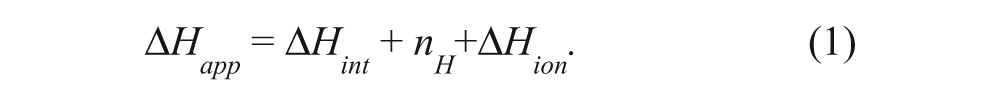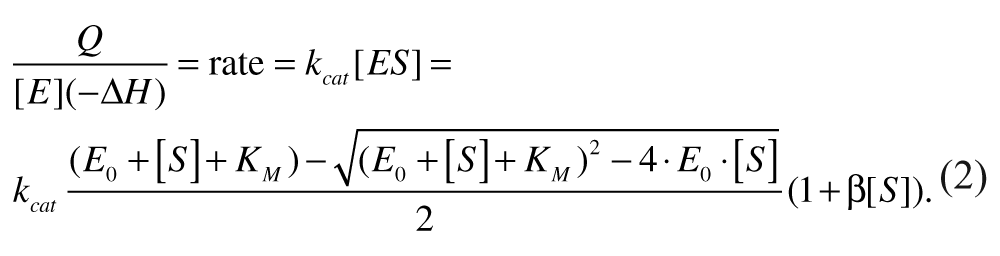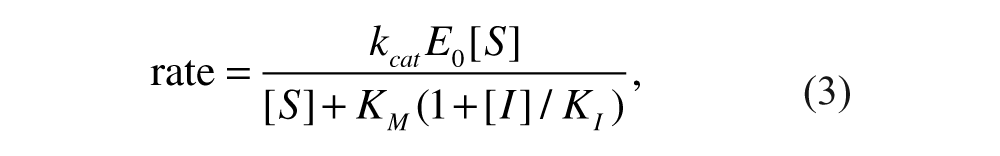Abstract
Fragment-based screening has typically relied on X-ray or nuclear magnetic resonance methods to identify low-affinity ligands that bind to therapeutic targets. These techniques are expensive in terms of material and time, so it useful to have a higher throughput method to reliably prescreen a fragment library to identify a subset of compounds for structural analysis. Calorimetry provides a label-free method to assay binding and enzymatic activity that is unaffected by the spectroscopic properties of the sample. Conventional microcalorimetry is hampered by requiring large quantities of reagents and long measurement times. Nanocalorimeters can overcome these limitations of conventional isothermal titration calorimetry. Here we have used enthalpy arrays, which are arrays of nanocalorimeters, to perform an enzyme activity-based fragment screen for competitive inhibitors of phosphodiesterase 4A (PDE4A). Several inhibitors with K I <2 mM were identified and moved to X-ray crystallization trials. Although the co-crystals did not yield high-resolution data, evidence of binding was observed, and the chemical structures of the hits were consistent with motifs of known PDE4 inhibitors. This study shows how array calorimetry can be used as a prescreening method for fragment-based lead discovery with enzyme targets and provides a list of candidate fragments for inhibition of PDE4A.
Keywords
Introduction
In fragment-based lead discovery (FBLD), small, low-complexity chemical fragments of 6 to 15 heavy atoms are screened for inhibiting the target. Hits are then linked and/or elaborated into tightly binding ligands, ideally yielding early lead compounds for drug discovery.1-3 Because of the simplicity of fragments, hit rates are high, typically 3% to 4%, so much smaller libraries are required than for high-throughput screening (HTS). 4 Because the average molecular weight of fragments is typically lower than for HTS, FBLD tends to identify weakly binding ligands with Kd in the ~100 µM to mM range. Because of this, ligand efficiency (LE 5 ) is often used to assess the binding of fragments rather than the binding affinity per se. This normalizes binding strength with compound size and better quantifies the effectiveness of a fragment as a building block for synthesizing tightly binding compounds. Ligand efficiency is defined as LE = –ΔG0/# heavy atoms = –RTln(K d )/# heavy atoms and is equivalent to the free energy of binding per heavy (i.e., nonhydrogen) atom. In practice, IC50measurements are often used as an approximation for K d . During fragment optimization, the expectation is that the ligand efficiency will be maintained, yielding a high affinity yet sufficiently small lead compound. Generally, ligand efficiencies of LE >0.3 kcal/mole/heavy atom are considered favorable. 5
Because hits arising from FBLD are usually low affinity, sensitive detection methods are required. A common approach to solving this problem has been to use X-ray 6 or nuclear magnetic resonance (NMR) 7 as the assay. These techniques have led to the identification of numerous potent and specific inhibitors and several drug candidates in clinical trials. 8 However, these techniques are expensive in terms of protein required, the time and expense of preparing either crystals for X-ray analysis or labeled protein for 2D NMR, the time needed to collect the massive amount of data necessary for analysis and structure determination, and equipment cost. Hence, for fragment screening, it is useful to have a high-throughput assay that can reliably detect ligand binding in the µM to mM range to be used as a prescreen for these more intensive methods.
Another approach, Tethering, 9 uses mass spectrometry to identify ligands that interact with a protein target at a specific site. The technique relies on the formation of a disulfide bond between the fragment and a cysteine residue (either native or engineered) in the region of interest of the target protein. The Tethering method provides some information about the binding mode of the fragment but requires moderate quantities of protein (0.5–5 mg) and specially synthesized fragment libraries that contain a disulfide moiety. 2
An alternative to the structure-based approaches is to use fragment screening methods based on high-concentration screening using HTS assays.10,11 The advantages of this approach are low protein requirements, high throughput, and cost savings for instrumentation. The major limitations with this approach include the large number of false positives due to compound interference or aggregation and the lack of information on binding motif or geometry. 2 In addition, a functional assay is required to initiate a screen with this method.
More recently, the use of surface plasmon resonance (SPR) methods to screen fragment libraries has been described.12,13 These assays are label free, so they do not suffer from the same limitations as standard HTS assays. Two limitations of the SPR method are that the assay is restricted to detection of binding, rather than activity-based measurements, and the binding affinity that can be detected is related to the molecular weight (MW) of the target protein. 12 Specifically, binding of a small fragment to a large protein will produce a smaller signal than binding of the same fragment to a smaller protein. Practically, this means that the affinity limit for larger proteins is lower than from smaller proteins. When properly executed, SPR assays can identify compounds that aggregate or bind promiscuously so that they can be removed from the library. 14
Isothermal titration calorimetry (ITC) directly measures the heat released or absorbed during a biomolecular reaction, and binding parameters determined by ITC are often referred to as the “gold-standard” values. 15 Because the measurement is label free and requires no immobilization of either the target or ligand, calorimetry can be used to perform a direct enzyme activity assay without modifying the substrate or using coupled reactions to produce the observed signal. 16 In addition, a calorimetric measurement is unaffected by spectroscopically opaque solutions that may occur with high concentrations of substrate, product, or fragments. Although the use of ITC to determine enzyme kinetics and inhibition has been described previously,16–18 the measurements are performed one at a time and require quantities of reactants that are considered quite large in biochemical studies, making high-throughput measurements or measurements with limited amounts of material unfeasible. Nanocalorimeters can in principle overcome these limitations of conventional ITC.
Enthalpy arrays are arrays of nanocalorimeters that allow scientists to measure thermodynamics and kinetics of molecular interactions using small sample volumes (250 nL) and short measurement times (typically 5–10 min). 19 Previously, we demonstrated that enthalpy arrays can be used to determine the kinetic parameters for enzymatic reactions and the mechanism of action of inhibitors. 20
In this study, we use enthalpy arrays in activity-based screening of fragments for discovery of inhibitors of phosphodiesterase 4A. Phosphodiesterase 4 (PDE4), which consists of four subfamilies (A–D), is a cAMP-specific phosphodiesterase for which selective inhibitors have been studied as anti-inflammatory therapeutics for the treatment of asthma, chronic obstructive pulmonary disease (COPD), and rheumatoid arthritis. 21 Although an array of inflammatory diseases might be treated with PDE4 inhibitors, narrow therapeutic windows have limited their clinical use. 21 To determine the utility of nanocalorimetry as a prescreening method in FBLD, we screened a 160-compound fragment library for competitive inhibitors of PDE4A.
Methods
PDE4A10 Cloning, Expression, and Purification
The catalytic domain of PDE4A10 (EC 3.1.4.53) was expressed and purified as described by Wang et al. 22 with modification. Because all isoforms of PDE4A have the same sequence in the catalytic domain, we will refer to the enzyme as PDE4A from this point forward.
PDE4A was cloned into a proprietary vector containing an N-terminal 6Xhis tag, cleavable by TEV protease. Positively identified clones were expressed in BL21(DE3)RIL cells. Cells containing target plasmid were grown in LB containing selection antibiotics at 37 °C until OD600 = 0.7–0.8. Cells were induced with 0.1 mM isopropyl-β-D-thiogalactopyranoside (IPTG) at 15 °C overnight. The cells were harvested and stored at −80 °C. Cell pellets were lysed in 50 mM Tris (pH 8.0), 500 mM NaCl, 0.1% (w/v) NP-40, and 10 mM imidazole (lysis buffer). The suspension was sonicated on ice for 2 min at 70% output with a 25% duty cycle. Clarified supernatant was loaded onto Ni2+-charged IMAC. Peak fractions containing target protein were cleaved overnight with TEV protease. Cleaved protein was isolated by running the sample over Ni2+ charged IMAC, collecting flow-through. Protein was characterized for relative oligomeric state by size exclusion chromatography in 20 mM HEPES (pH 7.5), 150 mM NaCl, and 5 mM dithiothreitol (DTT). Monomeric PDE4A was concentrated to 34 mg/mL.
Fragment Library
Select compounds from a commercial fragment library, ActiveSight Library 2, (ActiveSight, San Diego, CA) were used in this study. For the initial screen, individual compounds dissolved at 200 mM in DMSO and stored in 96-well plates were used. The average molecular weight of the compounds was 154.1, and the average number of heavy atoms was 10.4. Hits were verified by preparing fresh solutions from dry powder.
PDE4A Calorimetric Activity Assay
Hydrolysis of 3′,5′-cAMP was measured at 21 °C in 100 mM Tris-HCl (pH 7.5), 10 mM MgCl2, 1 mM tris(2-carboxyethyl)phosphine (TCEP), and 2% (v/v) DMSO. Each enthalpy array detector contains a sample region and a reference region, which are designated based on the material deposited on the region (see Fig. 1 of Recht et al. 20 ). The sample region materials consisted of a drop of PDE4A (10 µM) and a drop of substrate solution (4 mM 3′,5′-cAMP). The reference region used a drop of bovine serum albumin (BSA; 0.02 mg/mL, 0.30 µM) and a drop of the same substrate solution (4 mM 3′,5′-cAMP) used in the sample region. The purpose of the small amount of BSA is to give the reference drops wetting behavior similar to the sample drops, especially after merging. Immediately after merging, the combined drops in the sample region contained 5 µM PDE4A and 2.0 mM 3′,5′-cAMP, and the combined drops in the reference region contained 0.15 µM BSA and 2.0 mM 3′,5′-cAMP. Reactions with compounds from the fragment library (2 mM in the combined drops, except as noted) were performed as above, except that a single compound (4 mM, except as noted) was included with the substrate (4 mM 3′,5′-cAMP). Adenosine 3′,5′-cyclic monophosphate, pentoxifylline, and 3-isobutyl-1-methylxanthine (IBMX) were obtained from Sigma-Aldrich (St. Louis, MO) and used without further purification. The concentration of the 3′,5′-cAMP stock solution (adjusted to pH 7 with dilute NaOH) was determined by measuring the absorbance at 258 nm and using an extinction coefficient of 14 650 M−1 cm−1. 23
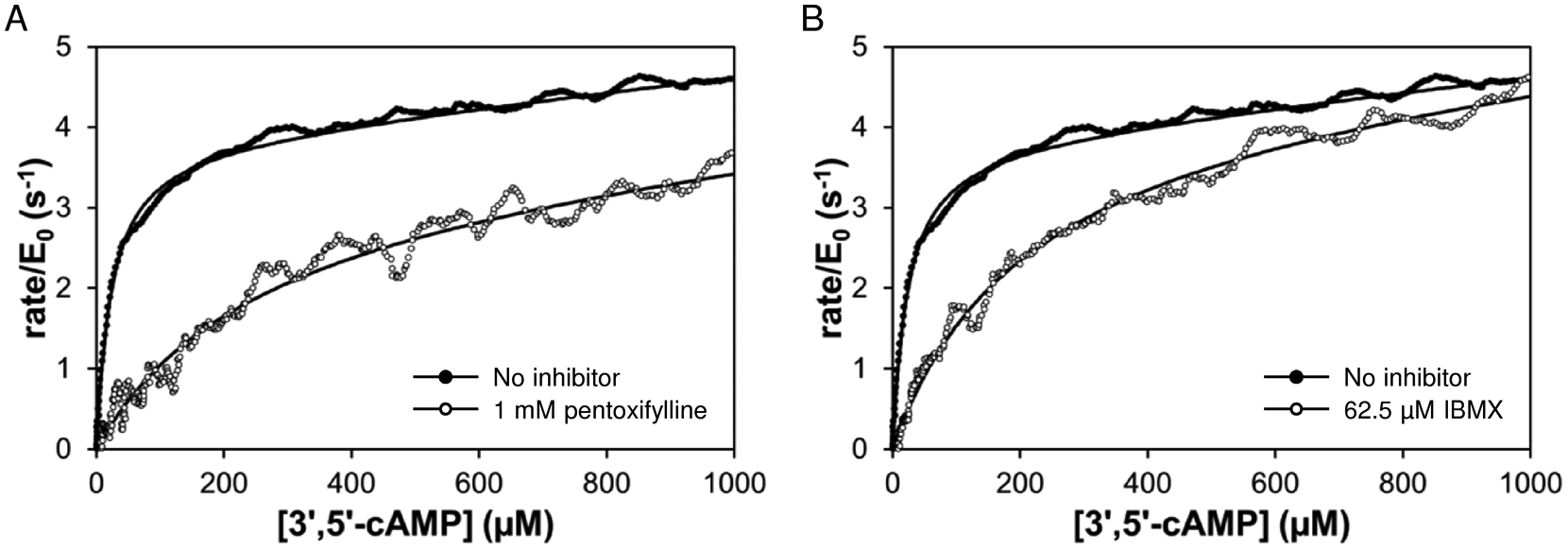
PDE4A hydrolysis of 3′,5′-cAMP in the absence (solid) and presence (open) of known general phosphodiesterase inhibitors. Reactions contained 5 µM PDE4A and 2 mM 3′,5′-cAMP. Solid curves are fit of data to equation 2. (
We used an enzyme concentration that would result in a good signal-to-noise ratio at Vmax, and the substrate concentration was then adjusted to produce the length of reaction desired. Typically, three replicates of each measurement were performed, and the average and standard error of the mean of the measurements are reported. Measurements with inhibitors had a corresponding set of control measurements without inhibitor performed at the same time.
The reactions were performed in Tris buffer to take advantage of the additional heat arising from protonation of the buffer by hydrogen ion release that accompanies hydrolysis of 3′,5′-cAMP to 5′-AMP (nH+ ≈ 0.86 per cAMP hydrolyzed at pH 7.3 24 ). The apparent enthalpy, ΔHapp, measured by calorimetry comprises the intrinsic enthalpy of the reaction (ΔHint) and ionization of the buffer (ΔHion) 25 so that
In Tris buffer, nH+ΔHion contributed an additional −9.75 kcal/mol, effectively increasing the signal 1.6-fold compared to the same reaction performed in phosphate buffer. 24
Follow-up studies on the inhibition mechanism of compounds 104 and 152 were performed using a Microcal iTC200 (MicroCal/GE Healthcare, Piscataway, NJ). The reactions were performed at 25 °C in 100 mM Tris-HCl (pH 7.5), 10 mM MgCl2, 1 mM TCEP, and 2% (v/v) DMSO and contained 47 nM PDE4A, 560 µM 3′,5′-cAMP, and compound at concentrations between 233 and 3720 µM. Control reactions in the absence of inhibitor yielded kinetic parameters similar to the enthalpy array measurements.
Data Analysis
For each measurement, we recorded the differential temperature as a function of time. This information was converted into an enthalpy by deconvolving the rate of heat generation (Q(τ)), which was subsequently transformed into the reaction velocity as a function of substrate as described in Recht et al. 20
One can determine kcat and K M for experiments with no inhibitor by fitting the data for Q/([E]ΔH) versus [S] to
In this equation, Q is the rate of heat generation, kcat is the turnover number, E0 is the total enzyme concentration, [S] is the substrate concentration, ΔH is the enthalpy per mole of substrate reacted, and β is the slope for Q versus [S] at [S] greater than K M . 20 The (1 + β[S]) term modifies the general rate equation. As a practical matter, the modification becomes significant at [S] » K M for the data we present below or, in other words, (1/β) » K M . Thus, it is clearly distinguishable from the effect of K M . The general rate equation was used because the concentration of enzyme in the reaction was high relative to K M (E0/K M > 0.1), and depletion of substrate from solution becomes significant under these conditions.26,27 We used the MATLAB function lsqcurvefit to perform a regression on equation (2), allowing for the possibility that the values of [S] may be off by a small constant in the regression, which helps to accommodate for noise near the [S] = 0 limit of the data.
We also examined inhibited reactions. We were able to observe a change in the apparent K M caused by a competitive inhibitor, allowing us to determine K I . To understand the meaning of an increase in apparent K M , it is useful to examine the Michaelis-Menten equation modified for competitive inhibition (equation 3.32 in Fersht 28 ):
in which [I] is the concentration of free inhibitor. The apparent K M is seen to be the true K M multiplied by (1 + [I]/K I ). At inhibitor concentrations [I] > K I , the shift in apparent K M becomes significant. When [I] >> E0 holds, as it does for all reactions with fragments here, the concentration of free inhibitor [I] is close to the total concentration of inhibitor I0, making it reasonable to use I0 in the above equation in place of [I], the standard practice in enzymology.
Protein Crystallization, Data Collection, and Structure Determination
For co-crystallization studies, PDE4A was incubated with 5 mM pentoxifylline on ice for 1 h. Crystals of PDE4 in complex with 5 mM pentoxifylline were obtained using the hanging-drop vapor diffusion method by mixing 2 µL of 34 mg/mL protein solution (in HEPES [pH 7.5], 150 mM NaCl, and 5 mM DTT) with 2 µL of 1.5 M ammonium sulfate and 0.1 M Bis/Tris propane (pH 7.0) at 20 °C. Diffracting crystals appeared within 3 to 5 days and grew to 0.4 mm in length. Prior to data collection, crystals were transferred into a cryoprotectant solution consisting of 25% (v/v) glycerol in crystallization buffer and then flash-frozen in liquid nitrogen.
Diffraction image data were collected at the Advanced Photon Source on beam line 21ID. Image data for the crystal containing pentoxifylline were processed using HKL2000. 29 The structure was solved by molecular replacement using the MOLREP program from the CCP4 program suite. 30 The protein component of an isomorphous crystal structure of PDE4, entry 3I8V from the Protein Data Bank (R. K. Y. Cheng, L. Crawley, J. Barker, M. Wood, B. Felicetti, and M. Whittaker, PDB ID: 3I8V), was used as the molecular replacement search model. Minimal refitting with the MIFit program 31 and refinement with the REFMAC5 program 32 were required to bring this model into good agreement with the data ( Table 1 ). Density corresponding to the pentoxifylline ligand was clearly visible in the PDE4A active site in protein copy A with a very evident “tail density” indicating the binding orientation. A somewhat less well-defined density is also present in protein copy B but with sufficient indications of tail density to show that the pentoxifylline molecule is bound in the same orientation as in protein copy A. In addition to visual examination, all structures were systematically and automatically checked throughout the refinement process for cis-peptides, various measures of covalent stereochemistry, close contacts, abnormal phi-psi angles, abnormal rotamers, and mismatched density features via output from the MIFit refinement interface. The final structure does not contain any significant abnormalities ( Table 1 ) and has been deposited with the Protein Data Bank as entry 3TVX.
Crystal and Refinement Parameters for PDE4A-Pentoxifylline
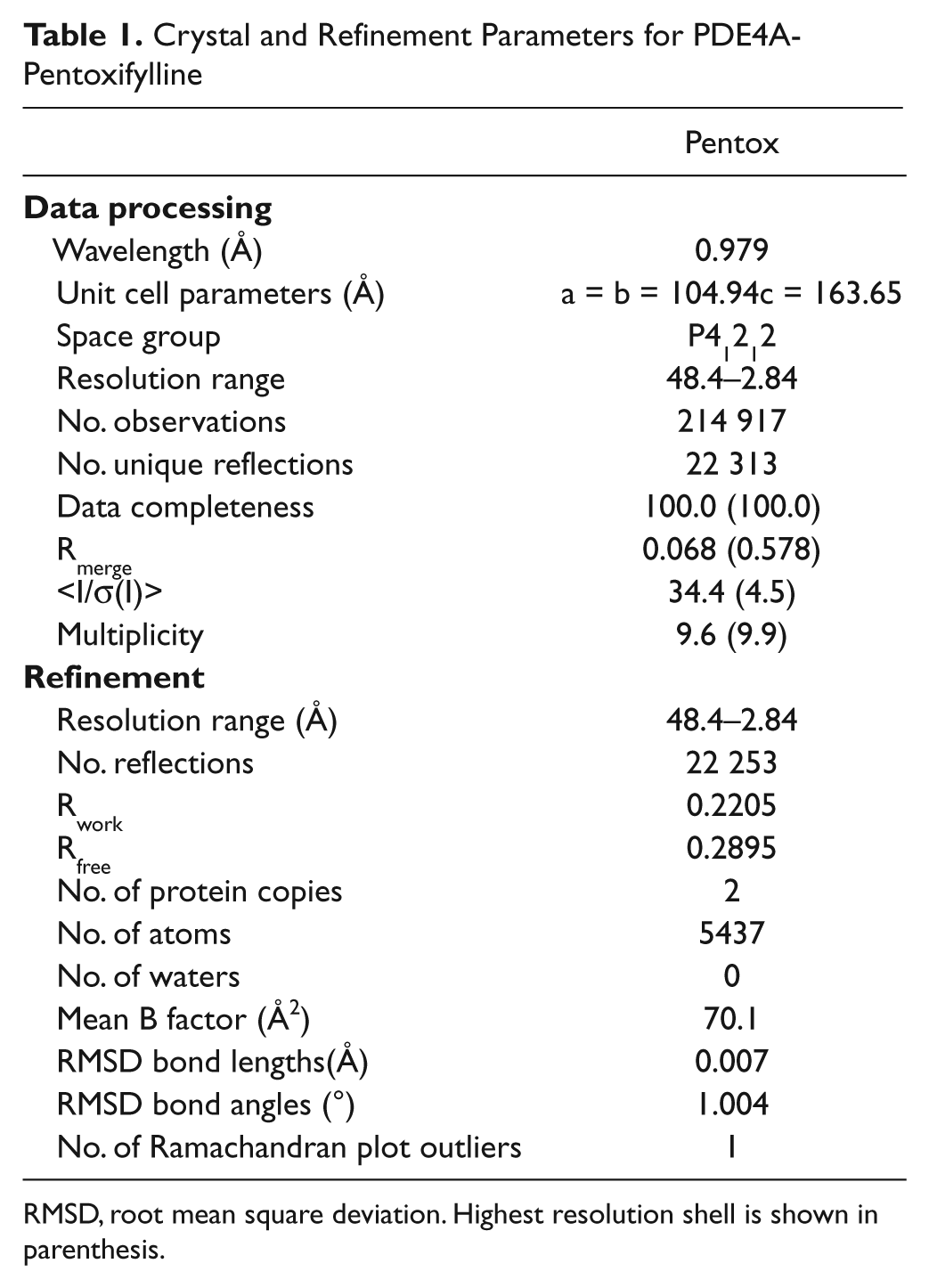
RMSD, root mean square deviation. Highest resolution shell is shown in parenthesis.
Results
The calorimetric activity assay was validated using two known general phosphodiesterase inhibitors: IBMX and pentoxifylline. As shown in Figure 1 , fitting the solid black points (no inhibitor) yielded kcat = 3.7 s−1 and K M = 24 µM, compared to kcat = 6.7 s−1 and K M = 5.1 µM for the catalytic domain of PDE4A using a radioactivity-based assay. 22 The open circles in Figure 1 show rate versus remaining substrate concentration for PDE4A hydrolysis of 3′,5′-cAMP in the presence of pentoxifylline ( Fig. 1A ) or IBMX ( Fig. 1B ). Both inhibitors display competitive inhibition of PDE4A, with K I values in good agreement with those expected based on IC50 values in the literature ( Table 2 ; pentoxifylline IC50 = 168 ± 105 µM 33 ; IBMX IC50 = 10.5 ± 0.3 µM 34 ).
Inhibition Constants and Ligand Efficiencies of Compounds Sorted Based on K I
The K
I
and corresponding ligand efficiency based on measurements using freshly prepared stock solutions of compounds. Compound number corresponds to the position along the x-axis in
The activity-based enthalpy array screen with the catalytic domain of PDE4A against a 160-compound fragment library was performed at a compound concentration of 2 mM. Corresponding control reactions in the absence of any inhibitor were performed for every five fragments tested and acted as the basis for comparison of K M values for those five fragments. Figure 2 displays the results of the fragment screen as the ratio of the K M in the presence of fragment (K M, app ) to the K M of the corresponding control reaction (K M, control ). Fourteen compounds produced a K M ratio ≥2. These were considered hits, and 11 of 14 were confirmed to have K I ≤2 mM using fresh stocks of fragments and protein. The K I and ligand efficiencies for all compounds that produced a confirmed K M ratio greater than 2 are presented in Table 2 .
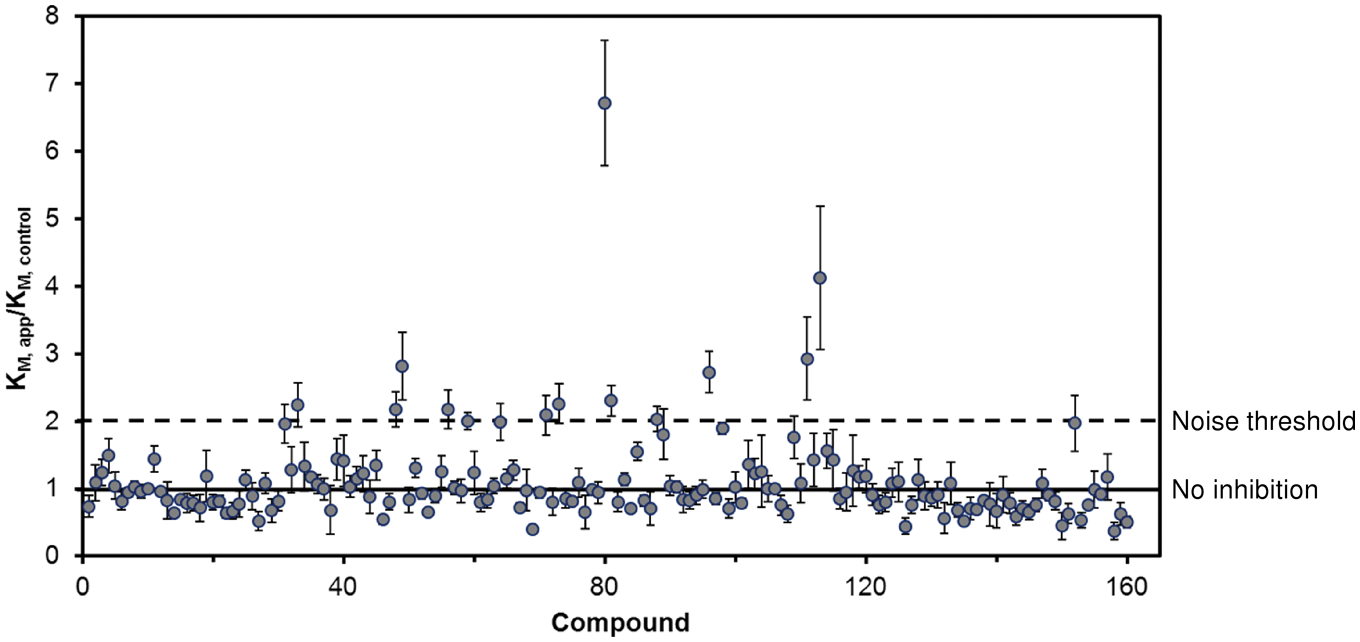
Results of the screen of 160 compounds with the calorimetric activity assay. The ratio of the K M in the presence of the fragment (KM, app) to the K M of an associated control reaction (KM, control) is shown. Compounds were tested at 2 mM concentration. For every five compounds, an associated control reaction (no inhibitor) was run and served as the KM, control for those compounds. The solid line indicates a K M ratio of 1, which means no effect compared to control. The dashed line indicates a K M ratio of 2, the noise threshold selected for a compound to be classified as a hit. The error bars indicate the standard error of the mean for the K M ratio.
We observed that two compounds, 104 and 152, caused inhibition that was not consistent with simple competitive inhibition. Compound 104 (at 2 mM) displayed mixed inhibition, reducing kcat by 25% and increasing K
M
by 2.7-fold. Compound 152 (at 2 mM) reduced kcat by 20% but did cause a significant change in K
M
. We investigated the inhibition mechanism of these two compounds using an ITC enzyme activity assay, which provides more sensitivity than the enthalpy array assay. Reactions were performed at several inhibitor concentrations, and the competitive (Kic) and uncompetitive (Kiu) inhibition constants were obtained for compound 104 (
Table 2
and
Examples of primary screening data are shown in Figure 3 . Figure 3A shows the data for the strongest inhibitor (80) and Figure 3B the weakest (33). Most compounds did not show any sign of competitive inhibition, as shown for compound 51 in Figure 3C .
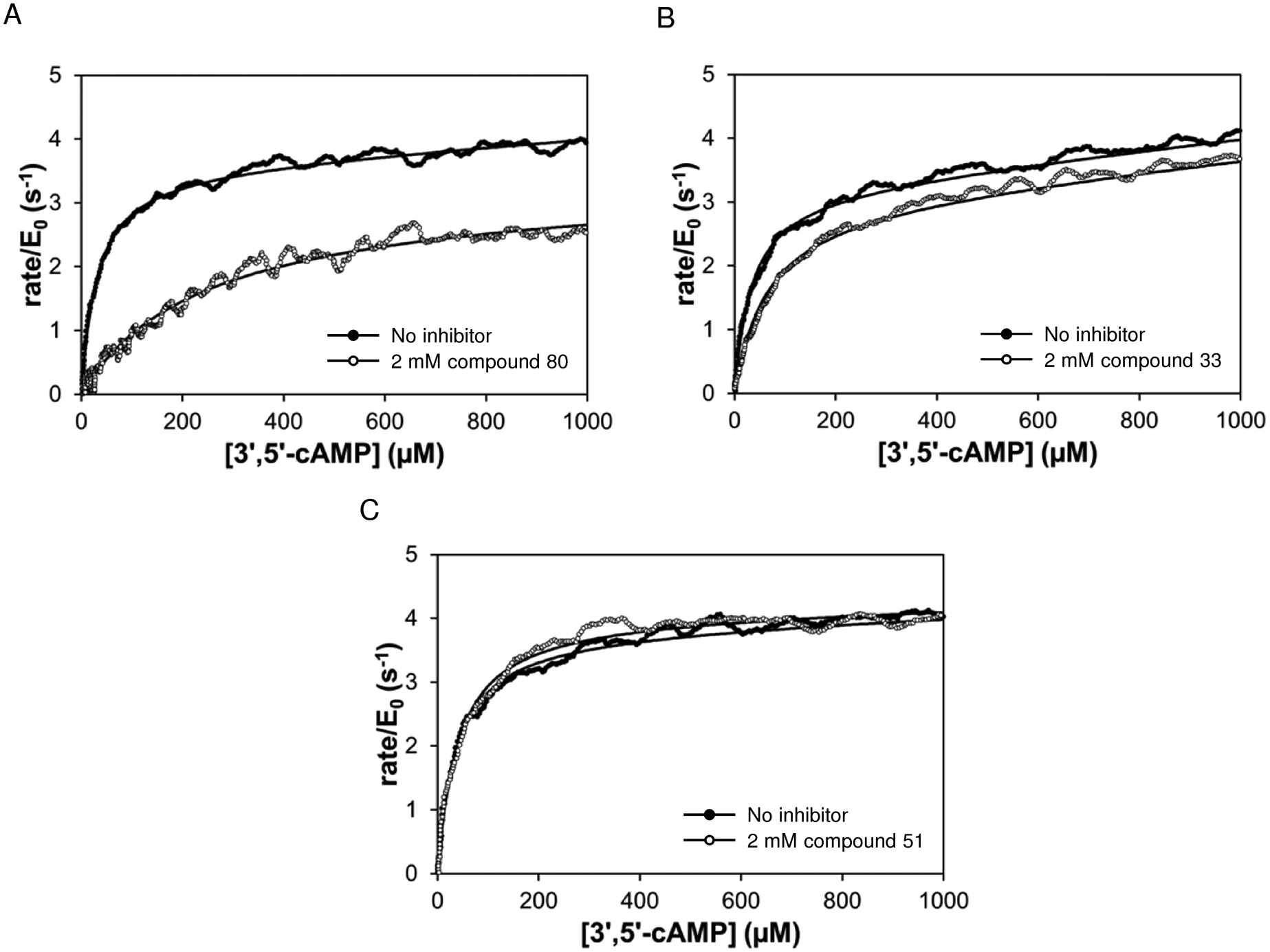
Rate versus remaining substrate concentration data for PDE4A hydrolysis of 3′,5′-cAMP in the absence (solid) and presence (open) of three compounds from the primary screen. Solid curves are a fit of data to equation 2. (
The 11 fragments listed in Table 2 were moved into X-ray crystallography studies. Initially, efforts were made to obtain high-quality apo-PDE4A crystals. Multiple crystal forms were observed and tested, but all diffracted only marginally >3.0 Å, and none were deemed suitable for high-throughput soaking experiments. Co-crystallization experiments followed. Although co-crystals could be obtained with the fragments in Table 2 , the diffraction quality of crystals appeared to vary with the potency of the starting ligand. High-quality diffraction was obtained with one potent ligand, pentoxifylline.
The crystal structure of PDE4A in complex with pentoxifylline is shown in Figure 4 . As stated above (in Methods), the ligand was bound in the same orientation in both molecules of the asymmetric unit and was clearly defined in the electron density map, particularly for molecule A ( Fig. 4A ). The purine ring of pentoxifylline is bound near the active site sandwiched between Phe 584 and Met 569 with the C18 methyl occupying a small subpocket. As shown in Figure 5A , the purine ring binds differently from that of adenosine monophosphate (AMP 35 ) and the PDE inhibitor IBMX. 36 In AMP and IBMX, the purine ring is bound in the primary pocket and accepting a hydrogen bond from Asn 321 (PDE4A numbering), although the hydrogen bonding patterns are different. In AMP, the hetrocyclic N1 is accepting the hydrogen bond from Asn-NH2, and in IBMX, the exocylic 6-oxy group is accepting the hydrogen bond as the N1 group is methylated. In the pentoxifylline structure, the aliphatic tail is bound in the primary purine pocket and interacting with Ile 548 ( Fig. 4B , C ). Although apparently not occupied by AMP, the area of the active site occupied by the pentoxifylline molecule does overlap with other small-molecule inhibitors of PDE4A such as NVP, 22 shown in Figure 5B . Pentoxifylline is not within standard hydrogen bonding distance of any PDE4A residues but may be making a long hydrogen bond with Gln 581 (3.8 Å).
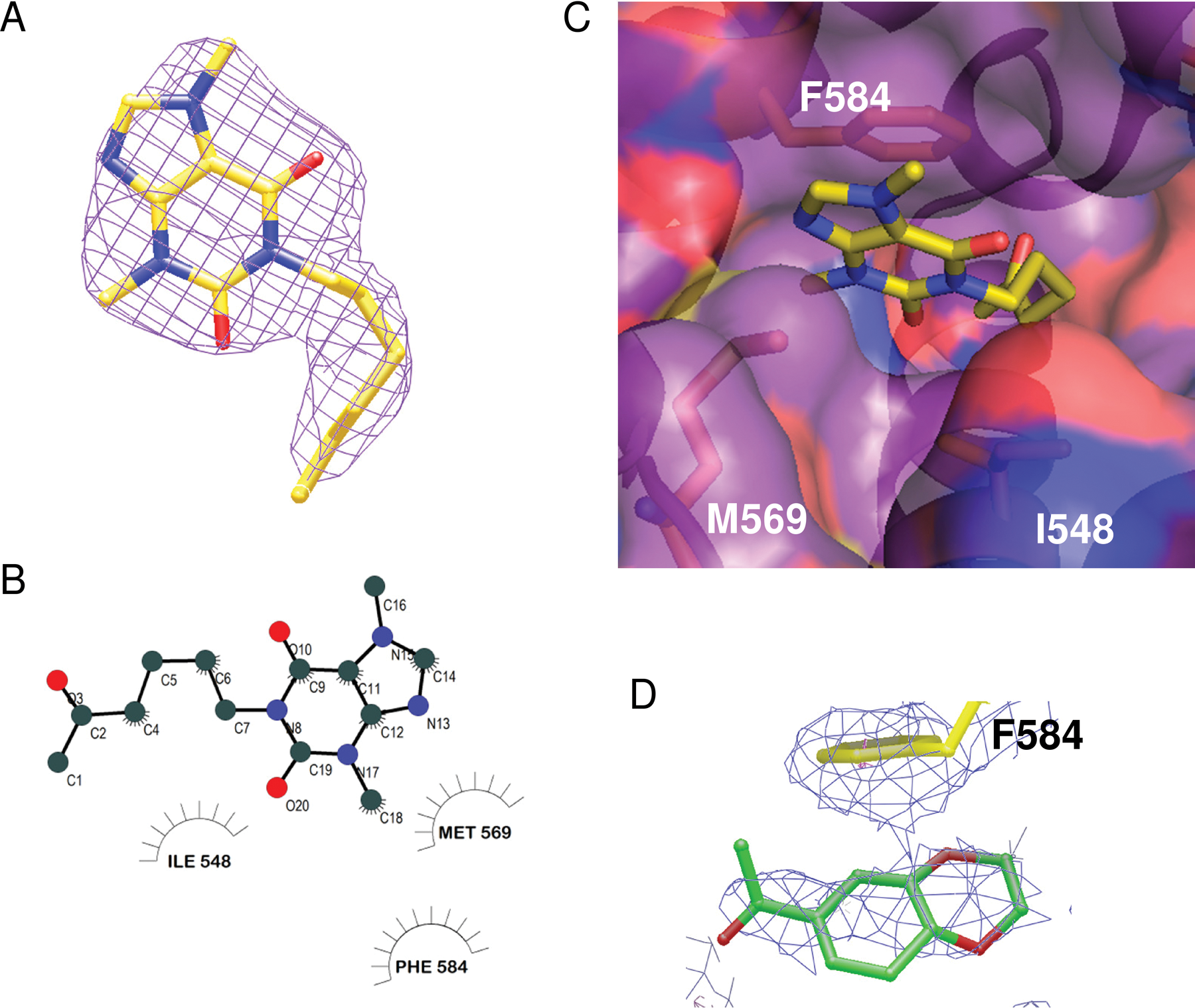
Crystal structure of PDE4A in complex with pentoxifylline. (
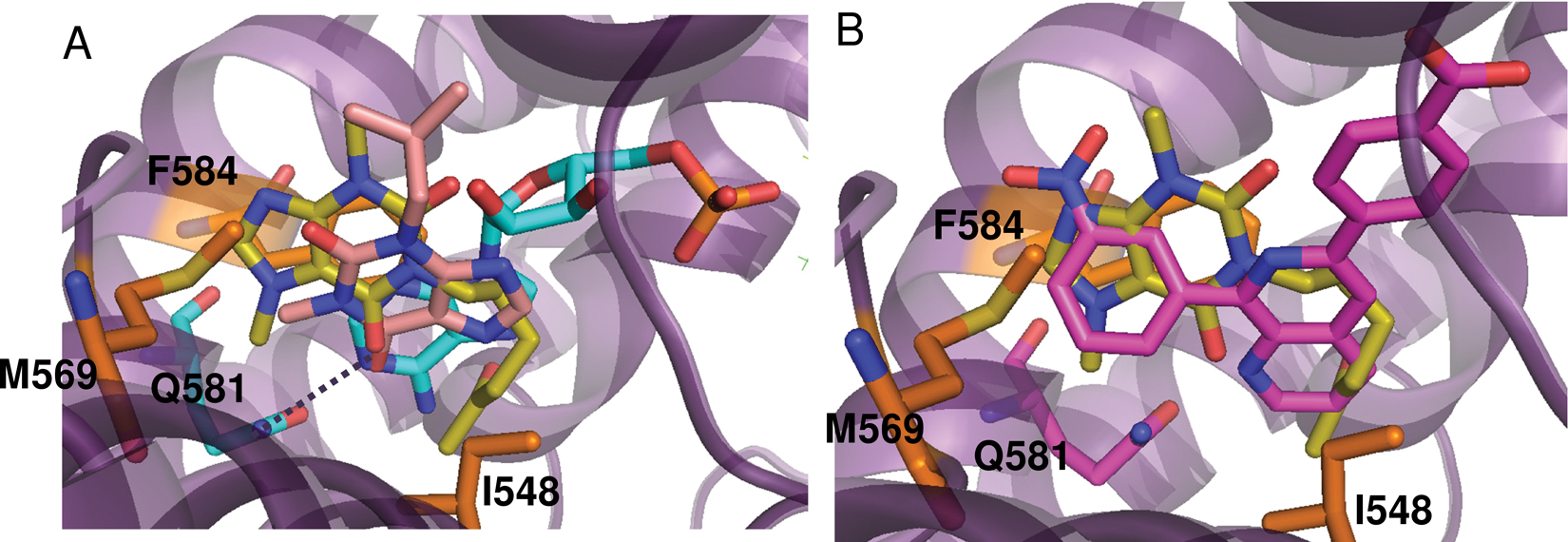
Overlay of pentoxifylline with published PDE4 crystal structures. Amino acid numbering corresponds to PDE4A. (
Lower resolution maps for co-crystal structures with other fragments showed evidence of binding but were not completed due to poor definition of ligand orientation in the active site. Figure 4D shows an example of these low-resolution data. Here, a tube of density at a peak height of 4σ consistent with fragment 113 was present in the unbiased electron density maps at 3 Å resolution. Because of the low resolution of the map, the compound was not included in the final structure.
Because a high-quality structure could be obtained in the presence of pentoxifylline, soak-out, soak-in experiments were completed to displace pentoxifylline with the fragment of interest. PDE4A-pentoxifylline crystals were soaked in the presence of fragment hits. However, surprisingly, the crystals were not stable to soaking times beyond ~5 min. To understand these data, crystals were exposed to the fragments individually and flash-frozen in liquid nitrogen after a 5-min incubation. Data were collected at high resolution (2.3–2.6 Å) and structures solved by molecular replacement. In all cases, pentoxifylline remained bound at the active site, and the fragment was found bound at a secondary binding pocket immediately adjacent to the pentoxifylline and partially composed of residues from a neighboring symmetry-related molecule. A second fragment molecule was found bound at a small groove at a close crystal contact. It is possible that high-occupancy binding at this site could result in disruption of the crystal lattice and lead to the crystal cracking observed with longer soaks. Details of these experiments will be published elsewhere.
Discussion
We demonstrate here that low-affinity competitive inhibitors of PDE4A can be identified using an enthalpy array enzyme activity assay. Screening at a compound concentration of 2 mM ensured that compounds with K I ≤ 2 mM could be detected above the noise threshold. The 2-mM cutoff corresponds to a ligand efficiency ≥0.3 for the average number of heavy atoms for the compounds in this library.
For phosphodiesterases, the direct label-free measure of substrate conversion using calorimetry provides a distinct advantage compared to the existing standard assays, which rely on coupled enzymes, 37 radioactivity, 38 or high-performance liquid chromatography (HPLC) separation. 39 In fragment screening with the coupled enzyme assay, there is the possibility that compounds could interfere with the coupling enzymes rather than the enzyme of interest. It is necessary that the appropriate controls be performed to ensure that inhibition is due to action on the desired target. The radioactivity and HPLC assays are discontinuous and labor intensive. In contrast, each calorimetric measurement is a rapid continuous assay yielding full enzyme kinetic parameters for reactions performed in the presence of each compound. For competitive inhibitors, this allows determination of K I for each compound from the primary screening data. A limitation of continuous assays is that product accumulates during the course of the reaction, which is an issue if product inhibition is significant. For the reactions presented here, the substrate concentration (2 mM) and therefore corresponding product concentration at the end of the reaction are below the reported Kd for AMP (7.1 mM 40 ), so any product inhibition will cause, at most, an ≈30% increase in the apparent K M for the control reaction.
The screen we describe here was used to identify inhibitors that act exclusively by a cAMP-competitive mechanism. By screening for compounds that alter k cat , the enthalpy array assay could be used to identify compounds that inhibit by a noncompetitive or uncompetitive mechanism with K I <2 mM. Activity-based assays in general exclude from the list of hits compounds that bind to the target protein but do not significantly inhibit the enzymatic reaction. These compounds could bind in pockets from which potent inhibitors could be developed; hence, some very weakly binding fragments that might lead to potent inhibitors may well be missed using activity-based assays.
Unlike PDE4B 41 or PDE4C, 22 PDE4A did not produce crystals in the apo form. Hence, co-crystallization was required. Crystals were obtained with multiple fragments, but all diffracted to low resolution (>3.0 Å) with the exception of pentoxifylline. Soak-out, soak-in experiments were attempted with the pentoxifylline crystals, but the crystals were unstable under these soaking conditions. Data collected with short soak times revealed both pentoxifylline and the fragment bound at the active site.
With a hard-to-crystallize protein, such as PDE4A, the calorimetric prescreen adds value because it provides data demonstrating that the compounds inhibit the enzyme by a particular mechanism. With the knowledge that the compounds act as competitive inhibitors, one could consider methods other than X-ray crystallography to obtain information about the specific interactions between the compound and the protein. For some targets, it may be possible to perform structural studies by NMR to determine the 3D structure of the PDE4A-compound complex. Although the X-ray and NMR methods are expensive in terms of labor, material, and equipment, they do provide structural information about the binding interactions between fragments and the biomolecular target, which is crucial for efficiently transforming fragment hits into leads via fragment elaboration or linking. Another option is to use docking methods based on crystal structures of related proteins (or crystal structures of other ligands bound to the protein of interest) to propose potential binding models for the fragments and use this information to guide the synthesis of modified or elaborated fragments, which can be tested for inhibition in the calorimetric assay. Additional attempts at X-ray crystallography perhaps using different protein constructs or protein mutants could be made, but this additional work was beyond the scope of this study.
Although we were unable to obtain crystal structures for all fragments bound to PDE4A, the chemical structure for many of the hits are consistent with either PDE4 substrates or published inhibitors. Specifically, fragments 109, 49, and 33 contain a hydrogen bonding motif present in the adenine core of AMP ( Fig. 6A ). In the PDE4D-AMP structure ( Fig. 5A ), the exocyclic nitrogen and adjacent ring heteroatom are within hydrogen bonding distance of Gln 581 (PDE4A numbering). Because the fragments are much smaller than AMP, it is possible they may maintain these hydrogen bonds but bind in a slightly different mode representing the lowest energy binding conformation. Similarly, fragments 73, 111, and 104 share a common core with published PDE4 inhibitors. Two of these, NVP (PDB code 2QYK 22 ) and 4-[3-(methoyphenyl)amino]-6-(methylsulfonyl)quinoline-3-carboxamide (PDB code 3FRG 42 ), have published crystal structures and are shown in Figure 6B . In these structures, the quinoline heterocyclic nitrogen is hydrogen bonding with Gln 581 (PDE4A numbering), and examination of the overlay shows that the core may shift in binding mode based on the substitution pattern on the ring without losing the hydrogen bond. As with the purine, it is likely that the quinoline rings of the fragment hits occupy slightly different relative orientations depending on their substitution patterns. Regardless of the binding modes, the chemical structures of these fragment hits are consistent with known PDE4 ligands, further substantiating them as valid hits.
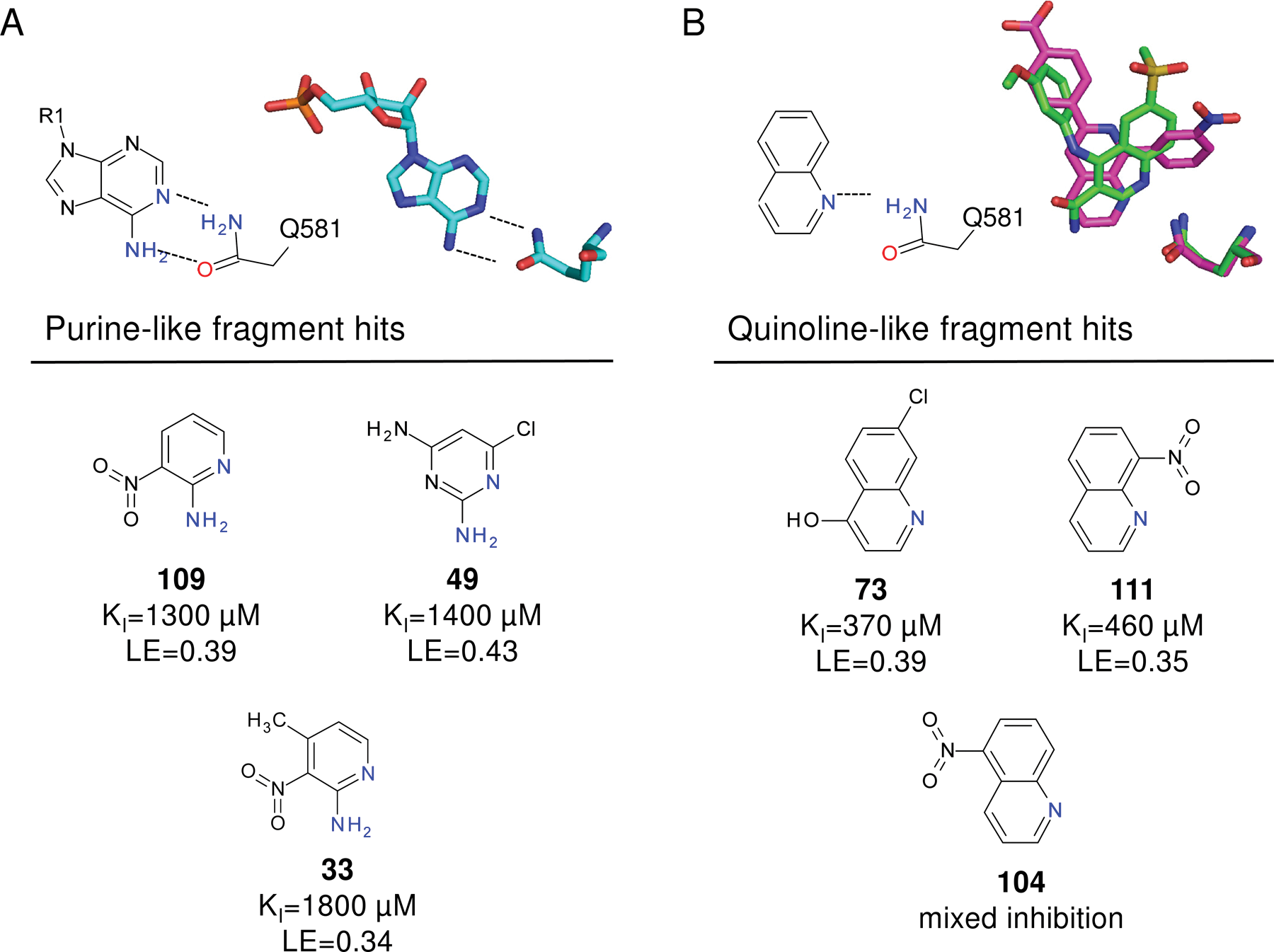
Summary of screening hits with purine and quinoline-like hydrogen bonding motifs. (
Although pentoxifylline was one of the more potent compounds tested (72 µM), it is the least ligand efficient (0.28). Examination of the crystal structure ( Fig. 4 ) provides a possible explanation for this observation. The aliphatic “tail” of the ligand inserts into the primary binding pocket rather than the molecule’s core, most likely because the core is highly substituted. The high degree of substitution could sterically interfere with binding in the pocket and also interferes with hydrogen bonding with Gln 581. Based on the crystal structure, the interactions between this ligand and protein appear to be purely van der Waals.
In contrast to the dearth of interactions observed here for pentoxifylline, IBMX forms an extensive network of interactions with PDE4 and has both higher potency (8.3 µM) and ligand efficiency (0.43). In the PDE4D-IBMX structure (Huai et al. 36 ; PDB 1ZKN, Figure 5A), the xanthine ring stacks against Phe-372 (Phe 584 PDE4A) on one side and contacts Ile-336 (Ile 548, PDE4A) and Phe-340 (Phe 552, PDE4A) on the other side. IBMX forms a hydrogen bond with the side chain of Gln-369 (Gln 581, PDE4A) and hydrophobic interactions with Leu-319 (Leu 319, PDE4A) and Asn-321 (Asn 533, PDE4A). In addition, there are interactions of the isobutyl group with Met-273 and Ile-376 (Ile 588, PDE4A) and unique interactions with Met-273 (Met 485, PDE4A) and Met-357 (Met 569, PDE4A).
The use of enthalpy arrays to prescreen fragments for inhibition of enzymatic activity has two benefits: (1) the number of structural characterizations is reduced, reducing labor and material cost, which is particularly important in cases where structural characterization is more difficult than expected, and (2) one knows that the compounds have an effect on the activity of the enzyme, indicating that the fragment is binding to a site of functional interest rather than simply binding. Moreover, calorimetric prescreening can reveal the inhibition mechanism, such as whether inhibition is competitive, mixed, or uncompetitive. In this study, the prescreening of PDE4A reduced the number of compounds on which structural characterization needs to be performed by more than a factor of 10. Because all but one of the compounds competitively inhibit the enzyme, there is increased motivation to try to obtain diffracting crystals, even if it is a difficult to crystallize target, or to put the investment into NMR characterization of structure.
Footnotes
Acknowledgements
We thank Richard Bruce, Alan Bell, and Dirk De Bruyker for their contributions to the enthalpy array technology, system automation, and array fabrication. We thank Duncan McRee for his contribution to the initiation of the PDE4A fragment screening project as well as helpful discussion. We thank G. Sridhar Prasad for his contribution during the early stages of the project.
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: M. Recht and F. Torres are employed by Palo Alto Research Center Incorporated. All other authors are employed by Zenobia Therapeutics.
The authors disclosed receipt of the following financial support for the research and/or authorship of this article: This work was supported by grant R01EB009191 from the National Institute of Biomedical Imaging and Bioengineering, National Institutes of Health (NIH). Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.