Abstract
An atypical Actinobacillus pleuropneumoniae serovar 12 strain, termed QAS106, was isolated from a clinical case of porcine pleuropneumonia in Japan. An immunodiffusion (ID) test identified the strain as serovar 12. However, the ID test also demonstrated that strain QAS106 shared antigenic determinants with both the serovar 3 and 15 reference strains. Strain QAS106 was positive in the capsular serovar 12–specific polymerase chain reaction (PCR) assay, while the PCR toxin gene profiling and omlA PCR typing assays indicated that strain QAS106 was similar to serovar 3. The nucleotide sequence of the 16S ribosomal DNA (rDNA) of strain QAS106 was identical with that of serovars 3 and 12, but it showed 99.7% identity with that of serovar 15. Nucleotide sequence analysis revealed that genes involved in biosynthesis of the capsular polysaccharide (CPS) of strain QAS106 were identical to those of serovar 12 at the amino acid level. On the other hand, strain QAS106 would express putative proteins involved in the biosynthesis of lipopolysaccharide (LPS) O-polysaccharide (O-PS), the amino acid sequences of which were identical or nearly identical to those of serovars 3 and 15. In conclusion, strain QAS106 should be recognized as K12:O3, even though typical serovar 12 strains are K12:O12. The emergence of an atypical A. pleuropneumoniae serovar 12 strain expressing a rare combination of CPS and O-PS antigens would hamper precise serodiagnosis by the use of either CPS- or LPS-based serodiagnostic methodology alone.
Actinobacillus pleuropneumoniae is the etiological agent of porcine pleuropneumonia, a severe respiratory disease that causes serious economic losses in the pig-rearing industry. 6 To date, 15 serovars have been recognized mainly based on the antigenic diversity of capsular polysaccharides (CPS) in the organisms, but also by lipopolysaccharide (LPS) O-polysaccharide (O-PS).3,6 However, O-PS antigens are shared by multiple serovars. For example, O-PS molecules of essentially the same or similar structure are shared by A. pleuropneumoniae serovars 1, 9, and 11; serovars 4, 7, and 13; and serovars 3, 6, 8, and 15,6,19 resulting in cross-reactivity among the serovars in each group. The antigenic types of CPS and O-PS antigens, which are expressed in A. pleuropneumoniae serovars, are usually inherent in each serovar, which has led to the proposal of a serotyping scheme with designations such as K1:O1, K2:O2, and K3:O3. 2 Rare isolates for which K and O antigens are atypical have also been reported. For example, strains of K2:O717 have been reported in Denmark, and strains of K1:O710 and K13:O1020 have been reported in Canada. The first isolation and genetic characterization of an atypical A. pleuropneumoniae serovar 12, which should be named serovar K12:O3 even though the typical capsular serovar 12 strain has been designated serovar K12:O12, is reported in the current study.
The field strain QAS106 of A. pleuropneumoniae was isolated in January 2011 in Japan from one of the lungs of a 139-day-old pig that showed no clinical signs on the day before death. The following Actinobacillus pleuropneumoniae serovar reference strains were also used: serovar 1, 4074; serovar 2, S1536; serovar 3, S1421; serovar 4, M62; serovar 5a, K17; serovar 5b, L20; serovar 6, Femø; serovar 7, WF83; serovar 8, 405; serovar 9, CVJ13261; serovar 10, D13039; serovar 11, 56153; serovar 12, 8329; serovar 13, N273; serovar 14, 3096; serovar 15, HS143. The serovars 1–13 reference strains were from the culture collection of the National Institute of Animal Health, whereas the serovar 1418 and 153 reference strains were kindly provided by Dr. Andresen (Denmark) and Dr. Blackall (Australia) via Dr. Nagai (Japan), respectively. All bacterial strains were grown as previously described.
Serotyping of the Japanese atypical strain QAS106 was carried out with serovar-specific rabbit antisera prepared against serovar reference strains by immunodiffusion (ID) tests as described previously. 16 Consequently, strain QAS106 was identified as serovar 12 by the ID test, based on the observation of a serovar 12–specific band that seemed to be derived from the CPS, when using rabbit antisera prepared against a serovar 12 reference strain (Fig. 1A). However, a cross-reactive band, which seemed to be derived from an O-PS common to strain QAS106 and the serovar 3, 8, and 15 reference strains, was observed with rabbit antisera prepared against the serovar 3 reference strain (Fig. 1B) even though the typical capsular serovar 12 strain expresses a serovar 12–distinct O-PS.2,6,19 In addition, a cross-reactive band was seen with rabbit antisera prepared against a serovar 15 reference strain in strain QAS106 and the serovar 3, 6, 8, and 15 reference strains (Fig. 1E). No cross-reactive band in strain QAS106 was exhibited with rabbit antisera prepared against the serovar 6 and 8 reference strains in the Japanese isolate (Fig. 1C and 1D, respectively). Analysis with the ID tests indicated that the atypical strain should be named serovar K12:O3 according to the previously proposed typing scheme. 2 It remains unknown why the atypical serovar 12 strain QAS106 does not exhibit similarity to serovar 8–specific antigenic determinants in the ID test using anti-serovar 8–specific antisera, despite the fact that the chemical structures of serovars 3, 8, and 15 have been reported to be identical.2,6,19
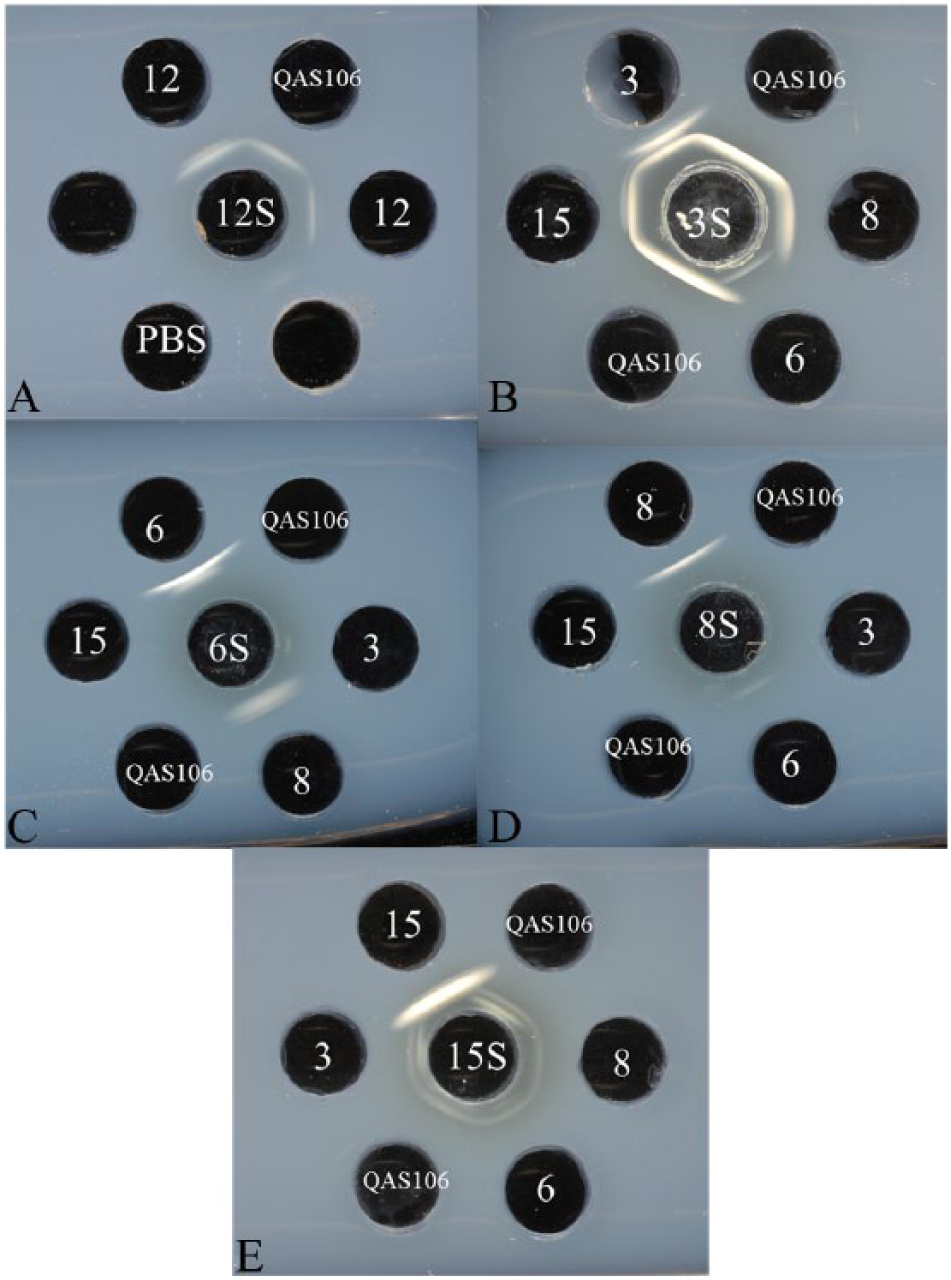

Immunodiffusion tests with antiserum prepared against the Actinobacillus pleuropneumoniae serovar 12 (
Genetic approaches, including PCR typing and nucleotide sequencing, were then used in order to confirm that the atypical serovar 12 strain QAS106 could be named serovar K12:O3. First, PCR typing including a capsular serovar 12–specific PCR that is based on genes linked to the CPS (cps12-specific PCR), 1 PCR toxin profiling, 5 and omlA PCR typing, 11 which facilitate rapid serotyping or serogrouping, were performed as previously described. The cps12-specific PCR was positive. The PCR toxin gene profiling was positive for apxIICA, apxIIICA, and apxIIIBD, but negative for apxICA and apxIBD, which is the profile seen in serovar 3. 5 The omlA PCR type corresponded to that seen in serovars 3, 6, 8, and 15, 11 indicating that strain QAS106 could be serovar 3.
As a second genetic study, the nucleotide sequence of 16S ribosomal DNA (rDNA) was determined in order to examine the phylogenetic relationship between the atypical serovar 12 strain QAS106 and A. pleuropneumoniae serovars. In order to amplify approximately 1,500 base pairs (bp) of the 16S rDNA, PCR was performed using 2 primers, FD1 (5′-AGAGTTTGATCNTGGCTNAG-3′) and RD1 (5′-AAGGAGGTCATCCAGCC-3′), the sequences of which were modified from the previous report. 22 Template DNA was prepared as described elsewhere. 14 The PCR was essentially performed as described previously. 22 The amplified product was purified using a commercially available purification kit, a and the nucleotide sequence was then determined by cycle sequencing using the synthesized oligonucleotides in order to determine the sequences of both strands as described previously. 13 The nucleotide sequence of the 16S rDNA of strain QAS106 (1,496 bp) was identical to that of the serovar 3, 6, 8, and 12 strains (EMBL/GenBank/DDBJ accession nos. CP000687, AF033060, AF033060, and AF033060, respectively), whereas it exhibited 99.7% identity to that of the serovar 15 strain (accession no. AY017472), revealing that the Japanese atypical isolate was more phylogenetically related to serovars 3, 6, 8, and 12 than to serovar 15.
As a third genetic study, the nucleotide sequence of the cps gene of strain QAS106 was determined and then compared with sequences deposited in the EMBL/GenBank/DDBJ DNA databases because the genetic organization of the cps genes of A. pleuropneumoniae serovars investigated to date correspond to the CPS structure. 15 In order to determine the nucleotide sequence of the cps gene of strain QAS106, the internal region of the cpxD gene of strain QAS106 was PCR-amplified with cpxD-F1 (5′-ACYTCAGCCCTAGCCATASTGC-3′) and cpxD-R2 (5′-CACACGATAAACCGTYGGTACATC-3′), which were designed based on the published sequences. 23 Purification of the amplified DNA and determination of the nucleotide sequence were performed as described previously. 13 Then, an inverse PCR 21 was performed in order to amplify the DNA regions flanked by cpxD with the primers invF (5′-GCAGTAGGCGGAACAACGGAAAACATT-3′) and invR (5′-ATATCCCGCACCGCCTACAGTACCTAAAAA-3′), which were designed based on the nucleotide sequences determined in the current study. Briefly, purified genomic DNA of strain QAS106, which was prepared as described previously, 14 was digested with restriction enzymes EcoRI b and HindIII, b and then self-ligated with T4 ligase b in order to generate circular DNA. An inverse PCR was then performed with the ligated DNA and primers cpxD-invUP2 (5′-GCAGTAGGCGGTACAACGGAAAATATT-3′) and cpxD-invLW2 (5′-ATAGCCCGCGCTGCCTACGTGCCTAAAAA-3′) and the PCR enzyme c as recommended by the supplier. The conditions for DNA amplification, or so-called stepdown PCR, were as follows: step 1, 94°C for 2 min (1 cycle); step 2, 98°C for 10 sec and 74°C for 20 min (5 cycles); step 3, 98°C for 10 sec and 72°C for 20 min (5 cycles); step 4, 98°C for 10 sec and 70°C for 20 min (5 cycles); step 5, 98°C for 10 sec and 68°C for 20 min (20 cycles); and final step, 68°C for 10 min. Purification of the amplified DNA and determination of the nucleotide sequence were carried out as described above. 13 The determined nucleotide sequences of the cps gene and the flanking region of strain QAS106 (8,551 bp) were subsequently analyzed and compared with those deposited in the EMBL/GenBank/DDBJ DNA databases. Two open reading frames (ORFs) that seemed to be related to CPS biosynthesis were found upstream of the cpxD gene of the atypical isolate. The nucleotide sequences of the 2 ORFs were identical to those of cps12A and cps12B of the A. pleuropneumoniae serovar 12 reference strain, respectively. 15 This finding supported the immunological result of the ID test that QAS106 was identified as serovar 12.
As a fourth genetic study, the nucleotide sequence of the o-ps gene of strain QAS106 (14,186 bp) was determined and compared with that of serovars 324 and 6, 23 as the ID test indicated that 1) the strain QAS106 might express an O-PS that was antigenically similar to that of serovars 3 and 15; 2) the O-PS structures among serovars 3, 8, and 15 are chemically identical2,6,19; and 3) serovars 3, 6, 8, and 15 often cross-react with conventional serotyping methods such as slide agglutination. 6 To amplify the o-ps genes, PCR was performed using 2 primers, erpA-F2 (5′-ACTCGTAAACGAAGATTCGGGTTATCYTCGCC-3′) and rpsU-R1 (GCGTGACGTTTAGCTAAAGATGCTTTTTCACGTTT-3′), which were designed based on the published data as described previously, 23 and a commercial PCR enzyme, c which was essentially used following the supplier’s protocol. Template DNA was prepared as described elsewhere. 14 The conditions for the stepdown PCR were as follows: step 1, 94°C for 2 min (1 cycle); step 2, 98°C for 10 sec and 74°C for 16 min (5 cycles); step 3, 98°C for 10 sec and 72°C for 16 min (5 cycles); step 4, 98°C for 10 sec and 70°C for 16 min (5 cycles); step 5, 98°C for 10 sec and 68°C for 16 min (30 cycles); and final step, 68°C for 16 min. Purification of the amplified DNA and determination of the nucleotide sequence were performed as described above. 13 In addition, the nucleotide sequences of the o-ps genes of serovars 8 and 15 (13,969 and 14,186 bp, respectively) were determined in the current study, for the first time, by the method used for strain QAS106. The o-ps gene of strain QAS106 contained 13 ORFs, the deduced amino acid sequences of which were identical or nearly identical to those of the serovar 3 strain JL0324 and serovar 15 strain HS143 (Table 1), suggesting that strain QAS106 would express O-PS, which shared antigenic determinants with those of serovars 3 and 15. The atypical strain QAS106 could therefore be named O3, as the O-PS molecules of serovars 3 and 15 are chemically identical.2,6,19 At the amino acid sequence level, 12 of 13 proteins encoded by the ORFs were identical or nearly identical (more than 98% identity) to those of serovar 623 and serovar 8 (Table 1). On the other hand, the Wzz-like protein (80.5% identity) and VI polysaccharide biosynthesis protein (66.7% identity) of strain QAS106 showed considerable but lower homology to those of serovars 8 and 6, respectively (Table 1).
Comparison of proteins involved in the biosynthesis of O-polysaccharide of lipopolysaccharide in Actinobacillus pleuropneumoniae strain QAS106 to those from A. pleuropneumoniae serovars 3, 6, 8, and 15 at the amino acid level.*
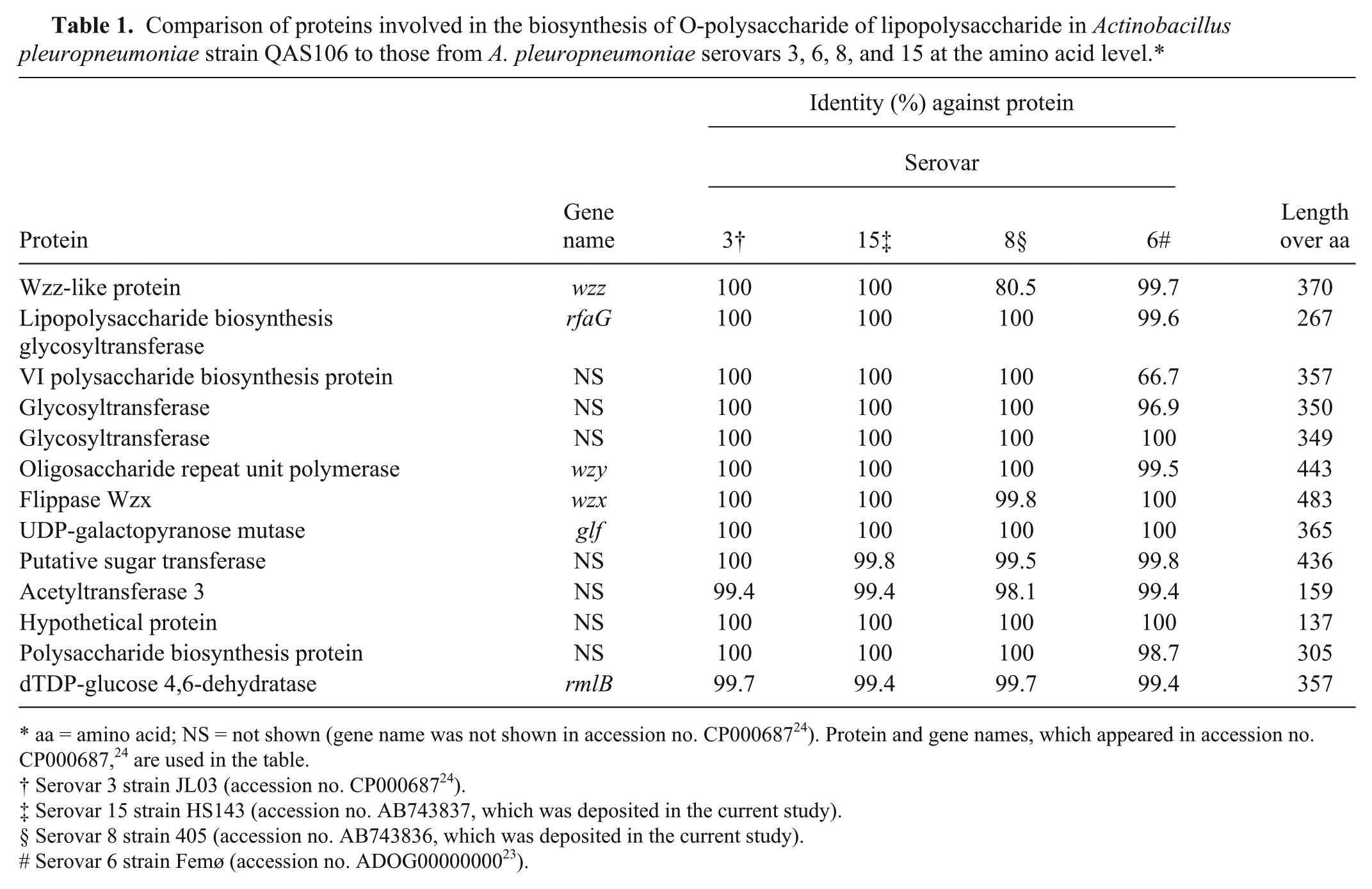
aa = amino acid; NS = not shown (gene name was not shown in accession no. CP00068724). Protein and gene names, which appeared in accession no. CP000687, 24 are used in the table.
Serovar 3 strain JL03 (accession no. CP00068724).
Serovar 15 strain HS143 (accession no. AB743837, which was deposited in the current study).
Serovar 8 strain 405 (accession no. AB743836, which was deposited in the current study).
Serovar 6 strain Femø (accession no. ADOG0000000023).
Nucleotide sequences of the 16S rDNA, and cps and o-ps genes of strain QAS106 were deposited in the EMBL/GenBank/DDBJ DNA databases under accession numbers AB635380, AB701757, and AB743838, respectively. Nucleotide sequences of the o-ps genes of the serovar 8 (strain 405) and serovar 15 (strain H143) reference strains were also deposited in the EMBL/GenBank/DDBJ DNA databases under accession numbers AB743836 and AB743837, respectively.
The genetic profiles in the current study revealed that almost all of the investigated genetic regions of strain QAS106 were the same as or the most similar to those of serovar 3 rather than those of serovar 12, with the exception that the cps12-specific PCR was positive. These observations indicated that strain QAS106 might have originally been serovar 3 and may have later evolved by acquiring genes for the biosynthesis of the serovar 12–specific CPS. In order to examine this hypothesis, full genome sequencing would be helpful. In practical application, the use of a CPS-based enzyme-linked immunosorbent assay (ELISA)4,12 would probably diagnose herds infected with the atypical isolate as infected by A. pleuropneumoniae serovar 12, whereas the use of an LPS-based ELISA7–9 would probably identify them as infected by serovars 3 or 15. In conclusion, the antigenic and genetic study of strain QAS106 in the current study revealed the presence of an A. pleuropneumoniae K12:O3 strain expressing a new combination of CPS and LPS O-PS antigens in Japan.
Footnotes
Acknowledgements
Drs. L.O. Andresen, P.J. Blackall, and S. Nagai are thanked for providing bacterial strains. The assistance of Dr. Kobayashi for critical reading of this article is acknowledged.
a.
QIAquick PCR purification kit, Qiagen GmBH, Hilden, Germany.
b.
Takara Bio Inc., Otsu, Japan.
c.
KOD-FX Neo, Toyobo Co., Osaka, Japan.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by grants from the National Agriculture and Food Research Organization, Japan.
