Abstract
Herpesviruses are found in free-living and captive chelonian populations, often in association with morbidity and mortality. To date, all known chelonian herpesviruses fall within the subfamily Alphaherpesvirinae. We detected a novel herpesvirus in 3 species of chelonians: a captive leopard tortoise (Stigmochelys pardalis) in western TX, USA; a steppe tortoise (Testudo [Agrionemys] horsfieldii) found near Fort Irwin, CA, USA; and 2 free-living, three-toed box turtles (Terrapene mexicana triunguis) found in Forest Park, St. Louis, MO. The leopard tortoise was coinfected with the tortoise intranuclear coccidian and had clinical signs of upper respiratory tract disease. The steppe tortoise had mucopurulent nasal discharge and lethargy. One of the three-toed box turtles had no clinical signs; the other was found dead with signs of trauma after being observed with blepharedema, tympanic membrane swelling, cervical edema, and other clinical signs several weeks prior to death. Generally, the branching order of the turtle herpesviruses mirrors the divergence patterns of their hosts, consistent with codivergence. Based on phylogenetic analysis, this novel herpesvirus clusters with a clade of viruses that infect emydid hosts and is likely of box turtle origin. Therefore, we suggest the name terrapene alphaherpesvirus 3 (TerAHV3) for the novel virus. This virus also has the ability to host-jump to tortoises, and previously documented herpesviral morbidity tends to be more common in aberrant hosts. The relationship between clinical signs and infection with TerAHV3 in these animals is unclear, and further investigation is merited.
Herpesviruses (Herpesvirales, Herpesviridae) are large, enveloped viruses with double-stranded DNA genomes.6,22 As viruses with intranuclear replication, herpesviruses usually display high host fidelity and appear to have often codiverged with the evolution of their host species. 31 Phylogenetic analyses suggest herpesviruses may have existed for >400 million y, prior to the reptilian–mammalian divergence. 24 All known reptilian herpesviruses are in the subfamily Alphaherpesvirinae and have been detected in squamates, chelonians, and crocodilians.23,25 More herpesviruses have been detected in the order Testudines than from any other order within the class Reptilia, and documented hosts include chelonians from the families Cheloniidae, Testudinidae, Emydidae, Chelydridae, and Pelomedusidae.22,23,25,36 The only official phylogenetic classification of chelonian herpesviruses is the genus Scutavirus, which includes Chelonid alphaherpesvirus 5 (ChAHV5; informally known as fibropapilloma-associated herpesvirus) and, more recently, Testudinid alphaherpesvirus 3 (TeHV3). 41 Additional phylogenetic information is needed to better understand the evolutionary relationships of these viruses.
In chelonians, signs associated with herpesviral infections have ranged from subclinical to severe, including cutaneous lesions such as fibropapillomas, ocular disease, and upper respiratory tract disease.18,22,26,35,44 These viruses have clinical and conservation significance. Chelonid alphaherpesvirus 6 (ChAHV6; informally known as lung-eye-trachea disease-associated herpesvirus) was associated with mortality as high as 38% in captive green sea turtles (Chelonia mydas). 17 High morbidity and mortality were also observed in captive Hermann tortoises (Testudo hermanni) and steppe tortoises (syn. Horsfield tortoise; Testudo [Agrionemys] horsfieldii) with TeHV3. 22 Although terrapene herpesvirus 1 (TerHV1) has been associated with severe necrotizing stomatitis and esophagitis in a rehabilitation center’s hatchling eastern box turtle (Terrapene carolina carolina), a survey of free-living eastern box turtles found no association of clinical signs with virus status.19,35 These remarkably different clinical presentations indicate the complex relationships among viruses, their host species, coinfections, environment, phylogeny, and pathogenicity. We describe here a novel herpesvirus that was detected in 3 different species of chelonians, serving as the first step in understanding this pathogen and improving our understanding of other chelonian herpesviruses.
Materials and methods
Animals
Animal 1: leopard tortoise
In 2013, a captive, adult, female leopard tortoise (Stigmochelys pardalis) in western Texas, USA began to display clinical signs (Table 1; Fig. 1). The animal was at least 30-y-old and was housed with other chelonians including ornate box turtles (Terrapene ornata). Despite an extended course of treatment for yolk coelomitis diagnosed via laparoscopic examination, the animal died, and postmortem examination was performed at Northwest ZooPath (Monroe, WA, USA). The tongue was sampled for PCR testing (Table 2).
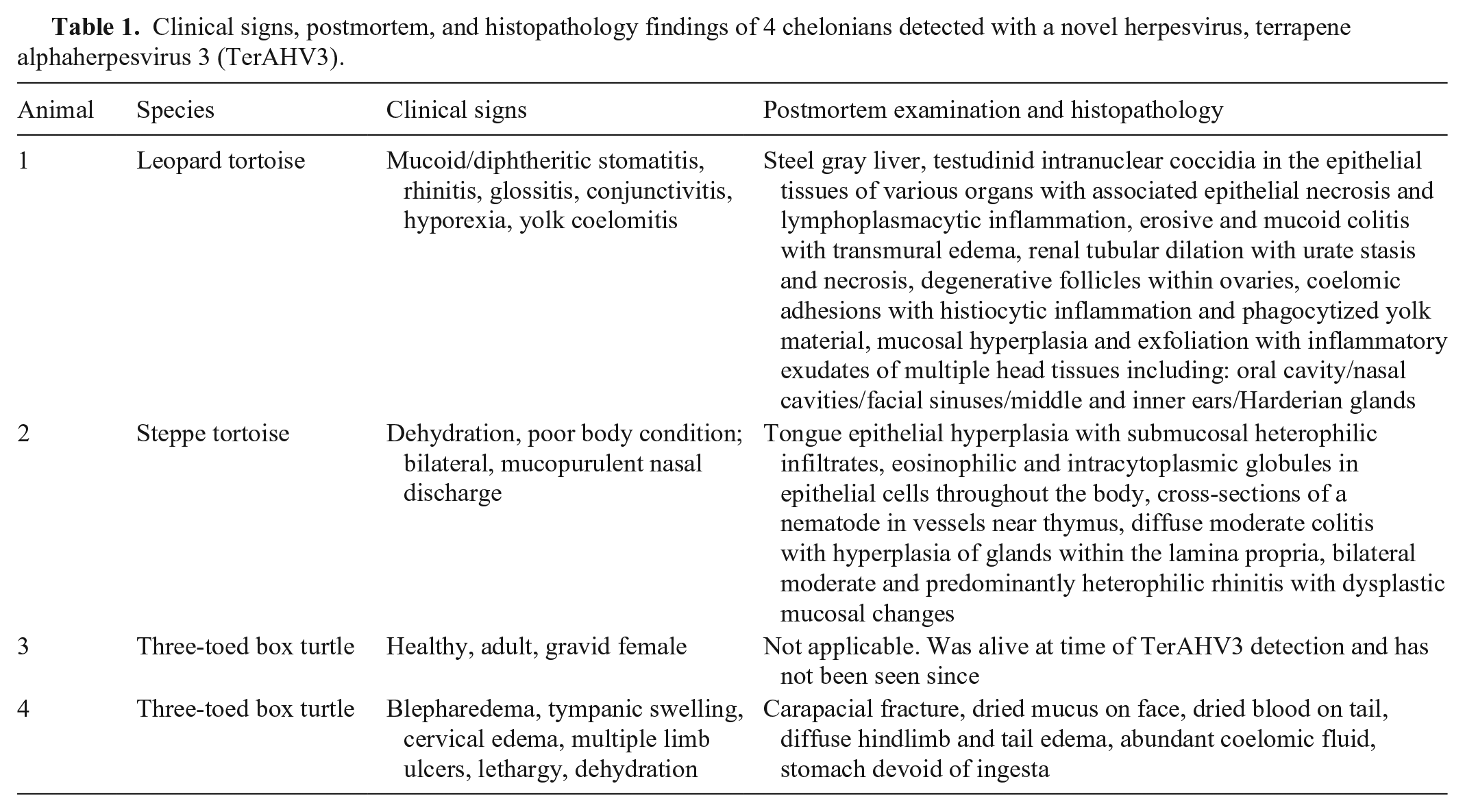
Clinical signs, postmortem, and histopathology findings of 4 chelonians detected with a novel herpesvirus, terrapene alphaherpesvirus 3 (TerAHV3).

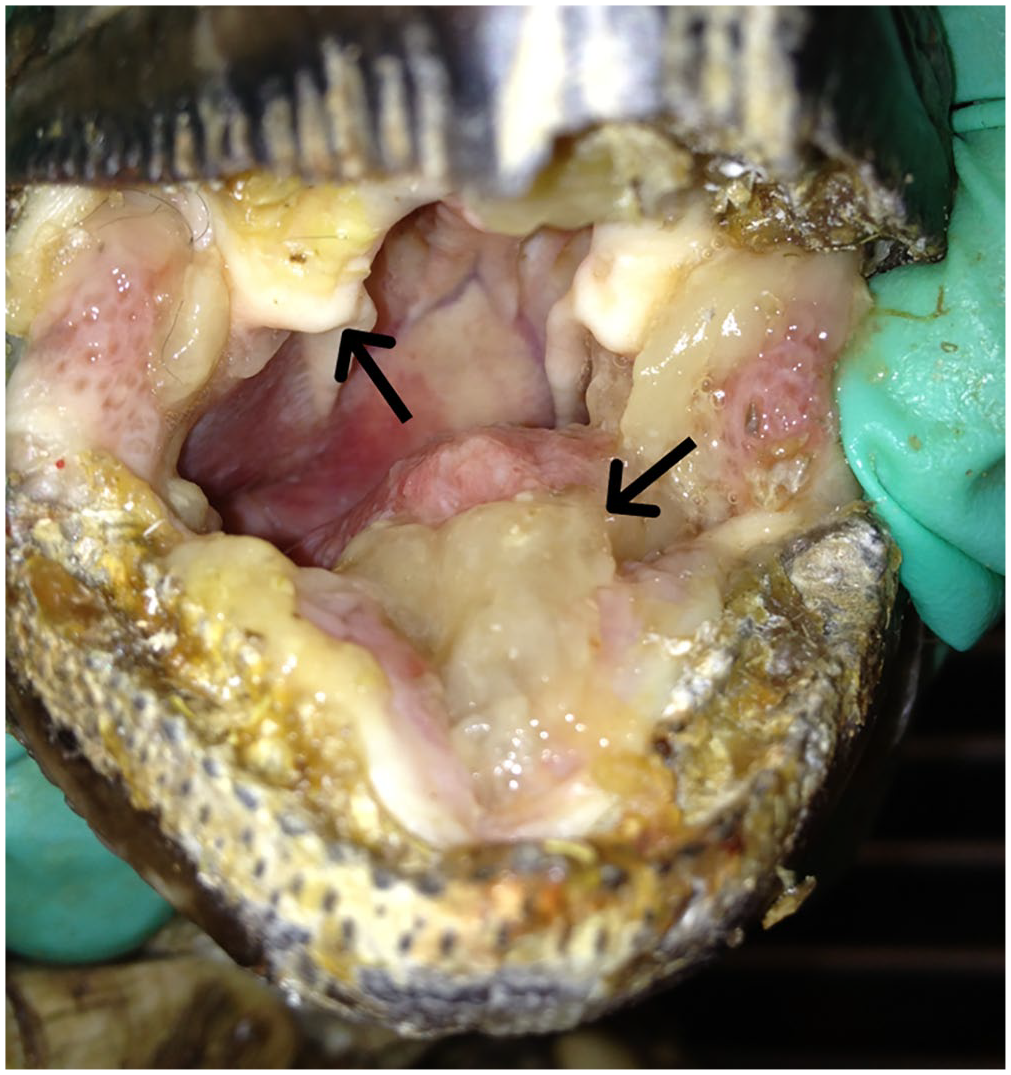
Oral cavity of the leopard tortoise (Stigmochelys pardalis; animal 1) with severe mucosal hyperplasia and exudate identified by arrows. The figure is oriented with the dorsal beak at the top and the ventral beak at the bottom.
Detection samples, methods, and PCR products of a novel herpesvirus, terrapene alphaherpesvirus 3 (TerAHV3), in 4 chelonians.
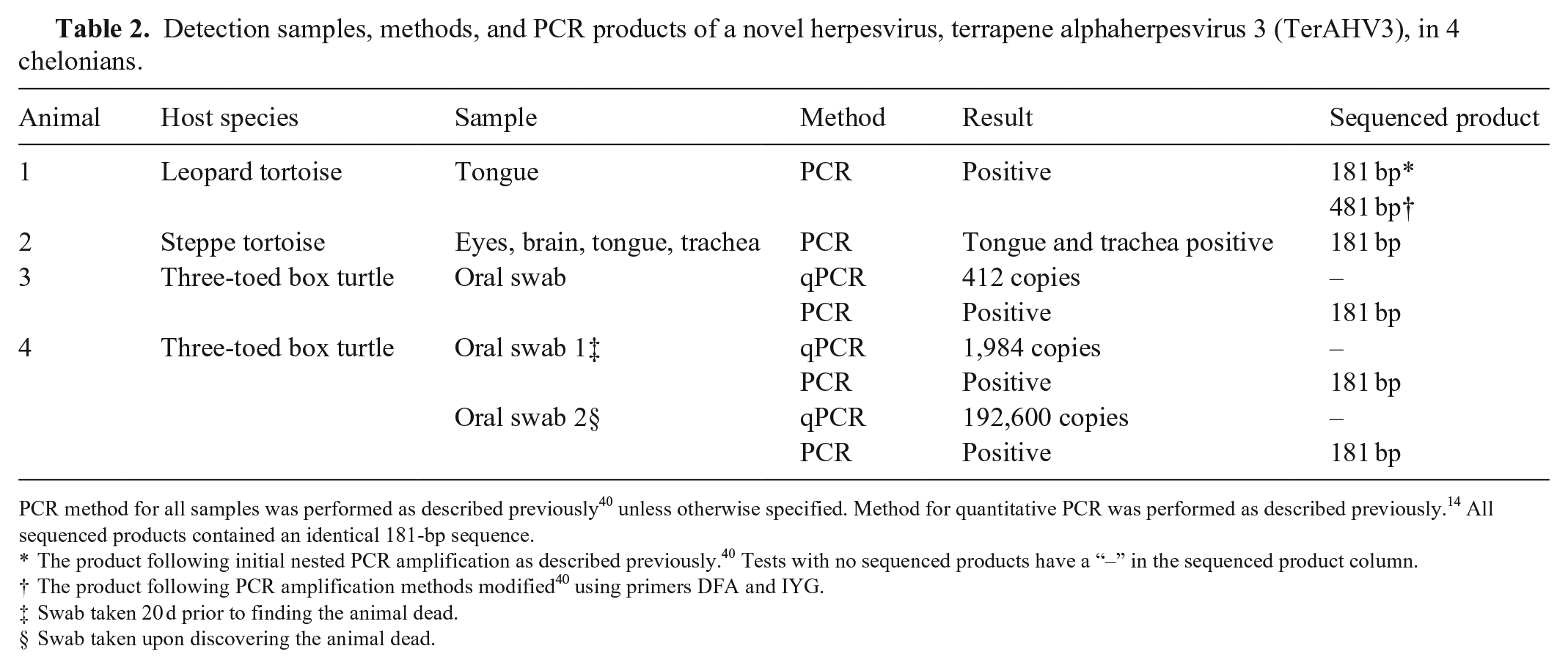
PCR method for all samples was performed as described previously 40 unless otherwise specified. Method for quantitative PCR was performed as described previously. 14 All sequenced products contained an identical 181-bp sequence.
The product following initial nested PCR amplification as described previously. 40 Tests with no sequenced products have a “–” in the sequenced product column.
The product following PCR amplification methods modified 40 using primers DFA and IYG.
Swab taken 20 d prior to finding the animal dead.
Swab taken upon discovering the animal dead.
Animal 2: Steppe tortoise
In 2013, a non-native, free-living, adult female steppe tortoise was found in poor condition on the Fort Irwin Western Expansion Translocation Area in the Mojave Desert, San Bernardino County, CA, USA. It was sent to the University of Florida (UF; Gainesville, FL, USA) for clinical evaluation where it was euthanized following several laboratory tests (Table 3). A full postmortem examination was performed by the UF Veterinary Diagnostic Laboratory, and tongue and trachea samples were collected for pan-herpesviral PCR.
Co-pathogen testing samples, methods, and results for 4 chelonians in which a novel herpesvirus, terrapene alphaherpesvirus 3 (TerAHV3), was detected.
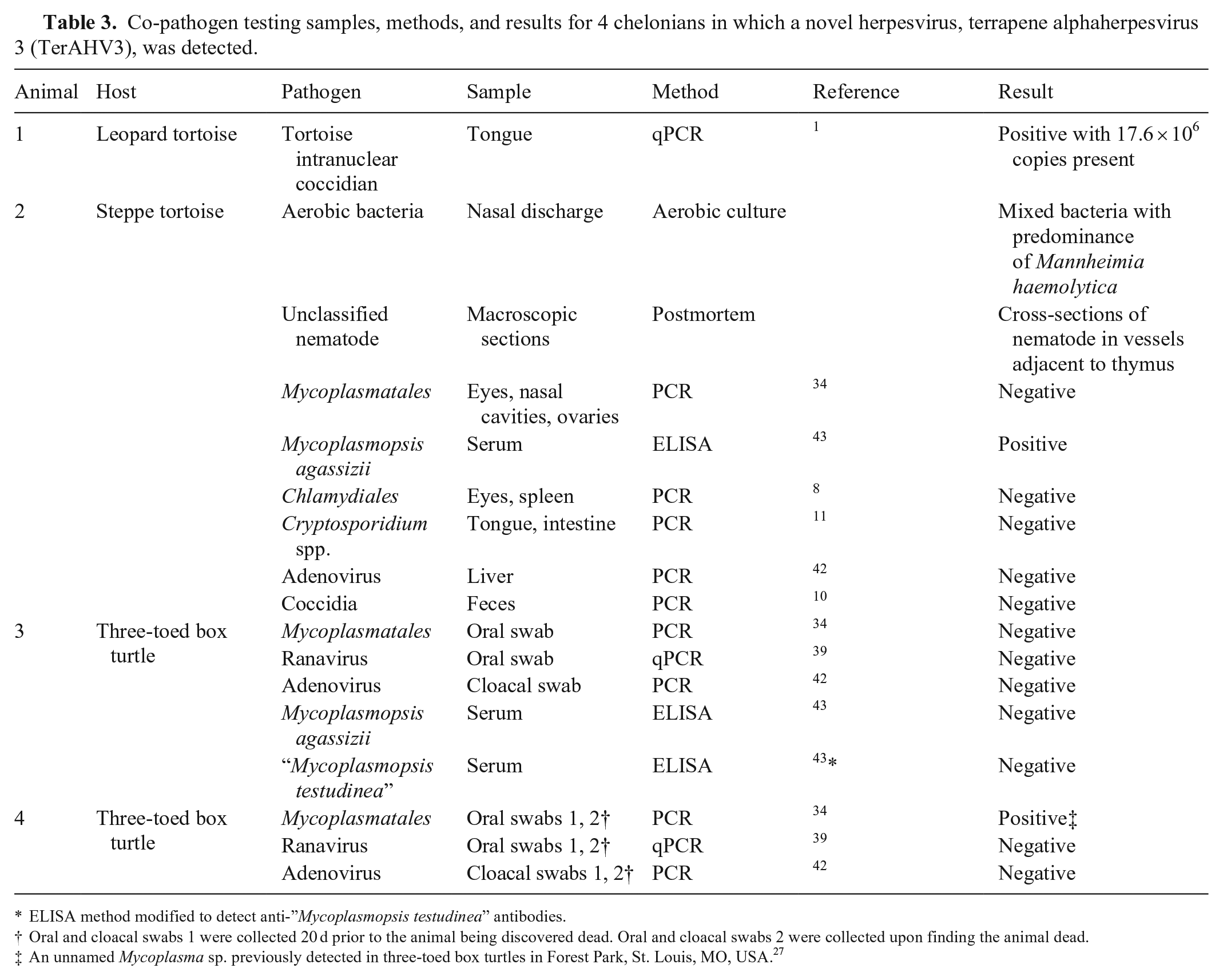
ELISA method modified to detect anti-”Mycoplasmopsis testudinea” antibodies.
Oral and cloacal swabs 1 were collected 20 d prior to the animal being discovered dead. Oral and cloacal swabs 2 were collected upon finding the animal dead.
An unnamed Mycoplasma sp. previously detected in three-toed box turtles in Forest Park, St. Louis, MO, USA. 27
Animals 3, 4: three-toed box turtles
Two free-living, adult female, three-toed box turtles (Terrapene mexicana triunguis) were captured multiple times using radiotelemetry or opportunistically in Forest Park, St. Louis, MO, USA as part of a long-term box turtle health survey. Oral and cloacal swabs for PCR pathogen screening for Mycoplasmatales, ranavirus, adenovirus, and herpesviruses (Table 3) were obtained using cotton-tipped applicators. The swabs were stored at −80°C until DNA extraction.
Box turtle 1 (BT1; animal 3) was captured and sampled twice, once in 2013 and once in 2016. This turtle has not been observed since its last capture in 2016.
Box turtle 2 (BT2; animal 4) was captured or sighted more than 150 times between 2013 and 2016. Serial health assessments and annual swab collection were performed. In 2016, 2 swab samples were collected within 2 wk of each other; BT2 was discovered dead upon the second sampling. A field postmortem examination was performed at the Saint Louis Zoo (St. Louis, MO, USA).
PCR assay and sequencing
DNA was extracted from samples (DNeasy kit; Qiagen). In all 4 animals, nested PCR amplification of a partial sequence of the herpesviral DNA-directed DNA polymerase gene (pol) was performed using methods described previously. 40 The product was resolved on 1% agarose gel and purified (QIAquick gel extraction kit; Qiagen). Direct Sanger sequencing was performed (Big-Dye terminator kit; Applied Biosystems). Primer sequences were edited out prior to sequence alignment and phylogenetic analyses. To obtain additional sequence for phylogenetic comparison from the leopard tortoise sample, the second round of amplification was altered from the initial PCR assay to use primers DFA and IYG.
In the box turtles, quantitative PCR (qPCR) with a standard curve for TerHV1 was initially performed using methods described previously. 14 Additional pathogen testing was performed (Table 3).
Phylogenetic analysis
A predicted 159 amino acid sequence of herpesviral pol identified in animal 1 was aligned with 66 other herpesviral pol sequences from GenBank (Fig. 2) using MAFFT. 20 Partial sequences from GenBank were included in the alignment, with ambiguities added for unknown amino acids. Elephantid betaherpesvirus 1 (GenBank AF322977) was used as the outgroup for both Bayesian and maximum-likelihood (ML) analyses. Bayesian analysis was performed using MrBayes v.3.2.7a on the CIPRES server with mixed amino acid substitutions, gamma-distributed rate variation, and a proportion of invariable sites, 4 chains, and unconstrained branch lengths. 32 The first 25% of 2,000,000 iterations were discarded as burn-in. ML analysis of the alignment was performed using RAxML-HPC2 v.8.2.12 on the CIPRES server using the best-substitution model as determined by model jumping in the Bayesian analysis, with a gamma-distributed rate variation and proportion of invariable sites. 37 Bootstrap analysis was performed to test the strength of tree topology using RAxML-HPC2 and Consense v.3.697 on the CIPRES server with 1,000 subsets. 9 The phylogenetic tree was visualized using FigTree software (http://tree.bio.ed.ac.uk/software/figtree/).
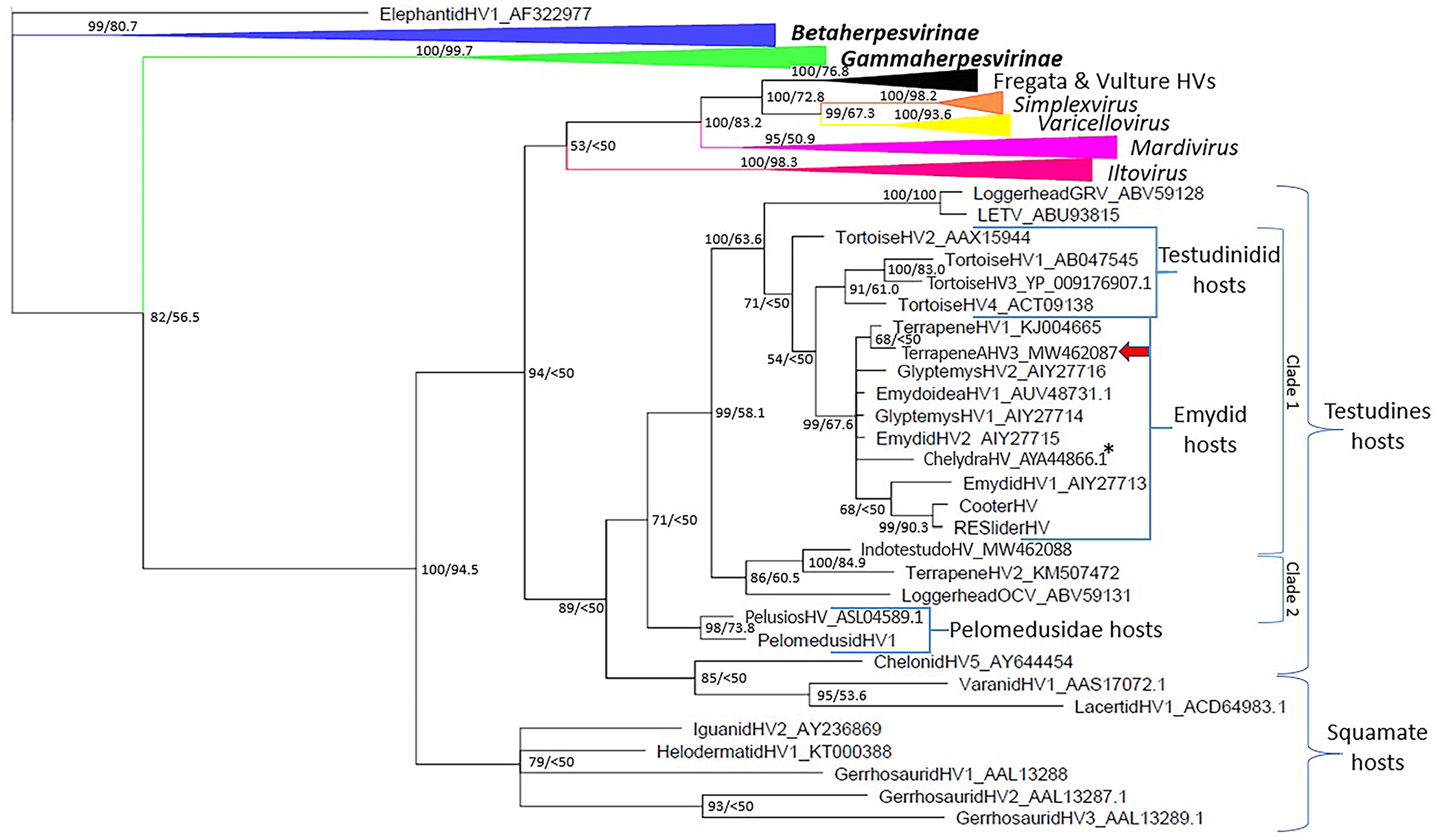
Bayesian phylogenetic tree based on DNA polymerase amino acid sequences of viruses within the family Herpesviridae. Posterior probabilities are the first number at each node, and maximum-likelihood bootstrap values are the second number.
Results
PCR assay and sequencing
Initial endpoint PCR amplification yielded a 181-bp product after primers were edited out from all animals. Viral sequences from all turtles were identical. Animal 3 (BT1) was negative on the 2013 sampling but positive on the 2016 sampling. The modified second round of the leopard tortoise sample returned a 481-bp product and was submitted to GenBank (GenBank MW462087).
At the second sampling of BT1 in 2016, the qPCR targeting TerHV1 detected 412 viral copies. At sampling 20 d before the death of BT2 in 2016, the qPCR targeting TerHV1 detected 1,984 viral copies. At postmortem examination of BT2, the qPCR targeting TerHV1 detected 192,600 copies.
Phylogenetic analysis
The highest score obtained by BLASTX and BLASTN (https://blast.ncbi.nlm.nih.gov/Blast.cgi) was with TerHV1 (GenBank KJ004665), with 96% predicted amino acid sequence homology and 92% nucleotide homology. Phylogenetic analysis revealed the sequence to be a novel alphaherpesvirus that groups with other chelonian herpesviruses within a clade of herpesviruses primarily using emydid hosts (Fig. 2). Support for this clade consists of a Bayesian posterior probability of 99% and a ML bootstrap value of 67.6% (Fig. 2). The virus is hereafter referred to as terrapene alphaherpesvirus 3 (TerAHV3).
Detection and identification of other pathogens
Multiple co-pathogens were detected with TerAHV3 (Table 3), including 17,610,000 copies of the tortoise intranuclear coccidian (TINC) in the leopard tortoise. Mannheimia haemolytica was cultured from the nasal discharge of the steppe tortoise, and the tortoise’s serum was also ELISA-positive for exposure to Mycoplasmopsis (formerly Mycoplasma) agassizii. 13 The 2013 and both 2016 samples from BT2 were PCR-positive for an unnamed emydid Mycoplasmopsis sp. described previously in three-toed box turtles. 27
Discussion
TerAHV3, the novel herpesvirus reported here, clustered most closely with herpesviruses detected previously in emydid turtles, making it likely that the natural host of TerAHV3 is an emydid, such as the three-toed box turtle. Detection of TerAHV3 in the leopard tortoise and steppe tortoise would indicate that this virus has the ability to jump hosts and replicate in aberrant species. Given the co-detection of multiple pathogens and the variable clinical signs observed in the 4 presented animals, the clinical importance of TerAHV3 remains unknown and deserving of further study.
With the exception of chelonid HV5, which clusters with squamate herpesviruses, all other herpesviruses using hosts in the testudines form a clade in our analysis, with 71% Bayesian posterior probability. The earliest divergence among extant testudines is between the side-necked turtles and the cryptodires. 28 Similarly, the first viral divergence in the testudines clade in our analysis is between those using side-necked turtle hosts and those using cryptodires. Within the cryptodires, after the divergence of soft-shelled turtles, the next divergence is between Americhelydia, which includes sea turtles and snapping turtles, and Testudinoidea, which includes tortoises and Emydidae. 28 Two subclades exist within the main clade of herpesviruses using cryptodire hosts. In the larger cryptodire viral subclade (Fig. 2, clade 1), the first divergence, again consistent with the host pattern, is a clade of viruses using sea turtle hosts (loggerhead genitorespiratory virus, and lung-eye-trachea virus [LETV]). Tortoise HV2 is not well resolved, but the remaining viruses of tortoise hosts (tortoise HV1, 3, 4) form a clade (posterior probability 91%, ML bootstrap 61%) also consistent with host divergence. The remaining 10 viruses, including TerAHV3, are found in emydid hosts, with the exception of chelydra HV, which is largely consistent with host divergence patterns and suggests that chelydra HV is a product of an ancestral host jump. Given that TerAHV3 is found within the larger emydid clade and clusters with TerHV1, a box turtle origin is consistent for TerAHV3, rather than being endemic in a tortoise host.
The smaller cryptodire clade (Fig. 2, clade 2), with a Bayesian posterior probability of 86% and an ML bootstrap value of 60.5%, shows that a sea turtle virus (loggerhead orocutaneous virus) branches before a clade containing viruses using tortoise (indotestudo HV) and emydid (terrapene HV2) hosts, consistent with host divergence patterns. Overall, this pattern suggests divergence of 2 viral lineages within the cryptodires after the hosts had diverged from the side-necked turtles (~220 million y ago) but prior to the Americhelydia/Testudinoidea divergence (~128 million y ago) followed by codivergence of these viruses within cryptodire hosts. 33 The divergence within the herpesviruses of turtles is consistent with multiple genera, as seen in avian and mammalian hosts.
The clinical importance of TerAHV3 is not clear. Of the 4 animals in which the virus was detected, 3 had evidence of ocular, upper respiratory tract, aural, or reproductive disease, were infected with co-pathogens, and were subsequently euthanized or discovered dead. The fourth (BT1) displayed no signs of disease and was negative for all potential co-pathogens. The most closely related virus, TerHV1, has been found in association with necrotizing hepatitis with intranuclear inclusions in a hatch-year eastern box turtle. 35 However, a qPCR survey of TerHV1 in 409 box turtles found 31.3% prevalence and no association of clinical signs with virus. 19 Often, herpesviruses have a high prevalence of infection without disease in their endemic hosts, with fatal disease being relatively uncommon. In endemic hosts, when fatal clinical disease is seen, it is typically in neonates. 21 Classic herpesviral lesions that included intranuclear inclusions and necrosis of infected cells were not seen in the chelonians of our report. Based on these findings, it is possible that this virus was not causing the clinical signs of disease observed in these animals.
TerAHV3 appears to have the ability to jump hosts and replicate in a variety of taxa, which is unusual for a herpesvirus. The history of the steppe tortoise (animal 2) is unknown prior to its capture in California. However, given that these tortoises are native to Asia and commonly sold as pets in the United States, it is likely that it was a captive animal that escaped or was released. The leopard tortoise (animal 1) was known to be housed with ornate box turtles, providing a potential source of exposure to TerAHV3. Although herpesviruses are considered relatively host-specific, a number of examples exist of alphaherpesviral infections causing severe disease or fatality in related aberrant hosts. 4 Although no known examples of this exist within chelonians, herpesviruses in their archosaur relatives (crocodilians and avian species), such as columbid herpesvirus 1 (Columbid alphaherpesvirus 1), are known to cause relatively minimal disease in the endemic host, usually young pigeons, but can cause rapid, fatal hepatitis in raptors.5,30 Although its pathogenicity is undetermined, the ability of TerAHV3 to infect chelonians in different families is a potential cause for concern.
The hallmark characteristic of Herpesviridae is the ability to enter latency with the potential to recrudesce during times of local trauma or systemic stress.3,12,21 In the animals presented here, stress may have precipitated reactivation of TerAHV3 and enabled viral detection via PCR. One potential stressor is coinfection with other pathogens, such as the TINC detected in the leopard tortoise (animal 1). TINC may be the most important known pathogen in captive leopard tortoises; there are reports of fatalities, and Koch’s postulates have been fulfilled for this organism. 15 TINC also appears to recrudesce after periods of inactive disease, with the triggers for this event unknown. 38 Stressors other than co-pathogens could also be considered. Thermal stress in experimentally infected rabbits leads to reactivation of human HV1, with high viral loads 4–5 d post-reactivation. 29 Factors found to increase shedding of gallid HV1 (Gallid alphaherpesvirus 1) in chickens include rehousing and egg laying. 16 Depending on a number of factors, including baseline prevalence, TerAHV3 detection may be able to serve as a biomarker for stress in turtle populations.
An emydid Mycoplasmopsis sp. was detected in the three-toed box turtles. This organism was previously isolated from 2, free-living, three-toed box turtles in Forest Park, displaying signs of upper respiratory tract disease, including nasal and ocular discharge, palpebral edema, lethargy, and weight loss. 27 The fourth turtle that we detected with TerAHV3 was the only turtle without clinical signs. Antibodies binding to M. agassizii were detected in the steppe tortoise through an ELISA, although PCR and culture for Mycoplasmatales spp. were negative, hence it is unknown whether this animal experienced concurrent infection with TerAHV3 and M. agassizii.
An organism identified as M. haemolytica, a known pathogen of mammals, was cultured from the steppe tortoise. Reports of M. haemolytica in other taxonomic classes are rare, and many avian isolates classified as M. haemolytica using biochemical methods have been reported to be misidentifications of Gallibacterium sp. 7 An organism isolated from sea turtles was reported to be M. haemolytica. 45 Misidentification is possible, and the clinical importance is unknown.
Co-pathogens previously detected with TerHV1 in eastern box turtles include a box turtle Mycoplasmopsis sp., adenovirus (terrapene adenovirus 1), and ranavirus (Frog virus 3).2,35 One study reported that during a ranavirus mortality event, fewer box turtles died when coinfected with TerHV1 and the box turtle Mycoplasmopsis sp., suggesting a potential protective effect.2,35
Although we detected a novel herpesvirus in 3 species of chelonians, the relationship between TerAHV3 and clinical disease remains unclear. Increasing surveillance in captive and wild chelonians, including routinely collecting samples, such as oral and cloacal swabs that are best stored frozen prior to submitting for molecular testing, may help identify new pathogens such as TerAHV3. We recommend further study to assess whether this virus poses health threats to chelonian species and to further improve understanding of its ecology, evolution, and clinical implications.
Footnotes
Acknowledgements
For information regarding the leopard tortoise, we acknowledge Cathy Minogue of Northwest ZooPath for data retrieval and Leroy Brown of Histologic Consulting Service for slide preparation. We thank Darryl Heard and Elliott Jacobson of the University of Florida College of Veterinary Medicine for postmortem examination of the steppe tortoise.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
