Abstract
Herpesviruses can be significant reptile pathogens. Herpesviral infection in a wild-caught, male spider tortoise (Pyxis arachnoides) under human care was detected during a routine wellness examination prior to transition between zoologic organizations. The tortoise had no clinical signs of illness. Oral swabs obtained during a physical examination as part of pre-shipment risk mitigation for infectious disease were submitted for consensus herpesvirus PCR assay and sequencing. Based on comparative sequence analysis, the novel herpesvirus identified is a member of the subfamily Alphaherpesvirinae. Studies of herpesviral phylogeny in chelonian species support branching patterns of turtle herpesviruses that closely mirror those of their hosts. The symmetry of these patterns is suggestive of close codivergence of turtle herpesviruses with their host species. The distribution of these viruses in both tortoises and emydids suggests a phylogenetic duplication event in the herpesviruses after host divergence of the Pleurodira and basal to the divergence of Americhelydia. Herpesviral infections have been documented to cause higher morbidity when introduced to aberrant host species, and significant consideration must be given to the presence of herpesviruses in the management of tortoise collections, particularly collections that include various species of testudines.
The Madagascar spider tortoise (Pyxis arachnoides) is a small tortoise species endemic to the coastal Mikea forests of southwest Madagascar. Listed by the International Union for Conservation of Nature as critically endangered, Madagascar spider tortoises are 1 of 4 extant endemic terrestrial chelonian species of Madagascar. Cheloniologists describe current threats to survival including habitat destruction, predation by humans, and collection for the exotic pet trade.29,30 Detection of a picornavirus from the lung of a spider tortoise has been reported, but otherwise, we found no reports regarding viruses in this species. 20
Herpesviruses are large, enveloped DNA viruses. 8 The family Orthoherpesviridae is divided into subfamilies Alphaherpesvirinae, Betaherpesvirinae, and Gammaherpesvirinae. All herpesviruses known to infect reptiles are in the Alphaherpesvirinae. 19 Phylogenetic study of herpesviruses suggests that each herpesvirus is closely associated with, and highly adapted to, a single host species.4,7,9 This suggests that the viruses have often coevolved with their hosts over long periods of time. Coevolution is supported by the corresponding branching patterns of mammalian herpesviruses and their hosts, although it is less well-studied in reptiles.4,18,21 A study of broader comparative host–herpesvirus evolution found that co-speciation events were less common in Alphaherpesvirinae than in Betaherpesvirinae or Gammaherpesvirinae. 6 However, the available datasets were biased, with 14 alphaherpesviruses from sauropsids (11 from birds, 3 from turtles [2 of which were the same virus species], and none from squamates or crocodilians), and 31 from mammals. Despite this concern, there were 6 co-speciation events more than a million years old found in mammal hosts and 5 in sauropsids. Broader examination of sauropsid herpesviruses suggests that the jump of alphaherpesviruses into mammals is fairly recent, and that mammalian alphaherpesviruses do not represent alphaherpesviral diversity well.9,21 Further knowledge of sauropsid alphaherpesviral diversity is needed to understand alphaherpesviral evolution.
Whole-genome analyses have confirmed turtles as a sister group to the archosaurs, which includes crocodilians and birds.22,25 Within the order Testudines, turtles are divided into 2 suborders: Cryptodira and Pleurodira, which differ in the way the animals retract their neck. Within the suborder Cryptodira, there is further division into a clade consisting of hardshell cryptodires, the Durocryptodira, and a clade for softshell turtles, the Trionychia. Durocryptodira is of particular interest in our report because the division of clades is mirrored in the phylogenetic analysis of herpesviruses that infect these hosts. Within the Durocryptodira, the next division is between the Americhelydia (sea turtles, snapping turtles, mud/musk turtles) and Testudinoidea (tortoises, New- and Old-World pond turtles). Testudinoidea is further subdivided into the families Emydidae (New World pond turtles), Geoemydidae (Eurasian pond turtles, Neotropical wood turtles), Platysternidae (big-headed turtle), and Testudinidae (tortoises). Understanding these relationships is needed to clarify the close association between virus and host codivergence in herpesviruses.
Among herpesviruses of non-avian reptiles, chelonian herpesviruses are the best characterized to date. 19 Significant morbidity is primarily observed in immunocompromised or young chelonians in hosts in which the virus is endemic, or as the result of infection in an aberrant host species. 19 Herpesviruses cause some of the most significant chelonian viral diseases, and reports of illness and death are common.14,16,19,26
The spider tortoise in our study was donated by a private collector to a zoologic organization in 2004 and was reported as wild-caught. This tortoise produced a male offspring (housed independently at the same institution) that was also sampled. When examined for a pre-shipment examination, both animals were clinically normal, with no evidence of oronasal discharge, rhinitis, or stomatitis. Oral cavity swabs were obtained using cotton-tipped applicators and shipped in a chilled container for PCR screening at the University of Florida (Gainesville, FL, USA). DNA was extracted from the samples (DNeasy kit; Qiagen). Samples were tested for adenovirus and intranuclear coccidiosis using described methods,2,31 with negative results. Nested PCR amplification of a partial sequence of the herpesviral DNA–dependent DNA polymerase gene was also performed using methods described previously (round 1: forward DFA 5′-GAYTTYGCNAGYYTNTAYCC-3′, forward ILK 5′-TCCTGGACAAGCAGCARNYSGCNMTNAA-3′, reverse KG1 5′-GTCTTGCTCACCAGNTCNACNCCYTT-3′; round 2: forward TGV 5′-TGTAACTCGGTGTAYGGNTTYACNGGNGT-3′, reverse IYG 5′-CACAGAGTCCGTRTCNCCRTADAT-3′). 28 The index case was herpesvirus positive, but the male offspring tested negative.
To obtain additional sequence for phylogenetic comparison, the second round of amplification was altered to use primers DFA and IYG. 28 Direct sequencing was performed (Applied Biosystems BigDye terminator kit; Thermo Fisher). Amplification yielded a 481-bp product after primer sequences were removed. The selected sequence was translated and compared to those within GenBank using BLASTX 1 ; the highest homology was with Emydoidea blandingii herpesvirus 2 (HV-2), with 78% amino acid identity. The virus that we detected is hereafter referred to as Pyxis alphaherpesvirus 1 (PyAHV-1). The sequence was submitted to GenBank (ON816017). The genetic distinction that can be observed between PyAHV-1 and other characterized herpesviruses is consistent with the placement of this virus as a novel species.
Homologous herpesviral partial polymerase amino acid sequences were downloaded from GenBank for comparison. Elephantid betaherpesvirus 1 (GenBank AF322977; Proboscivirus elephantidbeta1) was used as the outgroup. Bayesian analyses of each alignment were performed using MrBayes v.3.2.7a on the CIPRES server with mixed amino acid substitutions, gamma-distributed rate variation and a proportion of invariable sites, 4 chains, and unconstrained branch lengths. 24 The first 25% of 2 million iterations were discarded as a burn-in. The phylogenetic tree was visualized using FigTree (http://tree.bio.ed.ac.uk/software/figtree/; Fig. 1). The resulting tree was then compared with a phylogenetic tree for corresponding host species from TimeTree. 15 A comparative tanglegram was constructed using Dendroscope 3 (https://github.com/husonlab/dendroscope3) to visualize the similarities in the branching patterns of Testudines and their herpesviruses (Fig. 2). 11
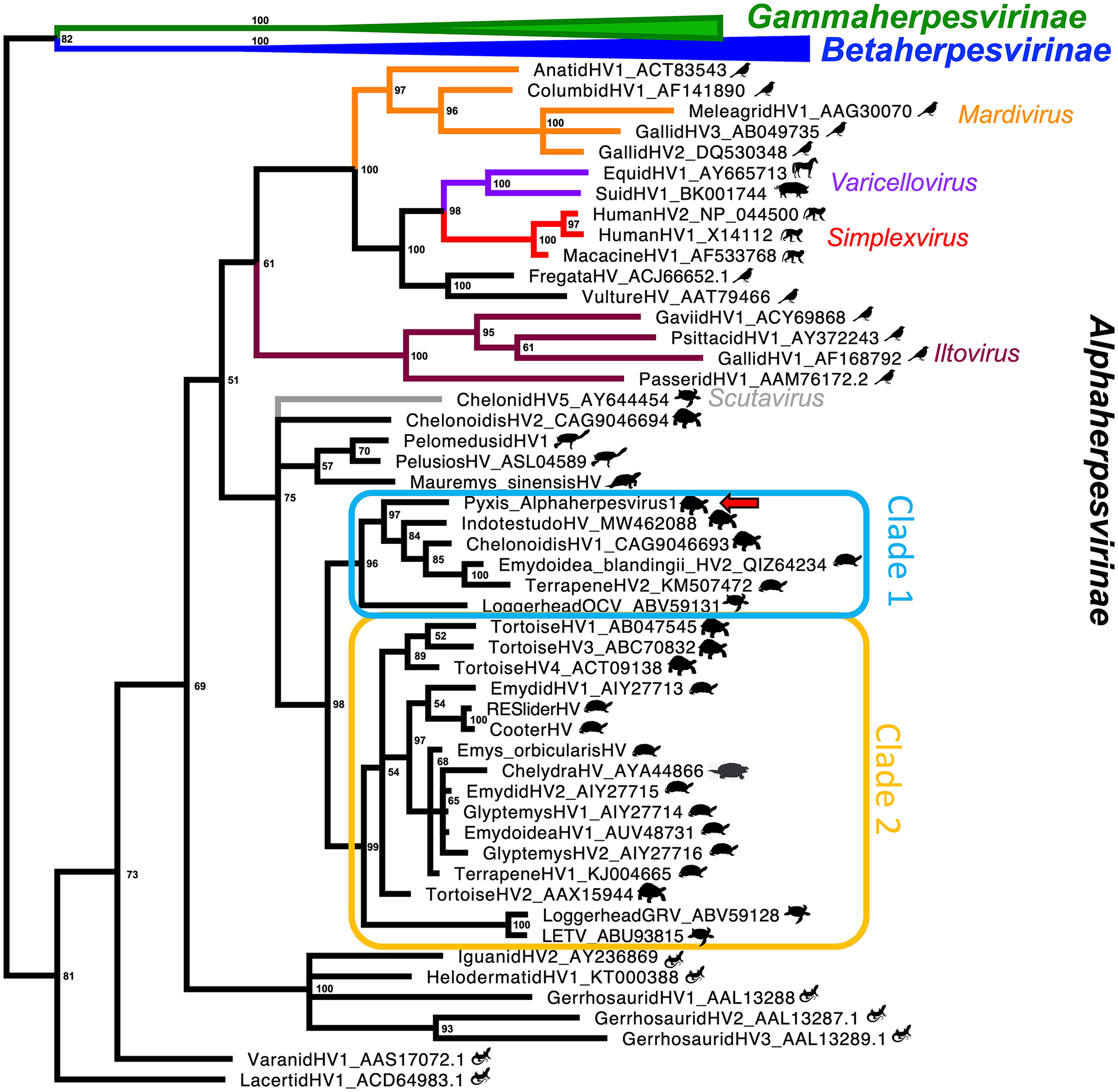
Bayesian phylogenetic tree based on DNA polymerase amino acid sequences of Orthoherpesviridae (posterior probability values at each node). Subfamilies and genera are indicated in different colors, and silhouettes of host families are given next to viruses. Clade 1 is outlined in blue and clade 2 in yellow. The red arrow indicates Pyxis alphaherpesvirus 1.
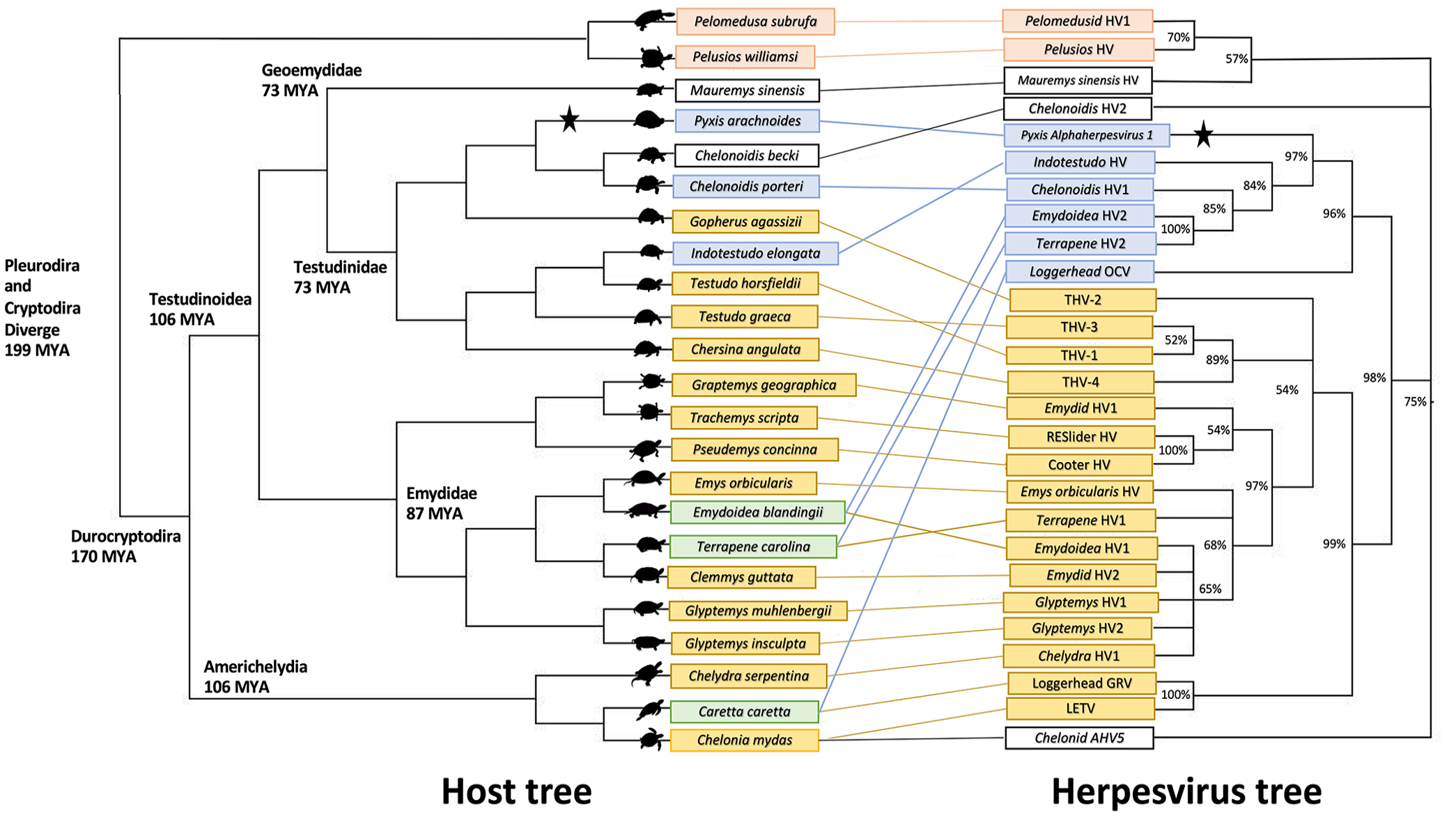
Tanglegram correlating cladogram of host species from the order Testudines with cladogram of Orthoherpesviridae DNA polymerase amino acid sequences (posterior probability values at nodes). The pink clade represents herpesviruses of the pleurodire “side-necked” turtles. The blue clade (clade 1) illustrates Pyxis alphaherpesvirus 1 and its closest viral relatives, along with corresponding hosts. The yellow clade (clade 2) shows the larger clade of herpesviruses infecting emydids, sea turtles, and tortoises, among others. Host species in green have > 1 corresponding herpesvirus. In the absence of host data, the host species of pelomedusid HV-1 (Pelusios castaneus) was substituted for another Pelomedusidae species, Pelomedusa subrufa. Cladogram branch lengths do not indicate distance.
Within the phylogenetic tree for turtle herpesviruses, the viruses that infect the suborder Pleurodira (side-necked turtles) branch basally. This mirrors the divergence of the host order Testudines into the suborders Cryptodira and Pleurodira. 22 Within Cryptodira, the hardshell turtles (Durocryptodira) divide into Americhelydia (which includes snapping turtles and sea turtles) and Testudinoidea (which includes emydids and tortoises), and there are 2 distinct clades of herpesviruses that each largely reflect host patterns. PyAHV-1 clusters with a clade containing Indotestudo HV, Chelonoidis HV-1, Emydoidea blandingii HV-2, and Terrapene HV-2 with a Bayesian posterior probability of 97% (Fig. 1). This clade of herpesviruses (called clade 1) infects hosts in Testudinoidea. Loggerhead orocutaneous virus, from a host in the Americhelydia, is basal to this group, clustering with a posterior probability of 96% (Figs. 1, 2). Viruses using hosts in the Emydidae cluster with a posterior probability of 100%. The branching order of viruses using hosts in Testudinidae does not mirror host divergence patterns as well; the confidence values of these groupings are less-well supported (posterior probability 84% and 85%). A similar pattern is seen in the other clade utilizing hosts in the Durocryptodira, shown in the tanglegram in yellow, and called here clade 2 (Figs. 1, 2). Viruses utilizing hosts in the Americhelydia cluster (posterior probability 100%) apart from Chelydra HV-1. A clade uses hosts in the Emydidae as well as Chelydra HV-1 (posterior probability 97%) and a clade using tortoise hosts contains tortoise HV-1, HV-3, and HV-4 (posterior probability 89%), with no clarity as to where tortoise HV-2 clusters. The phylogeny of clades 1 and 2 within Durocryptodira is suggestive of a duplication event in which 2 viral lineages emerged. This duplication would have occurred after the separation of Pleurodira and Cryptodira 199 million years ago (mya) and prior to the divergence of Testudinoidea and Americhelydia that occurred ~106 mya.22,25
Several viruses do not fit the expected divergence pattern. In clade 1, the herpesviruses of the Testudinidae are not found to be monophyletic; support values are low in this area, and this may be a spurious finding or may represent host-jumping events. In clade 2, Chelydra HV-1, from a host in the Americhelydia, clusters with a clade of herpesviruses of emydid hosts, which is well supported with a 97% posterior probability; this likely represents a host-jumping event. Outside of clades 1 and 2, chelonid alphaherpesvirus 5 (Scutavirus chelonidalpha5), in the genus Scutavirus, and Chelonoidis HV-2 may be host jumps or distinct lineages with earlier divergence from clades 1 and 2, which are well supported to be monophyletic with 98% posterior probability. Mauremys sinensis HV, from a host in the Geoemydidae, was found to have weak support for clustering with the pleurodire herpesviruses. If correct, this may represent a host-jumping event. Alternatively, this finding may be spurious and this virus may be a member of a distinct non-clade 1/2 lineage. Longer sequences and further taxon sampling, especially looking at Trionychia, Geoemydidae, and clade 1, are expected to help resolve these discrepancies. The phylogenetic distances between clades 1 and 2 are consistent with genus-level differences. There will likely be several genera present within the herpesviruses that infect testudinid hosts.
Although herpesviruses are typically host-specific, there are examples of alphaherpesviruses causing morbidity and mortality in related aberrant host species.14,23,26 A group of zoologically housed western pond turtles (Actinemys marmorata) that died acutely were infected with emydid HV-1; similarly infected red-eared slider turtles (Trachemys scripta) in the enclosure remained subclinical. 26 Tortoise HV-4 was first found in bowsprit tortoises (Chersina angulata) during quarantine examination in a zoologic institution. 5 This virus was later associated with mortality in leopard tortoises (Stigmochelys pardalis) in a mixed-species collection. 14 Mixed-species exhibits are of concern for cross-species infection.
Our patient’s wild-caught status suggests that this virus may be endemic in wild populations of spider tortoises in Madagascar. Herpesviruses have been detected in wild chelonians in the absence of clinical signs.3,13 Although the tortoise in our case had no signs of illness, the capacity of PyAHV-1 to cause clinical disease remains unknown. The relationship between external stressors and reactivation of latent herpesviral infection is well-established in mammals, including hyperthermia, hypothermia, fatigue, and immunosuppression.8,10 Given the shrinking habitat and steadily increasing temperature in southern Madagascar, anthropogenic stressors may render PyAHV-1 more dangerous.27,29,30 More than 60% of turtle species are either threatened or extinct, and anthropogenic influences can have catastrophic impacts. 17 Our finding underscores the importance of infectious disease screening to mitigate risk when transitioning animals between facilities. Pre-shipment testing supports animal welfare by ensuring that the animals are healthy and able to undergo transport and change of habitat, and also reduces the risk of introducing pathogens to recipient institutions.5,12,16
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
