Abstract
An outbreak of diarrhea in an outdoor group of captive Hermann’s tortoises (Testudo hermanni) was associated with fecal shedding of cryptosporidial oocysts, as determined by coproscopic and immunoassay examinations. With partial sequencing of the 18S ribosomal RNA gene, 2 different Cryptosporidium genotypes could be identified in the fecal samples. Cryptosporidium tortoise genotype has previously been found in tortoise and ophidian species, and Cryptosporidium ducismarci has been reported from a snake and a chameleon, and it has been linked to intestinal disease in tortoises. The Hermann’s tortoises described were also infected with oxyurid nematodes. Treatment specific for reptilian cryptosporidiosis was administered. The clinical signs and fecal shedding ceased, but 9 months later, diarrhea and fecal shedding were seen in 3 animals again. Either the oocyst shedding was temporarily suppressed below detection limits, or the animals were reinfected by oocysts still present in the environment. At least 1 of the detected Cryptosporidium genotypes was presumed to contribute to the clinical symptoms.
The clinical significance of cryptosporidiosis in captive reptiles is high. A key feature associated with cryptosporidial infection in reptiles is a chronic subclinical infection with shedding of oocysts at low levels, 7 although infected snakes and lizards can show chronically progressive disease, which is difficult to treat.14,16 Information on the disease course in chelonian species is scarce. Nine Cryptosporidium spp. were identified in reptilian feces in 1 study; the most common were Cryptosporidium serpentis and Cryptosporidium varanii (= saurophilum). 22 Cryptosporidium serpentis is found most commonly in snakes; the organism colonizes the gastric mucosa and can cause severe gastritis. The affected snake often regurgitates its prey.3,8 Cryptosporidium varanii occurs in lizards and tends to infect the mucous membrane of the small intestine and more rarely the stomach or large intestine. Cryptosporidium varanii has also been described in ophidian species, and a recent study found a 16% prevalence of C. varanii in fecal samples from corn snakes (Pantherophis guttatus). 18 Clinical signs have mainly been described for lizards and include emaciation, anorexia, and regurgitation.4,15 Similar signs such as weight loss and lethargy have been reported for tortoises. 3 Latent phases of cryptosporidial infections and the results of a study from Germany, which found that the proportion of Cryptosporidium-positive captive tortoises (tested by means of coproantigen enzyme-linked immunosorbent assay [ELISA] and carbol-fuchsin staining technique [CST] for oocysts) increased from 3.8% in 2006 to 11.1% in 2008, suggest that Cryptosporidium infections may have been underestimated in chelonians. 14 An Italian study found 7 of 21 captive Testudo spp. to be infected with Cryptosporidium by means of a polymerase chain reaction (PCR) assay targeting the Cryptosporidium oocyst wall protein (COWP) gene. 21
In an outdoor captive collection of 13 Hermann’s tortoises (Testudo hermanni) acquired from different sources, 2 animals suffered from chronic diarrhea with soft malodorous feces containing undigested food. The 2 tortoises were alert and had a normal appetite, but they gained weight slowly despite deworming with fenbendazole. a Bacterial culture of feces under aerobic conditions yielded colonies of Escherichia coli and Proteus spp. (sensitive to doxycycline only). Coproscopic examination showed no intestinal parasitic infection. However, large numbers of Hexamita spp. were present in the urine. Antibiotics, oral metronidazole b at 100 mg/kg of body weight once a week and doxycycline c at 25 mg/kg by intramuscular injection at 2-day intervals, as well as supportive therapy with probiotics d and nutritional supplements, e led to a transient improvement, although after 2 weeks, more severe and malodorous diarrhea and weight loss developed. Examination of pooled feces approximately 1 month after the first presentation revealed the presence of Cryptosporidium oocysts; a fecal flotation (262 mg/ml zinc chloride and 275 mg/ml sodium chloride with a specific gravity of 1.3) for nematode eggs was negative. The pooled sample and additional feces from each animal collected 5 days later were examined for Cryptosporidium oocysts with a CST for oocysts, 12 a commercial coproantigen ELISA f (enzyme immunoassay [EIA], previously described 5 and used as a commercially provided kit), and a PCR assay targeting a part of the 18S ribosomal RNA (rRNA) gene. 18 Four samples were positive with the CST, 5 samples were positive and 1 had a threshold value with the EIA, and 6 samples were positive and 2 had threshold values with the PCR (Table 1). The 2 initially presented tortoises with symptoms consistent with cryptosporidiosis (505110, 510210) tested positive with the EIA and PCR or only with the PCR. Two tortoises (504910, 510510) were examined for nematode eggs by fecal flotation, and oxyurid eggs (Pharyngodonidae family) were detected. The PCR amplicons of the Cryptosporidium-positive samples were further analyzed by nucleotide sequencing, 18 and sequence spans correlating in the respective forward and reverse sequences were submitted to a basic local alignment search tool (BLAST; http://www.ncbi.nlm.nih.gov/blast/Blast.cgi) for comparison with published GenBank sequences. The pooled sample and 2 single samples yielded a sequence similar to a part of the 18S rRNA gene of Cryptosporidium ducismarci11,17; the only inconsistency in the sequences was a stretch of 6 or 7 thymine nucleotides (top 5 sequences in Fig. 1). The nucleotide sequence of another sample was identical to the Cryptosporidium tortoise genotype (Table 1). Alignment of the generated sequences from the current study with previously published sequences can be seen in Figure 1. Two samples resulted in partial sequences of non-Cryptosporidium eukaryotes interpreted as nonspecific cross-reaction, and 2 samples yielded no evaluable nucleotide sequences.
Results of the examination of feces from Hermann’s tortoises (Testudo hermanni) specifically for Cryptosporidium spp. with carbol-fuchsin staining technique (CST), enzyme immunoassay (EIA), and polymerase chain reaction (PCR) targeting a part of the 18S ribosomal RNA gene with consecutive nucleotide sequencing of the amplicon, and comparison to published GenBank sequences.*
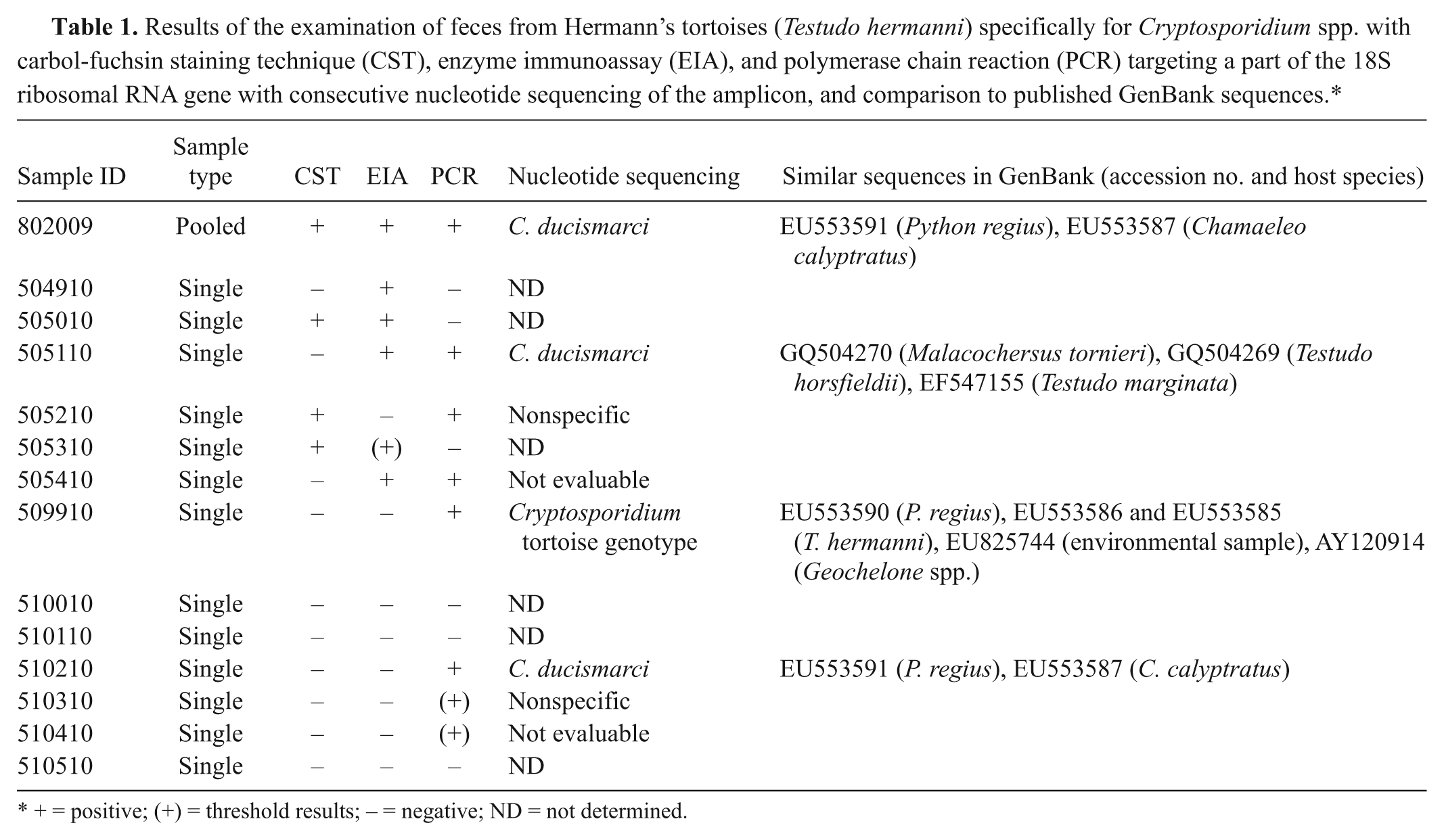
+ = positive; (+) = threshold results; − = negative; ND = not determined.

Alignment of the generated nucleotide sequences (specified by sample number) with exemplary similar partial sequences from GenBank (specified by GenBank accession no.). Details on the corresponding Cryptosporidium genotypes are given in Table 1. Not included are sequences that were not evaluable or belonged to non-Cryptosporidium eukaryotes. In these cases, the results of the polymerase chain reaction were interpreted as nonspecific.
All animals were treated2,15 in the clinic with paromomy-cin g (100 mg/kg body weight, orally, once a day for 7 days and then, at home by the owner, twice a week for 3 months). Additional supportive care with fluid therapy (Lactated Ringer’s at 20 ml/kg body weight) and an immunostimulant h was administered during the first week of therapy. Two weeks after the last paromomycin administration, fecal samples from all animals tested negative with the CST, EIA, and PCR, except for 1 animal that tested positive with PCR; however, the resulting nucleotide sequence was indicative of a nonspecific cross-reaction. Eight months later, a pooled fecal sample from the tortoises was negative for Cryptosporidium by CST and EIA, but a fecal flotation was positive for oxyurid eggs. Nine months after treatment, 3 tortoises developed diarrhea again. Fecal samples from these animals (505010, 505110, 510310) were negative with the CST, but 2 tortoises (505110, 510310) were positive for Cryptosporidium with the EIA. One of these animals (505110; tortoise with diarrhea, weight loss, and C. ducismarci detected at initial presentation) had lost 100 g.
The treatment against cryptosporidiosis led to a clinical improvement in all animals, but the reason for the recurrence of diarrhea and fecal excretion of Cryptosporidium oocysts remains unclear. Either reinfection occurred by long-term viable oocysts in the environment after the cessation of treatment, or the tortoises had a relapse with resistant persistent intestinal infection, and oocyst shedding remained below the detection limit for some months, leading to false-negative results in the applied tests. Clinical signs in leopard geckos improved with paromomycin (aminosidine, an aminoglycoside antibiotic), oocyst shedding was reduced, but the pathogen was not eliminated in all cases. 15
The varied results achieved with the 3 different detection methods used on the same samples in the current study indicate the need for repeated testing when dealing with reptilian cryptosporidiosis. Only the pooled sample was positive with all 3 techniques (Table 1). Other studies comparing different assays for Cryptosporidium detection have also shown a high degree of ambiguity1,21; the reason was suspected to be due to the low number of oocysts present in the samples for immunofluorescence and PCR, 17 which might also occur in chronic subclinical cryptosporidiosis in reptiles. All the techniques used have advantages and disadvantages. A combination of CST and EIA made a fast and simple screening tool for Cryptosporidium in reptile feces.14–16 Commercial immunoassays utilizing monoclonal antibodies were originally developed for the detection of Cryptosporidium parvum antigen, but in reptiles, other Cryptosporidium spp. predominate (e.g., C. serpentis, C. varanii, or various genotypes in chelonians).17,22 With commercial tests, the positive signal may be less evident with reptilian cryptosporidiosis, as has been implicated for oocysts from lizards. 9 Subjective interpretation of signals may lead to an increase of false-negative or inconclusive results, especially if the amount of the sample is not standardized. The EIA was more sensitive than Ziehl-Neelsen acid-fast staining and direct immunofluorescence detection, when used simultaneously to test feline fecal samples. 13 Nevertheless, a combination with a direct oocyst detection method is suggested. The main benefit of the PCR assay is the potential to further differentiate isolates to the species and genotype level by consecutive nucleotide sequencing.1,6 Cryptosporidium oocysts from infected food animals or from the environment can be shed in reptile feces. 10 With PCR and sequencing for species differentiation, interpretation of such pseudoparasites as false-positive results can be prevented. In a 2008 study, Cryptosporidium isolates of captive European tortoises (Testudo graeca, T. hermanni, and Testudo marginata) in Italy were examined genetically using 2 genes (COWP and/or small subunit [SSU] ribosomal DNA gene). 21 Six of these isolates showed 100% homology with the zoonotic and, in this case, possibly pseudoparasitic C. parvum (bovine genotype/Cryptosporidium pestis). Sequencing is also an important way to exclude false-positive PCR results, which consist of amplicons of the expected size unintentionally generated from organisms other than the target organism.
Only limited information is available about cryptosporidiosis in turtles, although a high prevalence of Cryptosporidium oocysts in feces from different tortoise species has been reported. 21 Data indicate the presence of 2 major unique Cryptosporidium spp. in turtles.17,21 In 2010, the histopathological description of 2 distinct Cryptosporidium spp., one in the stomach of a juvenile Russian tortoise (Testudo horsfieldii), and a second in the intestine of another Russian and a pancake tortoise (Malacochersus tornieri), was published. 11 Three partial 18S rRNA sequences (802009 [pooled], 505110, 510210) from the Hermann’s tortoises in the present study showed high similarity with sequences from GenBank databases attributed to C. ducismarci from a Russian and a pancake tortoise, showing inappetence and lethargy,11 and to a marginated tortoise (T. marginata), 21 a ball python (Python regius), 17 and a veiled chameleon (Chamaeleo calyptratus), 17 appearing healthy (Table 1). All these intestinal Cryptosporidium spp. differ in only 1 thymine nucleotide regarding the partial SSU rRNA gene, and they are considered to be from the same species with the proposed name of C. ducismarci. 19
The second commonly found Cryptosporidium genotype in tortoises has been designated the Cryptosporidium tortoise genotype. 22 It has been detected in 1 sample from the current study and from a star tortoise (Geochelone spp.), 22 2 healthy Hermann’s tortoises, 17 a healthy ball python, 17 and an environmental water sample. 23 This genotype shows 96% homology with the partial 18S rRNA gene with the unnamed gastric Cryptosporidium species or genotype from the juvenile Russian tortoise with chronically depressed growth and acute lethargy. 11
The present study describes 2 distinct Cryptosporidium genotypes in a single group of captive tortoises exhibiting clinical signs of disease. The data suggest the intestinal C. ducismarci may contribute to disease in tortoise species. The role of the oxyurid coinfection remains unclear, but generally, these nematodes are thought to be minimally pathogenic. 20 It remains unknown if snakes, lizards, and some tortoise species can be subclinically infected with these Cryptosporidium genotypes.
Footnotes
Acknowledgements
The scientific support of Herbert Weissenböck is gratefully acknowledged.
a.
Panacur®, Intervet Deutschland GmbH, Unterschleißheim, Germany.
b.
Drossapharm, Lörrach, Germany.
c.
Ratiopharm GmbH, Ulm, Germany.
d.
Bird Bene-Bac®, Albrecht, Aulendorf, Germany.
e.
Critical Care, Heusud, Albrecht, Aulendorf, Germany.
f.
ProSpect™, Cryptosporidium microplate assay, Remel, Basingstoke, United Kingdom.
g.
Humatin®, Pfizer Deutschland GmbH, Berlin, Germany.
h.
Zylexis®, Pfizer Ltd., Kent, United Kingdom.
The author(s) declared that they had no conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) declared that they received no financial support for their research and/or authorship of this article.
