Abstract
The objective of the present study was to compare the ability of 2 monoclonal antibodies (mAbs; SDOW17 and SR30) to detect types 1 and 2 Porcine reproductive and respiratory syndrome virus (PRRSV) in formalin-fixed, paraffin-embedded (FFPE) lung tissues by immunohistochemistry (IHC) and to compare the immunohistochemical results with in situ hybridization (ISH) and reverse transcription nested polymerase chain reaction (RT-nPCR) detection techniques. Lungs from 30 experimentally infected pigs (15 pigs with each genotype of PRRSV) and 20 naturally infected pigs (10 pigs with each genotype of PRRSV) with types 1 and 2 PRRSV, respectively, were used for the IHC, ISH, and RT-nPCR analyses. The SR30 mAb-based IHC detected significantly more type 1 PRRSV-positive cells in the accessory and caudal lobes from the experimentally infected pigs at 7 (P = 0.025) and 14 (P = 0.018) days postinoculation, respectively, compared to the SDOW17 mAb-based IHC. The results demonstrated that SR30 mAb-based IHC is useful for detecting both types 1 and 2 PRRSV antigen in FFPE lung tissues.
Porcine reproductive and respiratory syndrome (PRRS), caused by the PRRS virus (PRRSV), occurs in 2 clinical forms, reproductive failure and respiratory disease. Reproductive failure is characterized by abortion and delivery of stillborn near-term fetuses or premature and weak piglets. 27 The respiratory disease is characterized by pronounced hyperpnea, fever, and interstitial pneumonia in young pigs and mild flu-like signs in growing and finishing pigs. 27 PRRSV is a member of the Arterivirus genus (family Arteriviridae, order Nidovirales). 4 Genetic analysis has established 2 predominant PRRSV genotypes, type 1 (European genotype) and type 2 (North American genotype), which share approximately 60% nucleotide identity of the entire genome.1,18
The current diagnostic tests used to determine whether PRRSV is the cause of respiratory disease are viral isolation, light microscopy, immunohistochemistry (IHC), in situ hybridization (ISH), reverse transcription polymerase chain reaction (RT-PCR), and serology.8,15,17 Among these diagnostic methods, IHC, ISH, and RT-PCR can be applied to detect PRRSV in formalin-fixed, paraffin-embedded (FFPE) tissues.8,9
Pathologists prefer to use IHC or ISH to diagnose PRRSV infection because these 2 methods are useful for the simultaneous evaluation of histopathological lesions and the viral loads in lung tissues to determine the degree of severity of respiratory disease. SDOW17 and SR30 monoclonal antibodies (mAbs), which react with the 15-kD nucleocapsid protein of PRRSV by Western blotting analysis,19,25 are the prototypical PRRSV mAbs and are widely and routinely used in IHC to detect PRRSV in FFPE tissues.6-8,12,26 In addition to North America, 21 type 1 PRRSV has recently emerged in Asian countries, including Korea, China, and Thailand.2,5,16,22 Clinically, a type 2 PRRSV-based vaccine may not be effective for controlling type 1 PRRSV infection in pigs 23 (Kovacs F, Schagemann G: 2003, Efficacy of ingelvac® PRRS MLV against European isolates. In: Proceedings of the 4th Inter-national Symposium on Emerging and Re-emerging Pig Diseases, pp. 111–112. Rome). Hence, it is necessary to detect and differentiate between Korean type 1 and 2 PRRSV for proper selection of PRRSV vaccine to control both genotypes of PRRSV infection effectively in swine herds. The objective of the present study was 1) to compare the abilities of 2 mAbs (SDOW17 and SR30) in the detection of the 2 genotypes of PRRSV in FFPE tissues by IHC and 2) to compare the immunohistochemical results with ISH and RT-PCR techniques.
Korean type 1 (SNUVR100057 strain) and type 2 (SNUVR090851 strain) PRRSV used in the current study were isolated from lung samples from different postweaning pigs in a 1,000-sow herd in Chungcheung Providence in 2009 and 2010, respectively. This herd has shown severe respiratory problem in postweaning pigs, aged from 4 to 8 weeks old. The SNUVR100057 strain was identified as type 1 PRRSV based on the nucleotide sequences of open reading frame (ORF)5 (GenBank accession no. JN411262) and ORF7 (JN411261); the SNUVR090851 strain was identified as type 2 PRRSV on the basis of the nucleotide sequences of ORF5 (JN315685) and ORF7 (JN399072).
Thirty-six pigs were purchased from a PRRSV-free herd and were used at the age of 3 weeks old. All of the pigs were negative for PRRSV and Porcine circovirus-2 according to routine serological testing and real-time PCR performed prior to delivery and again on arrival.10,24 The pigs were randomly allocated into 2 infected and 1 control groups. In groups 1 and 2, 15 pigs each were inoculated intranasally with 3 ml of tissue culture fluid containing 106 50% tissue culture infective doses (TCID50)/ml of Korean type 1 PRRSV (second passage in the MARC [cloned African green monkey kidney cell line]-145 cells) or 106 TCID50/ml of Korean type 2 PRRSV (second passage in the MARC-145 cells). In group 3, 6 control pigs were exposed in the same manner to uninfected cell culture supernatants. Ten infected (5 each in groups 1 and 2) and 2 control pigs from each group were sedated by an intravenous injection of sodium pentobarbital and then euthanized by electrocution at 7, 14, and 21 days postinoculation (dpi) as previously described. 3 Tissues were collected from each pig at necropsy. Lungs were collected for viral isolation from all of the infected and negative-control pigs used in the study, as previously described. 8 The isolated PRRSV samples from the lungs and lymph nodes were further analyzed for ORF5 sequence. 20 The methods were previously approved by the Seoul National University, Institutional Animal Care and Use Committee.
Twenty pigs (10 pigs with each genotype of PRRSV) each naturally infected with type 1 or 2 PRRSV from 1 pig from each farm were also used for an evaluation of 2 mAbs (SDOW17 and SR30)-based IHC. Four blocks of lung tissue from 8 different portions of 4 lobes (cranial, middle, accessory, and caudal lobe) were collected, as previously described. 12 The mAbs, SDOW17 a and SR30 a were diluted 1:10,000 in phosphate buffered saline (0.01 M, pH 7.4) containing 0.1% Tween 20. The IHC was performed as previously described. 6
Probes used for the ISH assays were generated by RT-PCR using genomic RNA of isolates SNUVR100057 and SNUVR090851. For type 1 PRRSV, a 354-bp complementary (cDNA) fragment representing the 5’ region of ORF6 and ORF7 was used as a probe. The forward and reverse primers were 5’-CGCTGTGACAAAGCCCGG AC-3’ (nucleotides 14,482–14,501) and 5’-TCGATTGCA AGCAGAGGGAG-3’ (nucleotides 14,814–14,835), res-pectively. For type 2 PRRSV, a 349-bp cDNA fragmentrepresenting the 5’ region of ORF6 and ORF7 was used as a probe. The forward and reverse primers were 5’-TCGTTC GGCGTCCCGGCTCC-3’ (nucleotides 14,775–14,794) and 5’-TTGACGACAGACACAATTGC-3’ (nucleotides 15,122–15,141), respectively. 14 Polymerase chain reaction was carried out, as previously described. 14 The purified PCR product was labeled by random priming with digoxigenin–2’-deoxyuridine 5’-triphosphate (dUTP) with a commercial, kit. b The ISH was carried out as previously described. 6 Reverse transcription nested PCR (RT-nPCR) was also performed to detect type 1 and 2 PRRSV in the FFPE lung tissues from the experimentally and naturally infected pigs as previously described. 9
To obtain quantitative data, analyses of slides was performed with the NIH Image J 1.43m Program. c For each slide of lung tissue, 10 fields were randomly selected, and the number of positive cells per unit area (0.95 mm2) was counted, as previously described. 13 The order of the serial slides was the same for each test. The mean values were also calculated.
Porcine reproductive and respiratory syndrome virus was isolated from the lungs (30 pigs) of type 1 and 2 PRRSV-infected pigs at multiple dpi. The PRRSV isolated from all of the infected pigs was confirmed by sequence analysis to be the same propagating Korean type 1 and 2 PRRSV in the challenge stock. No PRRSV was isolated from the lungs of the negative-control pigs.
Type 1 PRRSV nucleic acid was detected in all 15 or 10 lungs of the pigs that were experimentally or naturally infected with type 1 PRRSV by RT-nPCR. Type 2 PRRSV nucleic acid was detected in all 15 or 10 lungs of the pigs that were experimentally or naturally infected with type 2 PRRSV by RT-nPCR.
Distinct positive immunohistochemical and hybridization signals by IHC and ISH, respectively, were detected at each day postinoculation among the lungs of infected pigs. Either type 1 or 2 PRRSV antigens or genomes were detected in the lung from 7, 14, and 21 dpi by IHC and ISH, respectively, in all of the infected pigs throughout the experiment (Table 1). Positive cells typically exhibited red reaction and dark-brown reaction product by IHC and ISH, respectively, in the cytoplasm, without background staining. The SDOW17- and SR30-based IHC detected PRRSV in the pigs experimentally infected with type 1 and 2 PRRSV, respectively.
Comparison of Porcine reproductive and respiratory syndrome virus (PRRSV)–positive cells detected by immunohistochemistry (IHC) and in situ hybridization (ISH).
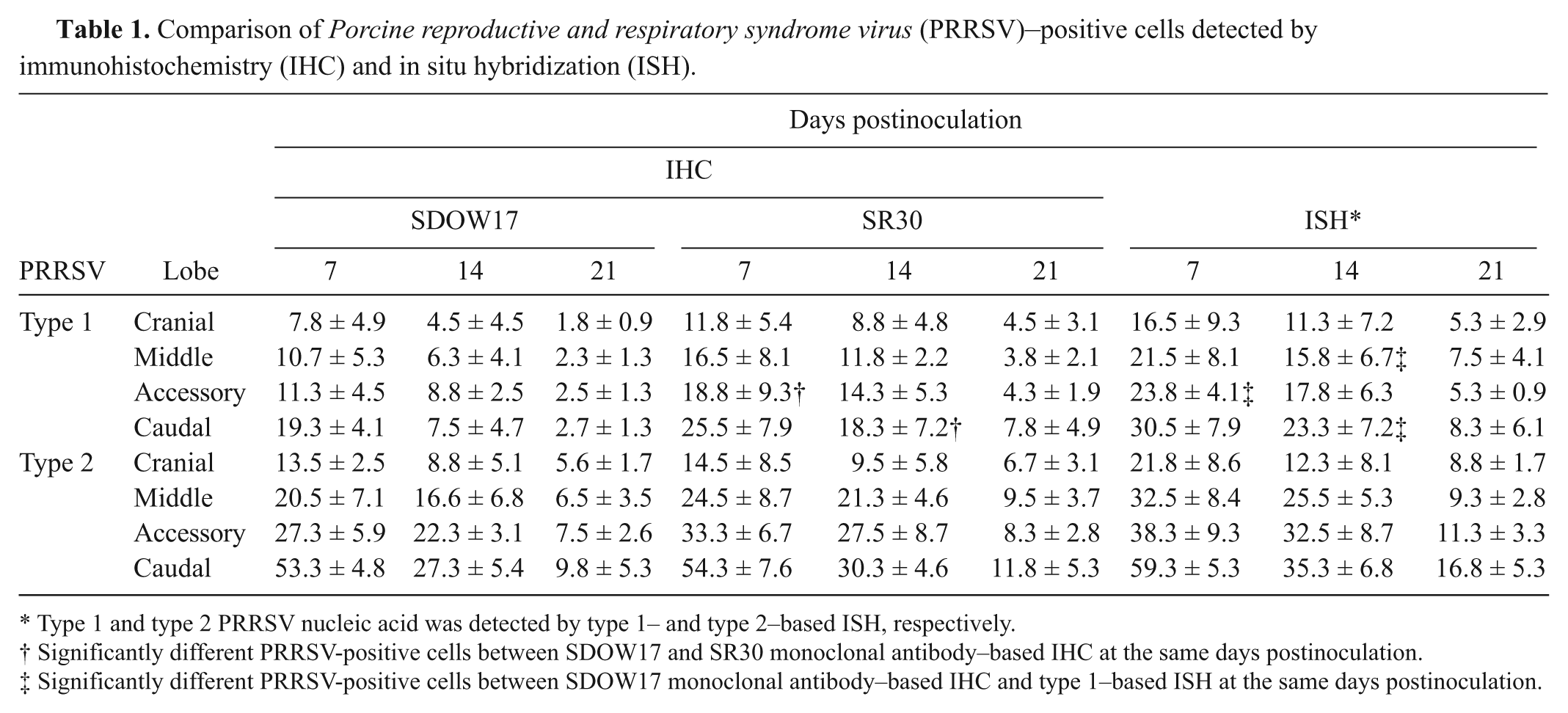
Type 1 and type 2 PRRSV nucleic acid was detected by type 1– and type 2–based ISH, respectively.
Significantly different PRRSV-positive cells between SDOW17 and SR30 monoclonal antibody–based IHC at the same days postinoculation.
Significantly different PRRSV-positive cells between SDOW17 monoclonal antibody–based IHC and type 1–based ISH at the same days postinoculation.
The type 1 PRRSV-based ISH detected the type 1 PRRSV from pigs experimentally infected with type 1 PRRSV. The type 2 PRRSV-based ISH detected the type 2 PRRSV from pigs experimentally infected with type 2 PRRSV. The type 1 PRRSV-based ISH does not detect the type 2 PRRSV from pigs experimentally infected with type 2 PRRSV and vice versa. The SDOW17- and SR30-based IHC and type 1 (or type 2)–based ISH of the serial sections of lung indicated that cells that stained positive for type 1 (or type 2) PRRSV antigen were also positive for type 1 (or type 2) PRRSV nucleic acid (Figs. 1, 2).
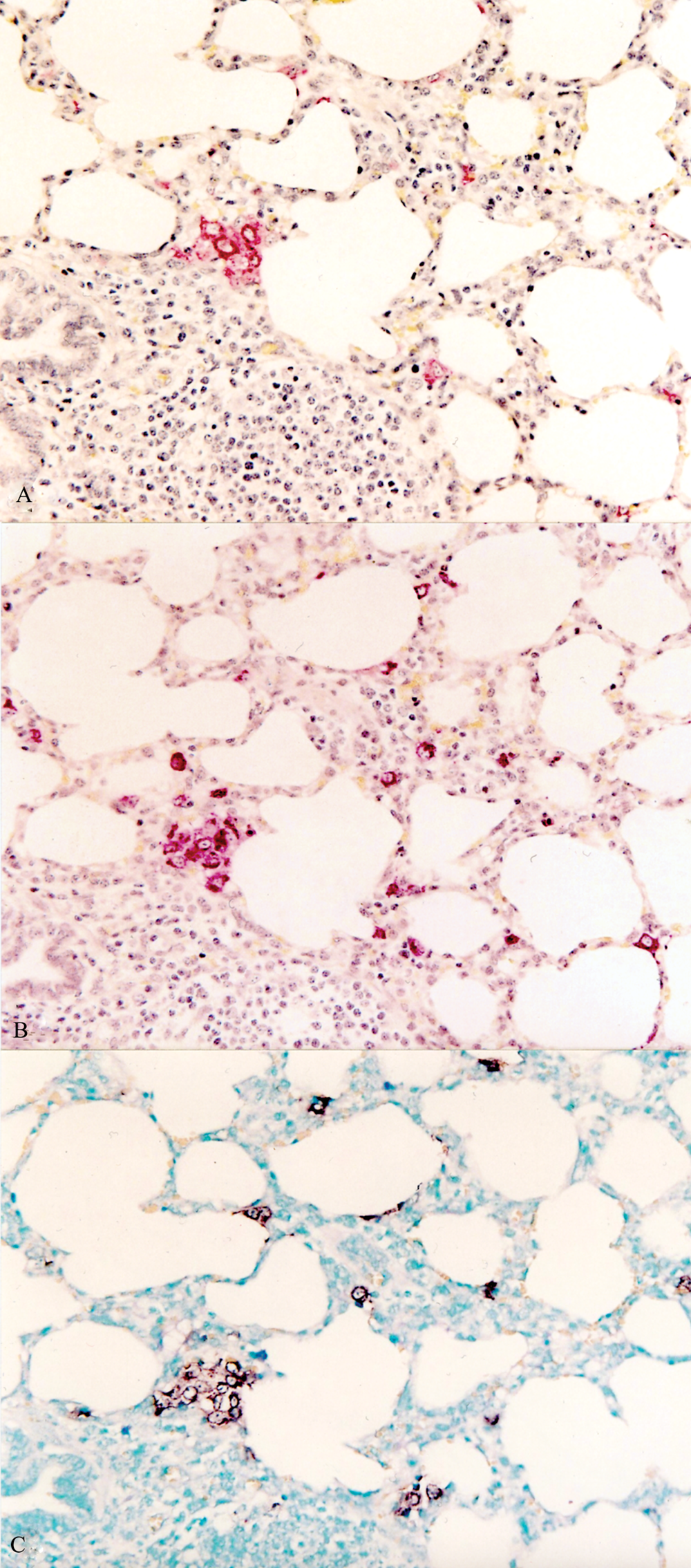
Serial section of a lung from a pig experimentally infected with type 1 Porcine reproductive and respiratory syndrome virus (PRRSV) at 7 days postinoculation.
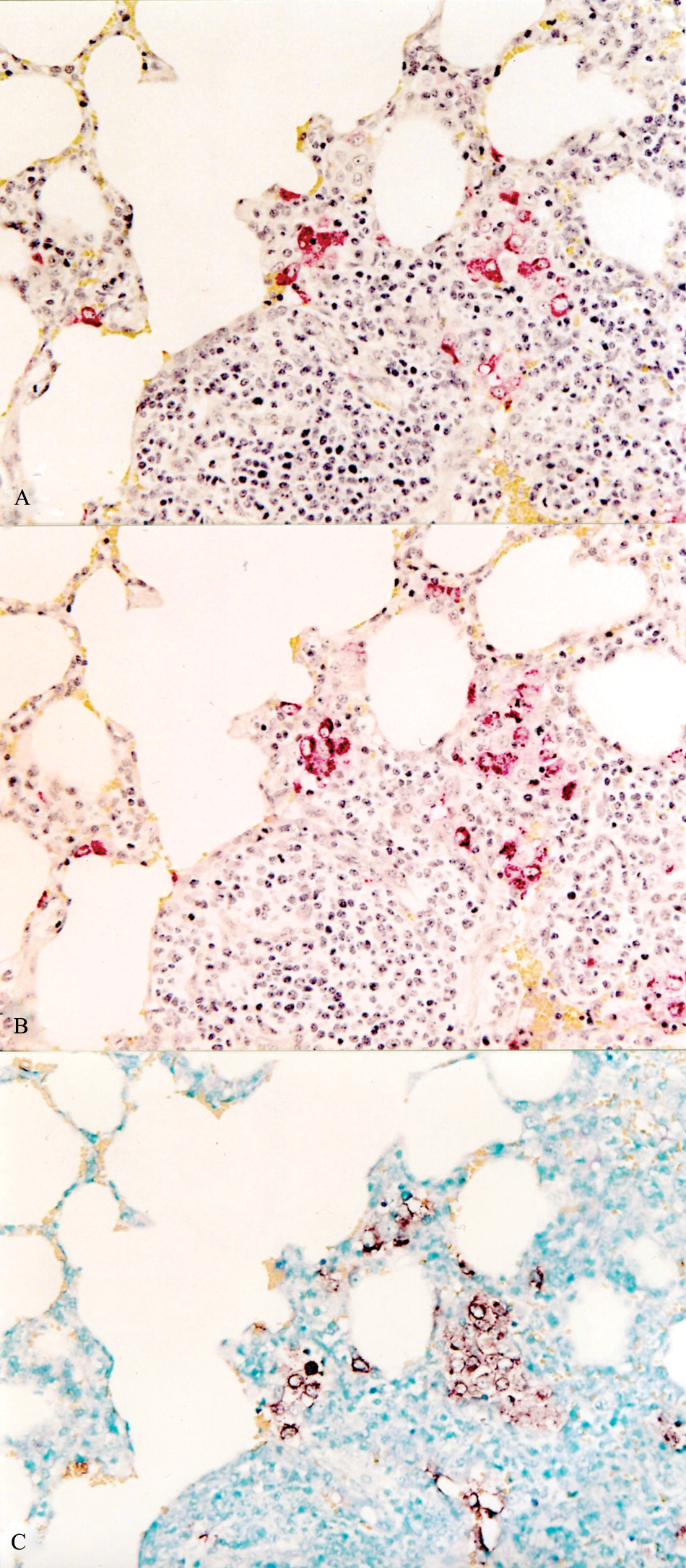
Serial section of a lung from a pig experimentally infected with type 2 Porcine reproductive and respiratory syndrome virus (PRRSV) at 7 days postinoculation.
The SR30-based IHC detected significantly more type 1 PRRSV-positive cells in the accessory and caudal lobes from the experimentally infected pigs at 7 (P = 0.025) and 14 (P = 0.018) dpi, respectively, compared to the SDOW17-based IHC. The type 1–based ISH detected significantly more type 1 PRRSV-positive cells in the accessory lobe (P = 0.007) at 7 dpi and the middle (P = 0.045) and caudal (P = 0.018) lobes from the experimentally infected pigs at 14 dpi, as compared to the SDOW17-based IHC (Table 1).
SDOW17- and SR30-based IHC detected both type 1 and 2 PRRSV from the 20 pigs (10 pigs in each genotype of PRRSV) naturally infected with type 1 and 2 PRRSV, respectively. The type 1 PRRSV-based ISH detected the type 1 PRRSV from pigs naturally infected with type 1 PRRSV. The type 2 PRRSV-based ISH detected the type 2 PRRSV from pigs naturally infected with type 2 PRRSV. There were no significantly different type 1 and 2 PRRSV-positive cells among the SDOW17- and SR30-based IHC and type 1 and 2–based ISH.
The results of the present study demonstrated that IHC and ISH are of potential value to detect both type 1 and 2 PRRSV in FFPE lung tissues. Because the SDOW17 and SR30 mAbs recognize a highly conserved epitope on the nucleocapsid (N) protein,19,25 these 2 mAbs are valuable diagnostic tools for the detection of 2 genotypes of PRRSV. However, there are clear differences between IHC and ISH for the detection and differentiation of the 2 genotypes of PRRSV in FFPE lung tissues from naturally and experimentally infected pigs. Although mAb-based IHC is specific for the detection of both 2 genotypes of PRRSV, none have been shown to differentiate type 1 PRRSV from type 2 PRRSV.
Reverse transcription nPCR alone cannot evaluate the severity of histopathological lesions induced by PRRSV, although RT-nPCR can differentiate between the 2 genotypes of PRRS and can measure the amount of PRRSV.8,9 ISH can differentiate between the 2 genotypes of PRRSV in FFPE tissues, as previously reported, 15 and provides histological architecture, thus, the detection of the causative organism and histopathological evaluation may be performed simultaneously in the same section using genotype-specific ISH. Routine use of ISH is largely restricted to diagnostic laboratories because the technique is technically more complex, cumbersome, and expense when compared with IHC. However, the ease of preparation of cDNA probes by RT-PCR compared with the generation of either monoclonal or polyclonal antibodies, combined with the increasing availability of probes and sequence databases, render the technique more applicable in diagnosis and research areas in the future.
The SR30 mAb–based IHC is more sensitive in the detection both type 1 and 2 PRRSV in FFPE lung tissues than SDOW17 mAb–based IHC. Similar to ISH, IHC also provides cellular detail and histological architecture so that the number of PRRSV-infected cells and lesions may be observed simultaneously in the same section. This advantage is critical for the evaluation of the PRRSV strain in infected pigs as a more virulent PRRSV strain can induce more severe interstitial pneumonia histopathologically than a less-virulent strain regardless of genotype of PRRSV.11,12
Presently, there is no ideal practical diagnostic method that allows detection and differentiation of the 2 genotypes of PRRSV in FFPE tissues. Based on the present results, IHC is the practical diagnostic method. However, it will be necessary to develop a genotype-specific PRRSV mAb to differentiate between the 2 genotypes of PRRSV for IHC in FFPE lung tissues.
Footnotes
a.
Rural Technologies Inc., Brookings, SD.
b.
Boehringer Mannheim, Indianapolis, IN.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by contract research funds of the Research Institute for Veterinary Science (RIVS) from the College of Veterinary Medicine and by the Brain Korea 21 Program for Veterinary Science in the Republic of Korea.
