Abstract
Despite significant advances in the management of myocardial infarction (MI), therapeutic options targeting upstream pathogenic mechanisms remain scarce. This study introduces a novel multiomics-to-drug discovery framework to identify and validate causal therapeutic targets for MI. We conducted a systematic two-sample Mendelian randomization (MR) analysis integrating expression quantitative trait loci (eQTL) and protein quantitative trait loci (pQTL) data from the IEU OpenGWAS database, with replication in the UK Biobank cohort. Causal inference was rigorously validated using HEIDI heterogeneity tests, Bayesian colocalization, bidirectional MR, and multivariate MR (MVMR) to account for potential confounders. Downstream applications were explored via protein-protein interaction (PPI) network analysis, phenome-wide association studies (PheWAS), and molecular docking simulations. Initial screening identified four candidate genes (BMP1, APOB, FABP2, and ALDH2) associated with MI risk in both discovery and replication cohorts. However, only BMP1 demonstrated consistent causal effects at both transcriptional and proteomic levels, passing all sensitivity analyses with no evidence of horizontal pleiotropy in PheWAS. Colocalization and bidirectional MR further confirmed BMP1 as a robust, independent causal driver of MI. Molecular docking revealed that UK-383367, a selective BMP1 inhibitor, exhibits high binding affinity to the BMP1 active site. While BMP1 is traditionally associated with extracellular matrix remodeling, this study provides the first genetic evidence establishing it as an independent causal risk factor for MI, distinct from conventional traits such as hypertension. By bridging causal genetic inference with structure-based drug prediction, we propose BMP1 inhibition, specifically via agents like UK-383367, as a promising therapeutic strategy to mitigate MI-related pathological remodeling.
Introduction
Myocardial infarction (MI) remains a leading cause of global mortality and disability. While current clinical strategies, including thrombolysis and percutaneous coronary intervention (PCI), have significantly improved survival rates, the management of postinfarction remodeling and the prevention of recurrent events remain challenging due to limited therapeutic targets and drug-related side effects.1–3 The success of drug development heavily relies on the precise identification of causal therapeutic targets; indeed, targets supported by human genetic evidence are twice as likely to succeed in clinical trials compared to those without such validation.4,5 Therefore, shifting from observational associations to causal inference is critical for discovering effective and safe treatments for MI.
Summary-data-based Mendelian Randomization (SMR) has emerged as a powerful tool to bridge this gap. By integrating genome-wide association study (GWAS) data with molecular quantitative trait loci (molQTL, eg, eQTL and pQTL), SMR leverages the random allocation of genetic variants to infer causal relationships between gene expression/protein levels and disease outcomes, effectively mimicking randomized controlled trials. 6 This approach minimizes confounding factors and reverse causation, which often plague traditional observational studies. While SMR has been successfully applied to various cardiovascular conditions,7,8 its potential to identify novel, causally driven drug targets specifically for MI remains underexplored. 9
Bone morphogenetic protein 1 (BMP1) is a key enzyme involved in collagen processing and extracellular matrix remodeling. Previous studies have reported elevated BMP1 levels in fibrotic tissues and observed its association with cardiovascular remodeling. 10 However, these findings largely rely on observational data or animal models, leaving the direction of causality in human MI unclear. It remains unknown whether BMP1 elevation is a primary driver of MI pathogenesis or merely a secondary consequence of tissue injury. Furthermore, no study has systematically evaluated BMP1 as a specific drug target for MI, distinct from its general role in fibrosis. 11
To address these critical knowledge gaps, this study leverages multiomics SMR analysis to definitively infer the causal effect of BMP1 on MI risk in humans. Crucially, we go beyond establishing causality by integrating drug prediction databases and molecular docking simulations. We explicitly identify and validate UK-383367, a specific small-molecule inhibitor, as a promising repurposing candidate. By transitioning BMP1 from a known fibrotic marker to a validated causal therapeutic target with a concrete drug hypothesis, our work offers a novel translational pathway that significantly exceeds current understanding of BMP1's role in cardiovascular disease.
Methods
Study Design
Figure 1 illustrates the research process. We performed SMR analysis on the MI cohort from the IEU-GWAS, using eQTL database from the GTEx and deCODE pQTL data as instrumental variables. Gene intersections were selected, and the target genes reaching the significance threshold were further validated in the UK Biobank cohort. A rigorous sensitivity analysis was conducted, and the consistency of the results across the discovery and replication phases was compared. To assess the combined causal effect of multiple risk factors, we employed multivariable Mendelian randomization (MVMR) analysis. To rule out potential confounding from linkage disequilibrium and heterogeneity, colocalization and HEIDI tests were also performed. Finally, PheWAS and drug prediction analyses were used to evaluate the potential clinical applicability of the identified drug targets.
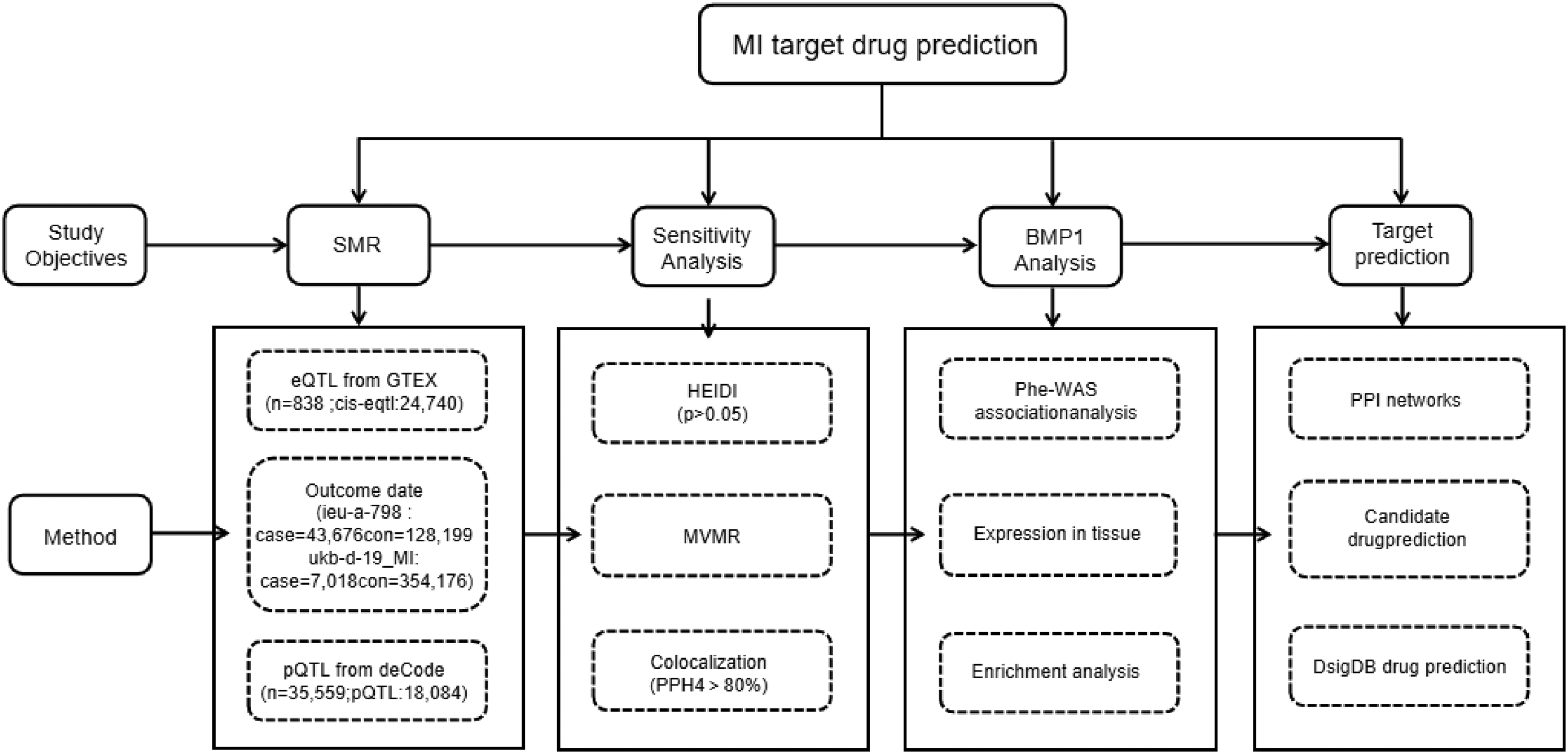
Overview of the study design.
eQTL
Expression quantitative trait loci (eQTL) data were obtained from the Genotype-Tissue Expression (GTEx) Consortium (v8; https://gtexportal.org/), a comprehensive resource characterizing the genetic regulation of gene expression and splicing across human tissues. 12 The GTEx v8 dataset encompasses 17,382 samples from 52 distinct tissues and two cell lines, derived from 838 donors. The cohort demographic composition includes 715 individuals of European American ancestry (85.3%), 103 African Americans (12.3%), 12 Asian Americans (1.4%), and 16 Hispanic or Latino individuals (1.9%), with a sex distribution of 557 males (66.4%) and 281 females (33.5%).
Given that myocardial infarction (MI) is fundamentally a pathology of cardiac tissue involving extracellular matrix (ECM) remodeling, we adopted a tissue-specific prioritization strategy to maximize biological relevance and minimize tissue-specific bias. While previous studies have occasionally utilized proxy tissues such as whole blood, such approaches risk diluting causal signals specific to cardiac pathogenesis. Since BMP1 is predominantly expressed in cardiac fibroblasts and plays a pivotal role in ventricular wall remodeling, we prioritized heart-specific tissues for eQTL extraction. Accordingly, our primary analysis focused on Heart Left Ventricle and Heart Atrial Appendage. To ensure robustness and explore potential systemic effects, we also included vascular tissues (Artery Aorta, Artery Coronary, and Artery Tibial) and Whole Blood as secondary contexts. This targeted selection ensures that the identified genetic instruments reflect the molecular mechanisms directly operative in MI-related cardiac remodeling.
pQTL
Protein quantitative trait locus (pQTL) data were sourced from the landmark study by Ferkingstad et al, which represents the largest large-scale integration of the plasma proteome with genetics to date. 13 This dataset encompasses 4907 proteins measured in 35,559 individuals of Icelandic descent, identifying 18,084 significant genetic associations. To ensure the validity of instrumental variables (IVs) for Summary-data-based Mendelian Randomization (SMR) analysis and to minimize bias from horizontal pleiotropy, we implemented a rigorous multistep filtering pipeline focused on cis-pQTLs: (1) We restricted our analysis to genetic variants located within 1 Mb upstream or downstream of the transcription start site of the corresponding protein-coding gene. This strategy prioritizes variants with a high likelihood of directly regulating protein expression while reducing confounding from distant regulatory effects. (2) Only SNPs demonstrating strong association evidence (genome-wide significance, P < 5 × 10−8) were retained. (3) Variants within the Major Histocompatibility Complex (MHC) region (chr6: 25-34 Mb) were excluded due to the region's exceptionally high gene density and complex linkage disequilibrium (LD) patterns, which frequently generate spurious associations. (4) SNPs with a minor allele frequency (MAF) < 0.01 were removed to avoid weak instrument bias and ensure statistical robustness. (5) To guarantee the independence of selected IVs, LD clumping was performed using a window of 10,000 kb and an r2 threshold of <0.001. This step ensures that only the most significant independent variant is retained for each locus, thereby minimizing the risk of correlated instruments violating MR assumptions. This stringent selection process yielded a high-confidence set of cis-pQTL instruments, optimizing the causal inference power for subsequent SMR analyses.
Outcome Data
The analysis utilized two different datasets to ensure consistency in exposure. The outcome data were all derived from individuals of European ancestry. The first dataset was obtained from the IEU OpenGWAS project, accessible at https://gwas.mrcieu.ac.uk/. This dataset published in October 2015, included 171,875 participants (43,676 patients and 128,199 controls) and analyzed 9,289,492 SNPs. 14 The replication dataset was sourced from the UK Biobank, published in 2018, and can be accessed at https://www.ukbiobank.ac.uk/. In this dataset, there were 7018 cases of MI and 354,176 healthy controls. The number of SNPs analyzed in this dataset was 12,640,541. 15 All MI cases met the diagnostic criteria for the fourth myocardial infarction adopted jointly by the European Society of Cardiology (ESC)/ American College of Cardiology (ACC)/American Heart Association (AHA)/ World Heart Federation (WHF) in 2018. 16 Importantly, there is no sample overlap between the two datasets. This multidataset synthesis method strengthens the validity of our findings across different populations.
Risk Factors
We identified five risk factors for MI, including dyslipidemia, hypertension, diabetes, arrhythmia, and coronary atherosclerosis (Table S1). Under the original GWAS agreement, all individuals gave informed consent. In addition, all ethical permissions related to GWAS are guaranteed by the original authors of GWAS.
SMR and HEIDI Tests
Summary-data Mendelian Randomization (SMR) is statistically more efficient than traditional Mendelian Randomization (MR), offering better control over potential confounders when exposure and outcomes come from independent samples with large sample sizes. 17 SMR enhances causal inference by incorporating gene expression data, such as eQTL or pQTL, to directly link gene effects to phenotypes. To ensure robust results, a strict P value threshold of 5.0 × 10−8 was applied, focusing on the gene center (±500 kb) and selecting relevant cis-QTLs. SNPs with median gene frequency differences >0.2 were excluded from multiple datasets, which included the 1,000 Genomes European population, QTL aggregate data, and GWAS aggregate data.
The heterogeneity in dependent instrument (HEIDI) test was employed to distinguish pleiotropy from linkage. SNPs with high (r2 > 0.9) or very low (r2< 0.05) linkage disequilibrium were excluded. The HEIDI test was conducted only when the number of cis-SNPs was ≥3, with a P value threshold of <.05 considered indicative of pleiotropy, prompting removal from the analysis. To control the false discovery rate (FDR), the Benjamini-Hochberg method was applied with a significance level of .05. SMR and HEIDI tests were conducted using the online tool Biowinford (http://biowinford.site:3838/trial/).
MVMR Analysis
Multivariable Mendelian Randomization (MVMR) analysis allows researchers to examine how multiple genotypic or phenotypic traits interact to influence the performance of a genetic trait, thereby revealing complex relationships between them. 17 In this study, five risk factors associated with MI were selected for MVMR analysis. The results revealed a significant independent association between BMP1 and MI (P = .001).
Colocalization Analysis
Colocalization analysis was conducted using Bayesian methods to identify shared causal variation between the eQTL or pQTL of the MI and APOB/FABP2/BMP1/ALDH2 genes. 18 Five posterior probabilities (PP) corresponding to different hypotheses were reported: (1) H0: Variation is not associated with either gene expression or MI; (2) H1: Variation is only associated with gene expression; (3) H2: Variation is only associated with MI disease; (4) H3: Variation is independently associated with gene expression and MI; and (5) H4: Variation is associated with both gene expression and MI. For colocalization between pQTL-GWAS and eQTL-GWAS, the colocalization window was set to ±1,000 kb. A posterior probability (PPH4) > 0.8 was considered evidence for colocalization. The analysis was performed using the “coloc” R package.
Phe-WAS Association Analysis
To assess potential horizontal pleiotropy of drug targets, we conducted a phenome-wide association study (PheWAS) using the AstraZeneca PheWAS portal (https://azpheas.com/). The original study utilized data from 15,500 binary and 1500 continuous phenotypes based on a large exome sequencing subset from the UK Biobank. Detailed methods can be found in the original article. 19 We applied a correction with a threshold of 2E−9 (the default value on the AstraZeneca PheWAS portal) to control for false positives. This comprehensive PheWAS analysis provides valuable insights into the genetic underpinnings of complex traits and aids in evaluating the safety and efficacy of drug targets.
PPI Networks
We used protein–protein interaction (PPI) networks to further evaluate relevant protein targets found in SMR analyses that were associated with MI risk (P < .05). The PPI network is available at String (https://cn.string-db.org), which describes functional protein association networks. In this study, a string with a confidence score of 0.4 was set as the minimum required interaction score, which was used to screen out protein pairs with strong interactions. If two proteins have an interaction score below 0.4, the interaction between them may be weak and therefore not included in the analysis. All other parameters are left at default levels. 20 PPI results are further processed via Cytoscape (V3.10.3) software. 21
Enrichment Analysis
To better understand the role of BMP1 and its causal mechanism in the interaction between genes and myocardial infarction (MI), we incorporated BMP1 and its interacting genes into the Metscape platform (https://metascape.org/gp/index.html#/main/step1). After setting parameters (Min Overlap = 1; P value cutoff = .01; Min Enrichment = 1.5), we performed Gene Ontology (GO) analysis. Data visualization and further bioinformatics analysis were conducted using the bioinformatics analysis platform (http://www.bioinformatics.com.cn).
Expression in Tissue
To analyze BMP1 expression in human tissues, we utilized the Human Protein Atlas (https://www.proteinatlas.org). 22 This atlas includes protein expression data from 44 normal human tissues, covering 15,323 genes (76%). Investigating both the mRNA and protein levels of BMP1 across different tissues can help identify potential therapeutic targets for myocardial infarction (MI).
Candidate Drug Prediction
DSigDB (https://dsigdb.tanlab.org/DSigDBv1.0/) is a comprehensive database that not only provides information on differentially expressed genes, but also integrates data on drugs and their gene associations. It contains 22,527 gene sets and 17,389 unique compounds linked to 19,531 genes. This extensive data allows researchers to explore the effects of drugs on gene expression and their potential connections to specific diseases. 23 Additionally, we investigated whether drugs target or interact with drug-related genes by searching the Comparative Toxicology Genomics Database (CTD, https://ctdbase.org/). We identified drugs interacting with these genes, supported by at least one study in the CTD database. 24
Molecular Docking
To elucidate the atomic-level interactions between candidate drugs and their target genes, and to evaluate their druggable potential, molecular docking simulations were performed. Three-dimensional structures of the target proteins were retrieved from the Protein Data Bank (PDB, http://www.rcsb.org/), 25 while ligand structures were obtained from the PubChem Compound Database (https://pubchem.ncbi.nlm.nih.gov/). 26 Prior to docking, protein structures underwent rigorous preprocessing using AutoDock Tools, which included the removal of crystallographic water molecules, addition of polar hydrogen atoms, and assignment of Gasteiger charges. Similarly, ligand molecules were optimized by adding hydrogen atoms and minimizing energy to ensure stable conformations.
Docking simulations were executed using AutoDock v. 4.2.6 software (http://autodock.scripps.edu/), 27 a widely validated algorithm for predicting binding poses and affinities. The docking grid box was meticulously defined to encompass the entire active site of the target protein, ensuring comprehensive sampling of potential binding modes. All resulting protein-ligand complexes were visualized and analyzed using PyMOL v2.6.0. 28 To prioritize compounds with significant therapeutic potential, we established a stringent threshold for binding affinity. Based on established criteria in structure-based drug discovery, a binding energy (ΔG) of ≤−7.0 kcal/mol was defined as the cutoff for “high-affinity” interactions.29,30 This threshold is scientifically grounded in thermodynamic principles, where values more negative than −7.0 kcal/mol typically correspond to a dissociation constant (Kd) in the micromolar to nanomolar range, indicating stable complex formation and strong spontaneous binding capability. 31
Results
Multiomics SMR Analysis and Sensitivity Test Showed That BMP1 Gene Was Causally Associated With MI
First, the MI research queue was selected from the IEU OpenGWAS database. SMR analysis was performed using eQTL data from the Artery Aorta, Artery Coronary, Artery Tibial, Heart Left Ventricle, and Whole Blood datasets (Table S2). To enhance the consistency of gene regulation and expression effects, SMR analysis was repeated at the protein level using plasma pQTL data from the MI study cohort (Table S3). Simultaneously, an independent MI replication cohort from the UK Biobank database was selected to ensure independence between the study and replication cohorts. The eQTL-GWAS and pQTL-GWAS analysis results for the replication cohort are detailed in Table S4 and Table S5. By integrating significant QTLs from both the discovery and replication cohorts, we identified four robust candidate genes associated with MI: BMP1, APOB, FABP2, and ALDH2 (Figure 2). These four genes demonstrated highly significant causal associations, with all SMR P-values remaining significant after stringent False Discovery Rate (FDR) correction (FDR < 0.05) (Table 1). Furthermore, the consistency of these signals across independent cohorts and multiple tissue types underscores the reliability of these findings as potential therapeutic targets.

Venn diagram of MI-related genes.
The Genes Associated With MI Identified by SMR Analysis.

Note: Every SNP that used was cis-acting. p_SMR: P < .05 suggests significant causal relationship between gene expression and target phenotype. p_GWAS: P < .01 indicates a statistically significant association between the SNP and the phenotype; p_eQTL: P < 5E-8 determine the SNP significantly affects the gene; FDR: False Discovery Rate, P < .05 indicates a lower proportion of false positive results in multiple hypothesis tests; SNP: single nucleotide polymorphism.
In the heterogeneity detection, we applied the criteria of p_HEIDI > 0.05 and n_SNP ≥ 3, excluding the APOB, FABP2, and ALDH2 genes. BMP1 (p_HEIDI = 0.08362655; n_SNP_HEIDI = 3) met the criteria for inclusion (Table S6-S7). To further assess the causal relationship between BMP1 and MI, we used the ggplot2 package in R 4.4.1, converted the SMR results, and generated a forest plot (Figure 3). The results demonstrated that BMP1 expression was positively correlated with the occurrence of MI, suggesting that high expression of BMP1 may increase the risk of myocardial infarction. This causal relationship was consistent across both the discovery and replication cohorts, as well as at both the gene expression and protein expression levels. Additionally, reverse MR analysis, which excluded the effect of MI on BMP1 expression, confirmed the protective role of BMP1 in MI (Figure S1).

The forest map shows the cause-effect of BMP1 genes in the combined discovery phase and replication phase.
To avoid correlated exposures and false positive results from two different causal SNPs, we conducted a colocalization analysis between BMP1 and MI (Figure 4). The PPH4 value of 0.999748510822785 supported the fifth hypothesis, showing that the rs73225842 variant is associated with both BMP1 expression and MI. To test the independent correlation between BMP1 and MI, we conducted MVMR analysis in six diseases related to MI. 32 The results showed that there was no causal association between BMP1 and these six diseases, which revealed that there might be an independent correlation between BMP1 and myocardial infarction (P = .05).
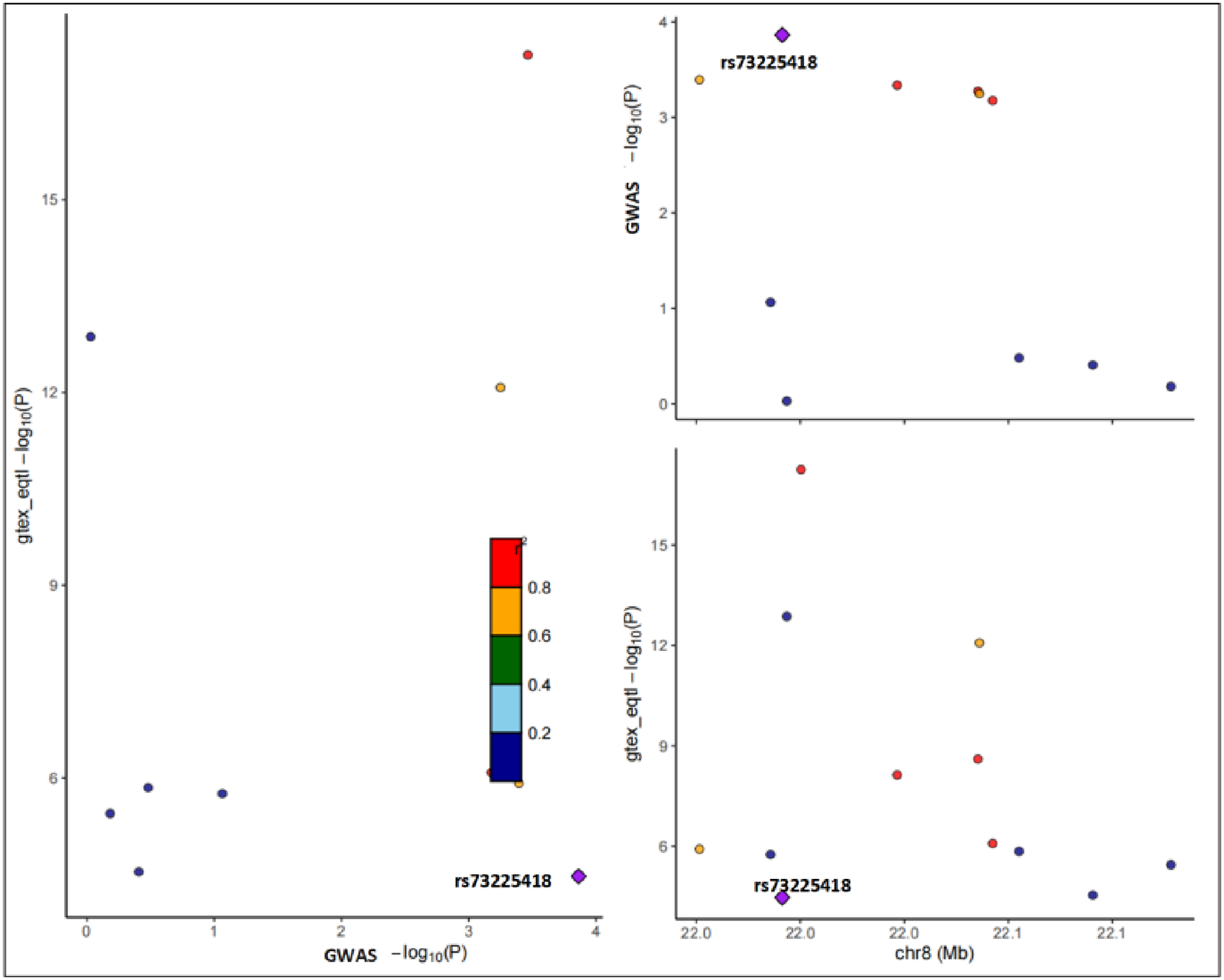
Regional plot of colocalization evidence of BMP1 and MI.
Feasibility Analysis of BMP1 Drug Target
To further assess potential beneficial or harmful effects and pleiotropy associated with BMP1 as a drug target, phenotypic association analysis was performed using the PheWAS portal database. PheWAS results provide insights into associations between gene-determined protein expression and specific diseases or traits. No significant associations were found between BMP1 and other phenotypes at the genome-wide significance level in either the PheWeb database (Table S8) or the PheWAS portal (Figure 5) (P < 5e-08). These findings support the potential of BMP1 as a therapeutic target and suggest a low risk of adverse drug reactions or unintended pleiotropic effects if BMP1 is targeted for therapy.

Phenotypic association diagram of BMP1 gene. (A) Binary phenotype map; (B) Continuous phenotype map.
We identified tissue-specific expression patterns of BMP1 using human protein mapping. The results showed that BMP1 was primarily expressed in the gut, placenta and myocardium (Figure 6A), while it exhibited low specificity in the brain (Tau specificity score = 0.14). Further analysis of core specificity cells revealed that BMP1 was highly enriched in fibroblasts of cardiac tissue, offering insights into the target tissues and pathways of BMP1 (Figure 6B).
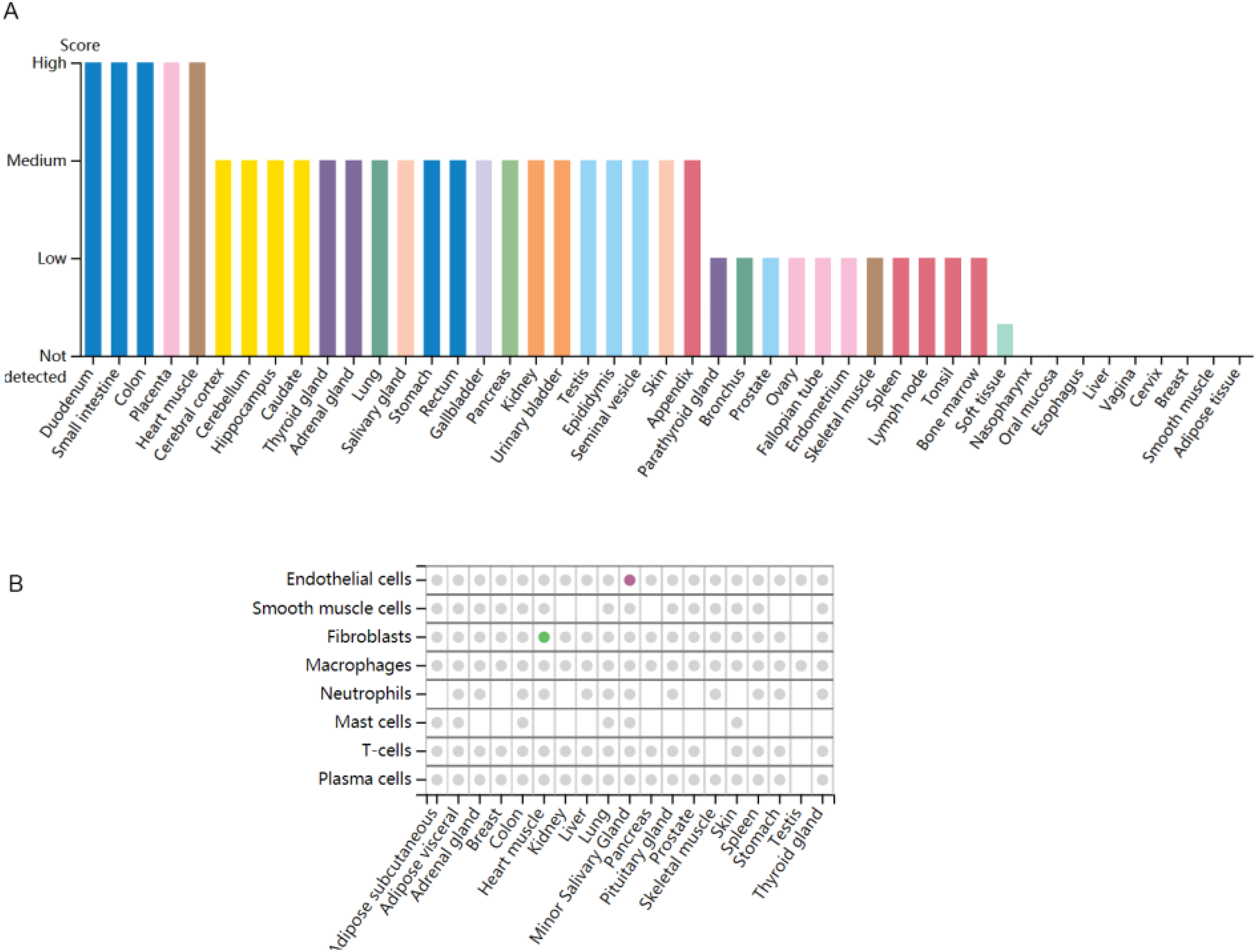
(A) Protein expression overview and (B) core cell types: colored dots indicate that the selected gene has core cell type specificity in the indicated tissue. Grey dots indicate that cell type is present within that tissue, but the selected gene is not predicted to be enriched there.
The BMP1 target gene was uploaded to the STRING database (https://cn.string-db.org) to construct a protein–protein interaction (PPI) network, and the resulting data were visualized using Cell Landscape. Figure 7 illustrates the interactions of BMP1 in an 11-node, 32-edge PPI network. These interacting genes were further analyzed for Gene Ontology (GO) enrichment. As shown in Figure 8, the most significant biological processes (BP) are related to cell growth, including cell differentiation, development and migration. In the cellular component (CC) category, drug target genes are enriched in the cytoplasm and in structures associated with cell growth, such as the extracellular matrix (containing collagen), BMP receptor complex and basement membrane. In terms of molecular function (MF), these genes are involved in activities related to cell growth regulation, protein production, and modification. These findings align with the pathological mechanisms of MI disease, supporting the protective role of BMP1 in the progression of MI.
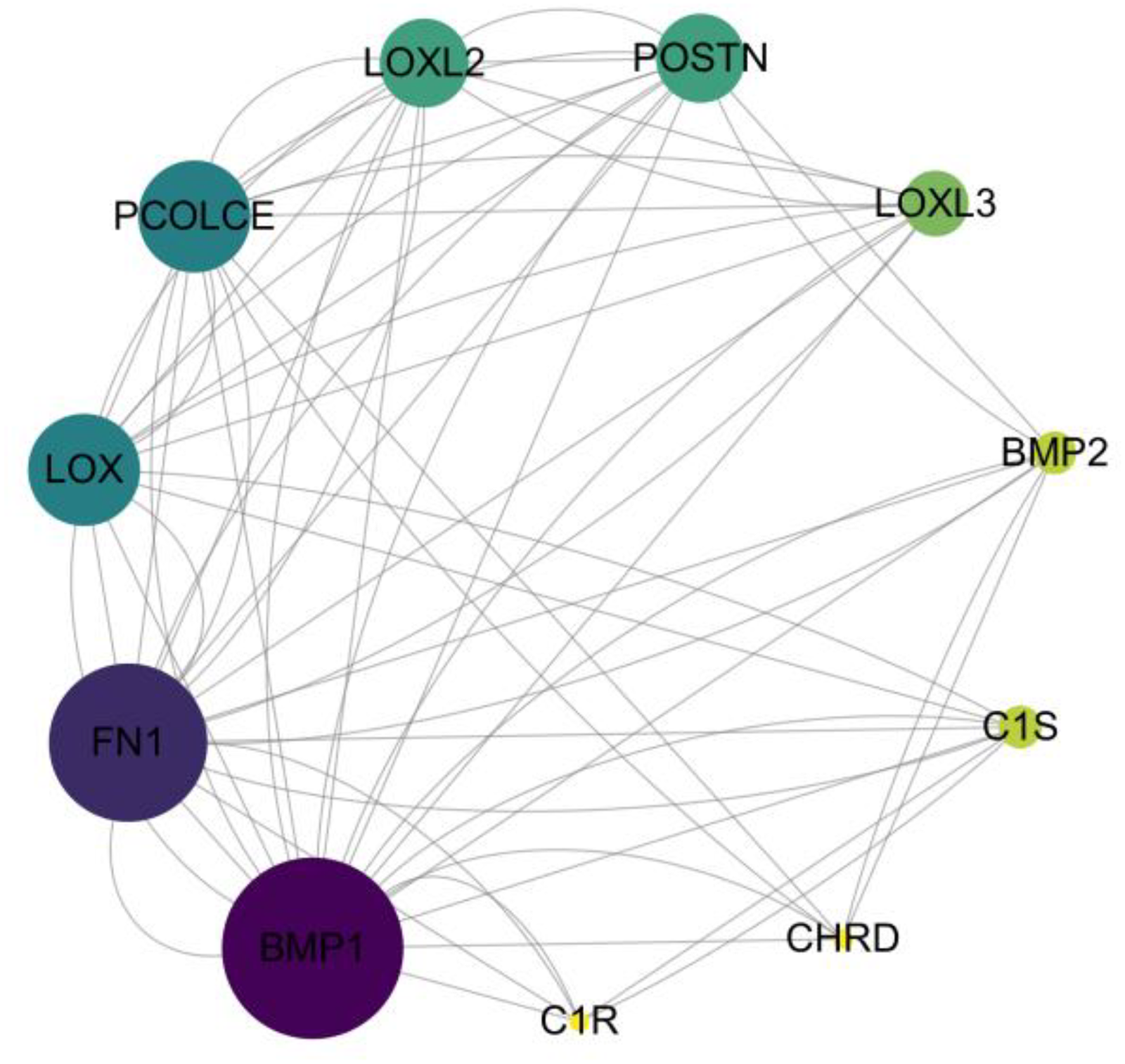
PPI network built with STRING. The size and color of nodes are related to the connectivity. The larger the connectivity, the darker the color and the larger the area.
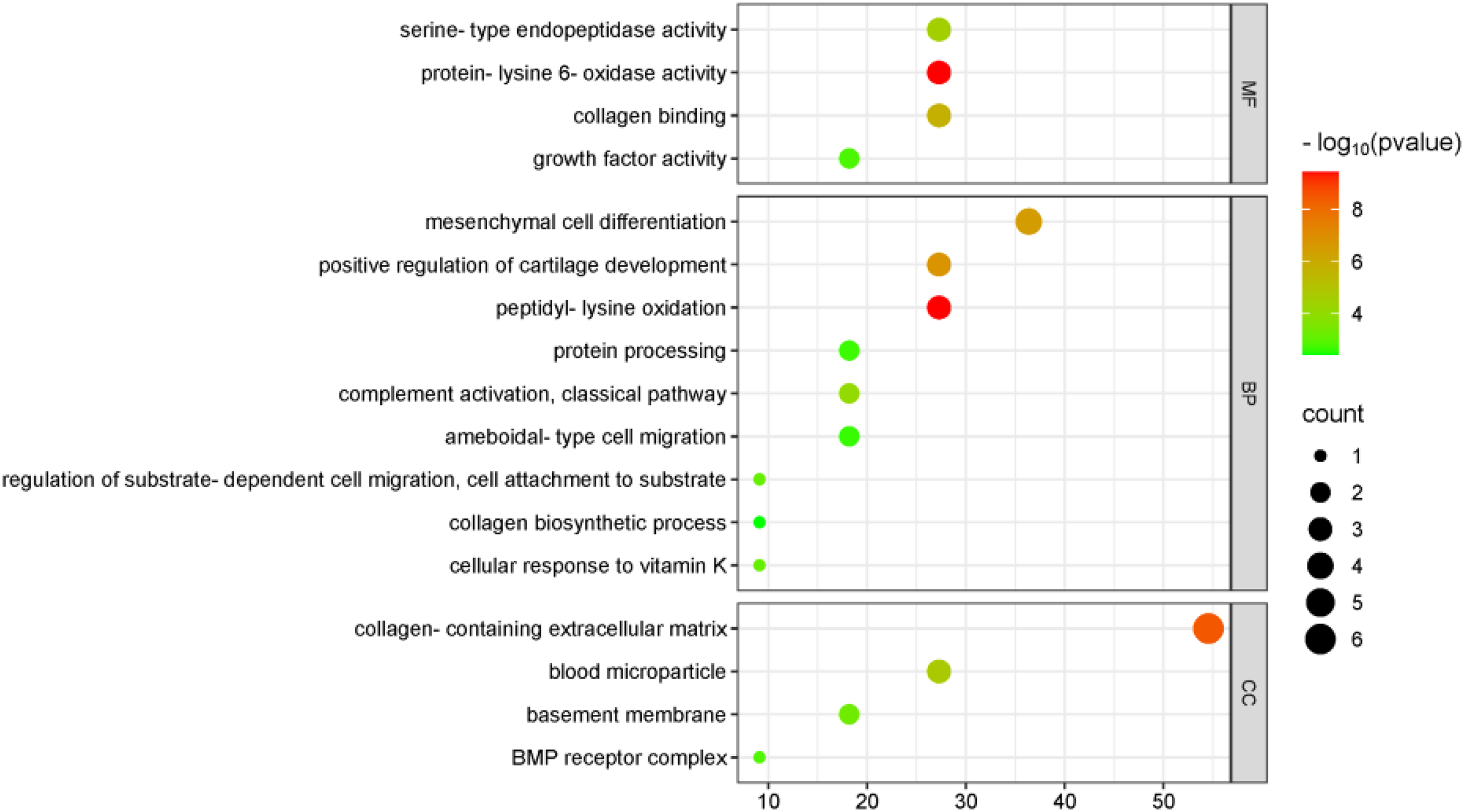
Go enrichment results for three terms.
The PPI network constructed using GeneMANIA (https://genemania.org/) was also employed to investigate the roles and functions of related genes. This network included an additional 20 potential interacting genes, with a total of 270 interaction links (Figure 9). Among these interactions, 77.64% were physical interactions, 8.01% were co-expression, and 5.37% were predicted. Functional analysis of this network highlighted the role of drug targets and their associated genes. The results were consistent with the previous enrichment analysis, showing a strong correlation with cell growth-related functions.

The PPI network established by GeneMANIA. Each node is colored to represent the functional pathway in which the gene is involved, and the node size represents the connection strength.
To further validate the functional mechanisms identified via STRING, we constructed an independent PPI network using GeneMANIA (https://genemania.org/). The functional enrichment analysis of this expanded network strongly corroborated our previous findings, consistently highlighting significant associations with cell growth-related processes and extracellular matrix components (Figure S3). This convergence of evidence from two distinct algorithms robustly supports the hypothesis that BMP1 exerts its protective role in MI by regulating cell growth and structural integrity.
BMP1 Drug Prediction and Analysis
To evaluate BMP1 as a potential drug target, protein-drug interactions were assessed using the DSigDB drug database to identify drugs associated with BMP1. The results revealed the top four drugs linked to BMP1: 2-Nonenal, 4-Hydroxy-, (2E,4R) (CTD 00006994), Phenytoin (CTD 00006994), Cobaltous chloride (CTD 00006994), and UK-383367 (CTD 00006994) (Table 2).
BMP1 Gene Candidate Drug Prediction.

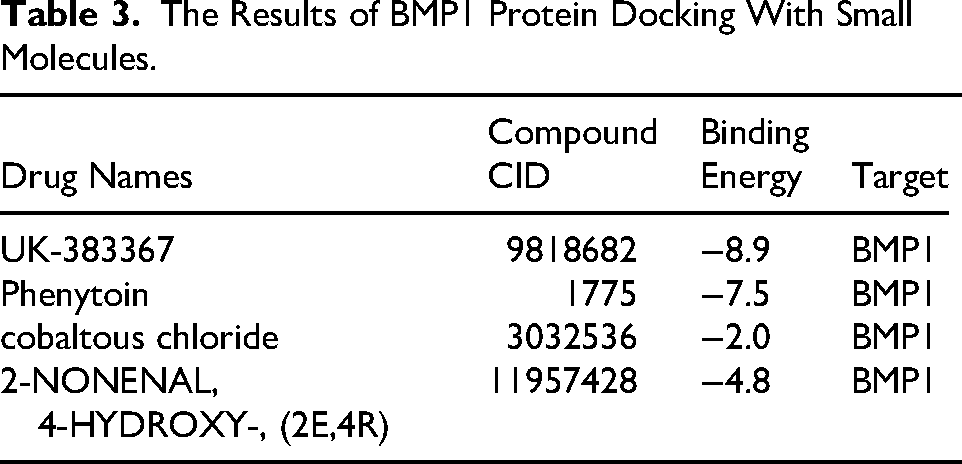
To evaluate the stability of the drug target and assess its potential properties, molecular docking was conducted in this study. AutoDock v4.2.6 was used to identify binding sites and calculate binding energies for interactions between the drug candidates and the proteins encoded by the corresponding genes (Table 3 and Figure 9). Each drug candidate formed visible hydrogen bonds and strong electrostatic interactions with its protein target. Additionally, the binding pocket for each target was successfully occupied by the drug candidates. Among them, BMP1 and UK-383367 showed the lowest binding energy (−8.9 kcal/mol), indicating a highly stable interaction.
The Results of BMP1 Protein Docking With Small Molecules.
Discussion
This study explores the potential causal relationship between eQTLs, pQTLs, and the risk of MI, as well as investigates potential therapeutic targets. First, we combined SMR analysis and colocalization analysis to establish the causal relationship between BMP1 and MI risk. However, in our bidirectional Mendelian randomization study, we did not observe any reverse causality, thus excluding the possibility of reverse causality or bias caused by linkage disequilibrium. Although phenotypic scans showed that BMP1 was associated with hypertension, acute ischemic heart disease, cardiomyopathy and atrioventricular block, further supporting the relationship between BMP1 and MI, we cannot completely rule out the potential causal role of hypertension and coronary artery disease in the association between BMP1 and myocardial infarction. MVMR analysis showed that there was no direct causal relationship between BMP1 and diabetes, hypertension, arrhythmia, dyslipidemia and atherosclerosis (P = .01), suggesting that there was an independent association between BMP1 and MI, and it might serve as a potential target for the treatment of myocardial infarction.
The biological plausibility of BMP1 as a therapeutic target lies in its central role in extracellular matrix (ECM) remodeling, a process characterized by a delicate balance between tissue repair and pathological fibrosis. As a member of the TGF-β superfamily, BMP1 functions as a procollagen C-proteinase, essential for cleaving pro-collagen into mature fibrillar collagen.33–35 This function creates a “double-edged sword” in the context of MI. On one hand, acute upregulation of BMP1 immediately post-MI is physiologically necessary to stabilize the infarct zone and prevent ventricular rupture by facilitating rapid scar formation. On the other hand, sustained or excessive BMP1 activity drives maladaptive fibrosis, leading to stiffening of the ventricular wall, impaired diastolic function, and adverse remodeling.36–39 This is consistent with our research findings. Preclinical studies in rat MI models have demonstrated that BMP1 inhibition significantly attenuates type I collagen deposition and left ventricular fibrosis, thereby preserving cardiac function and preventing adverse wall thinning. 40 Similar antifibrotic benefits of BMP1 inhibition have been observed in liver and kidney fibrosis models,41,42 although context-dependent effects remain a subject of debate, particularly in pulmonary fibrosis.43,44 However, translating BMP1 inhibition into clinical therapy requires careful consideration of its dual nature. While targeting BMP1 offers promise for attenuating pathological fibrosis, there is a legitimate concern that systemic or prolonged inhibition could compromise the initial wound healing process, potentially increasing the risk of cardiac rupture or impairing the structural integrity of the scar in the acute phase. Furthermore, given BMP1's role in processing various ECM components (eg, decorin, biglycan, laminin) across multiple tissues, off-target effects on connective tissue homeostasis in other organs cannot be ruled out. 40 Therefore, the therapeutic window for BMP1 inhibitors likely depends on timing and specificity.
PheWAS suggests that BMP1 has a relatively low potential for adverse effects as a drug target and significantly reduces the likelihood of pleiotropic biases. The drug targets analysis shows that compounds UK-383367, Phenytoin, cobaltous chloride, and 4-hydroxy-2-nonenal (4-HNE) can target BMP1, and molecular docking visually demonstrated the binding between these compounds and BMP1. UK-383367 (Binding energy: −8.9) and Phenytoin (Binding energy: −7.5) exhibited strong binding affinity and excellent binding modes with BMP1. Existing studies have shown that Phenytoin (PHT) influences atherosclerosis through cholesterol regulation,45–47 although the precise molecular mechanism remains unclear. BMP1 can regulate plasma LDL levels by cleaving human low-density lipoprotein receptor (LDLR),48–50 suggesting that BMP1 may serve as an intermediary in the effect of Phenytoin (PHT) on atherosclerosis.
UK-383367(5-[(3R)-6-cyclohexyl-1-(hydroxyamino)-1-oxohexan-3-yl]-12,4-oxadiazole-3-carboxamide), a specific BMP1 inhibitor, is a novel procollagen C-proteinase inhibitor used for treating postsurgical skin scars. 51 However, due to BMP1's critical role in collagen assembly, using BMP1 as a cholesterol-lowering drug target may carry potential risks of adverse effects. 49 Similarly, UK-383367 is a potent and specific procollagen C-proteinase inhibitor developed primarily for limiting cutaneous scarring. 51 While its high specificity is advantageous, its mechanism directly interferes with collagen assembly. As discussed earlier, systemic inhibition of BMP1 in the context of MI carries a double-edged risk—while it may attenuate pathological fibrosis, it could also compromise the structural integrity of the infarct scar, potentially leading to ventricular rupture or impaired healing. Furthermore, the pharmacokinetic properties of UK-383367 for systemic cardiac delivery remain unexplored. In conclusion, while our docking results provide valuable structural insights and confirm the druggability of the BMP1 pocket, neither Phenytoin nor UK-383367 currently emerges as an immediately feasible repurposing candidate for MI treatment without further optimization.
This study represents a paradigm shift by establishing the first robust human genetic evidence confirming a direct causal relationship between BMP1 and myocardial infarction (MI), thereby elevating BMP1 from a descriptive biomarker of fibrosis to a validated therapeutic target. While prior studies have largely relied on animal models or observational correlations to link BMP1 with cardiovascular remodeling, our work leverages a rigorous multiomics Mendelian randomization framework to demonstrate that genetically modulated BMP1 levels directly drive MI risk, independent of conventional confounders such as hypertension and lipid profiles. Most critically, we transcend the limitations of existing literature by delivering a concrete, safety-validated drug repurposing strategy. Unlike previous reports, we not only pinpointed UK-383367 as a high-affinity selective inhibitor through molecular docking but also performed a critical safety filtration to exclude theoretically binding yet clinically hazardous candidates. This dual approach of causal validation coupled with precision drug screening provides a novel, actionable roadmap for therapeutic intervention that significantly advances beyond the current state of knowledge, offering a specific and safe hypothesis for clinical translation.
However, several limitations must be acknowledged to contextualize these findings. First, our analysis relied exclusively on summary-level data from individuals of European ancestry. Given the variability in linkage disequilibrium patterns and allele frequencies across populations, our findings may not fully generalize to non-European groups (eg, Asian, African, or Hispanic populations), potentially introducing population-specific biases. Future efforts must prioritize cross-ancestry validation using emerging multiethnic biobanks to ensure the universal applicability of BMP1 as a target. Second, despite the robustness of our computational pipeline (including HEIDI tests, colocalization, and MVMR), our conclusions remain largely in silico. We currently lack wet-lab experimental validation (eg, in vitro binding assays or in vivo MI models) to confirm the actual therapeutic efficacy and pharmacokinetics of UK-383367 in a cardiac context. Consequently, our proposed “therapeutic window"—suggesting inhibition during the subacute-to-chronic phase to avoid impairing acute wound healing—remains a hypothesis requiring urgent empirical testing. Third, while MVMR adjusted for major risk factors, we cannot entirely rule out residual confounding by unmeasured environmental or lifestyle factors that might influence both BMP1 expression and MI risk. Finally, due to data availability constraints, we were unable to perform sex-stratified analyses. Given the known sexual dimorphism in cardiac fibrosis and remodeling, future studies utilizing individual-level data are essential to determine if BMP1 inhibition strategies require sex-specific tailoring.
Supplemental Material
sj-xlsx-1-cpt-10.1177_10742484261440344 - Supplemental material for Multiomics Mendelian Randomization of GWAS, eQTL, and pQTL Combined With Drug Analysis to Predict Drug Targets in Myocardial Infarction
Supplemental material, sj-xlsx-1-cpt-10.1177_10742484261440344 for Multiomics Mendelian Randomization of GWAS, eQTL, and pQTL Combined With Drug Analysis to Predict Drug Targets in Myocardial Infarction by Didi Yuan, PhD, Lian Hu, Yanqing Huang, PhD, Haotian Tang, Yuchen Ma, Sai Hu, and Junwen Liu, PhD in Journal of Cardiovascular Pharmacology and Therapeutics
Supplemental Material
sj-jpg-2-cpt-10.1177_10742484261440344 - Supplemental material for Multiomics Mendelian Randomization of GWAS, eQTL, and pQTL Combined With Drug Analysis to Predict Drug Targets in Myocardial Infarction
Supplemental material, sj-jpg-2-cpt-10.1177_10742484261440344 for Multiomics Mendelian Randomization of GWAS, eQTL, and pQTL Combined With Drug Analysis to Predict Drug Targets in Myocardial Infarction by Didi Yuan, PhD, Lian Hu, Yanqing Huang, PhD, Haotian Tang, Yuchen Ma, Sai Hu, and Junwen Liu, PhD in Journal of Cardiovascular Pharmacology and Therapeutics
Supplemental Material
sj-jpg-3-cpt-10.1177_10742484261440344 - Supplemental material for Multiomics Mendelian Randomization of GWAS, eQTL, and pQTL Combined With Drug Analysis to Predict Drug Targets in Myocardial Infarction
Supplemental material, sj-jpg-3-cpt-10.1177_10742484261440344 for Multiomics Mendelian Randomization of GWAS, eQTL, and pQTL Combined With Drug Analysis to Predict Drug Targets in Myocardial Infarction by Didi Yuan, PhD, Lian Hu, Yanqing Huang, PhD, Haotian Tang, Yuchen Ma, Sai Hu, and Junwen Liu, PhD in Journal of Cardiovascular Pharmacology and Therapeutics
Supplemental Material
sj-jpg-4-cpt-10.1177_10742484261440344 - Supplemental material for Multiomics Mendelian Randomization of GWAS, eQTL, and pQTL Combined With Drug Analysis to Predict Drug Targets in Myocardial Infarction
Supplemental material, sj-jpg-4-cpt-10.1177_10742484261440344 for Multiomics Mendelian Randomization of GWAS, eQTL, and pQTL Combined With Drug Analysis to Predict Drug Targets in Myocardial Infarction by Didi Yuan, PhD, Lian Hu, Yanqing Huang, PhD, Haotian Tang, Yuchen Ma, Sai Hu, and Junwen Liu, PhD in Journal of Cardiovascular Pharmacology and Therapeutics
Footnotes
Author Contributions
DY and LH contributed to conception and design of the study. QH and TT collected and analyzed experimental data. CM performed the statistical analysis. DY wrote the paper. LH prepared the figures and tables. JL and SH revised the paper for intellectual content. All authors contributed to manuscript revision and read and approved the submitted version.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by grants from the Education Reform Project of Hunan Province (No. HNJG-2022-04591), and from the Project of Top-notch Student Training Bases for Basic Disciplines (2.0) (No. 20232039), and Natural Science Foundation of Hunan Province (NO. 2025JJ50668),and Natural Science Foundation of Hunan Province (NO. 2025JJ50668).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statements
Supplemental Material
Supplemental material for this article is available online.
