Abstract
The fowl glioma-inducing virus prototype (FGVp) and its variants, which belong to avian leukosis virus subgroup A (ALV-A), induce cardiomyocyte abnormalities and gliomas in chickens. However, the molecular mechanisms underlying these myocardial changes remain unclear, and ALV-induced tumorigenesis, which is caused by proviral insertional mutagenesis, does not explain the early development of cardiac changes in infected chickens. We established a quantitative PCR (qPCR) assay to measure ALV-A proviral loads in the brains and hearts of FGV-infected Japanese bantam chickens and compared these results with morphologic lesions. Four of 22 bantams had both gliomas and cardiac lesions. Hearts with cardiac lesions had a higher proviral load (10.3 ± 2.7 proviral copies/nucleus) than those without cardiac lesions (0.4 ± 0.4), suggesting that the proviral load in hearts is correlated with the frequency of myocardial changes. Our qPCR method may be useful in the study of ALV-induced cardiomyocyte abnormalities.
Avian leukosis viruses (ALVs; Retroviridae, Alpharetrovirus) are major oncogenic retroviruses associated with neoplastic diseases, such as lymphoid and myeloid leukosis, in chickens. The fowl glioma-inducing virus prototype (FGVp; GenBank AB112960) and its variants (FGV variants), which belong to ALV subgroup A (ALV-A), induce astrocytomas, with disseminated nonsuppurative encephalitis, cerebellar hypoplasia, and nonsuppurative myocarditis.12,13,31,35–37 The Km_5666 strain, which was isolated from Japanese ornamental bantam chickens, is a unique FGV variant that causes cardiomyocyte abnormalities, such as lymphocytic myocarditis, cardiomyocyte hypertrophy, and cardiac rhabdomyoma, as well as astrocytoma in affected chickens.26,30
Oncogenic retroviruses are classified into 2 groups based on how they cause disease: acute transforming retroviruses, which carry viral oncogenes, and non-acute retroviruses. The latter cause neoplasms via 1 of 3 mechanisms: 1) oncogenesis by viral regulatory proteins (e.g., Tax), 2) proviral insertional mutagenesis, and 3) envelope protein (Env)-induced transformation. 20 ALVs lack viral oncogenes and regulatory proteins; therefore, in lymphoid leukosis (LL) caused by ALVs, the transformation of host cells is caused by proviral insertional mutagenesis, which activates a cellular oncogene by the nearby integration of the ALV provirus in the same host.20,28 In contrast to other well-studied retroviruses, such as human immunodeficiency virus 1 (HIV-1) and murine leukemia virus, ALVs randomly integrate into host gDNA without bias.2,18,21,27 Therefore, multiple rounds of infection are needed before a provirus inserts itself in the vicinity of a cellular proto-oncogene. 20 The earliest neoplastic changes in LL are detected 8 wk after infection, and lymphomatous foci develop in peripheral visceral organs after a latency period of ~14 wk. 7 In contrast, we have demonstrated that FGVs, including FGVp and FGV variants, induce fowl glioma, cerebellar hypoplasia, and cardiac rhabdomyomas (hamartoma) in as little as 50 d after in ovo inoculation25,26,29; the proviral insertional mutagenesis theory does not explain why FGV-related lesions develop much earlier than expected. Therefore, a distinct mechanism appears to be responsible for the tumorigenesis and pathogenesis of FGVs.
In human retroviral infections, such as HIV-1 and T-lymphotropic virus 1 (HTLV-1; Primate T-lymphotropic virus 1), an infected cell contains a single copy of the integrated provirus in the host genome.6,14 The infected cell rate explains the proviral load because the number of proviruses reflects the number of infected cells. A significant increase in the HTLV-1 proviral load is related to the development of HTLV-1–associated disease, 16 and the prevalence of the disease exponentially increases once the proviral load is > 1%.23,32 In contrast, difficulties are associated with calculating the ALV-infected cell rate because ALV proviral integration occurs at many sites in the host genome, and infected cells contain various numbers of viral DNA, namely, up to 20 copies of the provirus. 24 After reverse transcription (RT), the cDNA copies of ALV genomes are integrated into the host genome or circulate themselves. The circular forms contain either 1 or 2 copies of the long terminal repeat region and accumulate in infected cells because of abortive integration processes. However, the exact percentage of circular DNA products present is unknown in ALV infections. Additionally, it is technically challenging to distinguish these forms by quantitative PCR (qPCR). Circular DNAs may also participate in gene expression similar to integrated DNAs, followed by linearization, degradation, or accumulation as a reservoir. 8
We previously confirmed that the incidence of fowl gliomas was 17–100% 50 d after in ovo inoculation of FGVs,26,29 and the intracerebral proviral load was correlated with astrocytic proliferation following FGV infection. 25 Based on these findings, we hypothesized that the proviral load in tissues is correlated with pathogenicity. Given that multiple proviruses are integrated into a single cell in ALV infection, the signaling pathways of host cells may be activated more frequently with increases in the proviral load. However, there is no information on the copy number of FGV proviruses or the infected cell rate during the induction of glioma, to date. Furthermore, FGVs have the ability to integrate and replicate in the myocardium 35 ; however, the relationship between the pathogenicity of cardiomyocyte abnormality in FGV infection and the proviral load in the heart remains unclear. Therefore, we aimed to establish a qPCR assay to measure the ALV proviral load in the brains and hearts of FGV-infected Japanese bantam chickens and to clarify the relationship between the proviral load and pathogenicity.
Materials and methods
Animals and histopathology
Animal protocols were approved by the Animal Care and Use Committee of Iwate University (permits A201621, A201904, A202221) in accordance with the Japanese Act on the Welfare and Management of Animals. We examined 22 Japanese bantam chickens (Gallus gallus domesticus), known as Higo Chabo, in 2021 (Table 1). These birds were collected from the chicken population in Kumamoto, Japan. The birds were 0.5–5-y-old, and they were mainly culled at the breeders’ discretion because of torticollis, atrophic comb, emaciation, or old age (≥ 3-y-old). The chickens used for histologic examination were euthanized according to a procedure approved by Iwate University.
Summary of lesions and molecular analyses of Japanese bantam chickens and isolated viruses.
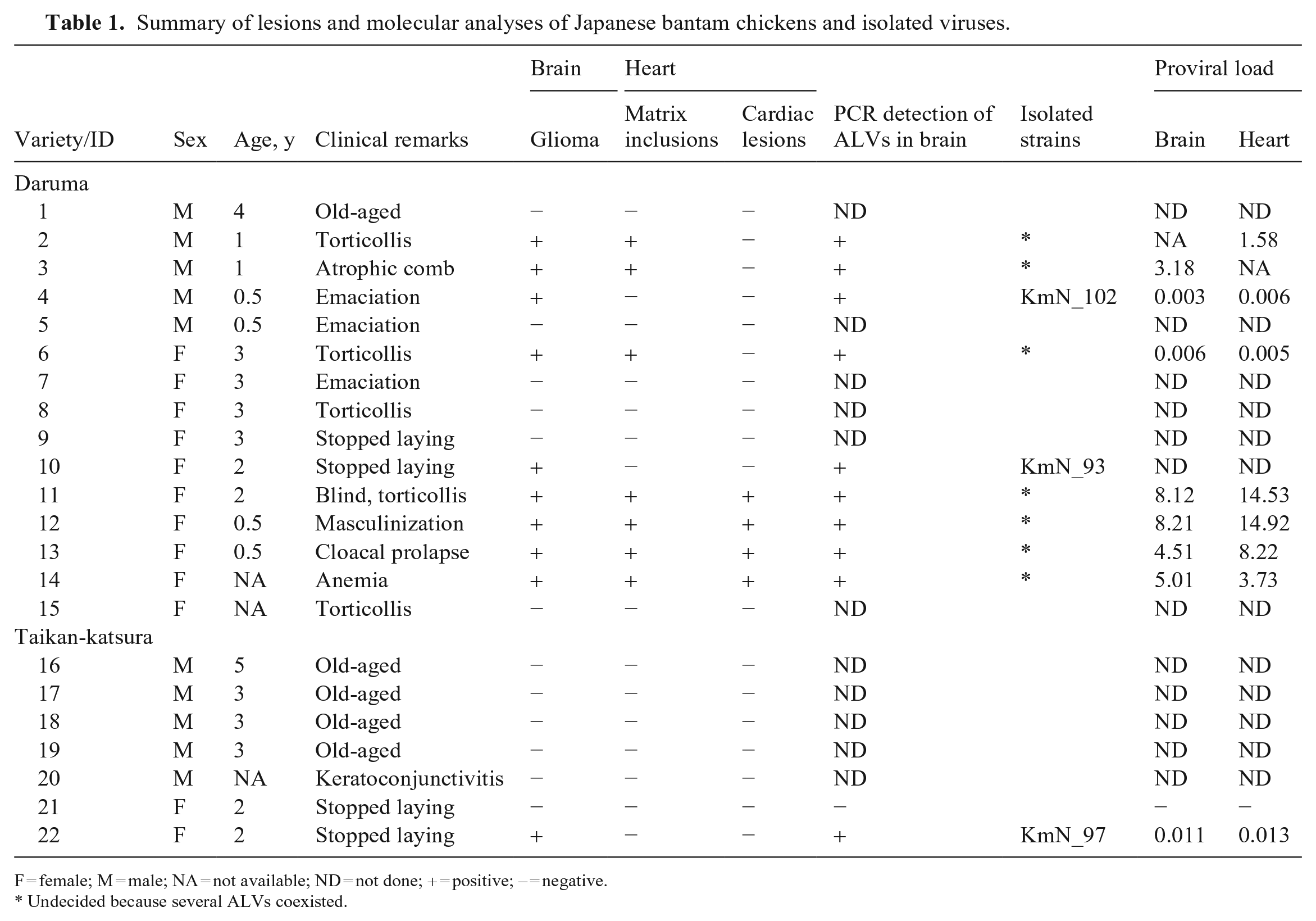
F = female; M = male; NA = not available; ND = not done; + = positive; – = negative.
Undecided because several ALVs coexisted.
Samples of liver, spleen, kidneys, lungs, heart, bones, and brain were fixed in 10% neutral-buffered formalin, processed routinely, and 4-µm sections were stained with H&E. Brain lesions were defined and classified as gliosis if there were clusters of < 10 astrocytes or as glioma (astrocytoma) if there was solid growth of > 10 atypical astrocytes distinguishable from the surrounding tissue. 25
Immunostaining was performed using antibody-labeled polymer methods (Histofine Simple Stain MAX PO (MULTI) kit, Histofine DAB substrate kit; Nichirei Bioscience) for glial fibrillary acidic protein (GFAP) and ALV antigens with the primary antibodies of rabbit anti-bovine GFAP (Dako) and anti-ALV antibodies at dilutions of 1:2,000 and 1:200,000, respectively. The polyclonal antibody against ALVs was generated via the immunization of rabbits with Rous-associated virus 2 (Tsukamoto K, pers. comm., 2001). 38 This antibody mainly recognizes the group-specific proteins p27 and gp85 of ALVs.
Virus isolation
The cerebrum of chickens was homogenized, and DF-1 cells (chicken fibroblast cell line) that had been cultured to a confluency of 80% were inoculated with the emulsion supernatant. DF-1 cells were obtained from the American Type Culture Collection and cultured in Dulbecco modified Eagle medium (Nissui Pharmaceutical) supplemented with L-glutamine (Gibco), penicillin–streptomycin solution (Gibco), and fetal bovine serum (lot 13K272; Sigma-Aldrich). The cell culture was performed as described previously. 11 The culture supernatant and cells were harvested 5 d after inoculation.
PCR amplification for ALV detection and DNA sequencing
We extracted total RNA from DF-1 cells (RNeasy mini kit, Qiagen; or NucleoSpin RNA plus, Takara Bio). We synthesized cDNA by performing RT as described previously. 10 Complementary DNA was then amplified by PCR. We used primers specific for chicken β-actin (5′-TATCCGTAAGGATCTGTATG-3′ and 5′-ATCTCGTCTTGTTTTATGCG-3′) as an internal control and the primer set ALV38 (5′-TTAGGTTCCCAGTCTCTCCC-3′) and ALV39 (5′-ATTGCGGGTGGTAGCGCTTT-3′) to target ALV-specific regions. 33 The primers were designed to target the variable region in the SU (gp85)-coding sequence in the env gene (envSU), which is often used for phylogenetic analysis and to estimate ALV subgroups.4,5,34 The reaction mixture contained 0.25 μL of PrimeSTAR HS DNA polymerase (2.5 U/μL; Takara Bio), 5 μL of PrimeSTAR buffer (5×, containing Mg++; Takara Bio), 7.5 pmol of each primer, and autoclaved Milli-Q water in a final volume of 25 µL. PCR was performed (ProFlex PCR system; Thermo Fischer) under the following cycling conditions: 94°C for 1 min, followed by 30 cycles of 98°C for 10 s, 60°C for 5 s, and 72°C for 40 s. The resultant PCR products were analyzed by 1.5% agarose gel electrophoresis using ethidium bromide staining. Additionally, amplicons were purified (NucleoSpin gel and PCR clean-up kit; Takara Bio), and DNA sequencing of the products was performed by a commercial laboratory (Eurofins Genomics). We named the strains isolated KmN_strains.
Real-time qPCR to measure the proviral load
Samples were total DNA extracted from DF-1 cells, specific pathogen–free chicken embryos, and pieces of brain and heart tissues from adult Japanese bantam chickens 2–4, 6, 11–14, 21, 22 (DNeasy blood and tissue kit; Qiagen). DNA concentrations were measured (NanoDrop ND-1000 spectrophotometer; Thermo Fisher). We quantified 7 candidate reference genes (ACTB, HMBS, TBP, HPRT1, PGK2, ACTG1, YWHAZ) to select 3 appropriate genes as biological replicates to calculate the number of nuclei. The primers for qPCR were designed using Primer 3 v.4.017,39 (Table 2). ACTB and HMBS have been reported as single-copy genes in chickens.40,42 TBP, HPRT1, PGK2, ACTG1, and YWHAZ are generally used as internal control genes in chickens.3,9,15,22 We then measured the absolute copy number of the ALV-A provirus utilizing a primer set designed to detect the ALV-A sequence (5′-TGGAAAGGTGAGCAAGAAGGA-3′ and 5′-CACACAAGACCAGGACACCAA-3′). 25 Standard plasmids were constructed by Taq-amplified cloning (Mighty TA-cloning reagent set for PrimeSTAR; Takara Bio) and transformed into JM109 competent cells by heat shock methods. After extraction (QIAprep spin miniprep kit; Qiagen), plasmids were linearized with the restriction enzyme Hind III-HF (New England Biolabs). Standard curves were generated by creating 10-fold serial dilutions of the standard plasmids ranging from 102–108 copies/reaction for the ALV provirus and 103–107 copies/reaction for the reference gene in duplicate. Quantitative PCR was performed (Applied Biosystems QuantStudio1 thermocycler, Thermo Fisher; TB green premix ex Taq II [Tli RNaseH Plus], Takara Bio) in a 20-μL reaction with 100 ng of DNA, 0.4 μL of ROX reference dye II (50×), and 6 pmol of each primer. Thermocycling conditions were as follows: initial denaturation at 95°C for 30 s, followed by 40 cycles of 95°C for 5 s and 63°C for 30 s, with a melting curve analysis step. Each sample was analyzed in duplicate. PCR efficiency and the correlation coefficient (R2) were calculated (QuantStudio Design and Analysis software v.2.6.0; Thermo Fisher). We calculated the number of nuclei from the average copy number of the 3 selected reference genes and defined the number of provirus copies per nucleus as the proviral load. We defined the number of all forms of viral DNA copies, including circular forms, as the proviral load. Sequence data are available in GenBank under accessions OP649596–OP649598.
Details of 7 candidate reference genes and primers used in the qPCR analysis.
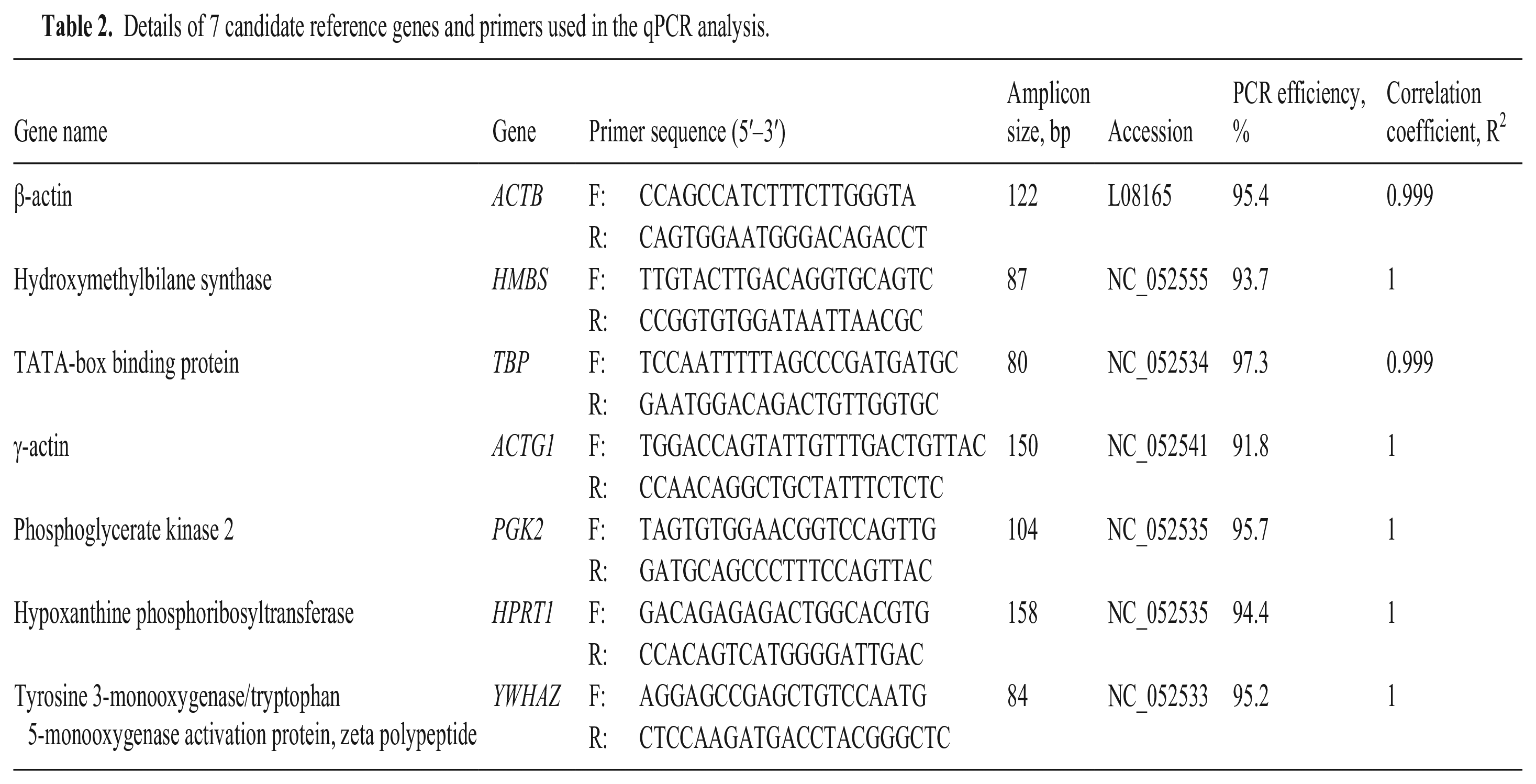
Results
Gross and histologic findings
We examined 22 Japanese bantam chickens (Table 1). At autopsy, as incidental lesions, birds 1 and 11 had small renal cysts; bird 14 had mild yolk peritonitis. The other 19 birds had no evident gross lesions. Histologically, 10 of 22 birds had gliomas (Figs. 1, 2). These astrocytes were predominantly gemistocytes and were immunopositive for GFAP (Fig. 3) and common ALV antigens (Fig. 4). Histologic findings were consistent with those reported in previous cases. 12
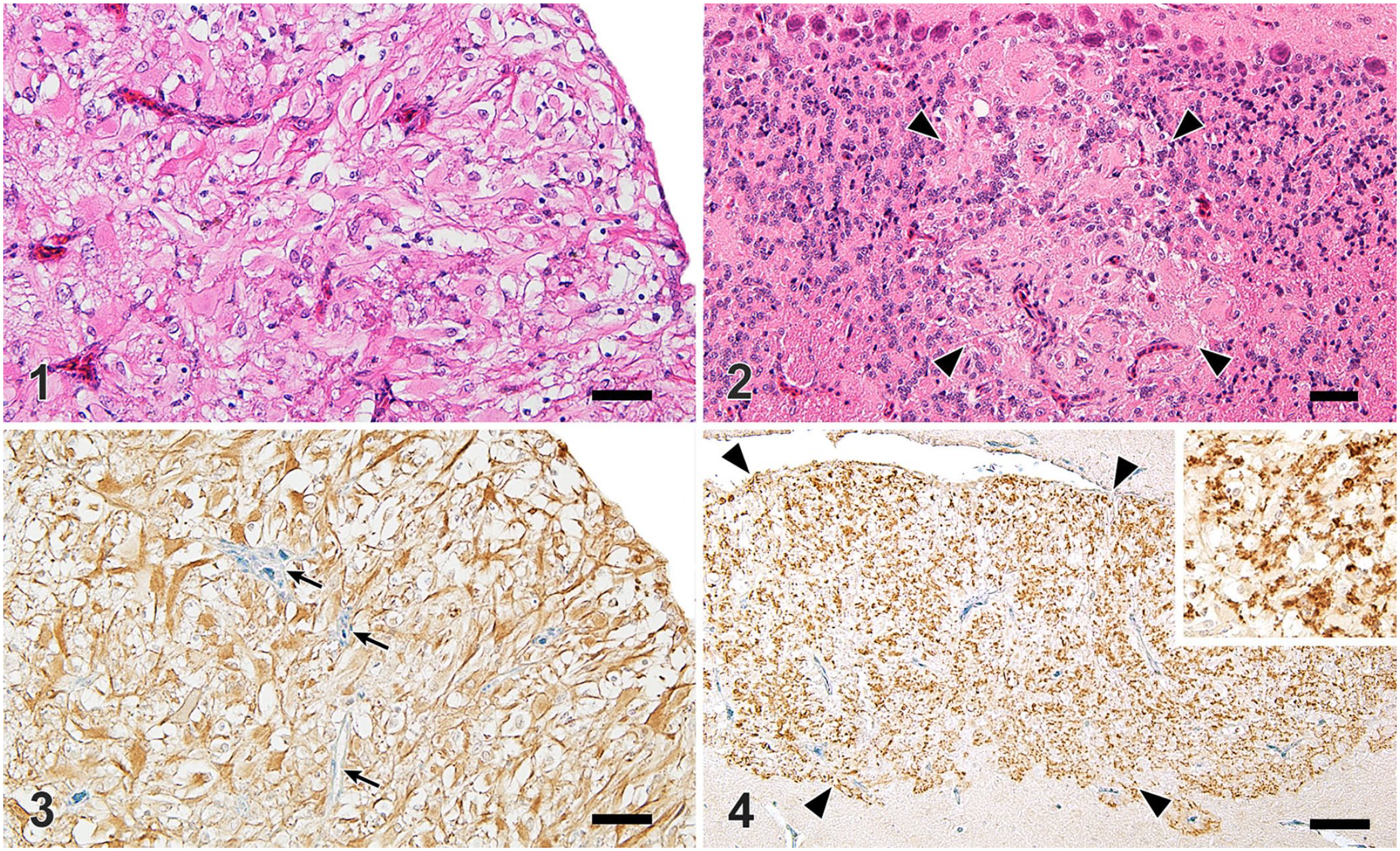
Fowl glioma in fowl glioma-inducing virus–infected Japanese bantam chickens.
Cardiomyocytes with intracytoplasmic matrix inclusion bodies were observed in 7 of 22 birds, of which birds 11–14 had multifocal lymphocytic myocarditis (Fig. 5) and myofiber disarray with cytoplasmic vacuolation in the left ventricular wall (Fig. 6). Occasionally, hypertrophic and multinucleate cardiomyocytes were observed that had numerous large intracytoplasmic matrix inclusion bodies (Fig. 7). These inclusions were immunoreactive for ALV antigens (Fig. 8). Significant lesions were not evident in any organ in any of the other examined chickens.
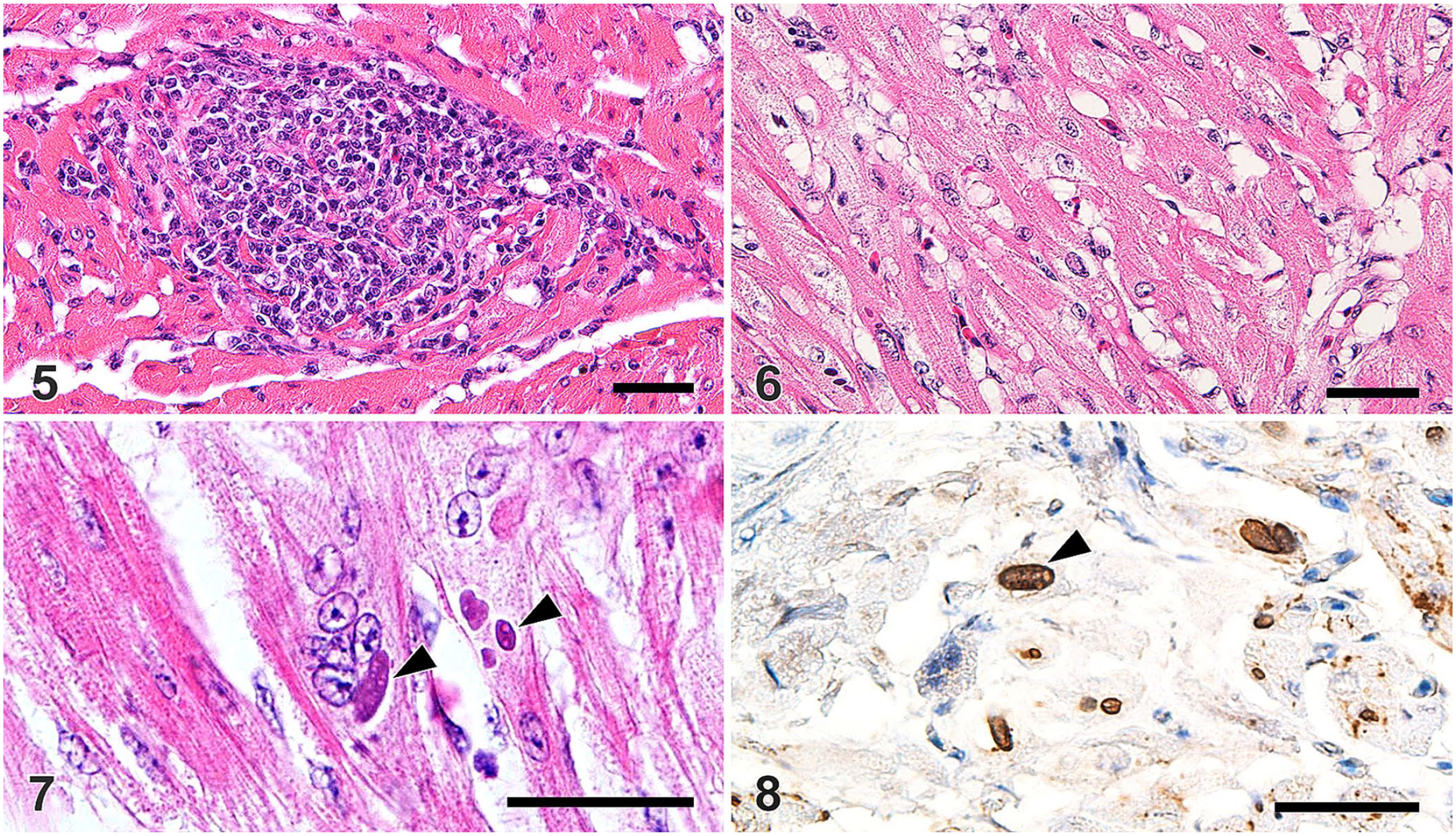
Myocardial abnormalities in fowl glioma-inducing virus–infected Japanese bantam chickens.
Virus isolation and detection and molecular analysis
ALVs were detected in 10 of 11 examined chickens by ALV-specific PCR assay (Table 1). Three FGV strains, KmN_93, 97, and 102, were isolated from glioma-affected chickens 4, 10, and 22. However, we were unable to isolate a single ALV strain from the 4 birds (11–14) with cardiac lesions and matrix inclusion bodies or from 3 chickens (2, 3, 6) with matrix inclusion bodies because multiple ALVs coexisted in these samples. The nucleotide identity of envSU among the 3 isolates was 99.5–100.0%. These regions shared > 99.5% nucleotide similarity with those of Km_6249 (AB764104), which is grouped in the Km/Oki_cluster and is considered to originate from avian endogenous virus ev-1 (AY013303). Overlapping sequence peaks were observed in chickens in which several viruses coexisted, and it was not possible to identify the dominant strains.
Evaluation of reference genes
We evaluated 7 reference genes (ACTB, HMBS, TBP, HPRT1, PGK2, ACTG1, YWHAZ). PCR efficiency was 91.8–97.3%, and R2 values were 0.999–1.000 (Table 2). Using the standard curve, we quantified the copy numbers of these reference genes in 100 ng of DNA extracted from DF-1 cells as well as from the hearts and brains of chicken embryos and adult chickens. No significant differences were observed in the copy numbers of the 7 genes (Fig. 9). Given that ACTB and HMBS have been identified as single-copy genes, we initially selected them as reference genes. Additionally, we used TBP as one of the biologic replicates because the gene set (ACTB, HMBS, TBP) successfully reflects the average of the 7 genes. Based on the results obtained, the number of nuclei was defined: DF-1 cells 2.50 × 103 nuclei/100 ng of DNA, embryonic hearts 1.65 × 104 nuclei/100 ng of DNA, embryonic brains 1.42 × 104 nuclei/100 ng of DNA, adult chicken hearts 6.92 × 103 nuclei/100 ng of DNA, and adult chicken brains 3.20 × 103 nuclei/100 ng of DNA. In contrast, by referring only to the sample of adult chickens (brain and heart), HPRT1 was the closest to the mean of the 7 genes.
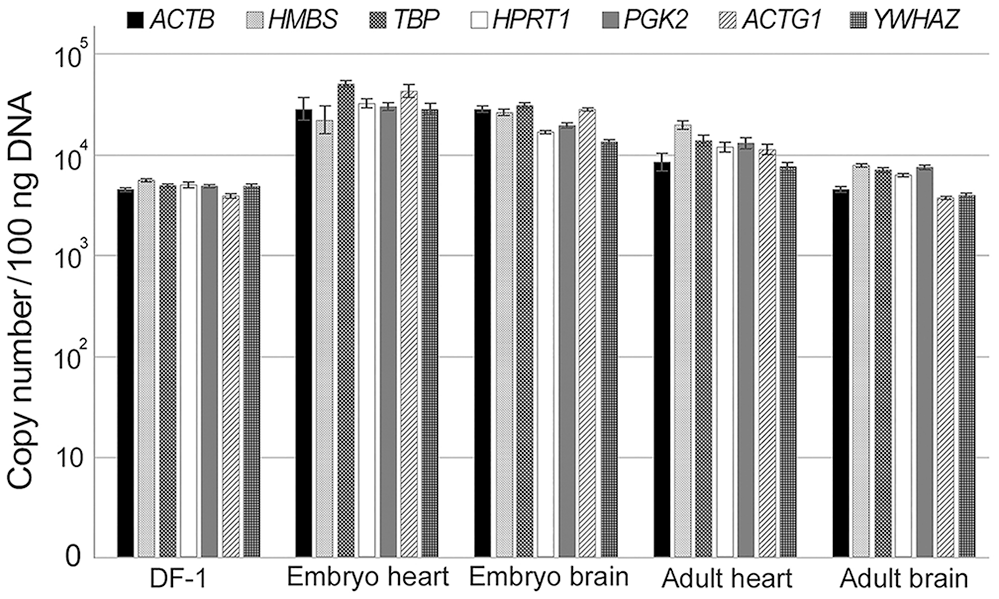
Comparison of copy numbers of 7 candidate reference genes in 100 ng of DNA extracted from DF-1 cells, hearts, and brains of chicken embryos and adult chickens. Data are expressed as the x̄ ± SEM (n = 3 per group). Logarithmic scaling was used for the y-axis.
Quantification of the proviral copy number per nucleus
The detection limit of provirus was 18.1 copies/100 ng of DNA. In the brains and hearts of 9 ALV-positive Japanese bantam chickens (2–4, 6, 11–14, 22) examined, the numbers of nuclei per 100 ng of DNA in the heart and brain samples were 5.74 ± 0.55 × 103 nuclei and 8.95 ± 2.63 × 103 nuclei, respectively (Table 1; Fig. 10). In the 4 chickens with cardiac lesions (11–14), the proviral load was 10.3 ± 2.7 copies/nucleus in the heart and 6.5 ± 1.0 copies/nucleus in the brain. In contrast, the 5 chickens without cardiac lesions (2–4, 6, 22) had 0.4 ± 0.4 proviral copies/nucleus in the heart and 0.8 ± 0.8 copies/nucleus in the brain.
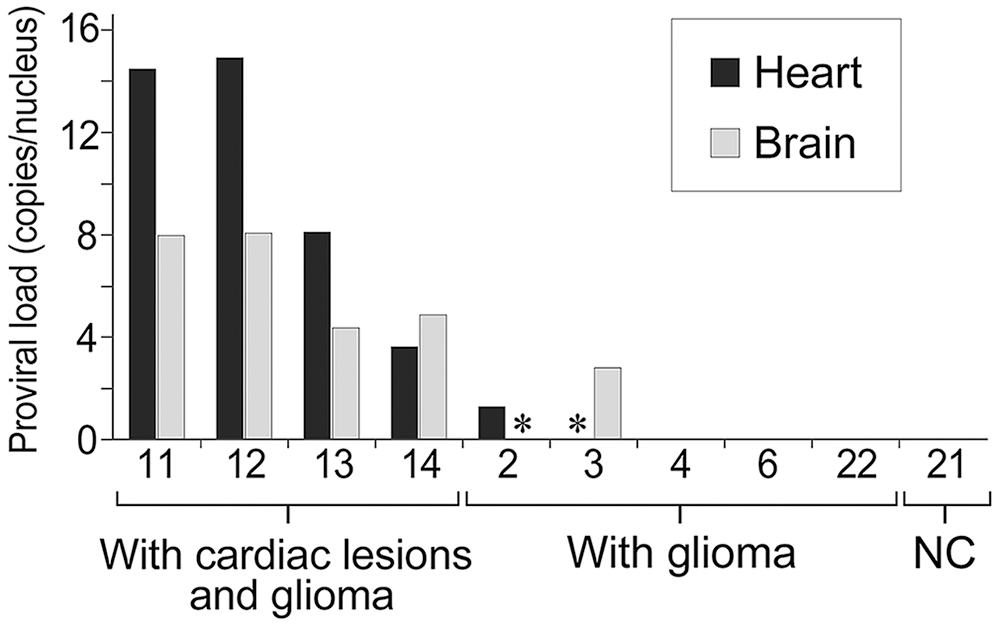
Comparison of the proviral load in 100 ng of DNA extracted from brains and hearts of 9 avian leukosis virus (ALV)-positive Japanese bantam chickens. ALV-negative Japanese bantam 21 was used as a negative control (NC). The proviral load was calculated by dividing the proviral copy number by the number of nuclei calculated from the average value of ACTB, HMBS, and TBP. The hearts of chickens 11–14 with cardiac lesions and glioma had a higher proviral load than those of chickens 2–4, 6, and 22 with glioma but without cardiac lesions. * Not done.
Discussion
We established a qPCR assay to quantify ALV proviral loads and compared proviral loads in the brains and hearts of FGV-infected Japanese bantam chickens with glioma and cardiac lesions. The myocardial lesions that we observed were characterized by atypical hypertrophic myocardial fibers with myofiber disarray. Our findings suggest that the disease involves either neoplastic changes, such as rhabdomyoma, or reactive changes caused by myocarditis. However, we have no previous experience with such atypical myocardium in myocarditis of chickens; we consider this lesion to be a neoplastic change resulting from ALV infection.
Reovirus and avian encephalomyelitis virus (Tremovirus A) can cause myocarditis in poultry. 1 Although these viruses cannot be ruled out without additional examinations, there are no reports of atypia and polymorphism of myocardial fibers caused by these viral infections. Moreover, no outbreaks of these viral infections have been confirmed in the past 10 y of examining the chicken flock in our study, suggesting that ALV is the cause of the current outbreak.
Because of the coinfection by several ALV strains, we could isolate only a few strains in our study. In addition, single viruses tended to be isolated only from chickens without cardiac lesions. The development of molecular clones of each coinfected ALV strain will be needed to clarify this detail. In general, coinfection among different subgroups, such as subgroups A, J, and K, has been reported. 19 However, viruses in the same subgroup are not thought to establish coinfection because of viral interference. Therefore, it is possible that the coinfecting viruses belong to different subgroups or that cells infected by each virus are mosaic-like in the host.
Although there was no significant difference in the copy number of each reference gene, there was some variation; therefore, we recommend that at least 3 reference genes be quantified to define the accurate number of nuclei. Further to examining the heart and brain samples, ACTB, HMBS, and TBP genes were selected in our study. In addition, when analyzing adult chicken brain and heart samples, we advocate the use of the HPRT1 gene, which had the smallest average value among the 7 genes, as a reference gene for more efficient analysis. Our study indicates that proviral loads are associated with the development of myocardial lesions in chickens with proviral loads exceeding 3.73. In the case of HTLV-1, which belongs to the same retrovirus family, it has been reported that high provirus levels are associated with a higher risk of developing HTLV-1–related diseases. 32 In the case of HTLV-1, increased proviral loads result mainly from the Tax-driven activation 41 ; however, ALV lacks a viral regulatory protein such as Tax. In contrast, there was no association between brain lesions and provirus levels, suggesting that the pathogenesis differs between myocardial abnormalities and gliomas.
Footnotes
Acknowledgements
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Our work was supported by the Japan Society for the Promotion of Science KAKENHI grants JP19K22356, JP21H02354.
