Abstract
A high-throughput online solid-phase extraction/tandem mass spectrometry (online SPE/MS/MS) system has been developed to support rapid evaluation of drug discovery compounds for possible drug-drug interaction (DDI). Each compound is evaluated for its DDI potential by incubating over a range of 8 test concentrations and against a panel of 6 cytochrome P450 (CYP) enzymes, 1A2, 2C8, 2C9, 2C19, 2D6, and 3A4. Previously, a postassay pooling and a 2-min gradient LC/MS/MS method had been reported to increase sample throughput, allowing for a 96-well plate of samples to be analyzed in under 4 h. The development of a new online SPE/MS/MS system has reduced the analysis time to less than 15 min per 96-well plate, translating to a 15-fold time savings compared to the 2-min LC/MS/MS method. Sampling precision without internal standard correction ranged from 3.1% to 5.6% relative standard deviation, and the carryover was determined to be between 1.0% and 4.1%. One hundred twenty in-house compounds were assayed and pooled for analyses using both the online SPE/MS/MS and LC/MS/MS, and the correlation coefficients ranged from 0.89 to 1.13, when comparing the IC50 results obtained from the 2 approaches for each of the CYP enzymes.
Introduction
V
Liquid chromatography/mass spectrometry (LC/MS) has gained popularity for CYP inhibition screening due to the specificity and sensitivity afforded by MS. The speed of LC/MS, however, is limited by the LC runtime, and this presents a major limitation to its use in high-throughput screening (HTS). A number of LC methods such as fast LC, 3 parallel LCMS, 4 staggered chromatography, 2 direct inject with online extraction, 5 sample pooling combined with fast LC gradient, 6 and offline solid-phase extraction (SPE) followed by flow injection analysis (FIA)–MS 7 have been developed and implemented to reduce analysis time. Recently, matrix-assisted laser desorption ionization (MALDI)–MS/MS and laser diode thermal desorption interfaced with atmospheric pressure chemical ionization mass spectrometry (LDTD-APCI-MS/MS) have also been evaluated for CYP inhibition screening. MALDI-MS/MS, although capable of achieving a sample throughput of about 5 min per 96-well plate, requires additional SPE steps prior to co-spotting the sample with matrix and internal standard. 8 LDTD-APCI-MS/MS is a new sample introduction technique combining thermal desorption and APCI. The method has been shown to analyze the samples without prior pretreatment with throughput ranging from approximately 30 min for manual acquisition to about 18 min in fully automated mode for a 96-well plate of pooled CYP samples (1A2, 2C9, 2D6, and 3A4). 9
We report here the implementation of an automated system coupling online SPE with triple quadrupole MS analysis for ultra-high-throughput analysis of CYP inhibition samples. The online SPE/MS/MS system uses a micro SPE cartridge with less than 4 µL bed volume but containing a large surface area for rapid online desalting of samples, and its core technology uses custom rotary valves with an electromechanical control system that facilitates valve actuation in 20 ms, resulting in an almost undetectable perturbation in the electrospray ionization (ESI) spray. The online SPE/MS/MS system has been used successfully in HTS assays involving kinase, 10 fatty acids, 11 and small molecules 12 with direct injection of samples and sustainable throughput as fast as 5 s per sample. The CYP assay results obtained from the online SPE/MS/MS system are compared to the currently used LC/MS approach, and the advantages as well as potential limitations using the online SPE/MS/MS system for CYP screening are discussed.
Materials and Methods
Materials
All recombinant CYP enzymes (1A2, 2C8-b5, 2C9, 2C19, 2D6, and 3A4) and the NADPH generating system were obtained from BD Biosciences (Woburn, MA). Substrates phenacetin, amodiaquine, and diclofenac were from Sigma-Aldrich (St. Louis, MO); S-mephenytoin, bufuralol, and midazolam were from Biomol (Plymouth Meeting, PA), BD Bioscience (Woburn, MA), and AllTech (Deerfield, IL), respectively. Standard inhibitors furafylline, sulfaphenezol, tranylcypromine, quinidine, and ketoconazole were from Sigma-Aldrich, and montelukast was purchased from Sequoia Research Product (Pangbourne, UK). Other reagents such as phosphate buffer and trichloroacetic acid (TCA) were also from Sigma-Aldrich. In-house Milli-Q water (Millipore, Billerica, MA) was used for all solution preparations.
Incubation conditions
The incubation conditions were based on established in-house assay protocols that were validated with optimized kinetic parameters; the Km of all 6 CYP isozymes was determined and substrate concentrations at or below Km were used for the assays; the assay protocols also ensured a linearity in metabolite production, with 20% or less turnover of substrates to metabolites, and a linear metabolite production relative to enzyme concentration.
The assay was automated using a Tecan EVO 2000 liquid handler (Tecan Group Ltd., Mannedorf, Switzerland) equipped with a multichannel arm and was programmed to provide IC50 data for 6 CYP isozymes up to 12 compounds at 8 concentrations per assay on 96-well plates. CYP isozyme concentrations were 12.5 nM for 1A2, 2C9, and 2C19; 25 nM for 3A4; 2.5 nM for 2C8; and 0.6 nM for 2D6 in a total incubation volume of 100 µL buffered with 100 mM potassium phosphate, pH 7.4. Eight concentrations of each test compound, starting at 50 µM and 5-fold serial diluted, were added during incubation to yield a final 0.5% DMSO and 0.5% acetonitrile in the reaction mixture. The substrates selected for 1A2, 2C8, 2C9, 2C19, 2D6, and 3A4 were phenacetin, amodiaquine, diclofenac, S-mephenytoin, bufuralol, and midazolam, respectively. Reactions were initiated with NADPH regenerating system and terminated by addition of 100 µL of 200 mM TCA after a 10-min incubation at 37°C for all isozymes, with the exception of 2C19, for which the incubation time was 20 min. The quenched samples from the 6 isozymes were pooled and filtered using a 0.45-µm Multiscreen 96-well filter plate (Millipore). A separate set of CYP assays was performed using standard inhibitors as a control assay; furafylline, montelukast, sulfaphenezol, tranylcypromine, quinidine, and ketoconazole were selected as standard inhibitors for 1A2, 2C8, 2C9, 2C19, 2D6, and 3A4, respectively. Control assays used the same protocol as described, but the concentration ranges and dilution factors were customized to bracket their known IC50 values.
Tacrine was selected as an alternate substrate for evaluating CYP1A2 inhibition. For tacrine, the assay conditions were similar to the assay using phenacetin as described above except that 3 µM of tacrine, a concentration slightly below the determined Km of 3.4 µM, was used for the assay.
Instrumentation
High-throughput online SPE/MS/MS experiments were performed using a RapidFire® (BIOCIUS Life Sciences, Inc., Woburn, MA) instrument coupled to an API4000 triple quadrupole mass spectrometer (Applied BioSystems, Concord, Ontario, Canada). Solvent A was 0.1% TFA in water, and solvent B was 0.1% TFA in acetonitrile. Sample was aspirated directly from 96-well plates to a 10-µL loop and onto a phenyl cleanup cartridge to remove nonvolatile buffer with solvent A with a 2-s wash cycle at 1.5 mL/min. The retained analyte was eluted to the mass spectrometer in 5 s using solvent B at 0.5 mL/min. The lower flow rate for the elution step was selected to facilitate better ionization efficiency in the ESI source. Reequilibration of the SPE cartridge was achieved with a 0.5-s wash with solvent A at 1.5 mL/min (equivalent to 3-column volume). The entire sampling cycle was 9 s per well, and the peaks detected by mass spectrometer were approximately 4 s wide at the baseline. The aspirator tip was washed using 50% acetonitrile in water between sample aspirations.
For LC/MS/MS analyses, the system comprised an API4000 triple quadrupole mass spectrometer (Applied BioSystems), 2 LC-20AD pumps with a CBM-20A controller (Shimadzu, Canby, OR), and a CTC HTS PAL auto-sampler (LEAP Technologies, Carrboro, NC). A 20-µL sample was injected onto a Pursuit Diphenyl 50 × 2-mm 3-µm column (Varian, Lake Forest, CA). Gradient elution at a flow rate of 0.7 mL/min was performed using the following mobile phases: solvent A was 0.04% formic acid in water, and solvent B was 0.04% formic acid in acetonitrile. The gradient was as follows: 17%B (0-0.3 min), 35%B (0.4 min), 50%B (1.5 min), 99%B (1.6 min), and 1%B (1.61-2 min). Flow was diverted to waste for the first 0.3 min of the gradient to remove nonvolatile salts, and the stop time was 2 min.
The mass spectrometer was operated in positive electrospray multiple-reaction monitoring (MRM) mode when coupled with either the RapidFire® or the Shimadzu LC system. MRM transitions, collision energy, and declustering potential (Q1/Q3/CE/DP) for each isozyme/substrate system were as follows: 1A2/phenacetin = 152/110/30/70 (or 1A2/tacrine = 215/181.8/40/60), 2C8/amodiaquine = 328/283/40/70, 2C9/diclofenac = 312/231/30/70, 2C19/mephenytoin = 235/150/30/70, 2D6/bufuralol = 278/159/30/70, and 3A4/midazolam = 342/203/45/70. A dwell time of 50 ms was allocated to each of the 6 simultaneous MRM channels, and the mass spectrometer ion spray was operated at 4500 V and a source temperature of 600°C.
Data processing
LC/MS/MS MRM data were processed using Analyst’s integration/quantitation module. Online SPE/MS/MS MRM data were processed using BIOCIUS peak integration software. In either case, the peak area versus inhibitor concentration data were fitted to dose-response curve model 205 using XLFit (IDBS, Guildford, UK) and IC50 values computed.
Results/Discussion
The online SPE/MS/MS system has enabled us to analyze CYP samples in ultra-high-throughput mode without prior sample pretreatment and without internal standard. For a full 96-well plate containing pooled samples from 6 CYP isozymes, we were able to reduce the time of analysis from 4 h using the 2-min LC/MS/MS per 96-well plate to less than 15 min. The analysis time using online SPE/MS/MS was further reduced to about 10 min per plate containing only discrete samples (data not shown). This was accomplished by shortening the elution time from 5 to 2 s and increasing solvent B flow from 0.5 to 1 mL/min.
The sampling reproducibility and extent of carryover for the online SPE/MS/MS instrument without internal standard was determined. Figure 1A shows the layout of pooled CYP samples and blanks in a 96-well plate. The CYP samples were prepared by incubating each CYP isozyme with its respective substrates over a 16-h period to maximize the production of metabolites. The samples from each isozyme were then pooled, the resulting mixture was “injected” repeatedly, and MRM signals were monitored to assess the precision of the instrument over all 6 CYP products simultaneously. Blanks (water) were injected between CYP mixtures to assess the extent of carryover. Figure 1B shows the row-to-row reproducibility of the online SPE/MS/MS system. Precision ranged from 3.1% relative standard deviation (%RSD) for 2D6 to 5.6 %RSD for 2C9. Sample carryover also appeared to be minimal despite a rather short wash time used for column cleanup. Overall, the carryover was approximately 4% or less across all 6 CYPs. More lipophilic compounds would be expected to be more susceptible to carryover, but this can be minimized by using a combination of longer wash time and stronger wash solvent. The total acquisition time for full 96-well plate samples with pooling was approximately 14 min.
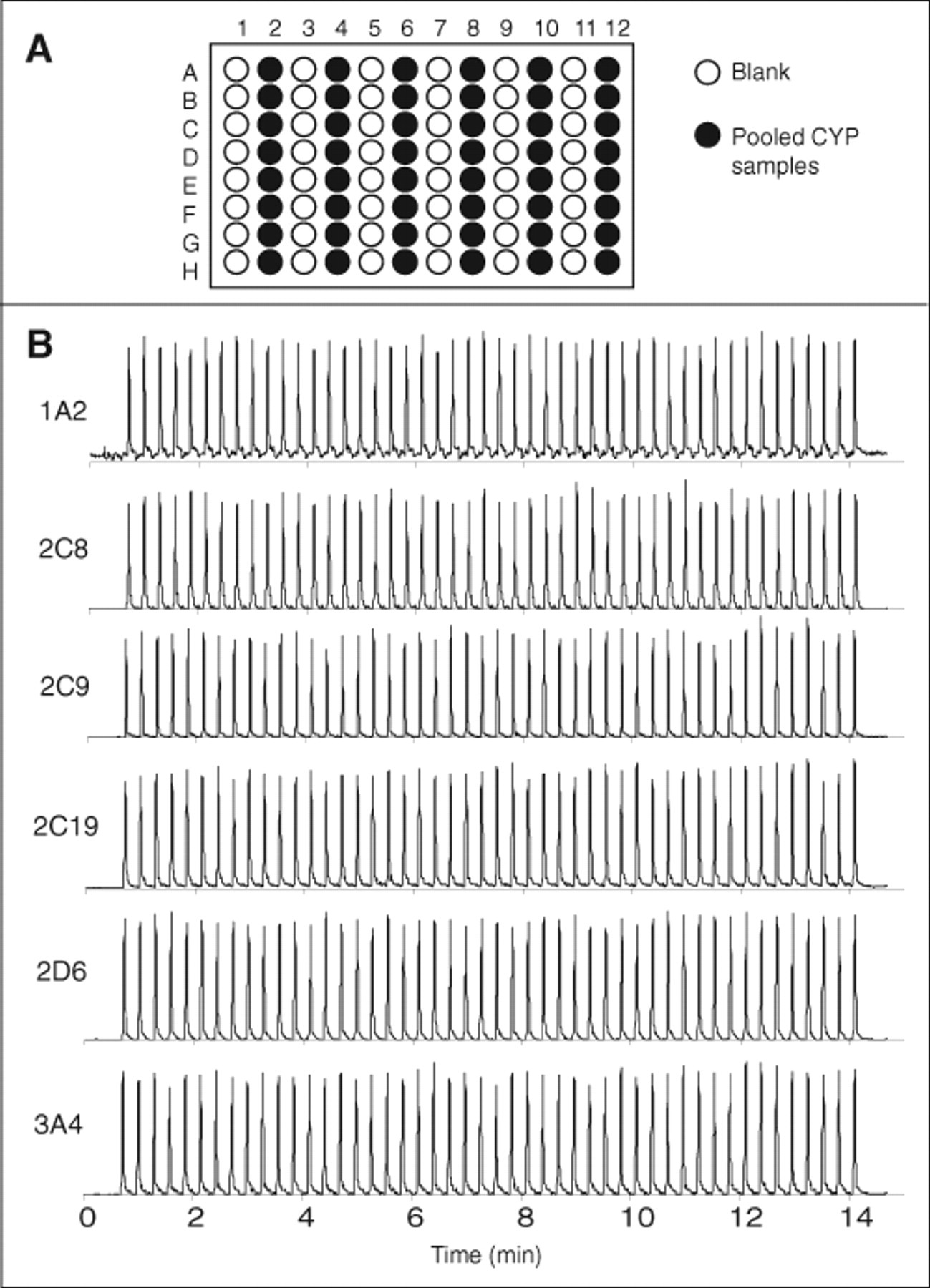
Experiment to determine sampling precision and carryover using online solid-phase
extraction/tandem mass spectrometry (SPE/MS/MS) with multiple-reaction mode (MRM)
detection. (
A series of assays using standard inhibitors as test compounds were analyzed next. To show reproducibility, the same samples were aliquoted into 8 identical sets and were analyzed 8 times. With the exception of 1A2, in which a relatively large background was observed across all extraction ion chromatogram (XIC), the peaks detected from all other isozymes were quite good and with baseline separation. The IC50 results obtained from the online SPE/MS/MS analyses of the pooled sample sets are summarized in Table 1 . The same table also lists IC50 results obtained from 2-min LC/MS/MS and those reported in the literature. With the exception of 1A2, results of the online SPE/MS/MS were roughly comparable to those obtained by LC/MS/MS.
Tabulated Results (Mean ± Standard Deviation) Comparing IC50 Values of Standard Inhibitors Obtained from Online SPE/MS/MS, LC/MS/MS, and Those Reported in the Literature
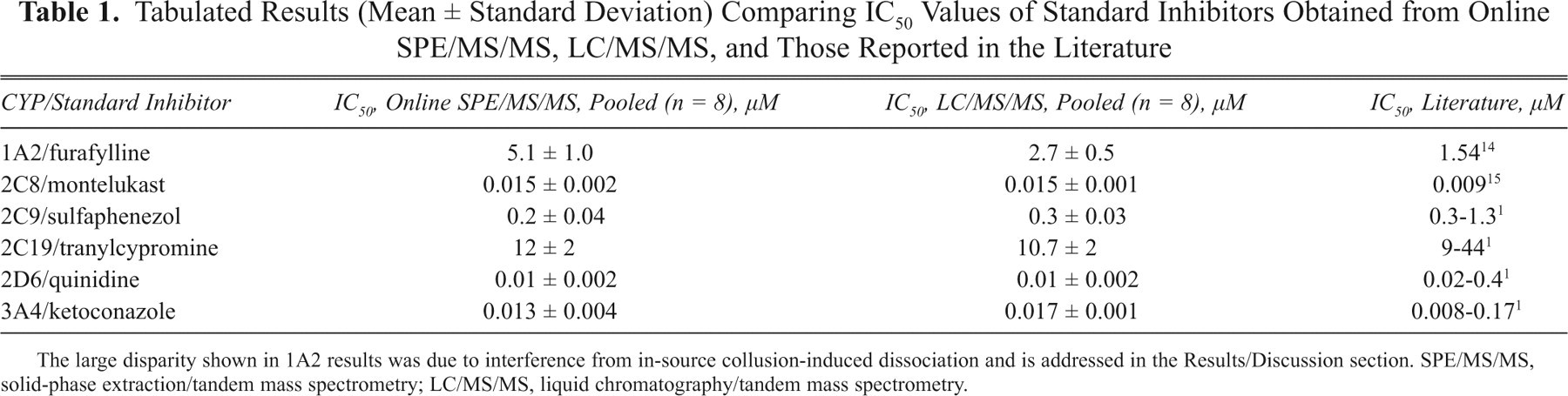
The large disparity shown in 1A2 results was due to interference from in-source collusion-induced dissociation and is addressed in the Results/Discussion section. SPE/MS/MS, solid-phase extraction/tandem mass spectrometry; LC/MS/MS, liquid chromatography/tandem mass spectrometry.
For online SPE/MS/MS, 1A2 presents a challenge in that the substrate phenacetin undergoes a substantial in-source fragmentation in ESI, 1 yielding an ion structurally identical to the metabolite product, acetaminophen. In LC/MS/MS mode, both acetaminophen and phenacetin are well separated chromatographically, and thus the in-source fragmentation of phenacetin to acetaminophen does not adversely affect the IC50 measurement, even though the 2 peaks are detected using the MRM transition tuned for acetaminophen. In the online SPE/MS/MS method, however, both peaks coelute, and the peak from in-source fragmentation leads to a significant overall background, which adversely affects the accuracy and reproducibility of IC50 data. This in-source fragmentation phenomenon for phenacetin is consistent as observed by others to various degrees of severity, including the MALDI and LDTD approaches. 9 One potential solution to the problem for the online SPE/MS/MS method of 1A2 would be to make a background adjustment to the raw signal prior to peak integration. However, a more permanent solution would be to use a different substrate, such as tacrine, 13 to avoid such interference from in-source fragmentation of the residual substrate.
Consequently, tacrine was selected to replace phenacetin as a substrate for 1A2. A total of 120 in-house compounds were randomly selected for the CYP inhibition assay, and the pooled samples were analyzed by both LC/MS/MS and online SPE/MS/MS. Figure 2 shows the IC50 correlation plots between the 2 methods for 1A2, 2C8, 2C9, 2C19, 2D6, and 3A4. A few compounds did not inhibit the enzymes at any of the tested concentrations, and therefore their IC50 values could not be calculated. Hence, data for these compounds were excluded in the correlation analyses. Excellent correlation was observed for the 2 data sets; the IC50 values obtained from online SPE/MS/MS were within 2-fold of those obtained from LC/MS/MS and with correlation coefficients ranged from 0.89 to 1.13.
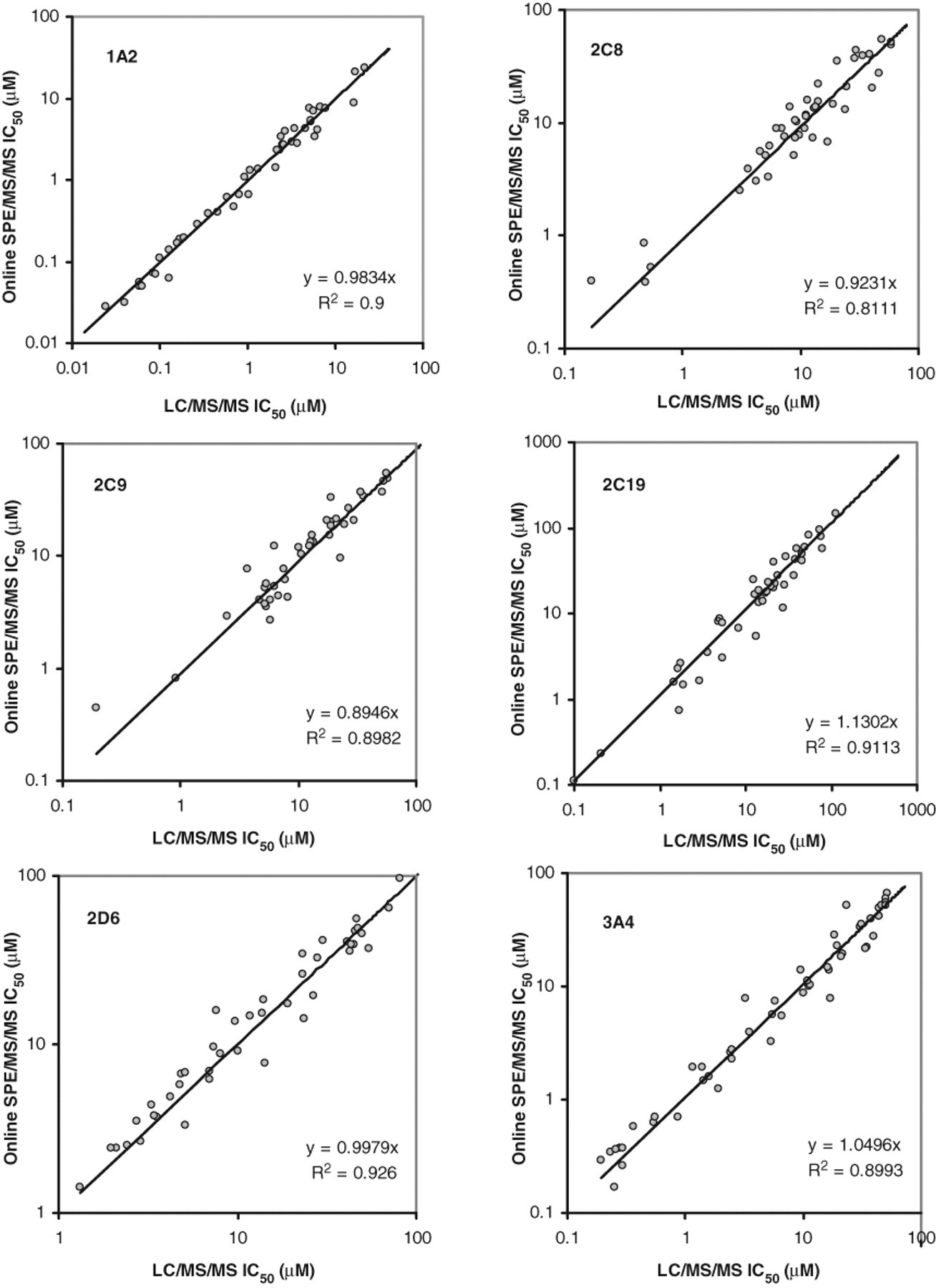
IC50 correlation plots comparing online solid-phase extraction/tandem mass spectrometry (SPE/MS/MS) and liquid chromatography/tandem mass spectrometry (LC/MS/MS) acquisitions using in-house synthesized compounds as inhibitors in 1A2/tacrine, 2C8/amodiaquine, 2C9/diclofenac, 2C19/mephenytoin, 2D6/bufuralol, and 3A4/midazolam CYP inhibition assay. Only compounds showing inhibitory responses were plotted here.
One major concern regarding the use of online SPE/MS/MS for pooled CYP analysis was how coeluting peaks might affect the measured IC50 values. Data summarized in Table 1 as well as results from studies shown in Figure 2 illustrate that coeluting peaks such as the ones from pooled analyses have little effect on the IC50, and the observed differences in IC50 were 2-fold or less.
Another concern was the dilution effect of pooling samples on detection sensitivity. By pooling equal volumes of 6 CYP samples into one for analysis, each CYP sample has been diluted effectively 6-fold. A higher pooling volume could be used for those samples that display weaker detection sensitivity. Consequently, we have doubled the pooling volume for 1A2 compared to other isozymes. Other considerations would be to increase the injection loop volume, but we found this option unnecessary with our assay conditions.
Conclusion
A high-throughput method for the analysis of samples from a CYP inhibition assay has been developed using online SPE/MS/MS. The method is sensitive and reproducible and has minimal carryover. An issue specific to in-source fragmentation of phenacetin has been addressed. With 1A2/tacrine replacing phenacetin, the system is able to analyze a pool of 96 samples containing 6 isozymes (1A2, 2C8, 2C9, 2C19, 2D6, and 3A4) in less than 15 min, resulting in a 15-fold time savings compared to the currently used 2-min LC/MS/MS method.
