Abstract
The lack of quick, accurate, and low-cost detection methods has hindered the active control strategies for bovine tuberculosis (bTB) in resource-limited countries with a high burden of disease. We developed a dry loop-mediated isothermal amplification (LAMP) assay for rapid and specific detection of Mycobacterium bovis, the principal causative agent of bTB, and evaluated the efficacy of the assay using suspected bTB samples collected during routine meat inspection at major regional abattoirs in Malawi. Template genomic DNA was extracted directly from the granulomatous bTB-like lesion (crude extracted DNA), as well as growth from the incubated mycobacterial growth indicator tubes (MGIT). Field results were visualized by the naked eye within 40 min following a color change of the amplified products. The sensitivity and specificity of the dry LAMP assay while using 152 DNA samples extracted from MGIT with confirmed M. bovis results were 98% and 88%, respectively. When 43 randomly selected crude DNA samples from lesions were used, the sensitivity and specificity of the dry LAMP assay were 100% and 75%, respectively. Our LAMP assay offers the potential to meet the demands for a low-cost and rapid field detection tool for bTB in resource-limited countries in which bTB is endemic.
Mycobacterium bovis is the causative agent of bovine tuberculosis (bTB), a chronic infectious disease of mammals.3,13 M. bovis is found mostly in cattle, in which it has caused considerable economic losses as a result of milk, carcass, or organ condemnation, as well as trade restrictions. 15 In addition, the zoonotic transmission of M. bovis to humans is a serious public health hazard, particularly in places where the disease is highly prevalent in animals. 12 In 2016, the World Health Organization reported that resource-limited countries, particularly those in Africa and Southeast Asia, had the highest burden of zoonotic TB. 15 Sadly, these areas are challenged by either inadequate or absent bTB surveillance and control techniques.1,4,9
In cattle, bTB diagnosis relies on the detection of characteristic lesions during routine postmortem examination, but this method is less sensitive and specific than culture and PCR. 8 On the other hand, advanced methods of detection of M. bovis, such as culture, PCR, and sequencing, require the use of standardized laboratory equipment and technically competent staff, thereby limiting their use in rural areas. 8 It is worth noting that the diagnosis of bTB in cattle gives crucial monitoring data that can aid in the development of effective control strategies to cut the transmission links between animals and humans. Therefore, there is a pressing need for the development of rapid, simple, and low-cost techniques for the detection of M. bovis that can be implemented effectively in resource-constrained areas where the bTB burden is still substantial.
Compared with the sophisticated bTB detection approaches, the loop-mediated isothermal amplification (LAMP) technique may be a better alternative for use in resource-constrained areas given the simplicity, speed, specificity, and use of low-cost equipment of the LAMP assay.5,8,11 In 2021, we reported our successful development of a conventional wet LAMP method for the specific detection of M. bovis by targeting a 12.7-kb genomic region (region of difference 4 [RD4]) in which a deletion occurs in M. bovis but not M. tuberculosis and other M. tuberculosis complex species. 8 However, the wet LAMP assay still requires standardized laboratory equipment, such as −30°C freezers to maintain a cold chain of reagents, especially enzymes. Unfortunately, factors such as purchase and maintenance costs and erratic power supply restrict the use of such equipment in resource-limited areas. Therefore, a dry LAMP assay is a better option because the use of dried reagents eliminates the need for cold-chain storage facilities. Dry LAMP assay kits can be prepared, transported, and stored easily without refrigeration.6,14
We based our dry LAMP assay on our validated M. bovis–specific LAMP reaction system. 8 The primers designed previously 8 to target the sequence flanking the 12.7 kb of RD4 were used in our dry LAMP assay. This primer set included the forward inner (F1) primer 3′ section that bridged the RD4 deletion junction that had shown high specificity and sensitivity for M. bovis. 8 A dry LAMP reaction mixture was adopted from other studies, with slight modifications.6,14 Briefly, the dry LAMP reaction mixture was comprised of 2 μL of primer mixture (100 μM primers: outer primers [F3 & B3], inner primers [FIP & BIP], and loop primers [FLP & BLP] mixed at 1:8:6 ratio), 1.4 μL of dNTPs (25 mM); 2.5 μL of 2 M trehalose; 1 μL of 25× LAMP buffer (500 mM Tris-HCl [pH 8.8], 250 mM KCl); 1.8 μL of 100 mM MgSO4; 8 U Bst 2.0 WarmStart DNA polymerase (New England Biolabs); and 1 μL of colorimetric fluorescence indicator (CFI). 8 The mixture was placed in the peripheral part of the lid of a 0.2-mL microtube and dried under a direct flow of clean air in a vacuum container (Sanplatec) overnight with phosphorus oxide and silica gel. Upon visual confirmation of complete drying, the kits were sealed in a light-shielding plastic bag with silica gel bags and kept at room temperature.
The sensitivity of the dry LAMP assay was assessed by diluting M. bovis BCG Tokyo 172 DNA solution to 5 pg, 500 fg, 300 fg, 200 fg, 100 fg, and 50 fg/reaction. We performed dry LAMP reactions by adding 2 µL of diluted DNA and 23 µL of double-distilled water (DDW) to the bottom of dry LAMP assay kit tubes. The tubes were then placed in an inverted position for 2 min to allow the reconstitution of reagents. The reaction mixture was mixed by inverting the tube 5 times. 14 The controls were DDW and M. tuberculosis H37Rv DNA. Isothermal amplification reactions were performed at 66°C for 40 min (LA-200 Loopamp real-time turbidimeter; Teramecs). The detection limit was determined as 200 fg/reaction (Fig. 1A).
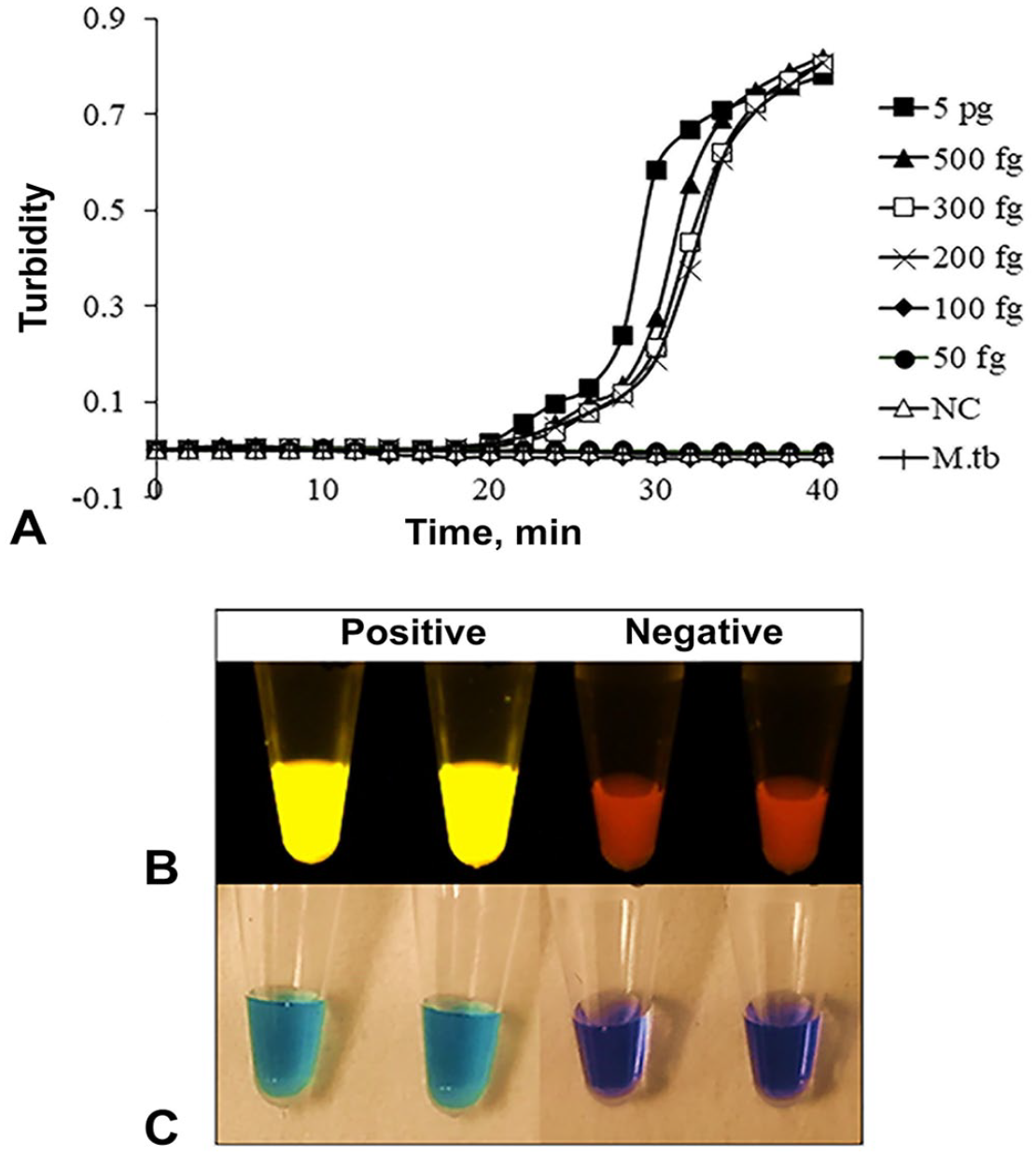
Sensitivity and observation of Mycobacterium bovis dry LAMP assay results.
Our study was approved (protocol 19/09/2398) by the National Health Science Research Committee (NHSRC), Ministry of Health and Population, Lilongwe, Malawi. We collected samples during routine postmortem inspections of cattle slaughtered at 3 major regional abattoirs in Malawi (Northern, Central, Southern regions) between August 2019 and February 2020. Meat inspection and sample processing have been described previously. 7 Briefly, one tissue with a bTB-like lesion was collected per animal, and the relevant demographic data were recorded. The sample was transported (in a cooler box with ice packs) to the National TB Reference Laboratory (NTRL; Lilongwe) for storage at −30°C until use.
For field evaluation, dry LAMP assay kits were transported from Japan to Malawi. We performed our dry LAMP assay on DNA that was either 1) extracted directly from granulomatous bTB-like lesions (crude extracted DNA) or 2) extracted from mycobacterial growth indicator tubes (MGIT) cultures. Approximately 5 g of tissue was trimmed of fat and ground to a paste, decontaminated by adding 4% NaOH, followed by neutralization with PBS and centrifugation at 3,200 × g for 20 min. The supernatant was discarded, and the pellet was re-suspended in 1 mL of PBS. For all of the samples, 500 μL of the re-suspended pellet was inoculated in the MGIT (BACTEC 960; Becton, Dickinson) following the manufacturer’s user manual. For 43 randomly selected samples, the remaining 500 µL of the pellet was subjected to crude DNA extraction. For both the MGIT cultures and processed pellets from granulomatous bTB-like lesions, genomic DNA was extracted by 2 cycles of heating at 95°C for 15 min and immediate cooling to −30°C for 30 min. 7 Dry LAMP reactions were performed as described above, and results were interpreted by observing the color change of CFI (Fig. 1B, 1C). Furthermore, we performed a standard multiplex PCR (mPCR) assay that targeted the RD4, as described previously,2,7 to confirm the identification of isolates as M. bovis. Statistical analysis was done (epiR package of R software; https://www.r-project.org/).
Of the 152 collected samples, 112 were positive on MGIT culture. Of these, we detected 85 isolates as M. bovis using our dry LAMP assay and confirmed by M. bovis–specific mPCR (Table 1; Suppl. Table 1). Based on the mPCR as a reference method to detect M. bovis, the sensitivity and specificity of our dry LAMP assay were 98% (85 positive of 87 samples tested) and 88% (57 negative of 65 samples tested), respectively, compared to MGIT culture results. There was excellent agreement (percentage agreement 93.4%, Cohen kappa = 0.86 [CI: 0.78–0.95]) between our dry LAMP assay and the standard mPCR assay for the detection of M. bovis (Table 1).
Comparative analysis of Mycobacterium bovis dry LAMP assay and the multiplex PCR (mPCR) based on DNA extracted from mycobacterial growth indicator tube (MGIT) positive isolates.
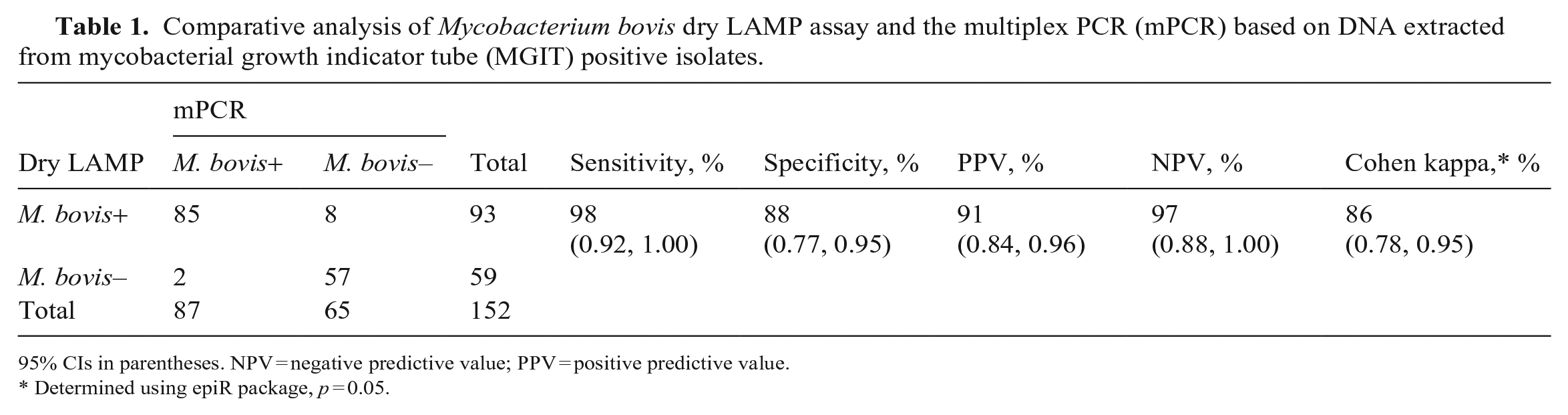
95% CIs in parentheses. NPV = negative predictive value; PPV = positive predictive value.
Determined using epiR package, p = 0.05.
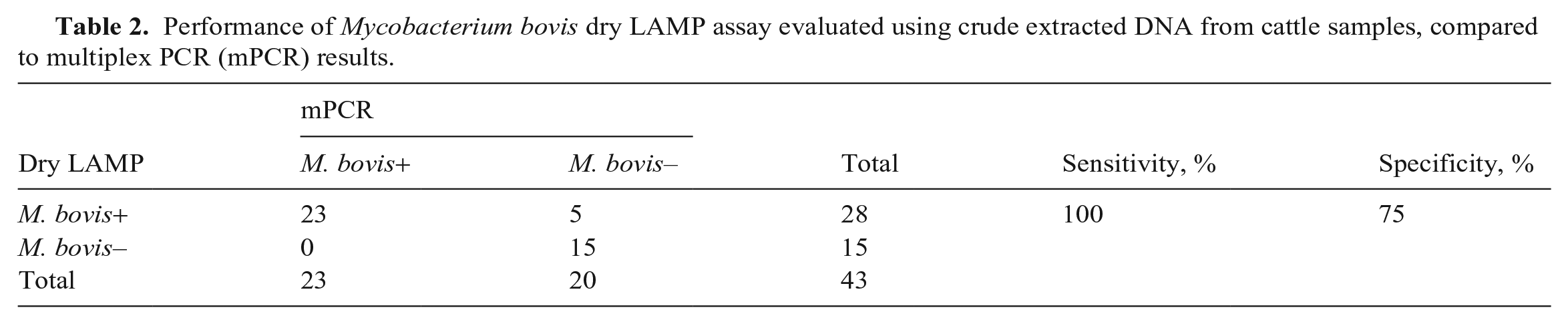
We used simple random sampling to select 43 samples from which crude DNA was extracted directly from the pellets before culturing. Of these selected samples, we detected M. bovis in 28 samples by our dry LAMP assay; M. bovis was confirmed in 23 samples by the M. bovis–specific mPCR assay (Table 2; Suppl. Table 1). Thus, the sensitivity and specificity of our dry LAMP assay with crude DNA extracted before culturing were 100% (23 of 23) and 75% (15 of 20), respectively (Table 2).
Performance of Mycobacterium bovis dry LAMP assay evaluated using crude extracted DNA from cattle samples, compared to multiplex PCR (mPCR) results.
Our dry LAMP assay may prove valuable compared to other traditional methods for detecting M. bovis. First, our LAMP assay identified M. bovis within a shorter time (40 min) than that taken for most PCR-based methods (typically ~3 h).2,10 Second, our LAMP assay procedures are simple and do not require standardized laboratory equipment such as thermocyclers. We used a previously developed battery-powered, hand-made portable LED illuminator 6 to visualize results (Fig. 1B). As well, dry LAMP assay results could be detected by the naked eye using color change (Fig. 1C). Third, the use of crude DNA extracted from samples without culturing removes the time taken for the mycobacteria to grow (~14 d) and hence expedites the surveillance for bTB. Hence, targeted control strategies can be implemented quickly to control the disease. Fourth, the use of dry reagents eliminates the need for cold-chain maintenance of reagents given that the kits are easy to transport and can be stored at room temperature for at least 6 mo. Our dry LAMP assay kits were made in Japan and transported to Malawi without any requirement for a cold chain. In Malawi, we used the dry LAMP assay tubes over 3 mo.
One major challenge that we faced was to optimize the dry LAMP assay to use crude extracted DNA. The specificity of our M. bovis dry LAMP assay was only 75% when DNA was extracted directly from lesions (Table 2). We hypothesized that the DNA extraction method and quality of DNA could have been a contributing factor. Hence, further optimization, especially of the DNA extraction process, will be required. Nevertheless, our M. bovis dry LAMP assay provides an alternative point-of-care detection method for bTB, especially in resource-limited areas because of its rapidity, sensitivity, and absence of a requirement for sophisticated laboratory tools.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387231164596 – Supplemental material for Direct detection of Mycobacterium bovis by a dry loop-mediated isothermal amplification assay in cattle samples collected during routine abattoir examination in Malawi
Supplemental material, sj-pdf-1-vdi-10.1177_10406387231164596 for Direct detection of Mycobacterium bovis by a dry loop-mediated isothermal amplification assay in cattle samples collected during routine abattoir examination in Malawi by Thoko Flav Kapalamula, Jeewan Thapa, Kyoko Hayashida, Joseph Chizimu, Wimonrat Tanomsridachchai, Mirriam Ethel Nyenje, Rajab Mkakosya, Chie Nakajima and Yasuhiko Suzuki in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was supported in part by a grant from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT), Japan, for the Joint Research Program of the International Institute for Zoonosis Control, Hokkaido University to Yasuhiko Suzuki, and in part by the Japan Agency for Medical Research and Development (AMED) under grants JP22jk0210005, JP22jm0110021, and JP22wm0125008 to Y. Suzuki.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
