Abstract
RNAscope in situ hybridization (ISH) detects target RNA in formalin-fixed, paraffin-embedded (FFPE) tissues. Protocols suggest that prolonged FFPE storage and formalin fixation may impact signal detection, potentially limiting the utility of RNAscope ISH in retrospective studies. To develop parameters for RNAscope use with archived specimens, we evaluated the effect of formalin-fixation time by measuring the signal of a reference gene (16srRNA) in selected tissues fixed in 10% neutral-buffered formalin for 1, 2, 3, 5, 7, 10, 14, 21, 28, 60, 90, 180, and 270 d. The signal intensity and percent area of signal decreased after 180 d. Tissues had detectable signal at 180 d but not at 270 d of formalin fixation. To assess target detection in paraffin blocks, we qualitatively compared the signal of canine distemper virus (CDV) antigen via immunohistochemistry and CDV RNA via RNAscope ISH in replicate sections from blocks stored at room temperature for 6 mo, 1, 3, 6, 8, 11, 13, and 15 y; RNA was detected in FFPE tissues stored for up to 15 y. Our results demonstrate that RNAscope ISH can detect targets in tissues with prolonged paraffin storage intervals and formalin-fixation times.
RNAscope (Advanced Cell Diagnostics) in situ hybridization (ISH) has become a popular tool to detect and measure RNA biomarkers in formalin-fixed, paraffin-embedded (FFPE) tissues. 22 Given its unique ability to amplify signal when probes are bound to target RNA with minimal nonspecific background, some researchers are replacing traditional detection methods, such as immunohistochemistry (IHC), with RNAscope ISH technology. Unlike other detection methods, RNAscope applies multiple pairs of target probes with separate tail sequences that hybridize to the target region—this “double Z” probe design controls for nonspecific hybridization. 22 Although RNAscope ISH was developed for and is mainly used in human testing and disease research using rodent and non-human primate animal models, veterinary research studies1,3,14,15,17,18 are increasingly utilizing RNAscope to target viral RNA in situ.
Archival materials represent an important sample source for studying natural disease. In veterinary medicine, particularly in the fields of zoologic and wildlife medicine, most pathology studies are retrospective rather than prospective and thus are limited to archival biomaterials. However, extended formalin fixation and FFPE tissue storage intervals can affect the tissue at a molecular level, potentially limiting the ability to target nucleic acid (DNA/RNA) in archival tissues. The most commonly used solution to preserve tissues, 10% neutral-buffered formalin (NBF), is a solution of 3.7% formaldehyde and 1% methanol. 21 By 24–48 h of tissue fixation, the formaldehyde in 10% NBF is hydrated to form methylene glycol, which penetrates cells, leading to the formation of protein cross-linkages in the tissue. 21 This initial phase is reversible and many techniques (e.g., IHC) utilize antigen or target retrieval to reverse these cross-linkages; the most common antigen-retrieval techniques involve heat, enzymatic degradation, and altering pH.21,23 After ~30 d, however, covalent bonds form in the tissue—this irreversible bond formation can damage RNA and DNA via strand fragmentation and molecular modification by adducts and other cross-links or cause decreased antigenicity through protein cross-links that alter or hide epitopes (i.e., epitope masking) or change the redox and membrane potentials of tissue.21,23 –25 Similarly, storing FFPE tissue samples at room temperature for extended periods might also harm RNA quality, likely as a result of the gradual fragmentation of RNA molecules over time and molecular modification through adducts and other cross-links.8,25
Some studies have concluded that RNA degradation over time in FFPE and formalin-fixed tissues stored at room temperature decreases the ability to detect RNA biomarkers using RNAscope.2,25 As a result, the user manual on FFPE tissue preparation for the RNAscope 2.5 HD Assay–Red (Advanced Cell Diagnostics) recommends that tissues be fixed for 16–36 h in 10% NBF, paraffin-embedded, and stored at 2–8°C with desiccants for proper assay performance. However, another study concluded that RNA quality does not decrease over time in FFPE tissues, allowing biomarkers to be detected using both RNAscope 4 and IHC. 23 Antigen detection in stored tissues by IHC is also dependent on the antigen being tested given that some antigens degrade faster than others. 24
We aimed to observe the trends of the RNAscope signal in samples stored for various times in formalin and in paraffin blocks to better evaluate the utility of using this technique in retrospective studies.
Materials and methods
Formalin-fixation time study
To examine the effect of 10% NBF fixation time on signal detection of the 16S ribosomal RNA (rRNA) reference gene using RNAscope ISH, various tissues were opportunistically obtained from an addax (Addax nasomaculatus), a critically endangered antelope species native to the Sahara Desert. 10 This individual addax had been euthanized because of chronic degenerative joint disease, and a postmortem examination was performed within 24 h of euthanasia. Individual tissue samples, < 1.0-cm thick, of brain, liver, spleen, intestine, testis, kidney, pancreas, thyroid gland, and lung were collected, separated for each time (1, 2, 3, 5, 7, 10, 14, 21, 28, 60, 90, 180, 270 d), and immersion fixed in a 10:1 ratio of 10% NBF (Thermo Fisher). All tissues were collected for each time, excluding intestines and spleen for 60 d, and spleen for 90, 180, and 270 d. Following the allotted formalin-fixation time, tissues were removed from formalin and placed in 70% EtOH (Thermo Fisher) for up to 60 d. Ethanol post-fixed tissue sections were processed routinely, paraffin embedded, sectioned at 5 µm, mounted on slides (Starfrost; Mercedes Medical), and left unstained for use with the RNAscope 2.5 HD Assay–Red. RNAscope ISH was performed at various times within 14 d of preparation of unstained slides. For each time, sections were incubated with the An-16srRNA probe or a negative control (DapB) probe at 1, 60, 180, and 270 d. A rRNA reference gene was selected as a target given that it is present in all tissues, should be expressed sufficiently to detect a significant change in signal over time, and has been utilized as a target to assess tissue quality and suitability for RNAscope assays. 22
RNAscope ISH was performed according to the manufacturer’s instructions. Slides were counterstained with 50% hematoxylin (Electron Microscopy Sciences) for 2 min at room temperature. To mount, each slide was quickly dipped into xylene, 1–2 drops of mountant (EcoMount; BioCare Medical) were added, and a coverslip was placed over each tissue section. Slides were left to dry at room temperature for ≥ 5 min before analyzing under a standard bright-field microscope.
Ten representative 200× images (.tiff files) were collected from RNAscope sections of each tissue and analyzed using ImageJ 19 (NIH). Similar corresponding areas of each tissue were collected for image analysis (i.e., intestinal mucosa [intestine], renal cortex [kidney], centrilobular hepatocytes [liver], seminiferous tubules [testis], thyroid follicles [thyroid], bronchioles and bronchi [lung], exocrine pancreas, cerebral gray matter [brain], and splenic white pulp [spleen]). Images were imported into ImageJ, the image channels were split for each image, and the “green” image was utilized because it highlighted the RNAscope signal amplification regions most effectively. The threshold of each image was adjusted to include colors of 0–136 to highlight only the positive signal. The image was “measured,” and the integrated density (signal intensity; i.e., the sum of all of the pixel values in an image) and percent area (%area) were recorded for each image (Suppl. Fig. 1).
To test variation in formalin-fixation time, signal intensity and %area of signal values from each image from a given tissue and time were averaged. Then, the difference in intensity and %area of signal between days 2–270 and day 1 was calculated for each tissue type. To account for the natural variation in architecture among tissue types, values for all dates were scaled to day 1. An ANOVA was run to determine the variance of signal intensity and %area between times. A p ≤ 0.05 was considered statistically significant. All statistical tests were performed, and graphs were made in R Studio (https://www.r-project.org/). GI tissue and spleen at 60 d and spleen at 90, 180, and 270 d were not available for analysis.
FFPE tissue storage time study
To determine whether RNA signal decreases over time in archival FFPE blocks, we compared RNAscope ISH to IHC for canine distemper virus (CDV; Morbillivirus canis). CDV is considered an important veterinary pathogen, 11 and it affects a wide variety of carnivores worldwide, including raccoons (Procyon lotor). 26
We chose 15 raccoon cases for inclusion in our study based on a range of FFPE tissue storage times and quality of tissues (i.e., no autolysis). FFPE tissue blocks with the following storage times were included in this study: 15 y (n = 2), 13 y (n = 1), 11 y (n = 1), 8 y (n = 1), 6 y (n = 1), 3 y (n = 5), and ≤ 1 y (n = 4). Initial tissue collection and fixation were performed within 4–24 h postmortem. Tissues from each case had been immersion fixed in 10% NBF and routinely processed and embedded in paraffin within 24 h to 10 d of collection. Representative tissues varied between cases and included brain, kidney, salivary gland, spleen, and lung. Five-µm thick tissue sections were cut from selected archived FFPE blocks and mounted to X-tra (Leica) or adhesive (Starfrost; Mercedes Medical) slides for use with the RNAscope 2.5 HD Assay–Red. RNAscope ISH was performed on all slides within a week of slide processing according to the manufacturer’s instructions using a custom probe developed to detect the L gene (V-CDV-L) and a negative control probe (DapB). Slides were counterstained and coverslipped as above.
To test the impact of paraffin block storage time on signal, RNAscope ISH–treated sections were qualitatively compared to IHC-treated sections for each raccoon to evaluate signal for tissue localization, pattern, and quality using bright-field microscopy. IHC to detect CDV utilizing antibody against CDV nuclear protein (DV2-12, 1:5,000 dilution; Custom Monoclonals International, Biocare Medical) had been performed previously (contemporaneously with the case submission). These IHC slides were preserved with mounting medium, coverslipped, and stored in the dark at room temperature from the time of initial case evaluation until further analysis for this study, ranging from < 1 to 15 y.
Results
Formalin-fixation time study
The negative control (DapB) had no signal in all tested tissues and at all times tested. Expression of the 16s rRNA reference gene was identified successfully by RNAscope in all tissues up to 180 d of formalin fixation using the An-16srRNA probe (Fig. 1). There was no significant difference in the change of RNAscope signal intensity (ANOVA, p = 0.115, df = 12, F = 1.56) and %area of signal (ANOVA, p = 0.128, df = 12, F = 1.52) in tissues across all times. There was, however, a significant decrease over time in the values of both signal intensity and %area of signal between days 1 and 270 (p = 0.034 and p = 0.024, respectively) tested with a t-test.
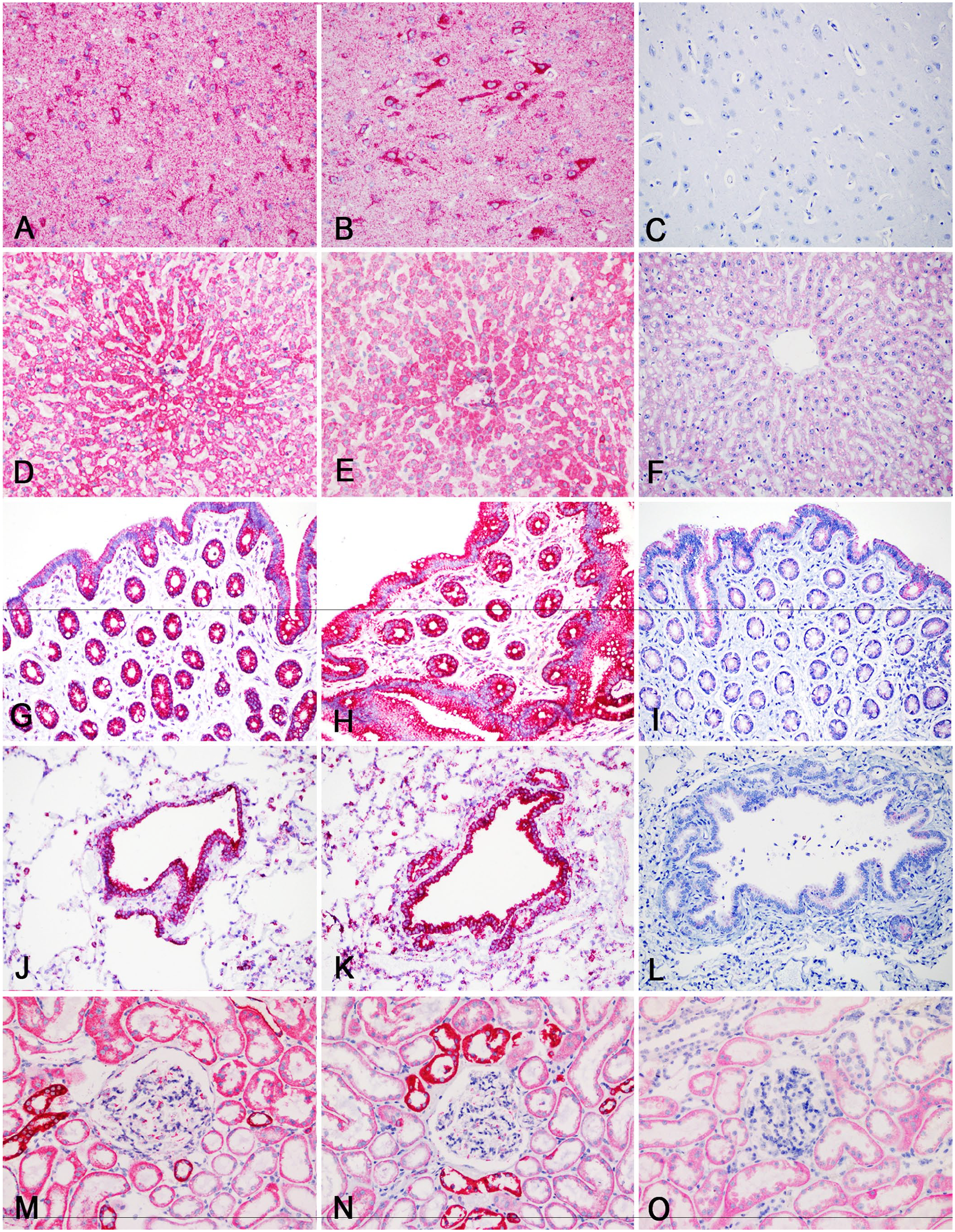
Comparison of RNAscope in situ hybridization signal amplification of an An-16srRNA probe in various addax (Addax nasomaculatus) tissues over days in 10% neutral-buffered formalin.
Signal intensity and %area of signal in some tissues began decreasing dramatically at different times (Figs. 2, 3). The 5-d formalin-fixation samples had decreased signal compared to some of the later times, and signal on days 10 and 14 for specific tissues (e.g., testis, lung, thyroid) was decreased (Figs. 2, 3). Overall, tissue signal intensity and %area of signal began to decrease significantly after 180 d. Positive signal was observed in all tissues up to 180 d in formalin.
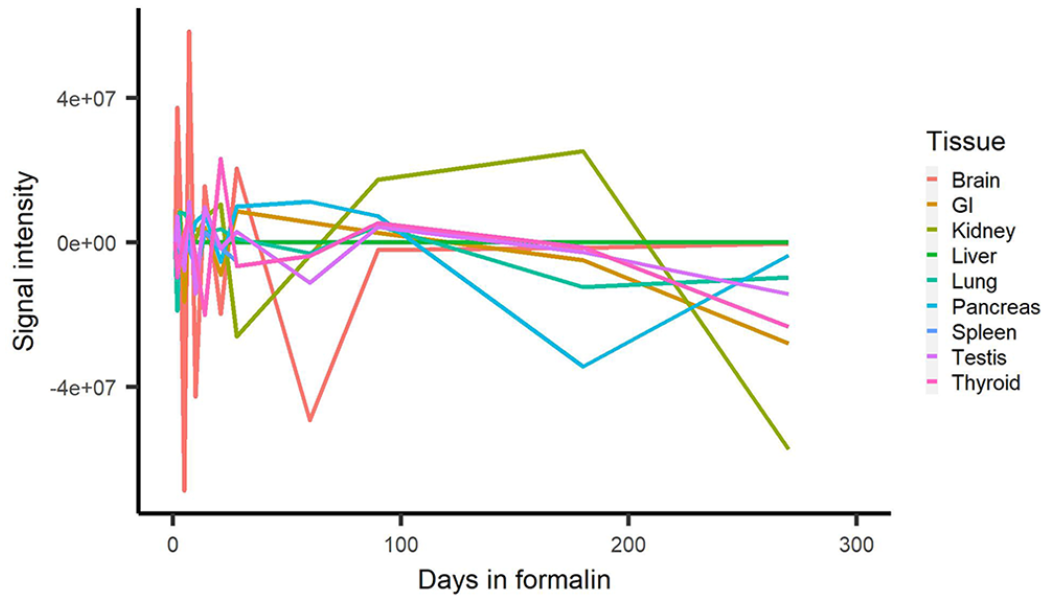
Comparison of the change in RNAscope in situ hybridization signal intensity (i.e., the sum of all of the pixel values in an image) of all tissues over days in 10% neutral-buffered formalin.
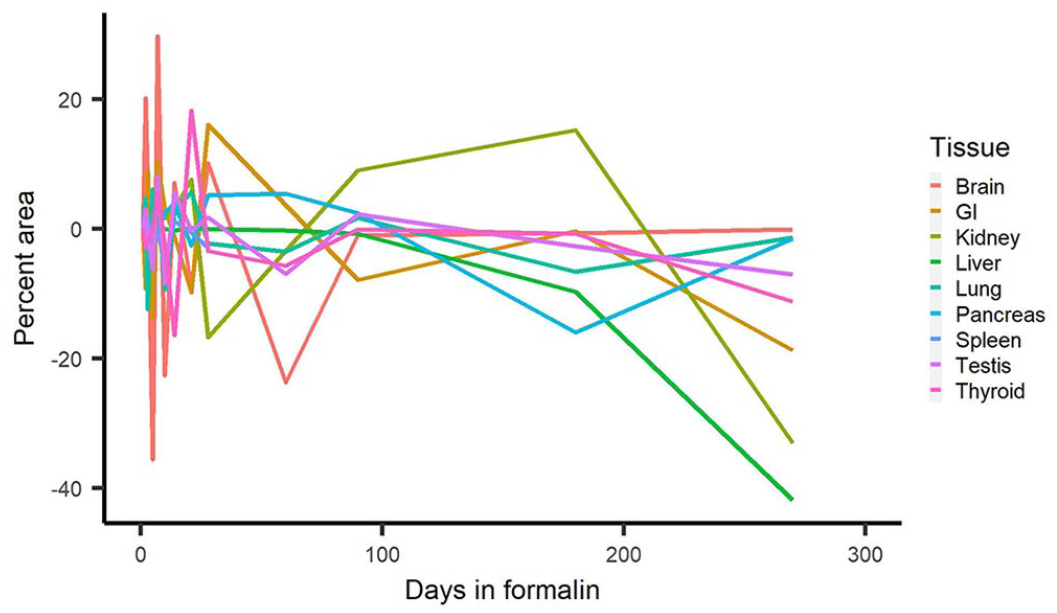
Comparison of the change in RNAscope in situ hybridization percent area of staining of all tissues over days in 10% neutral-buffered formalin.
FFPE tissue storage time study
RNAscope ISH successfully identified target CDV RNA in tissues at each time (Figs. 4, 5). Sections treated with the negative control probe (DapB) were negative in all cases. Tissues that had positive CDV signal using RNAscope ISH included: lung, spleen, brain, pancreas, and kidney (not all shown). In our qualitative visual comparison of RNAscope ISH and IHC signal in corresponding sections, there was very similar tissue localization, pattern, and quality of signal. Specifically, CDV ISH signal was most intense in the brain and lungs, in which characteristic CDV-induced lesions were found in routinely stained sections and corresponded strongly to IHC staining (Figs. 4, 5).
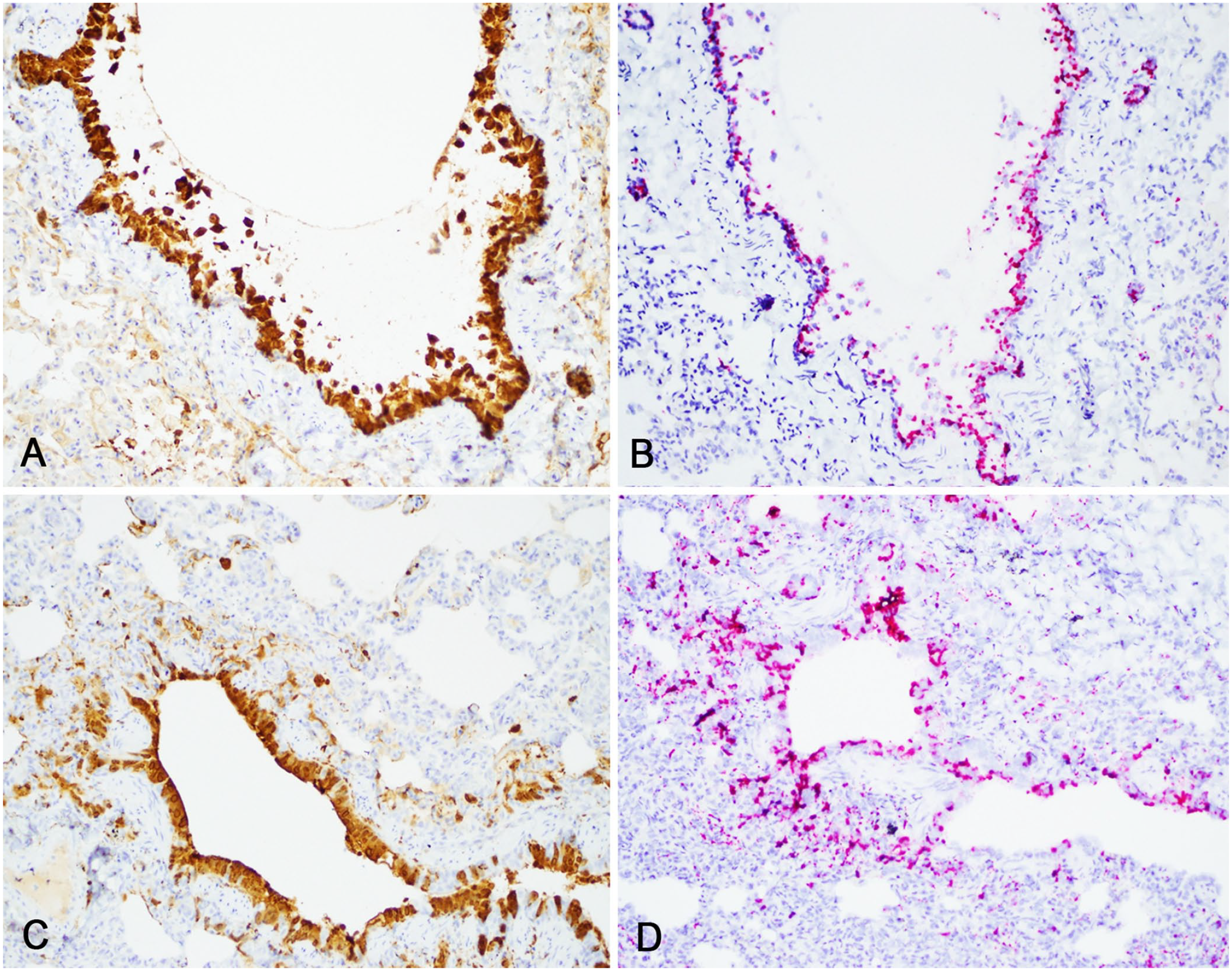
Comparison of immunohistochemistry (
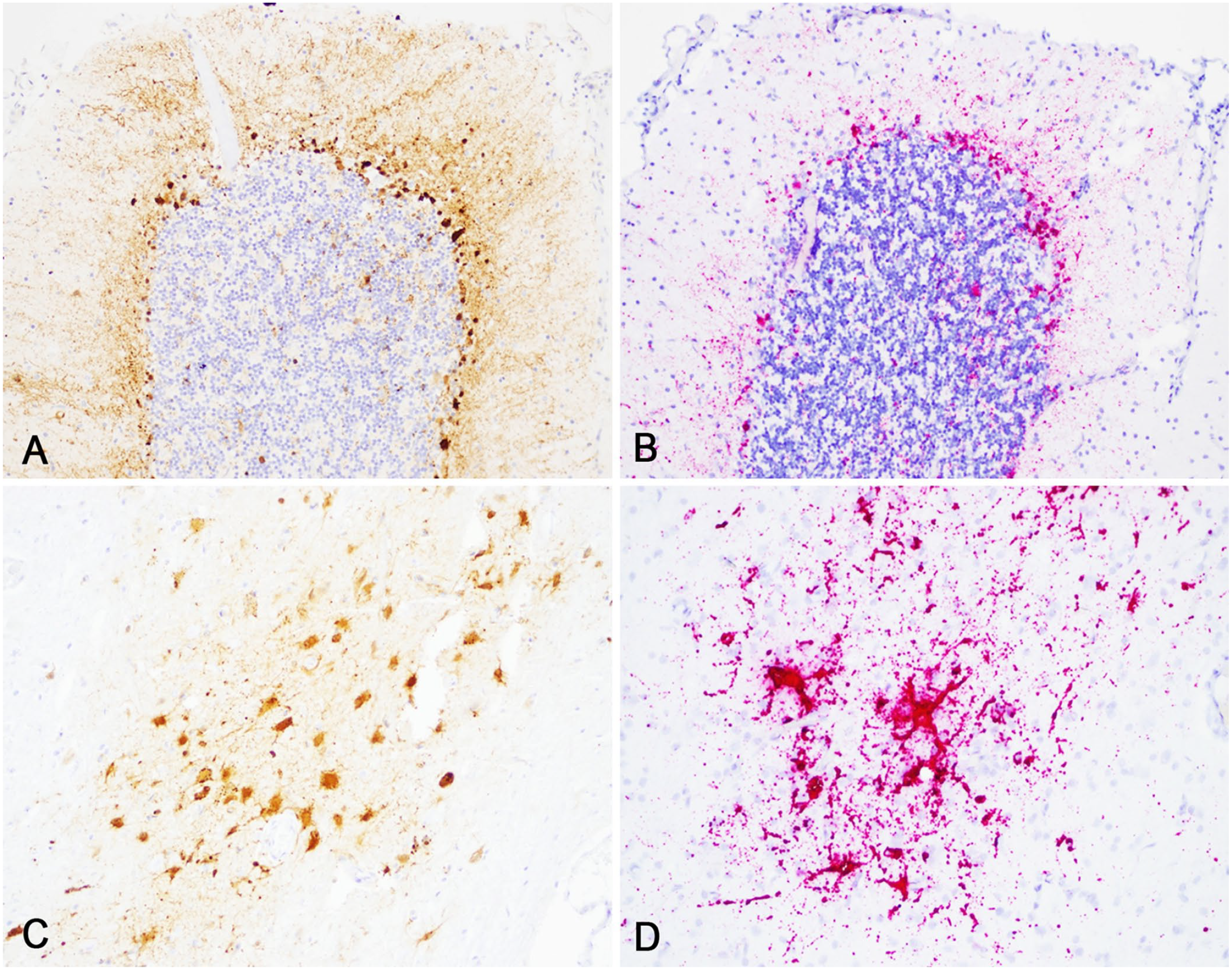
Comparison of immunohistochemistry (
Discussion
Signal intensity and %area of signal expression of the mRNA reference gene (An-16srRNA) changed over tissue-fixation time in formalin. Although the changes among times were not statistically significant, there was an observed decrease in RNAscope ISH signal after 180 d and a statistically significant difference in signal between days 1 and 270. These findings emphasize the importance of prompt processing of fixed tissue samples but indicate that the technique may not be as time-sensitive as suggested by the manufacturer’s protocols. We utilized a target reference gene with relatively high levels of expression. Additional studies utilizing targets with lower levels of expression are warranted to determine the sensitivity of detecting low levels of signal in case slight degradation not detectable in this reference gene occurs at an earlier time. Additionally, a previous study also suggested that ribosomal RNA is less prone to degradation, making it easier to detect in situ. 2 Therefore, although signal was still detectable in samples stored for > 180 d, RNAscope signal in these samples must be interpreted with caution and with appropriate controls, especially with low-expression targets.
Similar studies of the effect of formalin-fixation time on IHC report a wide variety of results, likely because of the difference in antibodies and antigen-retrieval methods used.13,23,24 One study reported the detection of African horse sickness virus in samples fixed in formalin for 365 d. 5 Two studies comparing 60+ IHC antibodies in canine tissues and common infectious diseases in domestic animals reported that moderate-to-strong signal was observed up to 7 wk for all antibodies studied and that one antibody (amylin) had strong signal following 13 wk of formalin fixation; more extended formalin-fixation times were not tested.23,24 Another study detected bovine viral diarrhea virus antigen in samples with 36 d of fixation, but little-to-no signal was detected in those fixed for 176 d. 13 Although these differing results are likely dependent on the target antigen, antibody used, or antigen-retrieval technique in the IHC protocol, it is clear that extended formalin-fixation time does affect the ability to detect target antigens.
We observed positive RNAscope ISH signal for CDV in archival FFPE tissues stored up to 15 y at room temperature. Importantly, corresponding RNAscope ISH and IHC sections had similar staining quality and overlapping tissue localization and patterns. Our findings suggest that appropriately curated archives could serve as a valuable resource for retrospective studies using this technology. Studies on the effect of FFPE storage time on RNAscope ISH signal amplification are limited and are mostly comprised of studies on cancer markers or positive control probes in human tissues. 4 Although the RNAscope ISH user manual suggests that the typical storage method of FFPE samples (room temperature and no desiccants) for > 1 y leads to mRNA degradation, various studies have utilized RNAscope successfully with archival tissues outside of this 1-y time period.1,2,4
A study evaluating the effect of FFPE storage times on RNAscope ISH using the human control probe Hs-PPIB found that signal was lost after 5 y of FFPE storage in prostatic cancer tissue sections, 2 although another study concluded that the same positive human control probe (Hs-PPIB) could be amplified using RNAscope in a variety of tumor tissues (colorectal, prostatic, ovarian, breast) stored in paraffin for up to 10 y. 4 One of the few veterinary studies utilizing RNAscope ISH detected amdoparvovirus in the lymph node of a 12-y-old FFPE tissue sample from a red panda (Ailurus fulgens). 1 Additionally, elephant endotheliotropic herpesvirus (EEHV) has been detected in FFPE tissue blocks stored for up to 5 y. 6 The ability to detect RNA targets in archival FFPE samples likely depends on several factors, such as the specific target, the gene expression level or viral load within the sample, and possibly the storage method (temperature, humidity, etc.). However, many archival tissues are sufficiently robust for retrospective studies involving RNAscope ISH. As always, appropriate positive and negative controls are necessary and may help interpretation in situations in which signal is not evident.
Statistical analysis identified a few outliers. For example, the 5-d formalin-fixation samples had decreased signal compared to some of the later times. Additionally, signal on days 10 and 14 for specific tissue types (i.e., testis, lung, thyroid) was decreased. Various factors could have contributed to this finding, including under-fixation of excessively large tissue samples, 23 an error in the RNAscope assay protocol that prevented saturation of the tissues in the probes and/or solutions, or inaccurate temperatures during processing. All RNAscope assays were processed manually, which could introduce error within a single run; for example, utilizing inadequate amounts of reagent or incomplete saturation of all tissues with reagents. Use of an automated stainer would limit intra- and inter-run variation. Large variation was also noted at 180 d. Additional biological replicates, including repeating ISH on multiple slide recuts of the same tissue, analyzing additional 200× images, or utilizing tissues from multiple individuals would potentially diminish the effect of individual slide errors and lead to a better understanding of differences in RNA stability among tissues.
In the FFPE study, obtaining duplicate representative areas was challenging when comparing signal (RNAscope vs. IHC) because sections were cut a few µm apart. Thus, representative 200× images between the 2 methods were not identical, resulting in slightly skewed results. In addition, the IHC technique highlights protein antigen, whereas RNAscope amplifies an RNA target in tissues. Therefore, signal intensity and %area cannot be directly compared statistically for the FFPE study, as expression of the signal intensity and %area of signal are not identical in comparison. The individual variation in disease expression and the infection timeline between CDV cases may have also affected the amount of viral mRNA present in each tissue section. In the FFPE study, we did not utilize a positive control RNA quality slide (raccoon reference mRNA), which could have demonstrated potential RNA degradation. Nonetheless, there was striking concordance in signal distribution and intensity among modalities.
Various studies have documented the superior DNA and RNA preservation capacity of alcohol-based fixatives over standard formalin fixation9,16,20,27; thus, EtOH was considered an appropriate fixative to “halt” formalin fixation and subsequent RNA degradation. Although formalin-fixation time was tested, it was not possible to control for identical 70% EtOH submersion times after formalin-fixation, thus variation in time in EtOH may have confounded interpretation. An additional limitation of our study was that tissues originated from a single addax. Additional investigations targeting specific times in other species and additional tissues should be conducted if this technology is to be applied more broadly. The amount of time the unstained slides sat at room temperature before being processed varied slightly among the times, thus introducing potential variation between times. Regardless, there is a measurable decrease in signal intensity in tissues that had little time in EtOH and as unstained slides (90–270 d), which makes this bias unlikely to have greatly affected the overall results.
Our findings suggest that RNAscope may have utility in some archived tissues. Although our results do not necessarily apply to all probes, given that expression patterns will differ depending on the desired target, our findings provide guidelines with which to establish inclusion and exclusion criteria for archived samples in retrospective studies. Qualitative studies on tissue distribution of signal may be less affected, but because there is some signal decrease over time, comparisons of signal intensity among samples with different fixation and storage times should be interpreted with caution. Given that RNAscope probes can be designed from easily obtainable sequences, the technique has enormous potential for use in non-domestic species for which validated antibodies may not be available or feasible.
Other methods of RNA analysis, such as NanoString and 10X Genomics, have been utilized in a limited number of veterinary species and laboratory animals to detect gene expression in FFPE tissues. NanoString technology allows spatial profiling of multiple genes in a single FFPE sample and is being utilized in the study of canine cancer pathogenesis.7,12 Although these technologies are becoming more popular in human medical research and laboratory animal studies, this technology is not yet widely available for veterinary diagnostic use. Future studies should focus on applying RNAscope methodology to tissues from a variety of taxa (i.e., avian, fish, reptile, amphibian, and invertebrate species) as well as different fixatives and processing techniques that may impact utility.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387241245777 – Supplemental material for Effect of formalin-fixation and paraffin-embedded tissue storage times on RNAscope in situ hybridization signal amplification
Supplemental material, sj-pdf-1-vdi-10.1177_10406387241245777 for Effect of formalin-fixation and paraffin-embedded tissue storage times on RNAscope in situ hybridization signal amplification by Megan E. Colburn, Martha A. Delaney, Gretchen C. Anchor and Karen A. Terio in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank the Zoological Pathology Program (ZPP) faculty and technical staff and Renee Walker and VDL Histology Laboratory staff for their assistance. We also thank Dr. Lois Hoyer at the University of Illinois–College of Veterinary Medicine for her support of our project.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
We thank the ZPP for funding this project and Student Support: Office of the Director, NIH, T35 OD011145.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
