Abstract
Three Actinobacillus pleuropneumoniae isolates from clinical cases of porcine pleuropneumonia were positive by capsular serovar 12–specific PCR assay, but not reactive to antiserum prepared against serovar 12 using the rapid slide agglutination (RSA) test. The isolates were positive for apxIICA, apxIIICA, apxIBD, apxIIIBD, and apxIVA in the PCR toxin gene assay, which is the profile seen in serovars 2, 4, 6, 8, and 15, and reacted with antisera against serovars 3, 6, 8, 15, and 17. Nucleotide sequence analysis revealed that genes involved in the biosynthesis of capsular polysaccharide of the 3 isolates were identical or nearly identical to those of serovar 12. However, genes involved in the biosynthesis of O-polysaccharide of the 3 isolates were highly similar to those of reference strains of serovars 3, 6, 8, 15, 17, and 19. In agreement with results from the RSA test, transmission electron microscopic analysis confirmed the absence of detectable capsular material in the 3 isolates. The existence of nonencapsulated A. pleuropneumoniae serovar K12:O3 would hamper precise serodetection.
Actinobacillus pleuropneumoniae is the causative agent of porcine pleuropneumonia, an economically important bacterial disease of swine. 6 Isolates of A. pleuropneumoniae can be separated into 2 biovars, the more prevalent biovar 1, which requires exogenous nicotinamide adenine dinucleotide (NAD) for growth, and biovar 2, which is NAD-independent. 20 To date, 19 serovars have been recognized, mainly on the basis of antigenic diversity of capsular polysaccharides (CPSs), but also by lipopolysaccharide (LPS) O-polysaccharides (O-PSs).3,4,26 The O-PS antigens are shared by many serovars (e.g., serovars 1, 9, and 11; serovars 4, 7, 13, 18, and 19; serovars 5 and 16; serovars 3, 6, 8, 15, 17, and 19),6,17 resulting in cross-reactions among the serovars in each group. The homologous O-PS antigens may be structurally similar or even identical. 17
A scheme using both CPS and O-PS antigens has been proposed for serotyping A. pleuropneumoniae.2,8,16 For example, serovars 1, 9, and 11 would be designated as serovars K1:O1, K9:O9, and K11:O1. 2 Rare isolates (serovar or subtype) have also been reported (e.g., strains of K2:O7, 16 K19:O4, K19:O3, 26 K1:O7, 8 K13:O10, 18 K4b:O3, 28 and K12:O3 10 ). We report here the characterization of 3 nonencapsulated A. pleuropneumoniae isolates of K12:O3.
We used field isolate 803 of A. pleuropneumoniae, obtained from the pneumonic lung of a diseased pig on farm A in central Japan in November 2015, and 2 other isolates, 2680 and 2725, obtained from the pneumonic lungs of 2 diseased pigs on farm B in southern Japan in January 2016 and February 2017 respectively. The set of A. pleuropneumoniae reference strains of serovars 1–18 (4074, S1536, S1421, M62, K17 (serovar 5a), L20 (serovar 5b) and Femø, WF83, 405, CVJ13261, D13039, 56153, 1096, N273, 3906, HS143, A-85/14, 16287-1, and 7311555, respectively) was from our laboratory stock culture collection (Nippon Institute for Biological Science, Tokyo, Japan). All bacterial strains were grown on chocolate agar (Becton, Dickinson) for 16 h at 37°C in the presence of 5% CO2.
Species-specific identification based on partial 16S rDNA sequencing, performed as described previously,1,29 showed that the nucleotide sequence of a 919-bp hypervariable region of the 16S RNA gene of the 3 isolates had homology of 99.0–100% with those of other A. pleuropneumoniae isolates available in GenBank. The PCR-based apx gene assay5,22 for the 3 isolates was positive for apxIICA, apxIIICA, apxIBD, apxIIIBD, and apxIVA, but negative for apxICA. The 3 isolates were positive by capsular serovar 12–specific PCR assay and negative by biovar (nadV)-specific PCR assay. 4 The negative result using the biovar (nadV)-specific PCR assay, and the fact that the isolates were found to be dependent on NAD in vitro, indicated that these 3 isolates were A. pleuropneumoniae biovar 1.
We serotyped the 3 field isolates in our study, as well as the serovar 12 reference strain 1096, with serovar-specific rabbit antisera prepared against serovar reference strains by using the rapid slide agglutination (RSA) test, as described previously.15,27 Isolates 803, 2680, and 2725 reacted to antisera raised against serovar 3, 6, 8, 15, and 17 reference strains but to none of the other serovar reference strains. After adsorption of the serovar 3 and 15 antisera with antigens prepared from the reference strains of serovars 15 and 3, 15 respectively, there was no reaction in the RSA test between the adsorbed serovar 3 and 15 antisera and antigens of any of the 3 field isolates and those of serovars 6, 8, and 17. However, the adsorbed serovar 3 and 15 antisera still gave a distinct reaction with antigens of serovars 3 and 15, respectively, suggesting that isolates 803, 2680, and 2725 have an O-PS similar to that of serovars 3 and 15. Taken together, these results provide very clear evidence that the 3 isolates of serovar K12:O3 were defective in CPS production, which differed from serovar K12:O3 isolate QA106, 10 a serovar K12:O3 isolate that produces both CPS and O-PS. Hence, cross-adsorption removed the cross-reactions without removing the specific homologous reactions.
To investigate the discrepancy between molecular and serologic typing of these 3 isolates, we PCR-amplified and sequenced the complete capsule and O-PS loci. The primer pairs used for amplification of the CPS and O-PS loci were designed based on previously published sequences (Table 1).28,29,32 The ingredients and conditions for the PCR assay (Tks Gflex DNA polymerase; Takara Bio) were as per the supplier’s protocol. Template DNA was prepared as described previously. 29 Briefly, the conditions for the PCR assay were as follows: 94°C for 1 min (1 cycle), 98°C for 10 s, and 68°C for 7 min 30 s (30 cycles). The PCR products were purified (NucleoSpin gel and PCR clean-up kit; Macherey-Nagel), and the nucleotide sequences were determined using a primer-walking procedure.
Primers used in the study of nonencapsulated Actinobacillus pleuropneumoniae isolates.
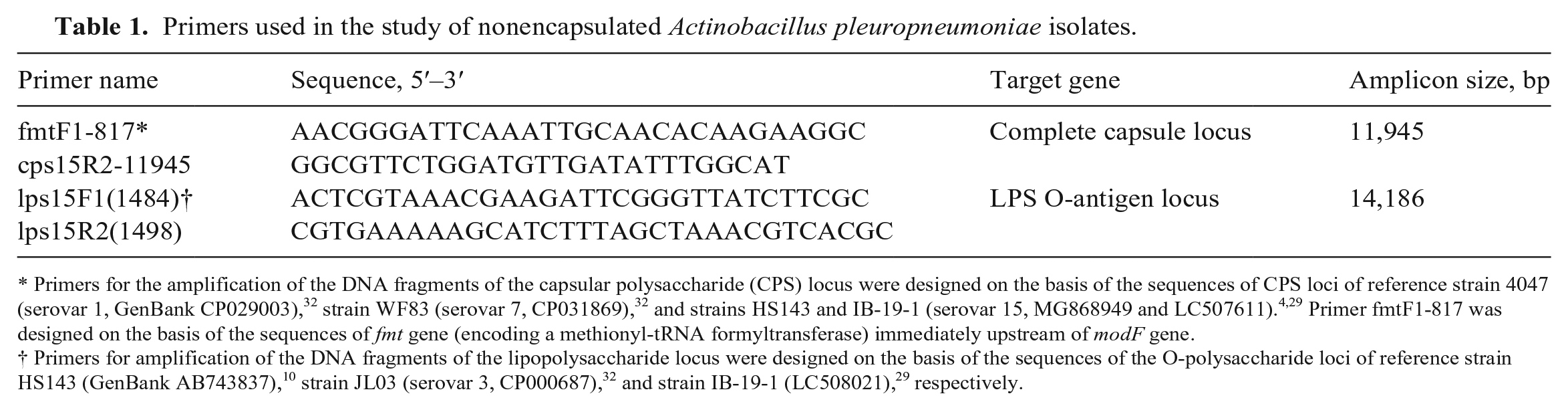

Primers for the amplification of the DNA fragments of the capsular polysaccharide (CPS) locus were designed on the basis of the sequences of CPS loci of reference strain 4047 (serovar 1, GenBank CP029003), 32 strain WF83 (serovar 7, CP031869), 32 and strains HS143 and IB-19-1 (serovar 15, MG868949 and LC507611).4,29 Primer fmtF1-817 was designed on the basis of the sequences of fmt gene (encoding a methionyl-tRNA formyltransferase) immediately upstream of modF gene.
By PCR amplification and sequence analyses, we found that the complete CPS loci (11,945 bp) of isolates 803, 2680, and 2725 were between the genes modF, encoding a putative molybdenum transport ATP-binding protein, and ydeN, a predicted serine hydrolase (Fig. 1A). The CPS export genes (cpxABCD) and CPS biosynthetic genes (cps12AB) in the 3 isolates were arranged in divergent directions similar to serovar 12 and other known serovars, except serovar 15, in which the export locus is downstream of the biosynthetic locus in the same orientation.4,10,12,32 The CPS loci identified in the 3 isolates were ≥ 99% identical at the nucleotide level and most closely resemble those of serovar 12 reference strain 1096 (GenBank ADOL00000000) and strain 8329 (CP031862), with ≥ 99% identity at the nucleotide level across 11,945 bp. The CPS loci of these 3 isolates and serovar K12:O3 strain QAS106 (AB701757) were > 98% identical at the nucleotide level across 6,879 bp.
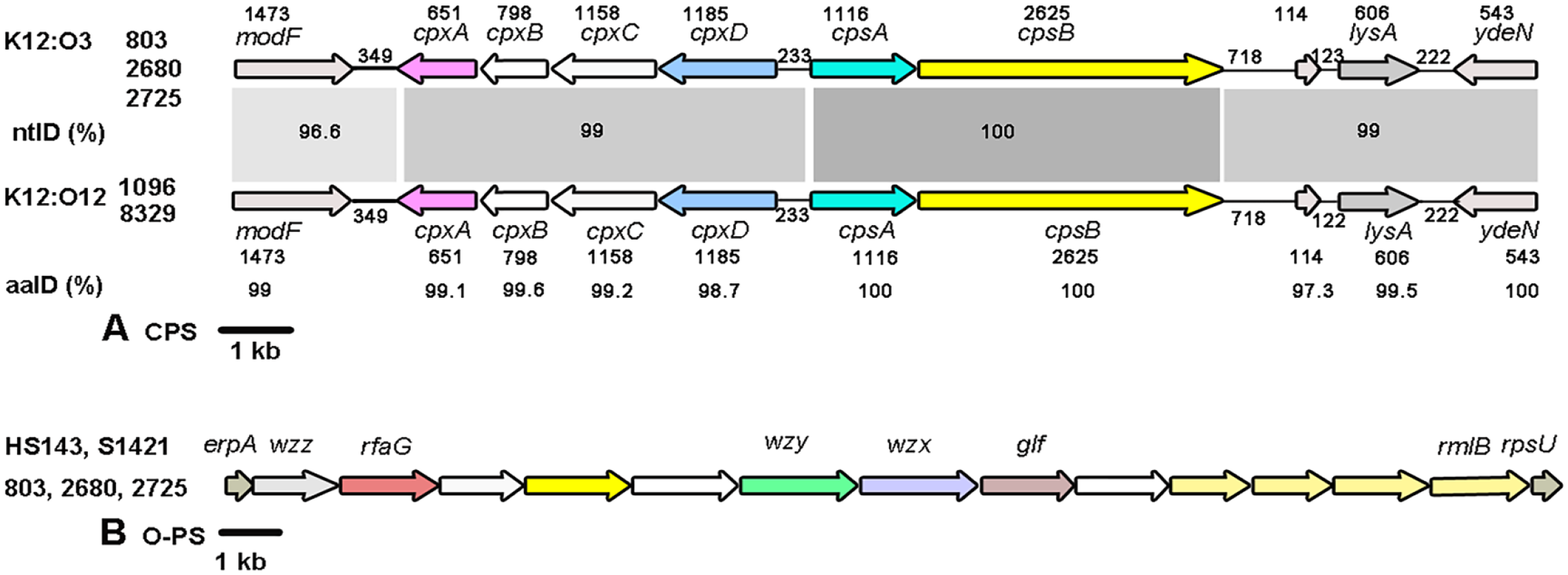
Schematic comparison of the gene clusters involved in the biosynthesis of (
The complete O-PS loci of isolates 803, 2680, and 2725 (14,186 bp) were between the genes wzz, encoding a Wzz-like protein, and rmlB, encoding a dTDP-glucose 4,6-dehydratase (Fig. 1B), sharing ≥ 99.9% identity with the O-PS loci in reference and field strains of serovars 3 (S1421), 15 (HS143), K12:O3 (QAS106), K4b:O3 (14-760), and K19:O3 (7213384-1) (GenBank CP031874, MG868949, AB743838, LC627113, and MT468889, respectively).4,10,26,28 Nucleotide sequences of the CPS and O-PS loci of isolate 803 were deposited in the DNA Data Bank of Japan (DDBJ)/European Molecular Biology Laboratory (EMBL)/GenBank as accessions LC740471 and LC740472, respectively.
To examine the capsular material surrounding cells of the reference strain 1096 (serovar K12:O12) and field isolates 803, 2680, and 2725 (K12:O3), the bacterial cells were prepared according to a previous study with some modification. 11 Briefly, agar-grown bacteria were harvested and washed with PBS (0.01 M, pH 7.2). The bacterial suspensions were adjusted to an optical density of 1.8 at 540 nm, and then exposed to rabbit antiserum prepared against the serovar 12 reference strain 1096 or to pre-immune serum for 1 h at 4°C. Bacterial cells were then washed with distilled water and fixed with 5% (v/v) glutaraldehyde containing 0.15% (w/v) ruthenium red for 2 h at room temperature. The cells were suspended in cacodylate buffer for 30 min. Next, the cells were immobilized in 4% agar, washed 5 times in cacodylate buffer, and post-fixed with 2% (w/v) osmium tetroxide for 2 h. Washings were repeated as above, and the samples were then dehydrated with a graded series of acetone washes. The samples were embedded in low-viscosity resin. Ultrathin sections of bacterial cells were stained (EM stainer; Nisshin EM) before examination with a transmission electron microscope (JEM-1400Flash; JEOL).
An irregular or thin layer of capsular material could be seen after immunostabilization of cells of the reference strain 1096 with the serovar 12 antiserum, which was not visible when the cells were exposed to pre-immune serum. On the other hand, no capsular material was observed when cells of field isolates 803, 2680, and 2725 were exposed to serovar 12 antiserum or to pre-immune serum, suggesting that these 3 field serovar K12:O3 isolates of A. pleuropneumoniae were defective in CPS production (Fig. 2). The findings, together with data reported by other investigators, support the idea that strains of A. pleuropneumoniae possess different amounts of extractable capsular material.11,19
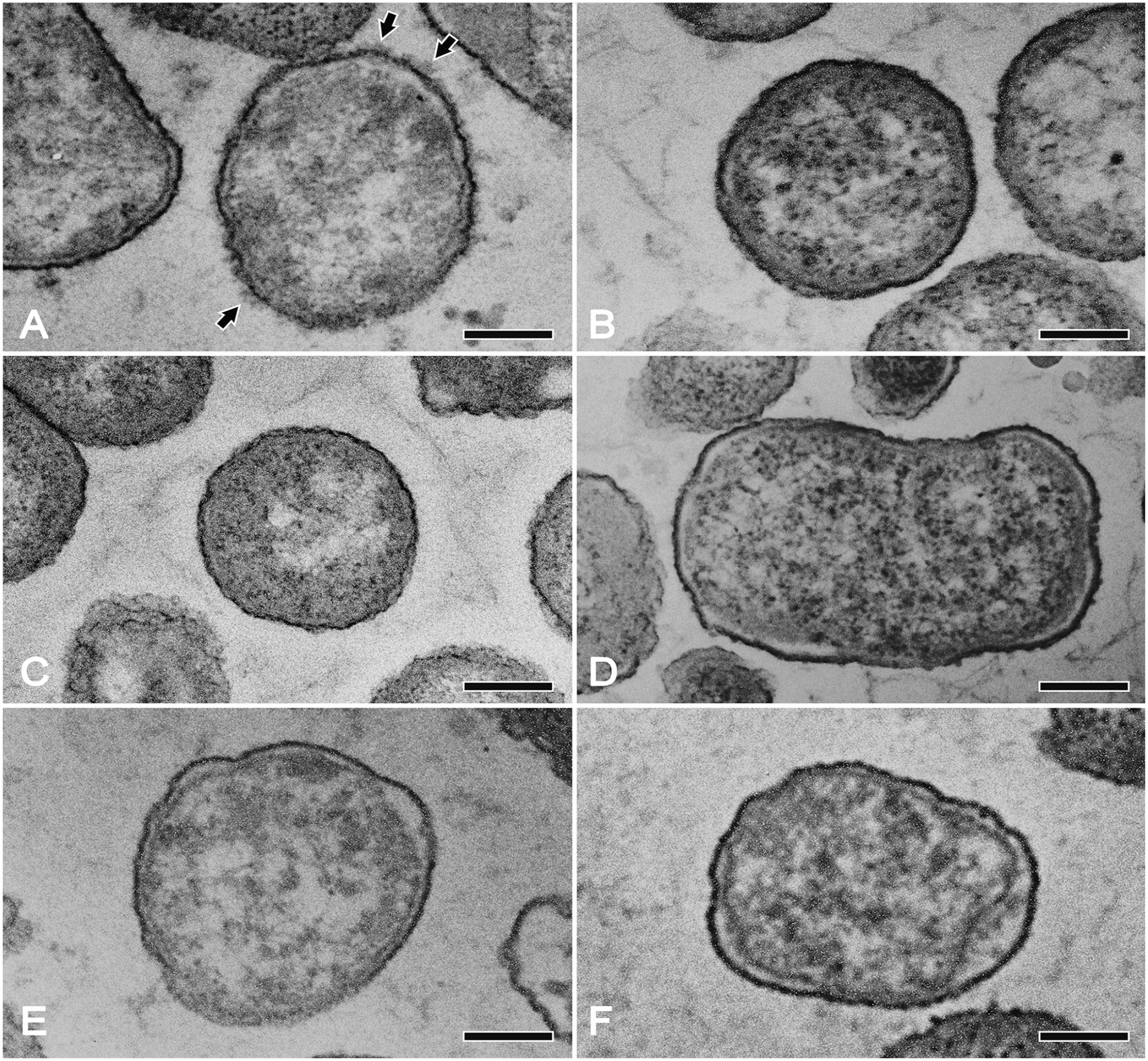
Transmission electron micrographs of ultrathin sections of Actinobacillus pleuropneumoniae cells exposed to whole-cell serovar 12 antiserum or to pre-immune serum. Cells of the reference strain 1096 (
Defects in the production of CPS or O-PS as a result of mutation, insertion, or deletion of nucleotides in structural genes for CPS or O-PS synthesis have been described previously.27–29 The cps gene cluster may include many of the genes specific to the capsule pathway, including the genes for polymerization, transport, and modification of the capsule. Data support the notion that the Rcs (the regulators of capsule synthesis) system, including RcsA, RcsB, and RcsC regulatory proteins, regulates the level of transcription of the genes involved in biosynthesis of CPS in Escherichia coli,9,25,31 Salmonella Typhi, 30 and Klebsiella pneumoniae. 31 The mutations in the regulatory genes may result in increased or decreased transcription of cps genes. 31
Sequence analyses of the CPS loci of the 3 field isolates revealed that there are no single-nucleotide variations (SNVs) in the cps12AB genes. Only missense and silent SNVs were found in the CPS export genes (modF, cpxA, cpxC) of these isolates. These SNVs do not appear to affect structure, stability, and function of enzymes encoded by the modF, cpxA, and cpxC genes. The reason for the defect in production of CPS in these 3 isolates is not clear but may be the result of mutations in the rcsA-, B-, and C-like genes of the regulatory system, a feature also seen in E. coli, Salmonella Typhi, and K. pneumoniae, as mentioned above.
Furthermore, the O-PS loci of isolates 803, 2680, and 2725 were identical or nearly identical to those of the serovar 3, 6, 8, 15, and 17 reference strains. This may be why antisera of these serovars (3, 6, 8, 15, 17) showed some cross-reactions to these 3 isolates, and why the antisera against serovars 3 and 15 absorbed with antigens of the serovar 15 and 3 reference strains, respectively, removed these cross-reactions. The cross-reactions observed with the O-antigens suggest that isolates 803, 2680, and 2725 could induce antibodies in infected pigs that would be detected by an ELISA based on serovars 3-6-8-15-17 LPS, as described recently for serovar 17–infected pigs. 7
The CPS and O-PS loci of isolates 803, 2680, and 2725 were highly similar to those of serovar K12:O3 isolate QAS106 isolated previously in Japan. However, the toxin gene profile of isolate QAS106 is typical of A. pleuropneumoniae serovar 3 (apxIICA, apxIIICA, and apxIIIBD only). 10 Findings from Apx toxin genotyping and DNA sequence analyses of the CPS and O-PS loci of the 3 field isolates in our study suggest that the K12 CPS locus may have arisen through recombination of the capsule-related genes acquired from a serovar 12 A. pleuropneumoniae isolate into the serovar 15 capsule locus. Evidence supporting serovar K12:O3 isolates 803, 2680, and 2725 as separate from isolate QAS106 includes the presence of the apxIBD export gene found in isolates 803, 2680, and 2725, and the loss or down-regulation of capsular expression. The absence of any detectable capsule was confirmed by our RSA test and ultrastructural study.
A. pleuropneumoniae serovar 2 strains are highly virulent in European and Asian countries owing to the secretion of 2 Apx toxins, ApxII and ApxIII, but serovar 2 strains in North America secrete only ApxII and are of low virulence.6,13 Hence, the 3 isolates of serovar K12:O3 producing both ApxII and ApxIII may be more virulent than isolates of serovar K12:O12 of A. pleuropneumoniae producing only ApxII. This aligns with the 3 isolates (803, 2680, and 2725) all being originally obtained from the lungs of diseased pigs with pleuropneumonia.
Inactivated vaccines are effective only against serovars present in the vaccine; subunit toxin– or bacterin toxin–based vaccines offer better cross-protection than bacterins.6,21 Most commercial vaccines successfully decrease clinical signs, but do not prevent colonization, infection, or transmission. 21 It seems that prevention and control measures lead to different selective pressures on the various A. pleuropneumoniae serovars, which in turn may result in a reduction of serovar diversity, and/or an increase in the frequency of yet unrecognized serovars or genetic changes in currently known serovars of A. pleuropneumoniae.13,14,23 This appears to be a country-specific phenomenon. 13 Three of 4 commercial, licensed vaccines in Japan, composed of major RTX toxoids (ApxI–III, capsule, and lipopolysaccharides from serovars 1, 2, and 5 as antigens) are effective against serovars 1, 2, and 5. 24 It remains to be seen whether the appearance of these new types reflects an emerging variant of A. pleuropneumoniae or if the present observation is a sporadic appearance, albeit it has already been observed on at least 2 different farms in Japan. This type of A. pleuropneumoniae, which contains K12 CPS and O3 O-PS loci but is defective in CPS production, may severely hamper serologic and bacteriologic characterization of porcine pleuropneumonia caused by serovars containing K12 and/or O3-6-8-15-17.
Footnotes
Acknowledgements
We thank Drs. Pat Blackall and Conny Turni (The University of Queensland, Australia) for providing A. pleuropneumoniae serovar 15 reference strain HS143, Dr. Fumiko Koike (SMC, Kanagawa, Japan) for providing field isolates 2680 and 2725 of A. pleuropneumoniae, and Atsushi Oshima (Nippon Institute for Biological Science, NIBS) for providing field isolate 803. We thank Dr. Masumi Sato (NIBS) for her expert advice on the preparation for transmission electron microcopy, and Ms. Michiha Kon, Mr. Yuta Akaike, and Ms. Kotoe Nogami (NIBS) for their excellent technical assistance during this work.
Declaration of conflicting interests
The authors declared no potential conflicts with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
