Abstract
Mycoplasma bovis DNA was detected in lung tissue of experimentally infected calves by in situ hybridization (ISH) with a nonradioactive, digoxigenin-labeled DNA probe. The 171–base pair DNA probe targeting part of the gene of the major immunodominant variable surface protein A, which is conserved among all vsp genes, was generated by polymerase chain reaction. Four calves between 57 and 63 days old were inoculated intratracheally with 30 ml of a suspension of M. bovis strain 1067 containing 7 × 104 colony forming units per milliliter. Two calves inoculated with 30 ml of sterile medium served as control animals. The calves were euthanized and then examined 21 days after inoculation. The ISH method developed in the current study was suitable for the detection of M. bovis DNA in formalin-fixed, paraffin-embedded lung tissue and may be a valuable tool for diagnostic purposes and for further investigating the pathogenesis of M. bovis infection.
Keywords
Mycoplasma bovis, excluding Mycoplasma mycoides subsp. mycoides small-colony type, which is the agent of contagious bovine pleuropneumonia, is one of the most important pathogens of bovine mycoplasmosis. The disease includes pneumonia and arthritis, especially in calves, and mastitis in dairy cows, as well as keratoconjunctivitis, otitis, decubital abscesses, metritis, abortion, and infertility, resulting in extensive losses for the cattle industry. 2,4,8,12 In several countries, an increasing prevalence of M. bovis infections can be recognized. 5,6,13 Commercial vaccines preventing the infection are not presently available, with the exception of recently approved, custom-designed, inactivated vaccines for use in North America. Increasing numbers of M. bovis strains are developing resistance to antibiotics, 18 and knowledge of pathogenesis and persistence of M. bovis, especially in the respiratory tract, is still insufficient.
In situ hybridization (ISH) is a very useful technique for studying the pathogenesis of infectious diseases by allowing the localization of organisms within the context of tissue morphology. In previous studies, M. bovis antigens were detected immunohistochemically in lung samples from calves with chronic pneumonia by using monoclonal antibodies to reveal the presence of antigens in different compartments of infected lungs. 1,7
Pathogenic mycoplasmas are equipped with sophisticated genetic systems, which allow these agents to change their surface antigenic makeup at a high frequency. These variable surface components have been implicated in providing the wall-less mycoplasmas with a means to avoid the host immune response and promote host colonization. In M. bovis, these variable surface components are the so-called Vsps, which represent the major immunodominant membrane lipoproteins and are encoded by a number of distinct, but related, genes. 11,14,17 The aim of the current study was to develop DNA probes for the detection of M. bovis by ISH in formalin-fixed, paraffin-embedded lung tissue of experimentally infected calves with different lung lesions.
Lung samples originated from experimentally infected calves. Four calves were inoculated intratracheally with 30 ml of a suspension of the virulent M. bovis strain 1067 15 containing 7 × 104 colony forming units per milliliter. Two calves inoculated with 30 ml of sterile medium served as control animals. The calves were euthanized 21 days postinoculation (PI), and bacteriological cultures and polymerase chain reaction (PCR) for the general detection of mycoplasmas and for the specific detection of M. bovis were performed on lung tissue. In total, formalin-fixed, paraffin-embedded lung samples of 6 calves obtained from 6 different lung locations each (left lung: lobus cranialis pars cranialis, lobus cranialis pars caudalis, lobus caudalis; right lung: lobus cranialis pars caudalis, lobus medius, lobus caudalis) were investigated. According to a method previously described, 3 lung samples from healthy, slaughtered cattle injected with a concentrated suspension of M. bovis type strain PG45 served as positive control tissue.
Samples of the lung were fixed in 4% neutral buffered formaldehyde for 72 hr, routinely processed, and embedded in paraffin according to standard laboratory procedures. Sections were cut 5-μm thick and mounted on positively charged glass slides. a One slide each was stained with hematoxylin and eosin (HE) for histologic examination.
The probe was synthesized by PCR using the sequence of the vspA gene of M. bovis type strain PG45. 11 The forward primer (vsp-69/26m [position −69], 5′-CGAAATATTTAGATAGTTATTTATAG-3′) and the reverse primer (vsp+100/24m [position +102], 5′-TGGTTTATTTTCTTCTTTGGTCTC-3′) were designed, resulting in a digoxigenin-labeled DNA probe of 171 base pairs (bp) that anneals to the highly conserved 5′, noncoding sequence extending 69 bp upstream of the ATG initiation codon, which exhibits 99% homology among all vsp genes, and the N-terminal region (93 bp) encoding 31 amino acids, which exhibit 98% identity in all Vsps and contain the prolipoprotein signal peptide. 11 The PCR was performed in 30 cycles with a denaturation temperature of 94°C for 1 min, an annealing temperature of 56°C for 70 sec, and an elongation temperature of 72°C for 20 sec. The concentration of magnesium was 2.5 mM. The PCR products were labeled by random priming using digoxigenin–2′-deoxyuridine 5′-triphosphate. b The specificity of the probe was tested by Southern blot using HindIII-digested DNA of M. bovis type strain PG45 and clinical isolates of other bacterial species commonly found as commensals or as infectious agents in the bovine respiratory tract, including Arcanobacterium pyogenes, Pasteurella multocida, Mannheimia haemolytica, and Mycoplasma bovirhinis.
The protocol for the ISH was adapted from published guidelines. 9 Tissue sections were deparaffinized and rehydrated in phosphate buffered saline (PBS; pH 7.4; 0.01 M) for 5 min immediately before use. Deproteinization was done in 0.2 N HCl for 20 min at room temperature. After rinsing sections twice in 2 x standard sodium citrate (SSC) with 5 mM ethylenediamine tetra-acetic acid (EDTA)–Na2 at 50°C, digestion was done at 37°C for 15 min with 7.5 μg/ml proteinase K b in Tris. Digestion was stopped by 0.2% glycine–PBS for 5 min, and tissues were fixed in 4% paraformaldehyde for 4 min. After rinsing twice in PBS, tissue sections were acetylated in 0.1 M triethanolamine–HCl buffer (pH 7.5) loaded with acetic anhydride (0.25%) for 10 min. The tissue sections were then rinsed in PBS. The sections were allowed to equilibrate for 60 min at 37°C in a prehybridization buffer consisting of 6 x SSC, deionized formamide 45%, and 1% Denhardt solution. Hybridization was carried out overnight at 37°C with a hybridization buffer consisting of deionized formamide 57%, c EDTA, piperazine-N,N'-bis(2-ethanesulfonic acid), d NaCl, Denhardt solution, heparin, b Triton X-100, e single-stranded DNA, d RNA solution, d and dextran sulfate. d Approximately 2 μl of the digoxigenin-labeled probe was added to 98 μl of hybridization buffer and layered over the tissue sections. The fluid was held in place by a coverslip, the edges were sealed with rubber cement, and the slides were heated for 10 min at 95°C. After overnight hybridization in a moist chamber, tissue sections were each rinsed twice in stringency solutions of 6 x SSC with 45% formamide e for 15 min at 42°C, in 2 x SSC for 5 min at room temperature, and in 0.2 x SSC for 15 min at 42°C. Tissue sections were then washed in buffer I (100 mM Tris, 150 mM NaCl; pH 7.5) for 1 min and in buffer II (1% sheep serum sterile filtered, 0.15% Triton X-100 in buffer I) for 30 min at room temperature. The sections were incubated for 120 min with anti-digoxigenin conjugated with alkaline phosphatase b diluted 1:200 in buffer II for the detection of hybrids. After 2 washes each of 15 min in buffer I and 1 of 2 min in buffer III (100 mM Tris, 100 mM NaCl, 50 mM MgCl2; pH 9.5), the sections came in contact with the substrate consisting of nitroblue tetrazolium and 5-bromocresyl-3-indolylphosphate d added to buffer III. Color was allowed to develop overnight in the dark, and the reaction was stopped by washing the slides twice for 10 min in buffer IV (10 mM Tris, 1 mM EDTA; pH 8). Tissue sections were counterstained with hemalaun f and, after rinsing in water, were covered with glycerogel. g
Other than M. bovis, no mycoplasmas were detected by PCR and culture, but in 2 infected calves (calves 1 and 2), an additional infection with A. pyogenes was found. Histologic examination revealed all 4 infected calves had suppurative bronchopneumonia. In addition, calf 1 had a severe, necrotizing bronchopneumonia, whereas obliterative bronchiolitis was present in 2 others (calves 1 and 4).
Testing of the digoxigenin-labeled probe in Southern blot analysis revealed a high specificity for M. bovis (Fig. 1). In situ hybridization produced a distinct positive signal for the M. bovis vspA gene in the lung. The morphology of the tissue was well preserved. The signal intensity varied within tissue sections and between calves. Positive hybridization typically exhibited a dark brown to violet reaction product, and there was only weak background staining. In lung tissues of calf 1, abundant and strong hybridization signals were detected in the center and at the borders of necrotic lesions (Fig. 2). Additionally, in areas of obliterative bronchiolitis, a weak focal signal was detectable. Weak signals within the intraluminal exudates of larger airways were also found in 3 of the 4 cases (except calf 3) with suppurative bronchopneumonia. Both extracellular and intracellular signals were detected. Identification of the cell types containing positive hybridization signals was occasionally difficult, but examination of adjacent sections stained with HE confirmed that positive cells resembled either macrophage-like cells, with large, oval nuclei and abundant cytoplasm, or neutrophil-like cells, with multilobulated nuclei. No hybridization signals were detected in the uninfected control calves. In the positive control lung tissue that was inoculated with a concentrated M. bovis suspension, strong hybridization signals were found mainly extracellularly in the interlobular interstitium.
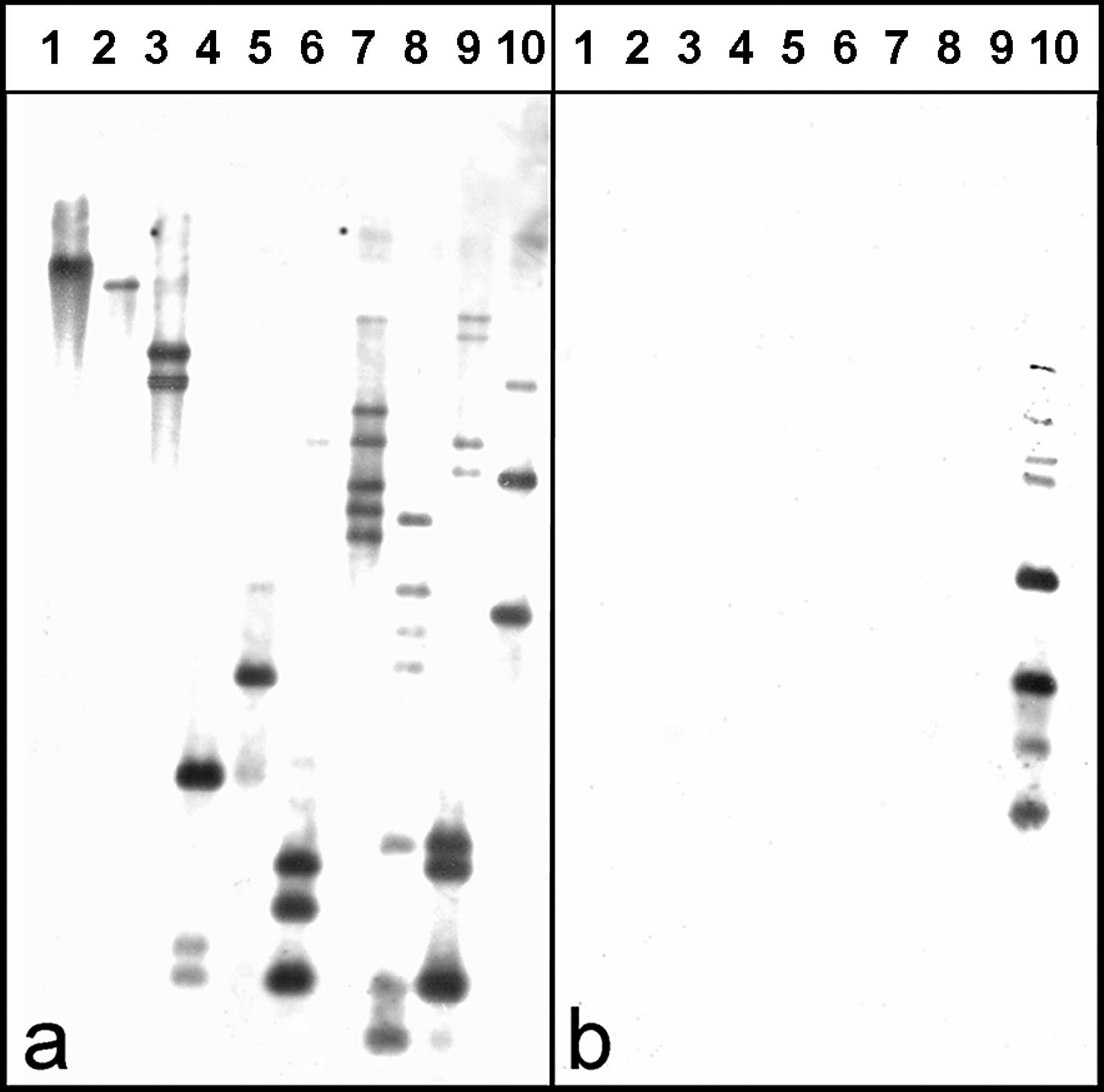
Southern blot of HindIII-digested genomic DNA probed with
The results of the current study demonstrate that DNA of M. bovis can be detected in formalin-fixed, paraffin-embedded lung tissue of calves using a digoxigenin-labeled vspA gene probe. A hybridization signal for M. bovis DNA was identified within necrotic areas, within the lumen of bronchi and bronchioles filled with suppurative exudates, and in areas of obliterative bronchiolitis.
The presence of strong hybridization signals within necrotic foci may indicate that these areas are reservoirs, in which M. bovis or at least M. bovis DNA can persist for several weeks PI. By using monoclonal antibodies in necrotic lung lesions of calves with natural M. bovis infection, strong deposits of reaction products in necrotic areas have been found, similar to the results seen in the present study. 7 The persistence of organisms in the exudates within bronchioli at least 21 days PI indicates that these calves still excrete M. bovis. Furthermore, this may be the reason for the proliferation of lymphatic tissue around the bronchioli described previously. 16
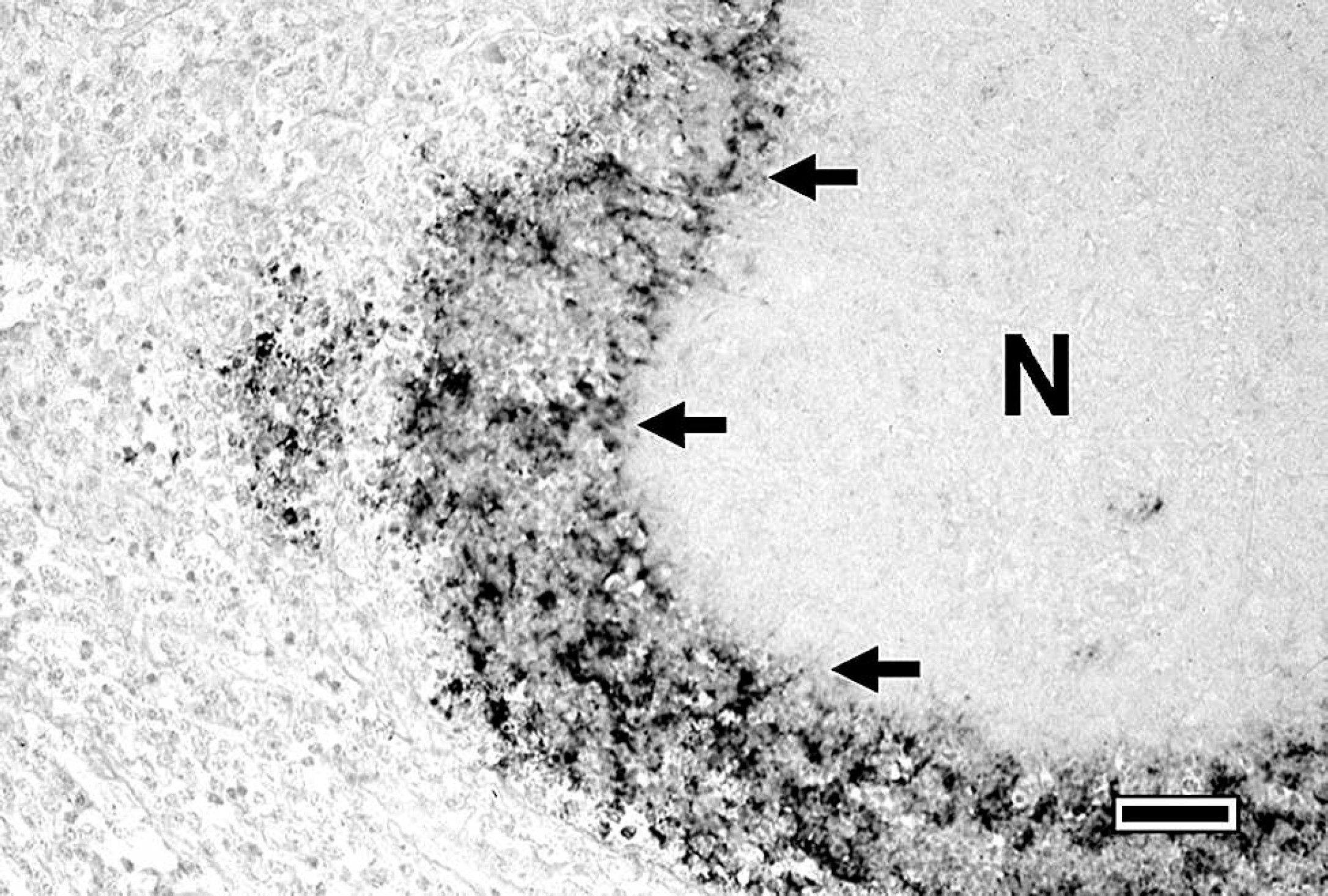
Section of the lung of a calf experimentally infected with Mycoplasma bovis. Hybridization signal at the border (arrows) of a necrotic area (N); in situ hybridization with a digoxigenin-labeled probe detecting vspA gene. Bar = 50 μm.
It can be postulated that M. bovis DNA persists in the respiratory tract of experimentally infected calves and is possibly excreted with the exudates in cases of suppurative bronchopneumonia. Because M. bovis was isolated from infected lung tissue, organisms detected by ISH were still viable.
The factors that enable M. bovis to persist in the lungs of infected cattle are poorly understood. The ISH method described herein was developed to detect the presence of the vspA gene during in vivo infection and provides an important tool for use in both diagnostics and pathogenetic studies. This method may also be used to detect other pathogenesis-related M. bovis–specific genes. Because the probe anneals to a region that induces the 2 highly homologous domains of all vsp genes found in different M. bovis strains, 11,14 any field strain of M. bovis can be reliably detected, even if it possesses modified versions of the vsp gene complex with extensive sequence variations occurring in the reiterated coding sequences of vsp structural genes.
Acknowledgements. Bjön Jacobsen was supported by a scholarship from the Friedrich Ebert Foundation, Germany.
Footnotes
a.
SuperFrost/Plus®, G. Menzel Glasbearbeitungswerk GmbH & Co KG, Braunschweig, Germany.
b.
Roche Diagnostics GmbH, Mannheim, Germany.
c.
Qbiogene, Heidelberg, Germany.
d.
Sigma-Aldrich Chemie GmbH, Taufkirchen, Germany.
e.
Merck KGaA, Darmstadt, Germany.
f.
Roth C. GmbH & Co. KG, Karlsruhe, Germany.
g.
Dako Cytomation GmbH, Hamburg, Germany.
