (2 µL). IC50 values were calculated as being the inhibitor concentration that increases nuclear accumulation of the reporter protein by 50% using Genedata Screener software (Genedata AG, Basel, Switzerland). Cells were treated with eight 2-fold serial dilutions of each compound spanning concentrations from 50 µM to 0.39 µM in 1% DMSO final. Compound activity was normalized to the in-plate negative and positive controls using DMSO and LMB, respectively. We chose 4 nM LMB as maximal inhibition and 1% DMSO as minimal inhibition of nuclear export in U2nesRELOC cells.
Immunofluorescence Analysis by Confocal Microscopy
U2OS cells were cultured as described above and exposed for 1 h to equal volumes of MDN-0105 (2 µL). The cells were 100% methanol fixed (5 min) and then incubated in 1% bovine serum albumin (BSA)/10% normal goat serum/0.3 M glycine in 0.1% phosphate-buffered saline with Tween (PBST) for 1 h to permeabilize the cells and block nonspecific protein-protein interactions. The cells were then incubated with nuclear factor (NF)–κB2 p100/p52 antibody (4882; Cell Signaling Technology, Danvers, MA) overnight at +4 °C. The secondary antibody was ab96921 DyLight 594 (Abcam, Cambridge, UK) donkey anti–rabbit IgG (H+L) for 1 h. DAPI was used to stain the cell nuclei (blue) at a concentration of 1.43 µM. Samples were imaged at 22 °C using 63× water immersion lenses on a Leica (Wetzlar, Germany) SPE confocal imaging system. Images were obtained using Leica software.
Results
The main objective of this study was the application of high-content imaging to identify novel nuclear export inhibitors from a collection of microbial natural product extracts enriched in secondary metabolites. To establish a system suitable to monitor CRM1-dependent nuclear export in an HTS format, we used a reporter protein previously described as extremely active in an in vivo nuclear transport assay. 6 This high-content screening assay is based on the human osteosarcoma cell line U2OS stably expressing GFP-labeled Rev protein, which contains a strong heterologous NES. The fluorescent signal of untreated U2nesRELOC cells predominantly localizes to the cytoplasm. Upon treatment with nuclear export inhibitors, the GFP-labeled reporter protein accumulates rapidly in the cell nucleus. LMB, an inhibitor of the CRM1–dependent nuclear export that covalently binds to a single cysteine residue of the CRM1 protein, 14 was used as a positive control. Consistent with previous reports, nuclear accumulation of the fluorescent signal in U2nesRELOC cells was detectable as early as 10 min after blocking nuclear export by LMB treatment and further increased for 1 h.6,19 Therefore, all the screening experiments presented in this study were performed using drug exposure for 1 h.
Screening Campaigns
MEDINA’s natural extract collection, employed in this assay, contains secondary metabolites from actinomycetes and fungal cultures. Microbial natural products have been one of the major sources of novel drugs, and the untapped chemical diversity in these extracts favors that novel molecules with potential therapeutic applications are still waiting to be discovered from these natural sources. Indeed, LMB is a secondary metabolite produced by Streptomyces spp.14,20 However, the discovery of novel nuclear export inhibitors from natural products has not been extensively pursued during the past decade. Once the assay conditions were optimized, 14,000 different microbial extracts were analyzed in triplicate for their capacity to induce nuclear accumulation of fluorescent signal in the reporter cells as described in the Materials and Methods. The standard deviations of the triplicates ranged from ±3% to ±5%. Of the 14,000 microbial extracts, 6000 were derived from fungi, 6000 from actinomycetes, and 2000 from marine actinomycetes. The results obtained from the primary screening procedure are summarized in Table 1 . A relatively high number of natural extracts (1.6%) displayed nuclear export inhibitor activity in the primary screening. Active natural product extracts were found more frequently in fungal cultures (81% of 151 total positive extracts) than in actinomycetes culture extracts (19%; Table 1 ).
Screening Campaign Results.
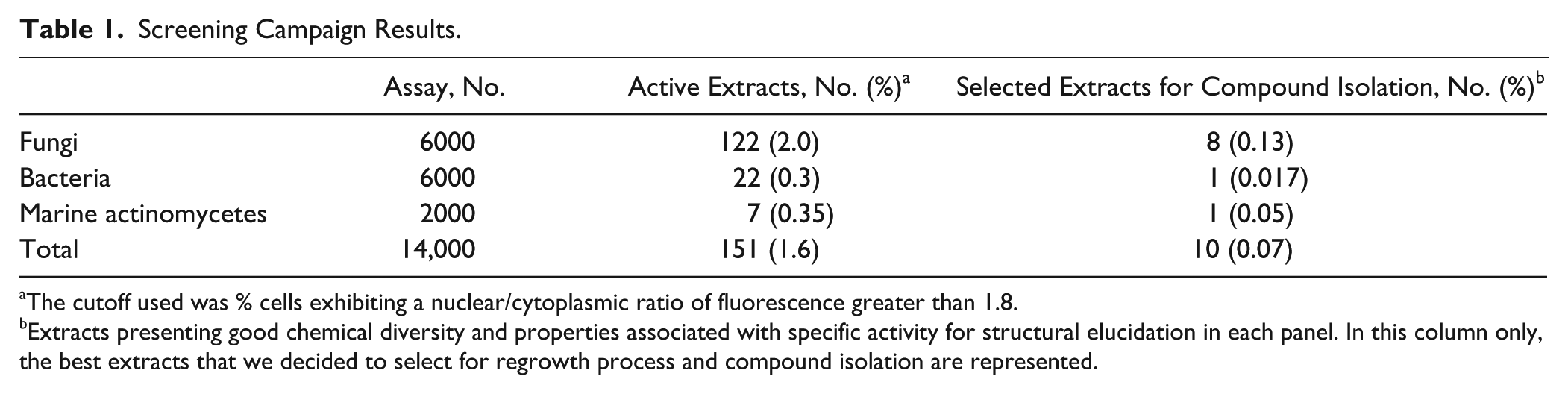
The cutoff used was % cells exhibiting a nuclear/cytoplasmic ratio of fluorescence greater than 1.8.
Extracts presenting good chemical diversity and properties associated with specific activity for structural elucidation in each panel. In this column only, the best extracts that we decided to select for regrowth process and compound isolation are represented.
A “cherry-pick” campaign was carried out to confirm the activity and the reproducibility of the natural extracts showing nuclear export inhibitor activity. To avoid reisolation of previously known natural products that often occur in the course of bioassay-guided fractionation,
21
active extracts or subsequently detected fractions were subjected to chemical dereplication (
Scale-up Fermentation and Bioassay-Guided Fractionation of Selected Extracts
Bioactive fermentation selected in the previous dereplication steps was optimized and reproduced at a sufficient scale (1L) to isolate and identify active compounds. Bioassay-guided fractionation of extracts to obtain the bioactive component for further biological testing was performed using high-performance liquid chromatography (HPLC) (
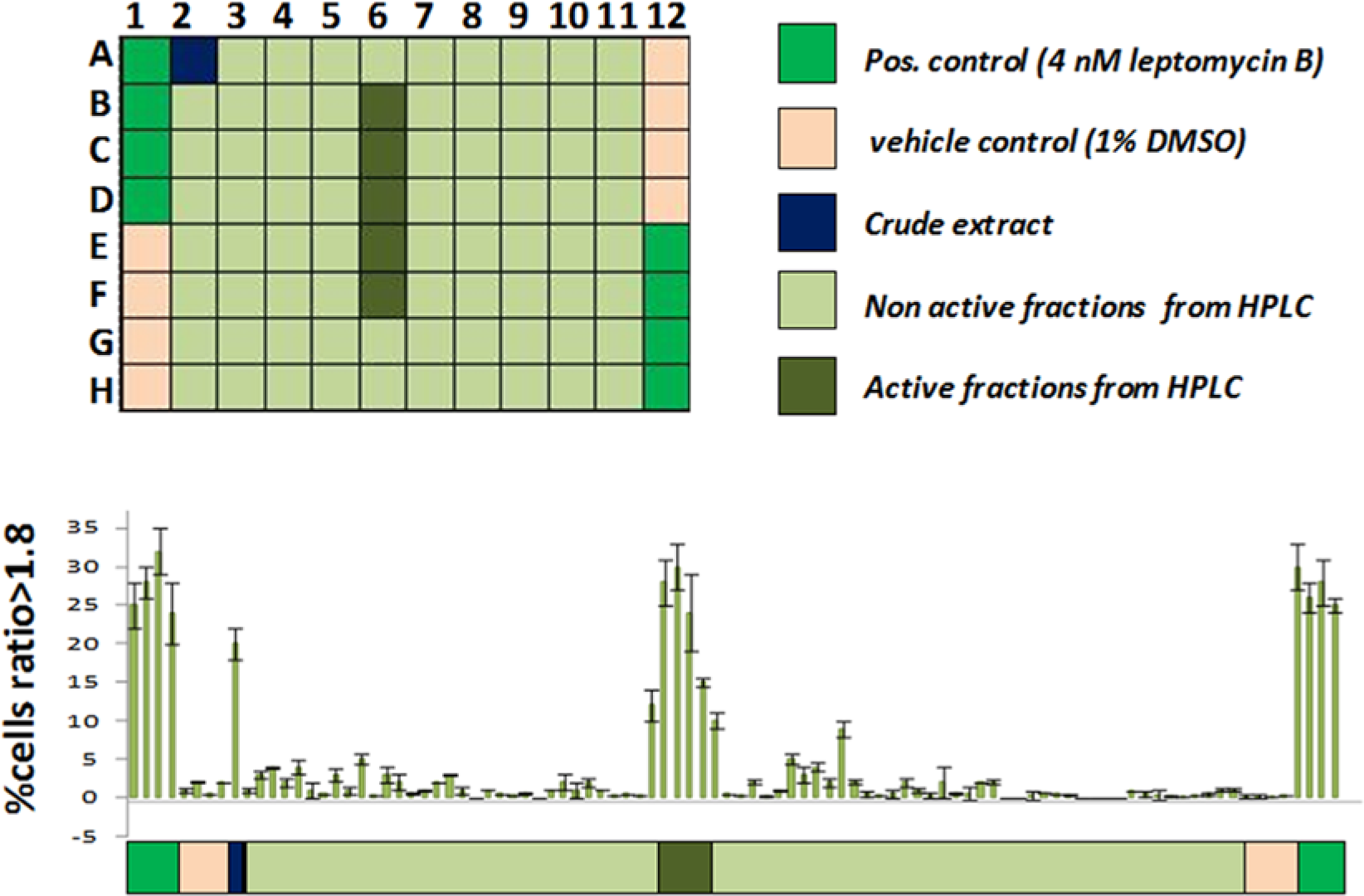
Identification of active extracts and fractions using the U2nesRELOC assay. U2nesRELOC cells were seeded in 96-well plates and exposed for 1 h to extracts or derived fractions. The nature of the samples in the assay plates is color coded below the graph. Each assay plate contained eight wells of cell samples treated with 4 nM leptomycin B (LMB) (positive controls, vivid green) and eight wells treated with 1% DMSO (negative control, light orange), crude extracts (purple), and high-performance liquid chromatography (HPLC) fractions. Nonactive fractions are represented in light green and active fractions in dark green. The nuclear/cytoplasmic ratios of fluorescence intensity were determined by dividing the fluorescence intensity of the nucleus by the cytoplasmic fluorescence intensity. A threshold ratio greater than 1.8 was employed to define nuclear accumulation of the fluorescent signal for each cell. On the basis of this procedure, we calculated the percentage of cells per well displaying nuclear translocation or inhibition of nuclear export. Compounds that induced a nuclear accumulation of the fluorescent signal greater than 60% of that obtained from wells treated with 4 nM LMB were considered hits.
Representative images of the compound-treated assay cell line using a high-content imaging platform are presented in Figure 2 . In untreated U2nesRELOC cells, GFP fusion protein was predominantly cytoplasmic ( Fig. 2A ). LMB inhibited the nuclear export of the reporter protein, leading to localization of the fluorescent signal in the cell nucleus ( Fig. 2B ). Active crude extract promotes localization of the fluorescent signal in the cell nucleus, indicating their inhibition capacity in U2nesRELOC cells ( Fig. 2C ). The same effect can be observed in active HPLC fractions ( Fig. 2D ). Several HPLC fractionation rounds were applied until pure compound purification and confirmation of inhibitory activity were achieved. The chemical structures of all isolated active metabolites were identified by spectroanalyses. None of these compounds had been shown to have nuclear export inhibitory activity. From the fungal extracts, we identified several molecules from the same chemical class as nuclear export inhibitors. We decided to analyze MDN-0105, one compound of this family.
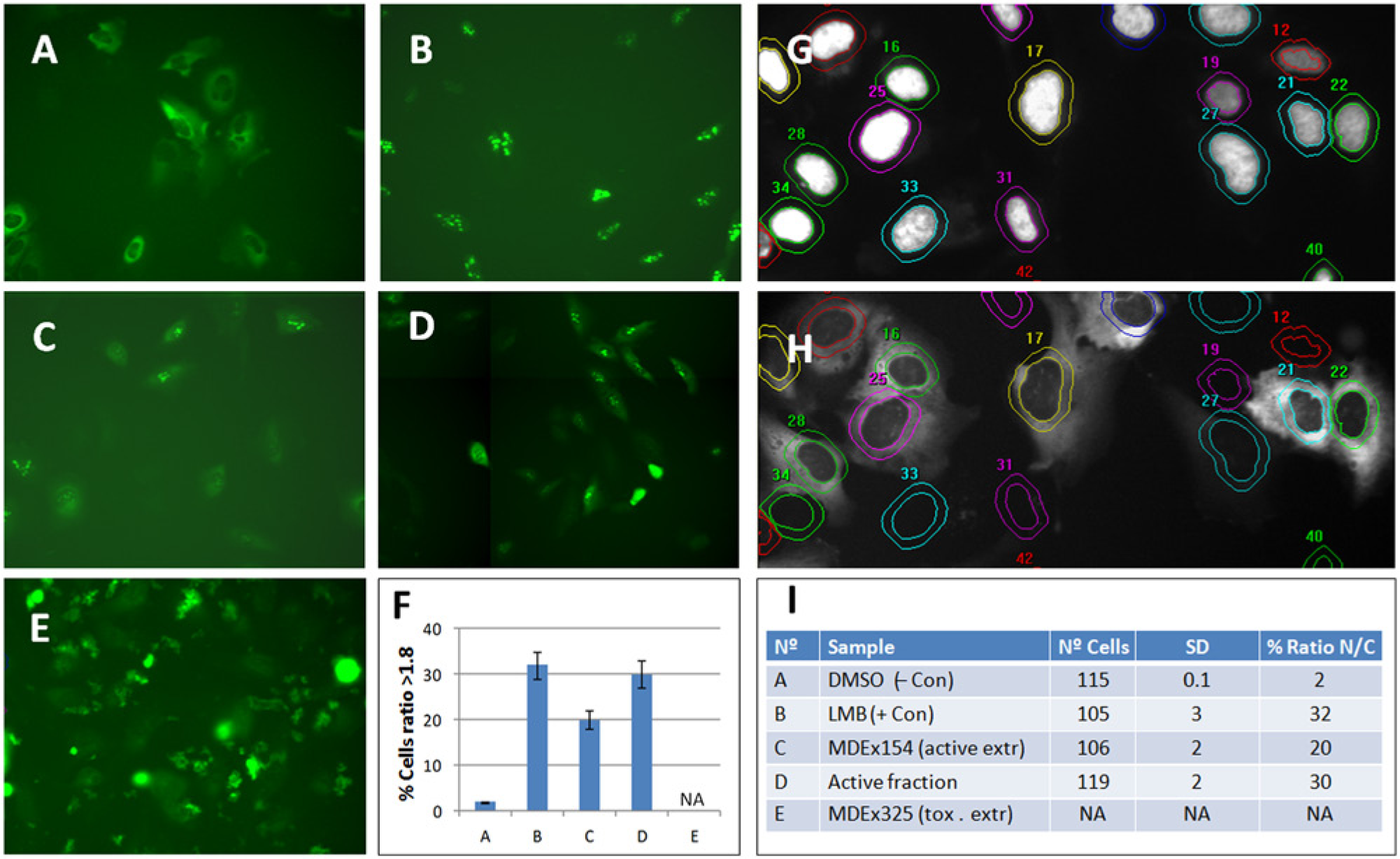
Representative images of the compound-treated assay cell line. Representative images of the compound-treated assay cell line using the high-throughput format (ABCDE) of the U2nesRELOC system: (
Pure Compound MDN-0105 Dose-Response Analysis Using U2nesRELOC Cells
We next determined the 50% inhibitory concentration (IC50) value of MDN-0105 in the U2nesRELOC assay. Cells were cultured as indicated in Materials and Methods and treated with different doses of MDN-0105 for 1 h. Figure 3 shows that U2nesRELOC is very sensitive to the presence of MDN-0105 at low concentrations. Our experiment demonstrated that the lowest concentration of MDN-0105 that interferes with the subcellular localization of the reporter protein was 0.3 µM, and a complete inhibition of nuclear export was observed at 15 µM. The IC50 of the reference compound LMB was 0.91 nM in this experiment (not shown). IC50 values for LMB-mediated inhibition of nuclear export in the low nanomolar range have been reported in several earlier reports for other cell types.6,22,23 IC50 for the MDN-0105 compound was 3.4 µM in this assay. These values demonstrate that MDN-0105 is indeed able to block the nuclear export pathway in concentrations that are higher than for LMB, which is a reference compound that presents a high toxicity.
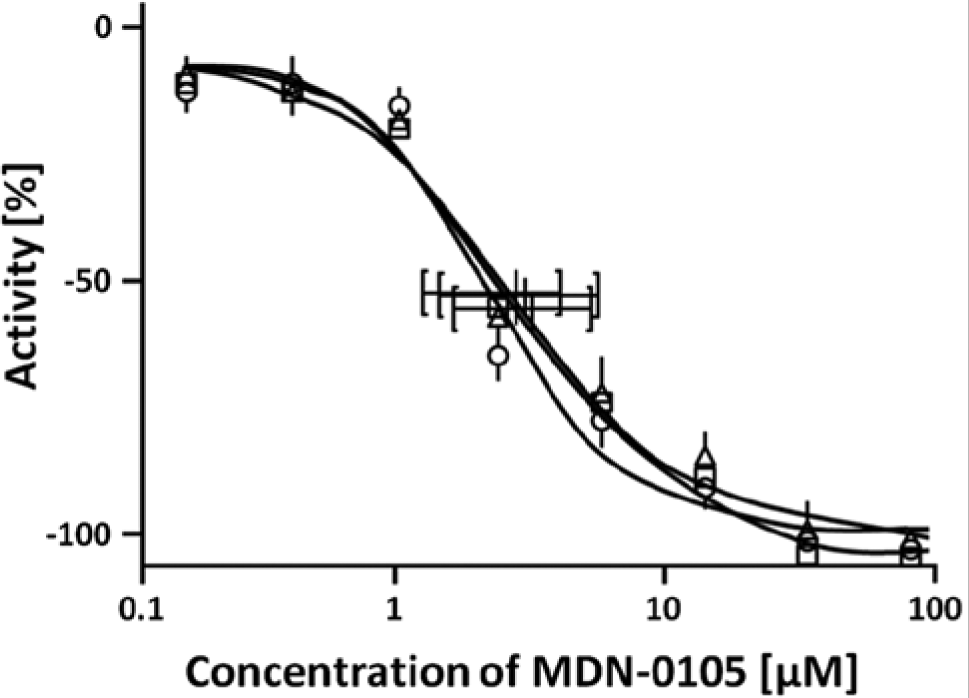
Dose response of the active compound inhibition of nuclear export measured using the U2nesRELOC assay system in the high-throughput screening format. Cells were seeded automatically at an appropriate density in 96-well black-wall clear-bottom tissue culture plates and allowed to attach overnight. Cells were then treated with different concentrations of leptomycin B (LMB) (100% inhibition control) or active compound for 1 h. Graphs represent the growth of the cells relative to cells treated only with carrier. Each curve is the average of triplicate samples performed independently. IC50 for each curve is, respectively, 3.512 µM, 3.002 µM, and 3.699 µM.
Here we described a newly purified natural compound (MDN-0105) that shows a nuclear export inhibition with an IC50 value of 3.4 µM ( Fig. 3 ).
MDN-0105 Inhibits the Nuclear Export of FOXO3a and Endogenous NF-κB2
We applied confocal microscopy to further characterize the effect of MDN-0105 on nuclear exports of proteins. The upper panel of Figure 4 (Rev-NES-EGFP) shows that the NES reporter protein was detected as nuclear speckles in the nucleus of U2nesRELOC cells treated with MDN-0105 for 1 h. To investigate if the effect of MDN-0105 is restricted to the NES reporter protein or also affects cellular proteins, we determined the subcellular localization of FOXO3a and NF-κB2 upon MDN-0105 treatment. FOXO3a and NF-κB2 are transcription factors known to be exported from the cell nucleus in a CRM1-dependent manner and have been shown to play a critical role in many human malignancies. 24 We analyzed the nuclear export of FOXO3a protein using the reporter cell line U2foxRELOC that stably expresses an EGFP-FOXO3a fusion protein.19,25 After treatment with MDN-0105 for 1 h, the fluorescently tagged reporter protein accumulated in the nucleus of U2foxRELOC cells ( Fig. 4 , EGFP-FOXO3a). To determine if this inhibition affects also the subcellular localization of endogenous proteins, we performed immunohistochemical detection of NF-κB2 protein using a specific NF-κB2 p100/p52 antibody. Figure 4 (Y NF-κB2) shows that MDN-0105 very efficiently inhibited the nuclear export of endogenous NF-κB2 transcription factor in U2OS cells. Taken together, these data indicate that MDN-0105 is a potent inhibitor of the nuclear export of cellular proteins involved in the formation and progression of human cancer. Complete characterization of this newly described compound and analysis of the mechanism of action are under evaluation.
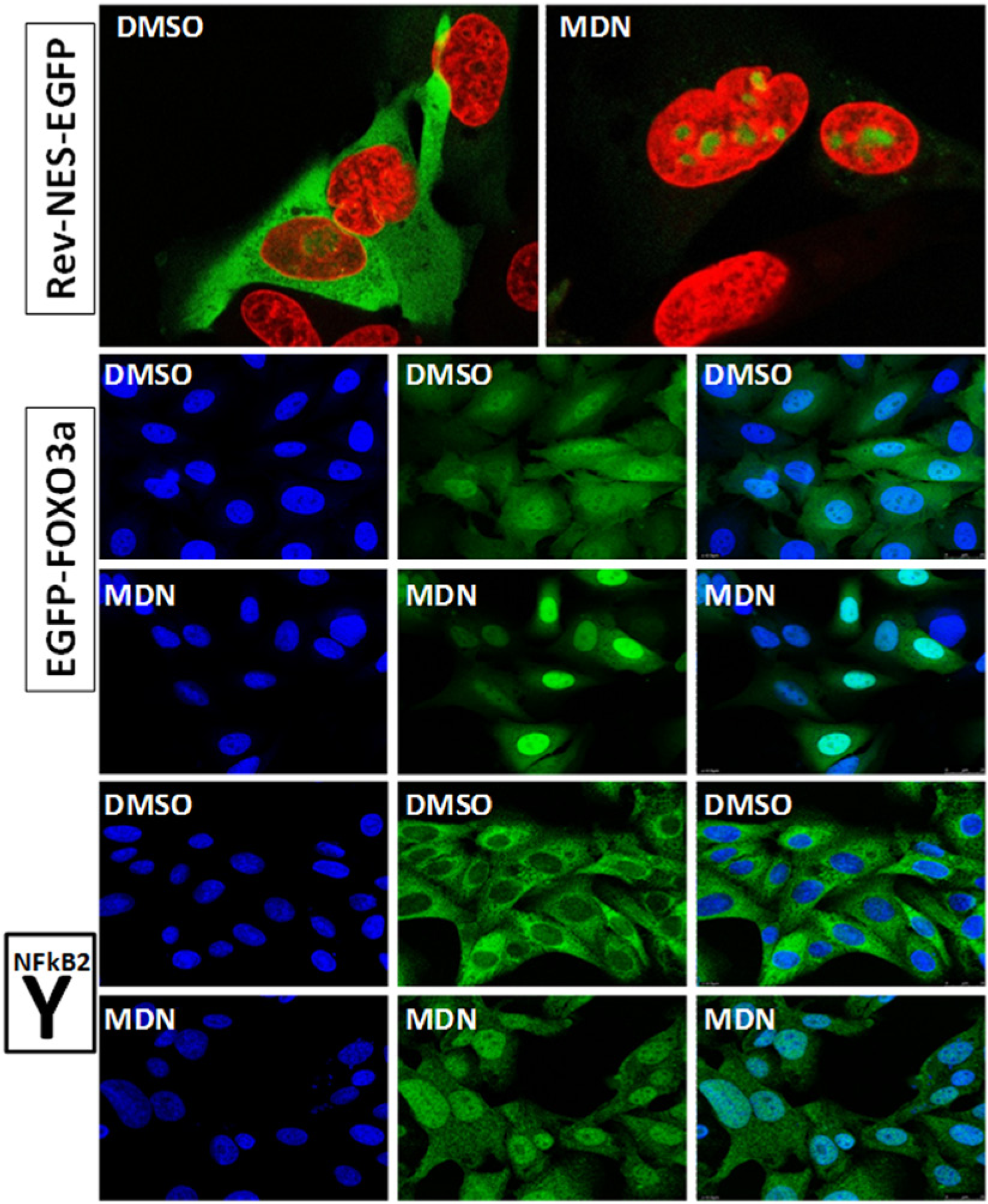
MDN-0105 inhibits nuclear export of reporter and cellular proteins. Representative images of the compound-treated cells using a Leica SPE confocal imaging system for nuclear export signal (NES) (Rev-NES-EGFP) and FOXO3a (EGFP-FOXO3a) reporter proteins and for endogenous nuclear factor (NF)–κB2 protein (Y). Cells were seeded automatically at appropriate density in 96-well black-wall clear-bottom tissue culture plates and allowed to attach overnight. Cells were then treated with MDN-0105 (MDN) for 1 h before paraformaldehyde (Rev-NES-EGFP and EGFP-FOXO3a) or methanol (NF-κB2) fixation and DAPI staining. The Rev-NES-EGFP panel presents a merge of DAPI (red) and green fluorescent protein (GFP) fluorescence (green) of U2nesRELOC cells after vehicle (DMSO) or MDN-0105 treatment. EGFP-FOXO3a and Y NF-κB2 present DAPI (blue), GFP (green), and merge, respectively, for EGFP-FOXO3a construct and ICC/IF for endogenous NF-κB2 (in U2OS wild-type cells) protein after vehicle (DMSO) or MDN-0105 treatment.
Discussion
The physical separation of the genome from the cytoplasm by the nuclear envelope (NE) is a hallmark of the eukaryotic cell and creates a requirement for the transport of macromolecules across the nuclear membrane as a fundamental aspect of normal cell function. The nucleocytoplasmic transport is a highly coordinated process and has been recognized as an important means of regulating cellular signaling. Spatial separation of proteins from their sites of function represents a recurrent motif to ensure the fidelity of signal transduction and to prevent spontaneous signal activation. Cancer cells use the aberrant localization of tumor suppressor proteins as a mechanism for their inactivation and to effectively evade the antineoplastic mechanism.11,13 In addition, the aberrant localization of oncoproteins can lead to their inadequate activation. 12 Therefore, the therapeutic manipulation of the nucleocytoplasmic shuttling of macromolecules has emerged as a promising approach to treat human diseases.
Natural products with industrial applications can be produced from primary or secondary metabolism of living organisms. Owing to technical improvements in screening programs, as well as separation and isolation techniques, the number of natural compounds discovered exceeds 1 million. 26 Of all the reported natural products, approximately 20% to 25% show biological activity, approximately 10% have been obtained from microbes, and of these, 45% are produced by actinomycetes, 38% by fungi, and 17% by unicellular bacteria. 26 The increasing role of microorganisms in the production of antibiotics and other drugs for treatment of serious diseases has been dramatic. Microbial metabolites are among the most important of the cancer chemotherapeutic agents. They started to appear around 1940 with the discovery of actinomycin, and since then, many compounds with anticancer properties have been isolated from natural sources. 27 More than 60% of the current compounds with antineoplasic activity were originally isolated as natural products or are their derivatives. 28 Although there are currently no inhibitors of the nuclear export in clinical trials, several different natural products 29 have been identified, and strategies are being developed to use them therapeutically.
In the present study, we established that U2nesRELOC, a quantitative cell-based high-content screening assay suitable for HTS, can be applied to perform a screening campaign with natural products of microbial origin. The assay strategy is based on a fluorescent reporter protein containing a heterologous NES that is efficiently exported from the nucleus into the cytoplasm. The reporter protein used in our screen was preselected with respect to the strength of the export signal based on the systematic comparison of Rev-type established by Henderson and Eleftheriou. 16 In untreated U2nesRELOC cells, the fluorescent signal is predominantly cytoplasmic. Compounds that interfere with the NES-mediated export lead to the nuclear accumulation of fluorescent reporter protein. Using robotic fluorescent microscopy and automated image analysis acquisition, the U2nesRELOC system monitors inhibition of nuclear export in a high-throughput manner. Applying this U2nesRELOC screening system to a small collection of microbial natural products extracts, we detected several positive extracts with nuclear export inhibition activity. It is interesting to note that extracts from fungi seem to present more putative active molecules than extracts from actinomycetes. Indeed, it has been suggested by others that metabolites from eukaryotes origin (fungi) present more activity against eukaryotic targets than metabolites from a prokaryotic origin.26,28 A high-content–based methodology was also used in further steps of the analysis of active fractions until a pure active compound, MDN-0105, was obtained. We showed that MDN-0105 inhibits the nuclear export of the reporter proteins Rev-NES-EGFP and EGFP-FOXO3a. Importantly, the endogenous NF-κB2 protein also accumulated in the cell nucleus after MDN-0105 treatment. The IC50 values for nuclear export inhibition of MND-0105 determined after 1-h drug exposure is 3.4 µM and can be compared with the IC50 of the reference compound LMB. LMB is a highly specific inhibitor of CRM1 with significant in vitro potency but limited in vivo efficacy due to toxicity.2,30,31 The potent activity of LMB stems from highly specific covalent binding to the nuclear export receptor CRM1, thereby blocking CRM1-mediated protein export from the nucleus.11–13 CRM1 belongs to the family of importin β-related nuclear transport receptors and is absolutely required for the nuclear export of a wide variety of cargo proteins, including many tumor suppressor and oncogenic proteins. The clinical development of LMB was discontinued due to the significant toxicity observed. 29 The relatively lower potency of MDN-0105 compared with the one of LMB can be an advantage if it is associated with decreased toxicity. Work is currently under way to investigate the mode of action of MDN-0105. It is particularly important to know whether the inhibition of nuclear export by MDN-0105 is mediated by covalent binding to CRM1. Reversible inhibition of CRM1 has been suggested as a strategy to limit the toxicity associated with the inhibition of the general nuclear export machinery. 13 Future studies will explore the therapeutic potential of rapid and/or prolonged inhibition of nuclear export by MDN-0105 in a broad panel of human tumor cell lines. In summary, our data demonstrate that U2nesRELOC is a sensitive and robust assay system that allows a rapid and simple screening of extracts containing complex mixtures of natural products. We showed the identification of a pure active compound that regulates the subcellular localization of proteins. Importantly, this method can be scaled up to an HTS assay format to test large collections of microbial natural product extracts.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the Fundación MEDINA, a public-private partnership of Merck Sharp & Dohme de España S.A./Universidad de Granada/Junta de Andalucía. In addition, the equipment used in this work was supported by MICINN (PCT-010000-2010-3).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
