Abstract
A fluorescent-based high-throughput screening (HTS) assay for small molecules that inhibit the interaction of MdmX with p53 was developed and applied to identify new inhibitors. The assay evaluated the MdmX-p53 interaction by detecting the quenching of the fluorescence of green fluorescent protein (GFP) fused to the MdmX protein, after its interaction with a p53 peptide labeled with a fluorescence quencher. In this report, the developed HTS assay was applied to about 40 000 compounds, and 255 hit compounds that abrogated the GFP quenching were selected. Next, the obtained hits were reevaluated by other assays. First, their effects on the diffusion time of a fluorescently-labeled p53 peptide after incubation with the MdmX protein were tested by measuring the diffusion time using fluorescence correlation spectroscopy, and six stable hit compounds with IC50 values less than 5 µM were selected. Next, we further confirmed their inhibition of the MdmX-p53 interaction by surface plasmon resonance. To indicate the efficacy of the hit compound as a candidate anticancer drug, we showed that the hit compound triggered apoptosis after p53 and p21 accumulation in cultured MV4;11 leukemia cells. Thus, the new HTS assay is effective for obtaining novel MdmX-p53 interaction inhibitors that are valuable as candidate compounds for cancer treatment.
Introduction
The tumor suppressor p53 family proteins are potent transcription factors that control a major pathway protecting cells from malignant transformation.1,2 Therefore, small molecules that activate the p53 pathway have been pursued by many groups over the past decade.3,4
The first drug target to activate the p53 pathway was Mdm2, which binds p53 and acts as its ubiquitin E3 ligase. 5 It keeps the p53 level very low because of rapid degradation through the ubiquitin-dependent protease system in unstressed, normal cells. 6 Nutlin-3, MI-219, and WK298 were discovered as potent small molecules that bind specifically to Mdm2 at the p53 interaction site and inhibit the Mdm2-p53 interaction.7–11 The compounds decrease the proteolysis of p53, increase the Mdm2 level, and trigger apoptosis in cultured cells with the normal p53 gene, such as SJSA-1 and MV4;11. They do not inhibit the MdmX-p53 interaction, and their combination with MdmX siRNA resulted in synergistic activation of p53 and growth arrest of cultured cancer cells.12–14
The second drug target, MdmX, has been attracting attention recently as a safe target for cancer therapy. 15 It shares significant sequence homology with Mdm2 and is another p53-binding protein inhibiting p53-dependent transcription.1,3 MdmX lacks intrinsic E3 ligase activity; however, it binds to Mdm2 through a C-terminal RING domain interaction, forms a supramolecular assembly, and stimulates the ability of Mdm2 to ubiquitinate and degrade p53.16–18 MdmX is overexpressed in many primary tumors or tumor cell lines with wild-type p53 19 and is believed to contribute to p53 inactivation during tumorigenesis. 20 In 2010, the first small-molecule inhibitor of MdmX, SJ-172550, was screened from 300 000 compounds by an HTS assay based on the fluorescence polarization measurement. 21 The compound binds reversibly to MdmX (EC50: 2.3 µM) and kills retinoblastoma-derived cells (EC50: 23 µM), in which the expression of MdmX is amplified. Unfortunately, SJ-172550 showed only weak inhibition of the MdmX-p53 interaction, and thus its effect on the p53 pathway was limited. In 2011, Wang et al. 22 described a new compound, XI-006, that inhibits MdmX expression in cancer cells and activates p53 pathways to increase the expression of proapoptotic genes and decrease the viability of cancer cells, thus clearly indicating the efficacy of inhibiting the MdmX-p53 interaction.
We now report a newly developed high-throughput screening (HTS) assay to screen for MdmX-p53 interaction inhibitors efficiently. We screened about 40 000 compounds by the new HTS assay, and evaluated the hit compounds by the fluorescence correlation spectroscopy (FCS)–based competitive binding assay 23 and the surface plasmon resonance (SPR)–based competitive binding assay. 5 We confirmed that the obtained hit potently increased the cellular level of p53 and downstream p21 and triggered apoptosis in cultured leukemia cells.
Materials and Methods
Preparation of MdmX(26-106), MdmX(26-106)-GFP, and Mdm2(17-124)
The DNA fragments encoding the SWIB domain of human MdmX(26-106; Uniprot accession No. O15151.2), Mdm2(17-124; Uniprot accession No. Q00987.1), and the green fluorescent protein (GFP)–fused MdmX(26-106) were cloned into the expression vector pCR2.1 (Invitrogen, San Diego, CA) as fusion fragments with an N-terminal native His affinity tag and a TEV protease cleavage site. The protein samples were synthesized by the cell-free expression system, 24 clarified by centrifugation at 16 000g for 20 min, and filtered through a 0.45 µm membrane (Millipore, Billerica, MA).
The protein samples were applied to a 5 mL HiTrap HP column (GE Healthcare, Uppsala, Sweden) equilibrated with 20 mM Tris-HCl buffer, pH 8.0, containing 100 mM NaCl and 20 mM imidazole, on an ÄKTA Explorer system (GE Healthcare). The protein samples were eluted with 20 mM Tris-HCl buffer, pH 8.0, containing 100 mM NaCl and 500 mM imidazole. After desalting on a HiPrep 26/10 desalting column with 20 mM Tris-HCl buffer, pH 8.0, containing 100 mM NaCl and 2 mM DTT, the eluted protein fraction was incubated at 4 °C overnight with TEV protease, to cleave the His-tag. The samples were passed through a HisTrap HP column and then desalted and concentrated with an Amicon Ultra-15 filtration device (Millipore).
Chemicals
Compounds were purchased from ASINEX Ltd. (Moscow, Russia), ChemBridge Corp. (San Diego, CA), and Maybridge Chemical Co., Ltd. (Cambridge, UK). The total set of 39 714 compounds has a mean molecular weight of 322.5 Da and an ALogP of 3.30. Nutlin-3 was purchased from Sigma-Aldrich. WK298 was synthesized according to Boettcher et al. 25 Black Hole Quencher Dye (BHQ-1) was purchased from Biosearch Technologies (Navato, CA). The compounds were dissolved in DMSO and stored at −20 °C as 10 mM stock solutions.
Fluorescent-Based HTS Assay
The interaction between purified MdmX-GFP and a BHQ-1–labeled p53 peptide (SQETFSDLWKLLPENSGSGK-BHQ-1: the p53-quencher probe), purchased from Toray Research Center (Tokyo, Japan), was analyzed. For the HTS analysis, the reaction mixture contained 100 nM MdmX-GFP and 1 µM of the p53-quencher probe, with or without a compound, in 10 mM HEPES-KOH buffer, pH 7.4, 150 mM NaCl, 0.005% Tween 20, and 1% DMSO. After an incubation at 25 °C for 4 h, the fluorescence of the fused GFP (ex. 485 nm/em. 535 nm) was measured, using Infinite 200 (Tecan Systems, Inc., San Jose, CA) or ARVO (PerkinElmer, Waltham, MA) in a 384-well plate.
FCS-Based Competitive Binding Assay
The FCS-based assay to detect protein-peptide interactions under solute conditions and to screen inhibitor compounds of the interaction was performed, as reported previously. 23 Briefly, the diffusion time was determined by an FCS measurement, and then the autocorrelation function was calculated from the fluorescence intensity fluctuations caused by fluorescent molecules diffusing in and out of the detection volume. FCS measurements were performed on an MF20 fluorescence detection system (Olympus, Japan), using a 543 nm He/Ne laser line for illumination. For each sample, at least five fluorescence time series of 10 s durations were recorded, and each autocorrelation function was calculated by the system.
The p53-derived peptides (SQETFSDLWKLLPEN: the p53-peptide, and SQETFSDLWKLLPENK-TAMRA: the p53-probe) were purchased from Toray Research Center. The peptides were diluted to a final concentration of 1 nM in 10 mM HEPES-KOH buffer, pH 7.4, containing 150 mM NaCl and 0.005% (v/v) Tween 20, to suppress glass surface interactions.
The probe peptide, the Mdm protein, and the compound were mixed and incubated at room temperature for more than 4 h before the FCS measurements. All FCS measurements were recorded at room temperature. To analyze the interaction between a probe peptide and the Mdm protein, we subjected the lateral diffusion time (τD) data to a two-component analysis by the MF20 and obtained the contribution percentages of the τD of the free probe peptide and the τD of the interacting probe peptide.
SPR-Based Competitive Binding Assay 5
Competitive binding experiments were performed at 25 °C using a BIACORE 3000 (GE Healthcare) SPR instrument. The biotinylated p53 N-terminal peptide (Biotin-CGCGSQETFSDLWKLLPEN), purchased from Toray Research Center, was attached to an avidin-coated gold surface (Sensor Chip SA; GE Healthcare). The peptide was immobilized to levels of approximately 400 to 600 RU in each flow cell.
The Mdm2 or MdmX protein (final concentration of 100 nM) was incubated with each of the inhibitor compounds identified in the first screen, in 10 mM HEPES-KOH buffer, pH7.4, 150 mM NaCl, 0.005% Tween 20, and 0.5% DMSO, for 4 h and then injected into the instrument. The binding of Mdm2 or MdmX to the immobilized p53 peptide on the SA Sensor Chip was monitored at a flow rate of 30 µL/min. In the absence of an inhibitor, the Mdm2 and MdmX proteins were captured at levels of approximately 170 to 280 RU. The binding levels of Mdm2 or MdmX were determined from the differences in the SPR signals between the flow cells immobilized with the p53 peptide and the reference cell. Additional instrumental contributions to the signal were removed by subtraction of the average signal of the blank injections from the reference-subtracted signal.
Cell Lines and Culture Conditions
The leukemia cell line MV4;11 (myelomonocytic leukemia, Nutlin-3 sensitive) and H1299 (non–small-cell lung cancer: p53 null) were obtained from the American Type Culture Collection (Rockville, MD). MV4;11 was maintained in Iscove’s modified Dulbecco’s medium (GIBCO, Carlsbad, CA), with 10% heat-inactivated fetal bovine serum (FBS; Invitrogen) and penicillin (5000 U/mL)–streptomycin (5 mg/mL; Sigma-Aldrich, St. Louis, MO), at 37 °C in 5% CO2. H1299 was maintained in RPMI1640 medium (Invitrogen) with 1 mM sodium pyruvate (Invitrogen) and 1/50 diluted penicillin (5000 U/mL)–streptomycin (5 mg/mL; Sigma-Aldrich). The normal human fetal lung fibroblast cell line WI-38 was purchased from the RIKEN Cell Bank (Ibaraki, Japan) and cultured in Minimum Essential medium (GIBCO) with 10% FBS and penicillin (5000 U/mL)–streptomycin (5 mg/mL), at 37 °C in 5% CO2.
After 24 h, various concentrations of the test compounds were added to the seeded culture wells. The cells were cultured further for 48 h, and then the numbers of surviving cells were determined by the Alamar Blue assay (Alamar Biosciences, Inc., Sacramento, CA).
Apoptosis was detected with ApoAlert AnnexinV (Clontech, Mountain View, CA). Cells treated with chemicals for 48 h were harvested and stained with FITC-conjugated Annexin V and propidium iodide, according to the manufacturer’s instructions. The stained cells were analyzed by an FACS Caliber, using the CellQuest software (Becton Dickinson, Franklin Lakes, NJ). The percentage of Annexin V–positive cells was quantified using the Flow Jo software (Becton Dickinson).
Cell Lysate Preparation and Western Blotting
The cells were inoculated and treated with or without compounds at each GI50 concentration for 5 h or overnight. The treated cells were harvested and rinsed with ice-cold phosphate-buffered saline. Ice-cold lysis buffer (50 mM Tris-HCl, pH 8.0, 5 mM EDTA, 150 mM NaCl, 0.5% Nonidet P-40, 1 mM phenylmethylsulfonyl fluoride) was added to the cell pellet, and the mixture was incubated on ice for 30 min, followed by centrifugation at 10 000g for 5 min. The protein concentrations of the lysates were determined by the Bradford method (BIO-RAD, Hercules, CA), and the lysates (10–50 µg of protein) were fractionated by SDS-PAGE and transferred to PVDF membranes (Millipore). The membranes were blotted with each primary antibody and developed using horseradish peroxidase–conjugated secondary antibodies and the ECL-Advance reagent (GE Healthcare). The primary antibodies were the following: DO-1 for p53 (Santa Cruz Biotechnology, Inc., Santa Cruz, CA), OP64 for p21 (Calbiochem, Billerica, MA), and an anti-actin antibody (Cell Signaling Technology, Danvers, MA). The goat anti-mouse IgG-HRP (sc2005) was purchased from Santa Cruz Biotechnology, Inc.
Results and Discussion
Fluorescent-Based HTS Assay for MdmX-p53 Interaction Inhibitors
We developed a new HTS method to screen for protein-protein interaction inhibitors, which is easily applicable to screen inhibitors against other protein-protein interactions. The MdmX SWIB domain fused with GFP (MdmX-GFP) and the p53-derived peptide labeled with BHQ-1 (the p53-quencher probe) were mixed in buffer. The extent of the interaction between the two molecules was detected by measuring the decreased intensity of fluorescence from the fused GFP ( Fig. 1a ). The enhanced interaction between MdmX-GFP (100 nM) and the p53-quencher probe with increasing amounts of added p53-quencher probe decreased the measured fluorescence intensity of MdmX-GFP ( Fig. 1b ). MdmX-GFP (100 nM), the p53-quencher probe (1 µM), and a 0.1 to 10 µM concentration of the p53-competitor probe were mixed, and after a 4 h incubation, their fluorescence intensities were measured. We confirmed that the p53-MdmX interaction was inhibited by the p53 peptide added to the mixture by the recovery of GFP fluorescence intensity in the assay mixture. The binding activity of the p53-quencher probe to the MdmX-GFP with the p53 peptide was calculated as the percentage ratio of quenching intensity in comparison with that without competitor ( Fig. 1c ). Figure 1c shows that the IC50 value of the competitive p53 peptide was 3.0 µM. The time-dependent changes of the fluorescent intensity with and without the inhibitors were examined. The fluorescent intensities measured after 1 to 5 h after the addition were constant, indicating that they were in equilibrium (data not shown).
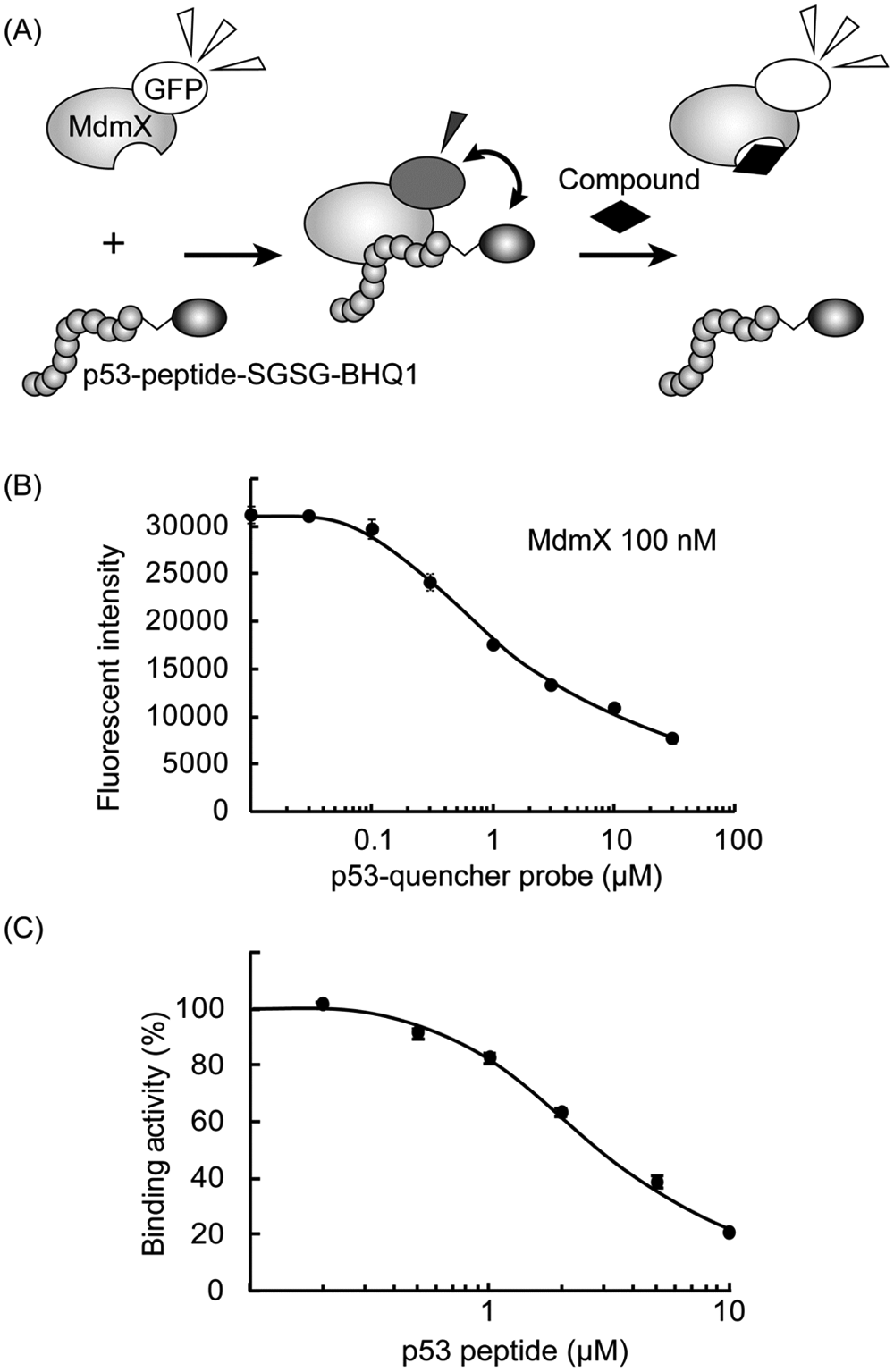
Fluorescent-based high-throughput screening assay for MdmX-p53 interaction inhibitors. (
We screened 39 714 commercially available compounds by this HTS method, using mixtures of 100 nM MdmX-GFP, 1 µM p53-quencher probe, and 20 µM compounds in 10 mM HEPES-KOH buffer, pH 7.4, containing 150 mM NaCl and 0.005% Tween 20. The mean Z′ value obtained per a 384-well plate in this screen was 0.73 ± 0.076. As a result, 435 compounds inhibited the interaction and weakened the fluorescent quenching by less than 30%. We omitted the compounds that increased the fluorescent intensities of the assay mixture by more than the mean intensities of the added MdmX-GFP plus three times its standard deviation, and thus 255 compounds were obtained as the hit compounds.
FCS-Based Assay for Hit Evaluation
To evaluate the obtained HTS hit compounds, we performed the FCS-based competitive binding assay. 23 As model systems of the interactions between the full-length p53 and Mdm proteins, we measured the p53-probe binding to the MdmX or Mdm2 SWIB domain. The protein-peptide interactions were detected as the increased τD of the p53 probe. The obtained τD values were fitted by the MF20 two-component analysis tool, and the percentage contributions of τD of the free p53 probe and τD of the p53 probe binding to the target protein were calculated. Obtained KD values were 60 nM for the Mdm2-probe interaction and 200 nM for the MdmX-probe interaction, respectively ( Figs. 2a , b ). Next, we measured the effect of Nutlin-3, as a positive compound that inhibits the interactions. The addition of a compound that competitively binds to the Mdm protein decreases the percentage contributions of the p53 probe binding to the Mdm protein, which can be detected as the decrease in the τD value of the p53 probe, as measured by FCS. In the assay, we applied the Mdm2 and MdmX proteins with concentrations of twice the KD values, as reported. 9 The IC50 values of Nutlin-3 on the Mdm2 and MdmX interactions determined by the FCS-based competitive assay, were 70 nM and 50 µM, respectively ( Fig. 2c ). These values are comparable to the results obtained previously by fluorescence polarization assays. 9
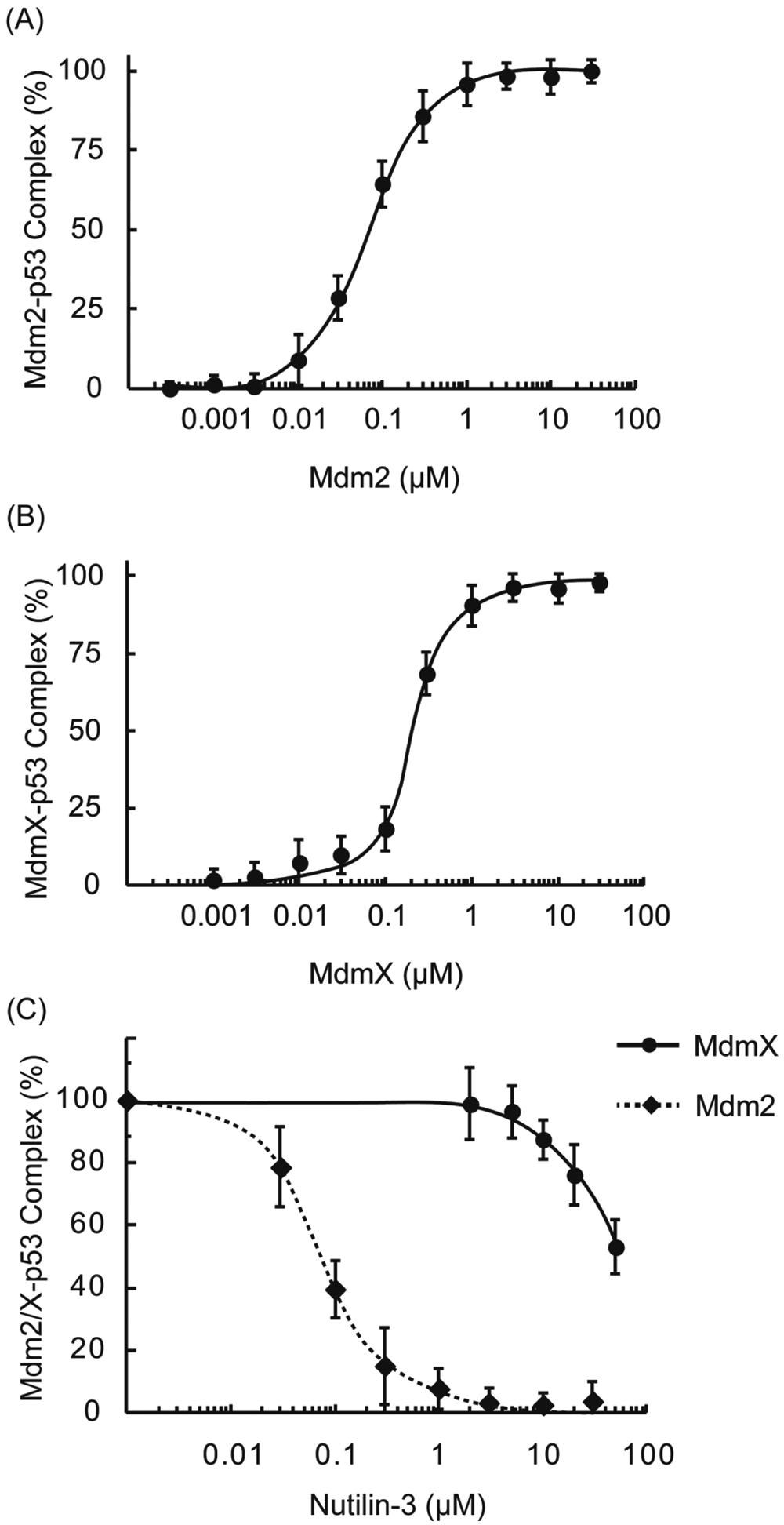
Fluorescence correlation spectroscopy–based competitive binding assay. (
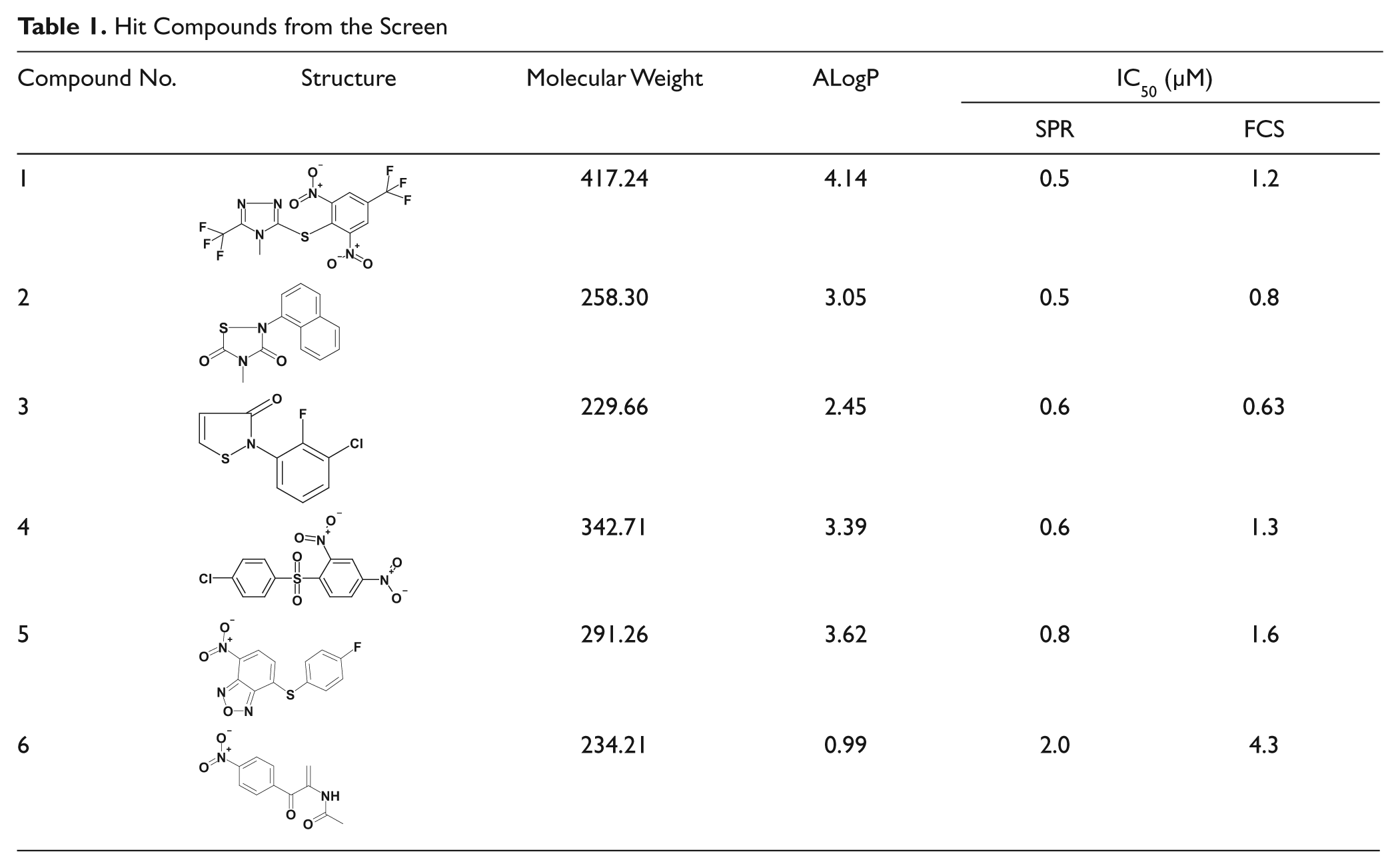
We further evaluated the hit compounds by the FCS-based competitive binding assay and selected eight hit compounds with IC50 values less than 5 µM. We measured their inhibitory activities after a 24 h incubation at 37 °C in the FCS assay mixture and selected six stable compounds that exhibited more than two-thirds of the initial inhibitory activity without incubation ( Table 1 ). To validate the obtained hits further, we performed the SPR-based competitive binding assay and confirmed their activities ( Table 1 ). The two IC50 values determined for each compound had an almost linear relationship, and those determined by the SPR-based assay exhibited lower values.
Hit Compounds from the Screen
Cell-Based Assay for Hit Evaluation
We tested the cytotoxic effects of the compounds on normal human cultured cells, WI-38, and Nutlin-sensitive and p53-normal leukemia cells, MV4;11, 26 after a 48 h co-culture. All compounds except one exhibited potent leukemia cell–specific killing activities, although compound 5 exhibited strong activity even on normal cells.
We selected compound 1 and compared it with Nutlin-3 and WK298. We measured their inhibition of the MdmX-p53 and Mdm2-p53 interactions by the FCS-based and SPR-based competitive binding assays. The results (summarized in Table 2 ) showed that compound 1 exhibited specific inhibition of the MdmX-p53 interaction. We tested the cytotoxic effects of the compounds on WI-38, MV4;11, and p53-deficient H1299 cells. The cells were co-cultured with various concentrations of the compounds for 48 h, and their viability was measured. Although the compounds showed only weak toxicity on WI-38, the GI50 values for compound 1, Nutlin-3, and WK298 on MV4;11 were 1.6, 2.2, and 1.9 µM, respectively ( Table 2 ). Compound 1 exhibited a remarkable effect on leukemia cells, similar to those of the known Mdm2-p53 inhibitors. 26 Moreover, compound 1 induced moderate cell death (GI50: 5.5 µM) in H1299 cells as reported before with an MdmX inhibitor, XI-006. 22 It is plausible that the p53 binding site of MdmX is known to interact with p63 and p73 that can alternatively activate proapoptotic genes in the absence of functional p53. 27 To test the cytotoxicity, MV4;11 cells, co-cultured with 1.6 µM compound 1 for 48 h, were harvested, stained with FITC-conjugated Annexin V and propidium iodide, and analyzed using a flow cytometer. Almost all of the cells stained with propidium iodide were Annexin V positive, which indicated that the observed cell death was caused by apoptosis, as reported in the case of XI-006 22 (data not shown).
Characterization of Compound 1
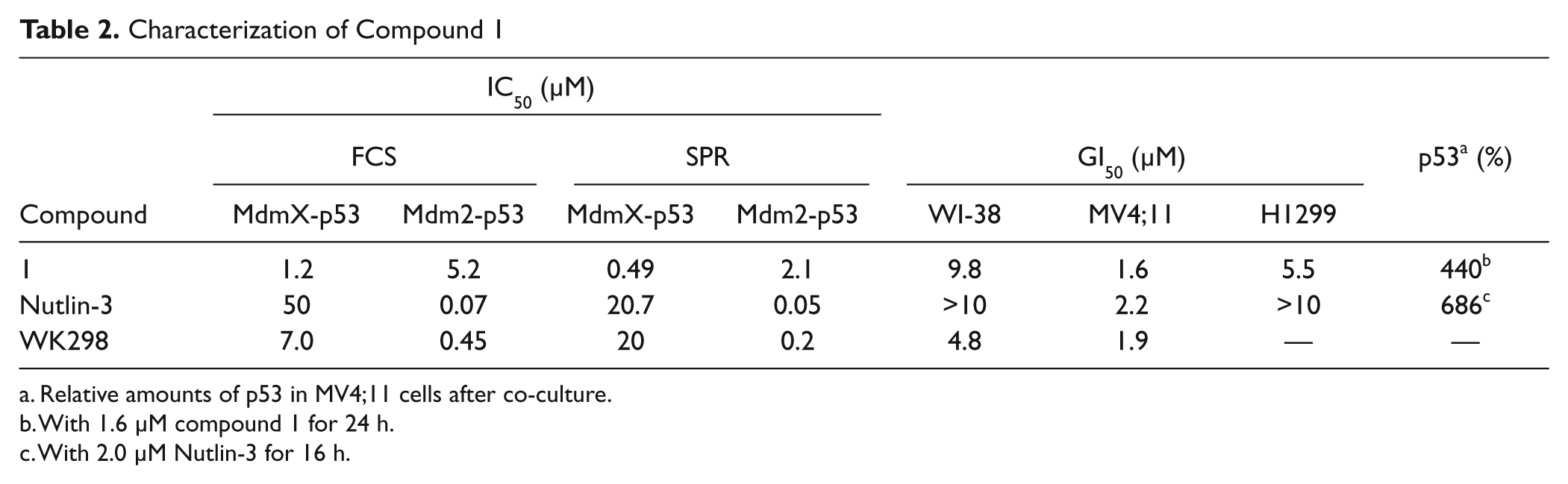
Relative amounts of p53 in MV4;11 cells after co-culture.
With 1.6 µM compound 1 for 24 h.
With 2.0 µM Nutlin-3 for 16 h.
We cultured MV4;11 cells with compound 1 and Nutlin-3 overnight and studied their effects on the cellular amounts of p53 and p21, a critical p53 target, by Western blotting. Compound 1 strongly increased the amounts of p53 (4.4-fold), as also observed with Nutlin-3 (6.9-fold;
Table 2
;
Through the HTS screening by the fluorescence polarization assay from 300 000 compounds, Reed et al. selected representative 11 hit compounds. 21 They reported that half of them were unstable in a buffer solution and some of them covalently modified the MdmX protein. In 2012, they published that SJ172550 alkylated Cys76 of MdmX protein to block p53 binding. 28 As concerns our hit compounds, compounds 2 and 3 have typical structures to bind proteins covalently. We also detected their irreversible inhibition after centrifugal filtration to wash out the compounds from treated MdmX protein. These results indicated that screening a large-scale compound library is required to obtain a lead compound with more specificity and efficacy for further development. Our newly developed HTS assay to screen for MdmX-p53 interaction inhibitors with a high Z′ value will be useful for the future large-scale screening.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by the Development of Fundamental Technology for Protein Analyses and the Targeted Protein Research Programs; from the Ministry of Education, Culture, Sports, Science and Technology of Japan; and the RIKEN Program for Drug Discovery and Medical Technology Platforms.
