Abstract
Background
The differential diagnosis of ovarian cancer is important, and there has been ongoing research to identify biomarkers with higher performance. This study aimed to evaluate the diagnostic utility of combinations of cancer markers classified by machine learning algorithms in patients with early stage ovarian cancer, which has rarely been reported.
Methods
In total, 730 serum samples were assayed for lactate dehydrogenase (LD), neutrophil-to-lymphocyte ratio (NLR), human epididymis protein 4 (HE4), cancer antigen 125 (CA125), and risk of ovarian malignancy algorithm (ROMA). Among them, 53 were diagnosed with early stage ovarian cancer, and the remaining 677 were diagnosed with benign disease.
Results
The areas under the receiver operating characteristic curves (ROC-AUCs) of the ROMA, HE4, CA125, LD, and NLR for discriminating ovarian cancer from non-cancerous disease were .707, .680, .643, .657, and .624, respectively. ROC-AUC of the combination of ROMA and LD (.709) was similar to that of single ROMA in the total population. In the postmenopausal group, ROC-AUCs of HE4 and CA125 combined with LD presented the highest value (.718). When machine learning algorithms were applied to ROMA combined with LD, the ROC-AUC of random forest was higher than that of other applied algorithms in the total population (.757), showing acceptable performance.
Conclusion
Our data suggest that the combinations of ovarian cancer-specific markers with LD classified by random forest may be a useful tool for predicting ovarian cancer, particularly in clinical settings, due to easy accessibility and cost-effectiveness. Application of an optimal combination of cancer markers and algorithms would facilitate appropriate management of ovarian cancer patients.
Keywords
Highlights
Risk of ovarian malignancy algorithm (ROMA) is a widely used ovarian cancer marker. ROMA consists of cancer antigen 125 (CA125), human epididymis protein 4 (HE4), and menopause state. Lactate dehydrogenase (LD) related to cancer is a routinely prescribed biomarker. The diagnostic utility of combined cancer markers with LD for ovarian cancer, which has been seldom reported, was evaluated. Machine learning algorithms were applied to differential diagnosis of early stage ovarian cancer from benign conditions for better performance. Areas under the receiver operating characteristic curves (ROC-AUCs) and sensitivities at 75.0% specificities were measured. The combination of ROMA with LD classified by random forest showed the best ROC-AUC (.757, 68.8% sensitivity at 75.0% specificity), indicating acceptable usefulness for differential diagnosis of ovarian cancer.
Introduction
A total of 295 414 new ovarian cancer patients were diagnosed and 184 799 cancer-related deaths occurred worldwide in 2018. 1 Ovarian cancer is the eighth most common cause of cancer-related death in women in South Korea. 2 The age-standardized incidence rate of ovarian cancer has increased progressively from 5.0 to 6.3 based on the Korea Central Cancer Registry. 3 Unfortunately, the physical inaccessibility of the ovaries and the lack of specific symptoms in the early stages of ovarian cancer make differential diagnosis difficult. Several patients undergo extensive surgical staging, such as oophorectomy, without a definite diagnosis of malignant cancer, leading to increased morbidity. 4
There has been ongoing research to identify biomarkers with higher diagnostic performance in differentiating ovarian cancer from other benign conditions. Among diverse biomarkers, cancer antigen 125 (CA125) is a representative marker for detecting and guiding treatment in patients with ovarian cancer. However, CA125 showed a wide range of sensitivity (27–66%) regarding detection of early stage ovarian cancer due to high false-positive rate among premenopausal women with benign diseases.5,6 Therefore, there have been attempts to find other biomarkers that can complement or replace CA125. Among them, human epididymis protein 4 (HE4) has been reported to have better specificity than CA125 in discriminating benign from malignant ovarian masses. 7 The combination of these markers and the menopausal status of patients led to the proposition of the risk of ovarian malignancy algorithm (ROMA) to predict ovarian cancer. 8 However, there have been discrepancies regarding the reported diagnostic performances of CA125, HE4, and ROMA in previous studies. 7 In addition, multiple markers other than these representative ovarian markers and their combinations of them based on machine learning algorithms have been also investigated and showed inconsistent results.9,10
In this study, we evaluated the diagnostic value of combinations of conventional ovarian cancer markers with routinely prescribed markers such as lactate dehydrogenase (LD) and neutrophil-to-lymphocyte ratio (NLR), which were rarely reported, to identify practically useful tools for differentiating early stage ovarian cancer from benign diseases in clinical settings. We also applied machine learning algorithms including bagging, boosting, classification tree, random forest, support vector machine, and K-nearest neighbor algorithms10,11 to investigate the optimal diagnostic performance of combinations of multiple ovarian cancer markers.
Materials and Methods
Study Population
A total of 743 samples from patients who visited Kangnam Sacred Heart Hospital for ROMA testing were collected consecutively to demonstrate the diagnostic performance of ovarian cancer markers between June 2014 and October 2016. We excluded 35 patients with malignant diseases other than ovarian cancer. Three patients diagnosed with advanced stage (n = 2 for stage III and n = 1 for stage IV) were also excluded to investigate only patients with early stage ovarian cancer. Additionally, 25 patients diagnosed with early stage ovarian cancer between November 2016 and December 2020 were included for more thorough analyses. The 730 samples without duplicated patients were classified according to patients’ diagnosis as follows: ovarian cancer group (n = 53) and control group (n = 677) (Supplementary Figure 1). All patients were diagnosed by specialized gynecologists and pathologists in their clinics at Kangnam Sacred Heart Hospital based on the criteria of the International Federation of Gynecology and Obstetrics12,13 for ovarian cancer. The control group included patients with benign pelvic masses such as simple cysts of ovary, and leiomyoma of uterus, reflecting actual clinical laboratory conditions. The dataset analyzed in this study is provided in Supplementary Table 1. The procedures for the determination of major laboratory parameters used for ovarian cancer markers were described as follows. The medical technicians and researchers were blinded to the test results.
HE4, CA125, and ROMA
HE4 serum concentration was determined using a commercially available Alinity i HE4 Reagent kit (Abbott Diagnostics, Abbott Park, IL, USA), which was used according to the manufacturer’s instructions. A two-step chemiluminescent microparticle immunoassay was used for quantitative analysis of HE4. Serum samples were incubated with 2H5 anti-HE4-coated paramagnetic microparticles. After non-bound antibodies were washed out, an acridinium-labeled 3D8 anti-HE4 conjugate was added. After another wash, pretrigger and trigger solutions were combined with the reaction complexes. The resulting chemiluminescent reaction was measured as relative light units (RLUs). The amount of HE4 antigen in the serum and the RLUs detected by the Alinity i HE4 assay exhibited a direct relationship and the results were calculated automatically by the analyzer. CA125 was also detected using a two-step chemiluminescent microparticle immunoassay with Alinity i CA125 II Reagent kit (Abbott Diagnostics). Serum samples and paramagnetic microparticles coated with ovarian cancer 125 were incubated for binding to CA125 reactive determinants to the particles. After washing, a M11 acridinium-labeled conjugate was added to the mixture. The followed steps were similar to the protocol for Alinity i HE4 Reagent kit. ROMA was calculated according to a study by Moore et al. 8 as follows.
Premenopausal: PI (predictive index) = −12.0 + 2.38 * LN (HE4) + .0626 * LN (CA125).
Postmenopausal: PI = −8.09+1.04 * LN(HE4) + .732 * LN(CA125).
Then, ROMA value (predictive value) was calculated using the following equation: ROMA (%) = ePI/(1 + ePI) * 100
LD and NLR
The AU LD reagent kit (Beckman Coulter, Inc, Brea, CA, USA) was applied on Beckman Coulter AU5800 to quantitate LD levels. Lactate and nicotinamide adenine dinucleotide (NAD) were converted to pyruvate and NADH catalyzed by LD. NADH strongly absorbs light at 340 nm, whereas NAD does not. The rate of change of absorbance at 340 nm is directly proportional to LD activity in serum samples. Values were calculated automatically by the analyzer. The Siemens Advia 2120i Hematology System (Siemens Health care Diagnostics, Deerfield, IL, USA) was used to count total and differential white blood cells (WBCs). This flow cytometry–based system uses a combination of reactions that occur within the peroxidase and the basophil/nuclear lobularity channels. A cluster analysis of the cells within each channel was used to generate a cytogram in which the x-axis reflected nuclear complexity and the y-axis reflected cell size. We calculated NLR by dividing the neutrophil counts to the lymphocyte counts provided by this hematology system.
Statistical Analysis
Statistical analyses were performed using Analyse-it Method Evaluation Edition, version 2.26, software (Analyse-it Software Ltd., Leeds, UK), PASW version 18.0 (SPSS Inc, Chicago, IL, USA), and R statistical software (version 3.6.3, R Foundation for Statistical Computing, Vienna, Austria). Comparisons of nominal variables and continuous variables between groups were assessed with Pearson’s chi-square and Mann–Whitney U tests, respectively. The adjusted P-values were calculated using the Benjamini–Hochberg method 14 for multiple tests. Variables satisfying the Benjamini–Hochberg method were included for receiver operating characteristic (ROC) analysis. ROC curves were plotted for ovarian cancer markers and their combinations with LD in order to assess their diagnostic ability to differentiate between ovarian cancer and control groups. The areas under ROC curves (AUCs) of ovarian cancer markers and their combinations with LD were compared. The numbers on the curve present the degree of accuracy as follows: no discrimination (AUC < .5), acceptable (.7 < AUC < .8), excellent (.8 < AUC < .9), and outstanding (.9 < AUC). 15 Binary logistic regression analysis was used to calculate the predicted probability values of the combinations of ovarian cancer markers and LD, and these values were used to estimate ROC-AUCs, similar to the previously described method. 16 The presence of ovarian cancer as the outcome and the results from ovarian cancer marker identification were used as predictor variables. P-values less than .05 were considered statistically significant. In addition, widely used and available machine learning algorithms used for ovarian cancer markers such as bagging, boosting, classification tree, random forest, support vector machine, and K-nearest neighbor algorithms10,11 were applied to our datasets for better diagnostic performance in differentiating ovarian cancer from control. The ratio of independent datasets used for training and testing, which were randomly separated, was 7:3. Three-fold cross-validation was performed for machine learning analyses. When these machine learning algorithms were applied to our data, the values of markers as they stand were used for analyses.
Results
Study Population Characteristics
Basic Characteristics and Laboratory Results Related to Ovarian Cancer of the Study Population.
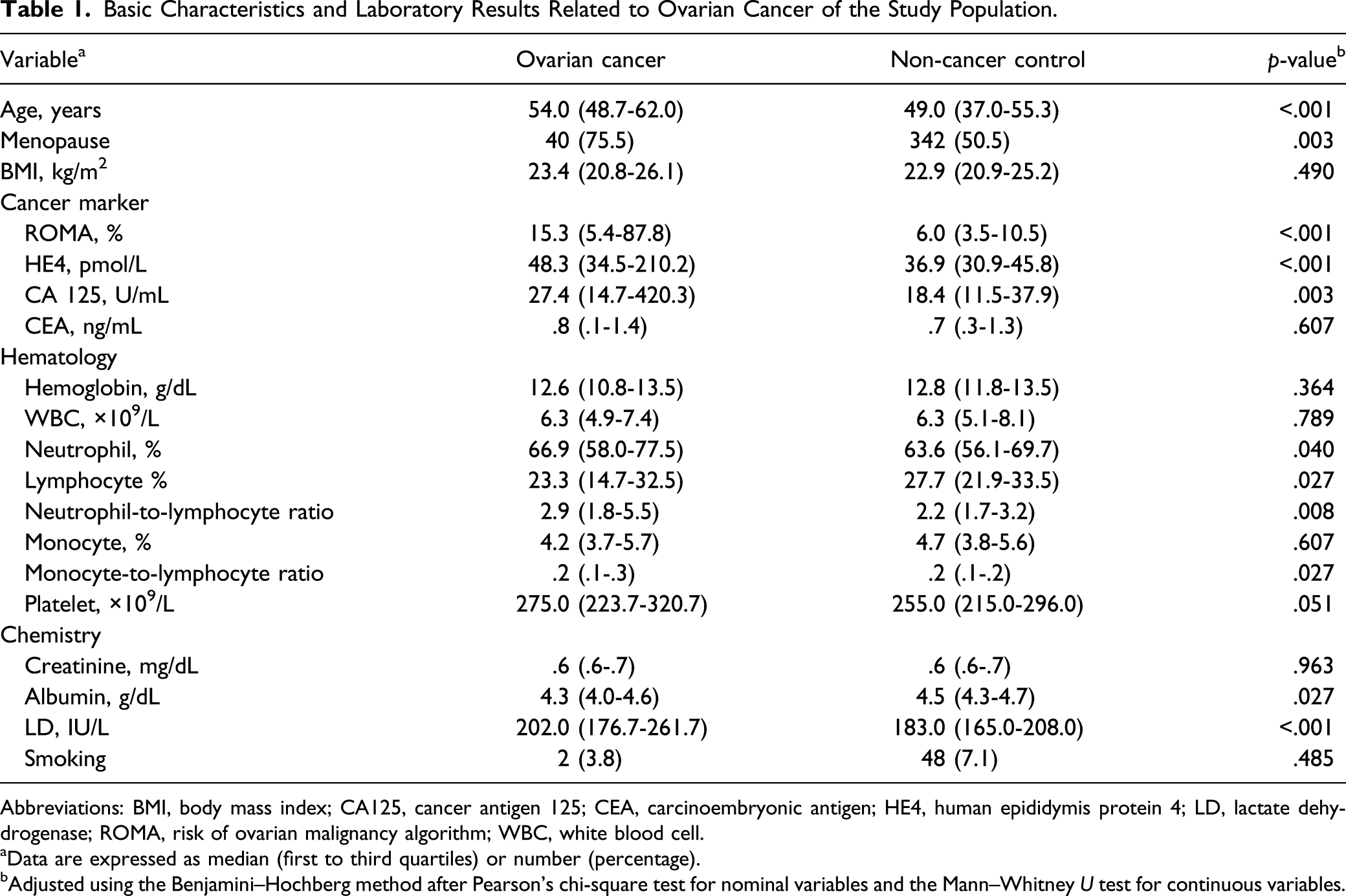
Abbreviations: BMI, body mass index; CA125, cancer antigen 125; CEA, carcinoembryonic antigen; HE4, human epididymis protein 4; LD, lactate dehydrogenase; ROMA, risk of ovarian malignancy algorithm; WBC, white blood cell.
aData are expressed as median (first to third quartiles) or number (percentage).
bAdjusted using the Benjamini–Hochberg method after Pearson’s chi-square test for nominal variables and the Mann–Whitney U test for continuous variables.
Performance of Single Ovarian Cancer Markers
AUCs of ROMA, HE4, CA125, LD, and NLR for differentiating ovarian cancer from all other conditions were .707, .680, .643, .657, and .624, respectively. Among single markers, only ROMA showed acceptable performance based on AUCs. The study cohort was subsequently divided into premenopausal (n = 348) and postmenopausal (n = 382) groups. In sub-group analysis, the AUCs were .580 for ROMA, .589 for HE4, .540 for CA125, .586 for LD, and .609 for NLR in the premenopausal group, while AUCs were elevated in the postmenopausal group (.685 for ROMA, .684 for HE4, .693 for CA125, .635 for LD, and .623 for NLR).
Performance of Single Ovarian Cancer Markers. a

Abbreviations: CA125, cancer antigen 125; HE4, human epididymis protein 4; LD, lactate dehydrogenase; NLR, neutrophil-to-lymphocyte ratio; ROC-AUC, areas under the receiver operating characteristic curve; ROMA, risk of ovarian malignancy algorithm.
aData are shown as value (95% confidence interval).
bSensitivities at 75.0% specificities are presented.
Performances of Combined Ovarian Cancer Markers
The performances of the combinations of ovarian cancer markers were evaluated because these 5 markers showed overlapping ROC curves. In particular, the combination of conventional cancer markers (ROMA, HE4, and CA125) with LD was examined because LD showed better performance than NLR among routinely prescribed laboratory results. The AUCs of ROMA with LD, HE4 with LD, CA125 with LD, and NLR with LD for differentiating ovarian cancers from all other conditions were .709, .692, .698, and .690, respectively. Regarding the combination of more than 3 markers, the AUCs for distinguishing ovarian cancers from other conditions were .708 for ROMA + NLR + LD, .705 for HE4 + CA125 + LD, .698 for HE4 + NLR + LD, .690 for CA125 + NLR + LD, and .696 for HE4 + CA125 + NLR + LD. In sub-group analysis, the AUCs in the premenopausal group (.556 to .600) were lower than those in the postmenopausal group (.687 to .718).
Performance of Ovarian Cancer Markers in Combination. a
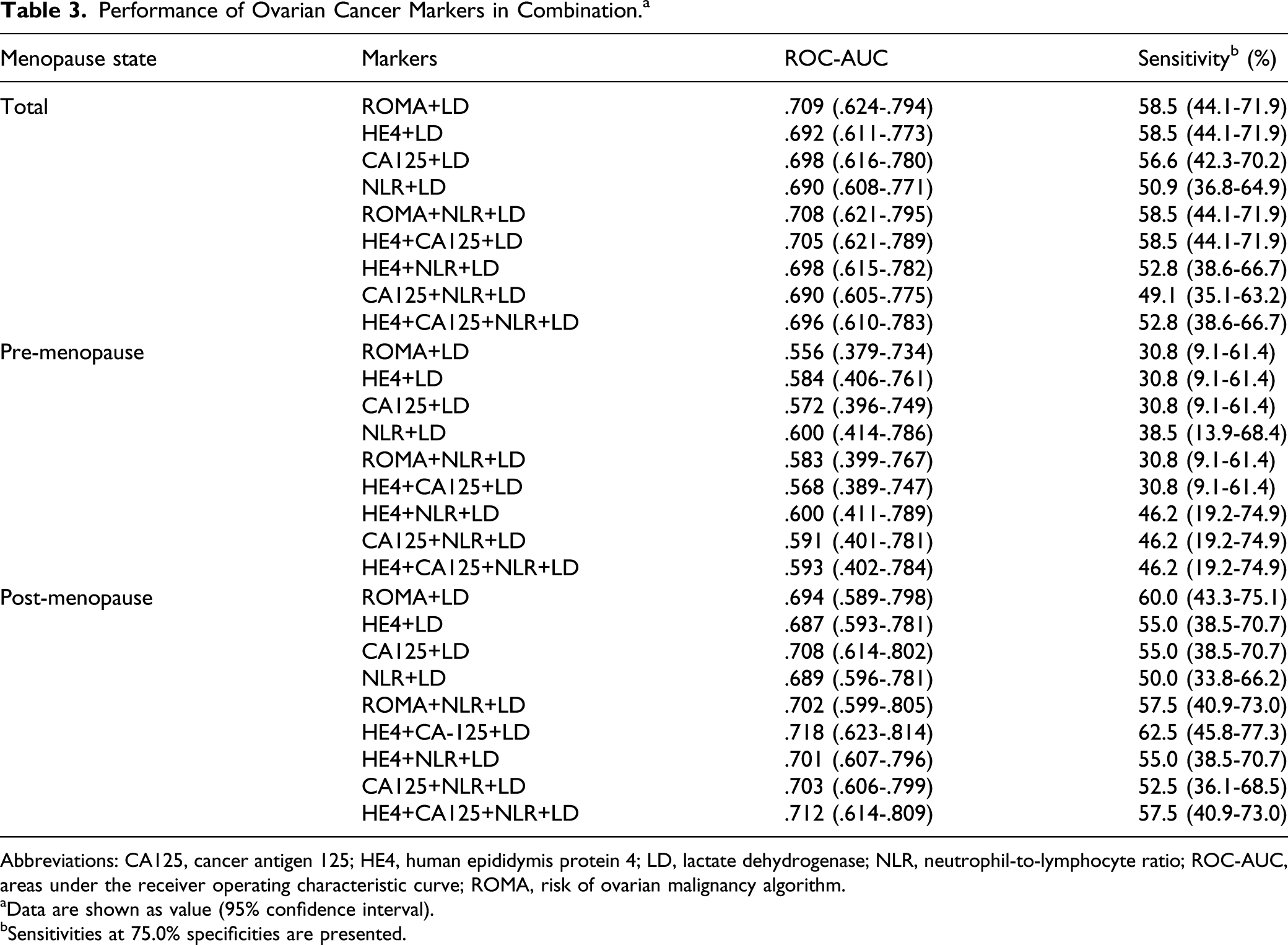
Abbreviations: CA125, cancer antigen 125; HE4, human epididymis protein 4; LD, lactate dehydrogenase; NLR, neutrophil-to-lymphocyte ratio; ROC-AUC, areas under the receiver operating characteristic curve; ROMA, risk of ovarian malignancy algorithm.
aData are shown as value (95% confidence interval).
bSensitivities at 75.0% specificities are presented.
Machine Learning Analysis of Ovarian Cancer Markers
Machine learning analyses, including classification tree, bagging, random forest, adaptive boosting (AdaBoost), support vector machine, and K-nearest neighbor algorithms, were performed. The presence of ovarian cancer was considered the dependent variable. The estimated values of each machine learning algorithm from ROMA + LD for the total cohort and HE4 + CA125 + LD for the postmenopausal group presenting the best AUCs with acceptable performances in conventional combinatorial marker analysis were predictors (Figure 1(A) and (B)). We found that ROMA + LD classified by random forest showed the best AUC (.757, 95% confidence interval [CI] = .615-.898) among machine learning sets. Three-fold cross-validation was performed, and its sensitivity at 75% specificity was 68.8%. Additionally, ROMA for total cohort was analyzed using these machine learning algorithms based on the acceptable AUC value of conventional analysis. The best AUC of single ROMA (.681 by AdaBoost) was lower than that of ROMA + LD (.757 by random forest) (Supplementary Figure 2). Regarding the postmenopausal group, HE4 + CA125 + LD classified by these 6 algorithms revealed AUCs ranging from .500 to .648, which were not higher than those of conventional logistic regression. Machine learning analyses of these algorithms showed AUCs less than .500 for the premenopausal group. Performance of combined markers classified using machine learning for predicting ovarian cancer. (A) ROC curves of ROMA + LD determined by classification tree, bagging, random forest, adaptive boosting, support vector machine, and K-nearest neighbor analyses for distinguishing ovarian cancer from the non-cancer controls, (B) ROC curve of HE4 + CA125 + NLR + LD in the postmenopausal group. The areas under the ROC curves (AUCs) of combined markers are presented in brackets. Abbreviations: AdaBoost, adaptive boosting; CA125, cancer antigen 125; HE4, human epididymis protein 4; KNN, K-nearest neighbor; LD, lactate dehydrogenase; ROC, Receiver operating characteristic; ROMA, risk of ovarian malignancy algorithm; SVM, support vector machine; Tree, classification tree.
Discussion
Here, diagnostic applications of single cancer markers and their combinations with LD were evaluated in Korean patients with early stage ovarian cancer. The diagnostic values of machine learning algorithms for these combinations of cancer markers were also examined.
In terms of single cancer markers, our data showed that ROMA incorporating CA125, HE4, and the menopausal status was the best marker (AUC = .707) for discriminating epithelial ovarian cancer from benign disease. Many studies supported that HE4 was likely more specific than CA125, the conventional ovarian cancer marker.17,18 Consistent with our study, ROMA has been also suggested to be an effective diagnostic tool for the detection of ovarian cancer.8,17,19 In contrast, ROMA and HE4 were not superior to CA125 in postmenopausal groups. The reported diagnostic performance of these markers has been controversial. Some studies revealed no benefit for ROMA.20-22 A prospective validation study showed that ROMA and HE4 alone revealed similar performance to CA125 alone in the premenopausal group, whereas their performance was worse in the postmenopausal group. 23 In another retrospective study, perioperative CA125 alone was superior to ROMA and HE4 in predicting ovarian tumors based on ROC analysis. 24
In addition to these markers, it was also investigated whether LD, which can be obtained in routine chemistry, can complement HE4, ROMA, and CA125. LD is an enzyme that plays a major role in anaerobic glycolysis and is related to the prognosis of patients with various cancers. 25 Serum LD levels in ovarian cancer patients were significantly elevated and were correlated with shorter survival time in previous reports.26,27 Special AT-rich-binding protein 1, a global genome organizer, may reprogram energy metabolism in ovarian cancer by mediating LD levels, thus promoting metastasis. 27 Although there were only a few studies covering LD in ovarian cancer patients, LD was considered as a potential biochemical marker due to diagnostic accuracy with relatively high specificity.26,27
NLR has been reported as a potent prognostic biomarker for progression-free survival and overall survival in ovarian cancer.28,29 Regarding diagnostic utility, some studies demonstrated that preoperative NLR could differentiate ovarian cancer from benign ovarian masses.30-32 In a recent study investigating NLR in a Korean cohort, the AUC for NLR was .709, which was slightly higher than AUC reported in our study (.624, 95% CI = .533-.716). The higher proportion of patients with advanced stages (52.9%) compared to our study cohort with early stages may account for this difference. 30
A recent review supported that combination of HE4 with CA125 has been a highly efficient tool for the diagnosis of ovarian cancer. This combination can bypass variations in HE4 derived from smoking or contraception drugs. 7 The diagnostic performance of this combination with LD has seldom been reported, while combination with NLR has been evaluated in some previous articles. These studies demonstrated that preoperative CA125 in combination with NLR would be more sensitive and cost-effective. Furthermore, this strategy could be conducted routinely for identifying ovarian cancers.33,34
There have been a few studies applying machine learning algorithms for the detection of ovarian cancer.9,10,35 A recent study demonstrated that machine learning algorithms enhanced biomarker specificity for several types of tumors, including ovarian cancer. K-nearest neighbor and classification tree were used to improve specificity, which is challenging in early detection of cancer by conventional serum biomarkers. 35 In another study, machine learning was applied to preoperative diagnosis and prognosis prediction in ovarian cancer based on blood biomarkers. They found that machine learning systems provided critical diagnostic and prognostic prediction before initial intervention. 9 Song et al 10 also adopted machine learning algorithms such as linear discriminant analysis and K-nearest neighbor for the early detection of ovarian cancer. The 3 or 4 combinations, which included transthyretin and prolactin, revealed outstanding performance ranging from .91 to .95 for cancer detection. The study cohort included healthy controls, and the choice of serum biomarker, which was not routinely used in clinical settings, generated differences compared to our study. The utilization of machine learning algorithms may facilitate personalized management and increase the number of treatment options for better outcomes through early stratification of patients.
This study had some limitations. Only a small number of ovarian cancer patients were available from collected samples. In addition, early stage patients would bias toward lower diagnostic performance compared to other studies. However, evaluation of early stage patients is important because early differentiation is correlated with a better outcome. Furthermore, to the best of our knowledge, this is the first study to apply machine learning algorithms to combinations of ovarian cancer-specific markers with LD and NLR, which could be utilized in clinical practice. Additional studies with large sample size are necessary for the validation of our algorithm in ovarian cancer patients.
Conclusion
In conclusion, we evaluated the diagnostic benefit of HE4, CA125, and ROMA combined with LD to identify early stage ovarian cancer patients. Although a few published studies have discussed the usefulness of machine learning algorithms, no study has assessed the diagnostic performance of combinations of ovarian cancer markers with LD using machine learning algorithms for early stage ovarian cancer. The combination of ROMA and LD was acceptable for ovarian cancer patients, and classification by random forest was effective for the differential diagnosis of cancer. Our study provides information on the application of machine learning to combinations of practical biomarkers for patients with early stage ovarian cancer to facilitate appropriate patient management. Because our study results are based on a relatively small sample size of cancer patients, further studies including a larger number of ovarian cancer patients are needed to confirm our study findings.
Supplemental Material
sj-pdf-1-ccx-10.1177_10732748211033401 – Supplemental Material for Evaluation of Combined Cancer Markers With Lactate Dehydrogenase and Application of Machine Learning Algorithms for Differentiating Benign Disease From Malignant Ovarian Cancer
Supplemental Material, sj-pdf-1-ccx-10.1177_10732748211033401 for Evaluation of Combined Cancer Markers With Lactate Dehydrogenase and Application of Machine Learning Algorithms for Differentiating Benign Disease From Malignant Ovarian Cancer in Neuroblastoma by Seri Jeong, Dae-Soon Son, Minseob Cho, Nuri Lee, Wonkeun Song, Saeam Shin, Sung-Ho Park, Dong Jin Lee and Min-Jeong Park in Cancer Control
Footnotes
Acknowledgments
The authors would like to thank staff members of Abbott Diagnostics, Beckman Coulter, and Siemens Healthcare Diagnostics for their technical support.
Abbreviations
AdaBoost, adaptive boosting; BMI, body mass index; CA125, cancer antigen 125; CEA, carcinoembryonic antigen; HE4, human epididymis protein 4; LD, lactate dehydrogenase; NAD, nicotinamide adenine dinucleotide; NLR, neutrophil-to-lymphocyte ratio; PLR, platelet-to-lymphocyte ratio; RLUs, relative light units; ROC-AUCs, areas under the receiver operating characteristic curves; ROMA, risk of ovarian malignancy algorithm; WBC, white blood cell.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Research Foundation of Korea (NRF) grant, funded by the Korean government (Ministry of Science, ICT & Future Planning) [NRF-2017R1C1B2004597].
Ethics Statement
This study was approved by the independent Institutional Review Board of Kangnam Sacred Heart Hospital (HKS 2020-05-009) and was conducted in accordance with the Declaration of Helsinki. Moreover, the need for informed consent was waived because anonymity of personal information was maintained.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
