Abstract
Canine parvovirus 2 (CPV2) causes one of the most serious canine viral infections, with high mortality in young dogs. In 2014, 2019, and 2021, we determined genetic sequences of CPV2 strains obtained from 39 fecal samples collected from the Yanbian Korean Autonomous Prefecture in the Jilin Province of China. Sequence alignments were performed using the major capsid protein (VP2) gene; protein sequences of these samples had high nucleotide (>97.4%) and amino acid (>95.6%) identity. All of the amino acid sequences contained Ser297Ala and Tyr324Ile mutations. Our survey indicated a high prevalence of CPV2 variants in Yanbian Prefecture, with the new CPV2a variant (26 of 39; 67%) being the most frequent. CPV2c, identified in 9 of 39 (23%) samples, had not been detected in this region previously, indicating the potential risk of CPV2 mutation. The sequences of our 39 CPV2 samples were more highly homologous to the published Chinese strains than to the CPV2 variant strains found in other countries.
Canine parvovirus 2 (CPV2; Parvoviridae, Carnivore protoparvovirus 1) first emerged as a novel pathogen in 1978, causing acute hemorrhagic enteritis and myocardial disease in young dogs. CPV2 is a nonenveloped virus with a single-stranded 5,232-bp DNA genome containing 2 open-reading frames (ORFs) 11 encoding the major viral capsid proteins VP1 and VP2, as well as nonstructural proteins NS1 and NS2. VP2 is the major capsid antigen of CPV2 and plays an important role in determining host range and cell tropisms. The canine parvoviruses are canine minute virus (also known as canine parvovirus 1; Parvoviridae, Carnivore bocaparvovirus 1), and CPV2. 1 CPV2 causes serious economic losses to the canine industry, including the purebred pet industry, especially in Yanbian, China.
Three pathogenic mutant variants of CPV2 arose between 1980 and 1996. CPV2a, the first CPV2 to be isolated, contains 4 substitution mutations, namely Met87Leu, Ile101Thr, Ala300Gly, and Asp305Tyr. In 1984, CPV2b was identified with an Asn426Asp substitution in addition to those of CPV2a. In 1996, CPV2c was identified in Germany as the third CPV2 antigenic variant, and contained an Asp426Glu substitution in the capsid surface of its CPV-VP2. CPV2c was the predominant variant observed through genotype analysis in Europe, America, Africa, and Asia. Since the emergence of CPV2 in 1980, various CPV2 subtypes have affected dogs in China. Investigations have shown that CPV2a is widely distributed throughout China; CPV2b is prevalent mainly in the southern regions of the country. 20 CPV2c was initially identified in Changchun, Jilin Province, in northeast China. Given that a further substitution, Ser297Ala, was seen in these CPV2a/2b variant strains, they were designated new CPV2a and new CPV2b and have been reported worldwide.2,3,6,8
Yanbian Korean Autonomous Prefecture, Jilin Province, borders North Korea and Russia. Yanbian is in the center of northeast Asia and has close trade with South Korea. The canine breeding industry, especially for pet dogs, in Yanbian is economically important, and the prevalence of CPV2 has seriously hindered the development of the regional economy. Therefore, we assessed the prevalence of CPV2 for 3 discrete years in the Yanbian region and identified the main CPV2 variants.
We collected fresh fecal swab samples from 39 dogs with clinical signs of parvoviral infection from various hospitals in 8 counties in Yanbian Prefecture in 2014, 2019, and 2021. A CPV2 antigen colloidal gold test (CCV/CPV Ag test kit; Asan Pharm) was used to confirm CPV2 infection in all 39 dogs. CPV2 infection in some dogs was accompanied by vomiting, diarrhea, and dehydration; others had dyspnea and exercise intolerance. After a positive result in the CPV2 Ag test, 1 mL of whole blood from each dog was centrifuged at 2,000 × g for 3 min; 10 μL of serum was added to 100 μL of diluent in the antibody test kit (CPV2, ICH, CDV triple Ab test kit; Wondko) and mixed well. Then 75 μL was injected into the sampling well, and the antibody level was read in an immunofluorescence detector (YG-101; Wondko). The demographics of the 39 dogs were documented, including their age, sex, breed, vaccination status, antibody status, and collection counties (Suppl. Table 1).
Subsequently, each collected swab was immersed in 5 mL of PBS (pH 7.2). The fecal swabs were pressed against the wall of the centrifuge tube to remove residual fecal material; the resulting suspension was frozen (−80°C) and thawed 3 times. We centrifuged 200 μL of the fecal suspension at 12,000 × g for 1 min at 4°C, discarded the supernatant, added proteinase K (CWBIO), and dissolved the precipitate. The final step was to extract genomic DNA from the dissolved precipitate (Mini BEST viral RNA/DNA extraction kit, v.5.0; TaKaRa) following the manufacturer’s instructions, and store the DNA at −80°C before PCR amplification.
Using the VP2 region of the CPV2 (JN867607) GenBank sequence, a primer pair (For: 5′-GAATTCATGAGTGATGGAGCAGTTCAAC-3′; Rev: 5′-CTCGAGATAATATAATTTTCTAGGTGCTAG-3′) was designed using Oligo 6.0 software (Molecular Biology Insights). The VP2 fragments were amplified using a thermocycler (MyCycler; Bio-Rad) with the following protocol: pre-denaturation at 94°C for 5 min, 30 cycles of 30 s at 94°C, 1 min at 52.2°C, 2 min at 72°C, and a final extension at 72°C for 10 min. Each purified, positive PCR product (1,755 bp) was cloned into the pMD19-T recombinant plasmid and directly sequenced (Invitrogen). The VP2 sequence insertion was ~1,755 bp and encoded a protein of 584 amino acids. We uploaded our 39 sequences to GenBank as accessions KP749837–KP749855, KM014812, MW715593–MW715601, MW570769, OM057673–OM057681. In the sequenced samples, 26 of 39 (67%) were positive for the new CPV2a variant, 4 of 39 (10%) were positive for the new CPV2b variant, and 9 of 39 (23%) were positive for the CPV2c variant (Fig. 1).
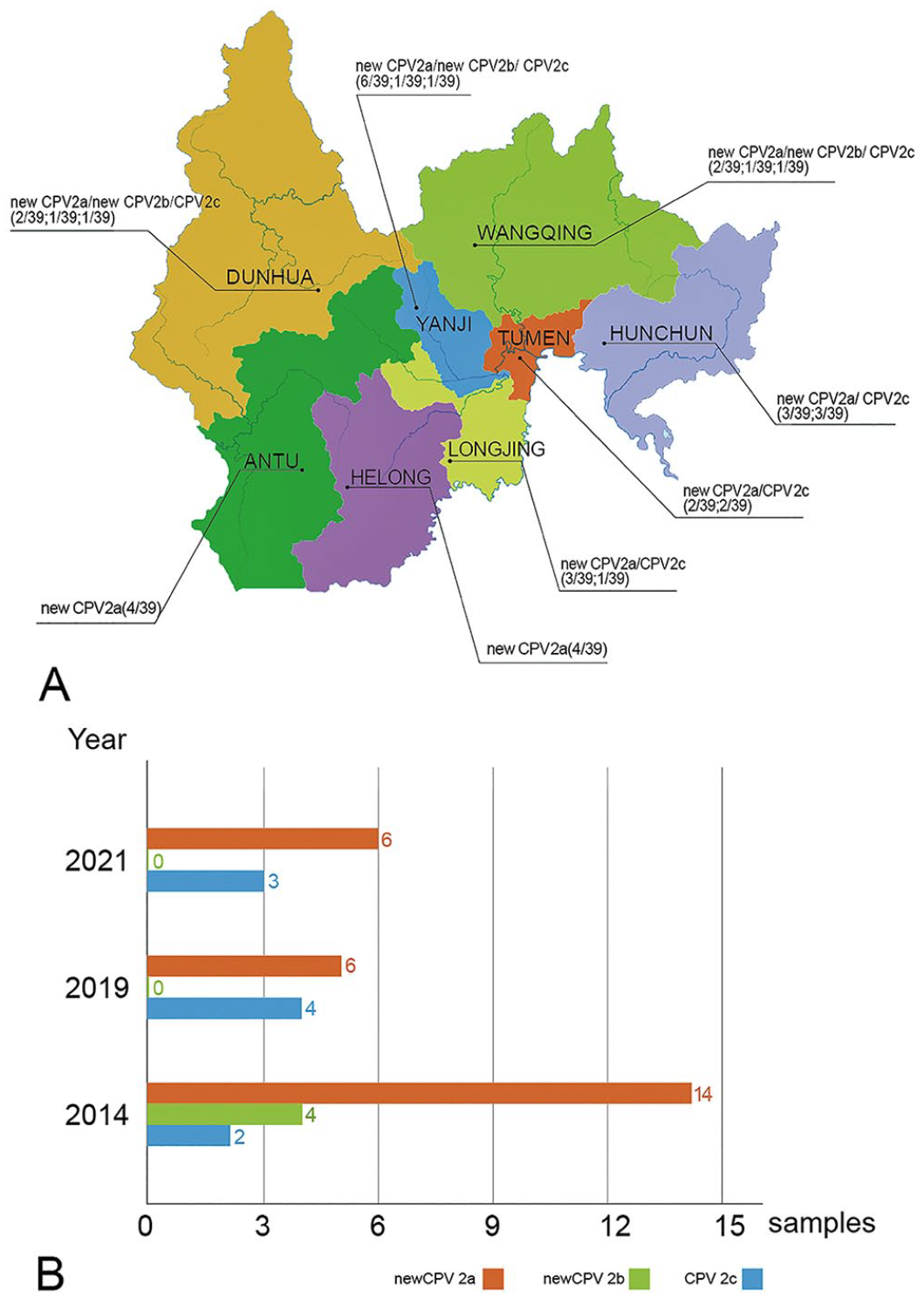
Molecular epidemiology of canine parvovirus 2 (CPV2) in Yanbian, China in 2014, 2019, and 2021.
The distribution of CPV2 variants indicated that the age of the dog had a substantial effect on the infection risk. Puppies 2–6-mo-old were more susceptible to CPV2 infection than older dogs, with 2-mo-old puppies most affected. Additionally, the CPV2c variant was detected in 2 dogs >1-y-old. There was no significant difference in the CPV2 detection rate between sexes; 24 of 39 (62%) positives in male dogs, and 15 of 39 (38%) positives in female dogs. The clinical histories of dogs in our investigation suggested that animals living in poor sanitary environments were more susceptible to infection. Furthermore, the timing of vaccination was important. We estimate that the dams of ~13 puppies had been vaccinated with CPV2 vaccine. There were significant differences in antibody status between vaccinated dogs (vaccinated at least once; 27 of 39, 69%) and non-vaccinated dogs (12 of 39, 31%; Suppl. Table 1), with antibody status significantly higher in vaccinated dogs (vaccinated at least once) than in unvaccinated dogs (Suppl. Table 2). The chi-square test resulted in a χ2 value of 17.0, and p = 0.009. CPV2c was detected in 5 dogs after 3 full doses of vaccine, and we suspect that the vaccine used did not provide adequate immunity against this circulating strain of CPV2.
We analyzed the homology of CPV2 strains isolated from 39 canine fecal samples by phylogenetic analysis and determination of amino acid mutations and compared them to reference strains from GenBank (Fig. 2). Nucleotide and amino acid sequence prediction was performed following analysis of mutations at key amino residues, including Lys87Arg, Phe267Tyr, Ser297Ala, Asp305Tyr, Tyr324Ile, Gln370Arg, Asn426Asp/Glu, Thr440Ala, and Ala555Gly (Suppl. Table 3). To facilitate the characterization of genotype distribution, 98 amino acid sequences of VP2 regions in CPV2 were used to construct a phylogenetic tree that contained all of the CPV2 variants; variation was strong within branches. The GenBank sequences used in our analysis included our 39 samples, 4 vaccine sequences (containing the vaccines used for dogs in our study), and VP2 sequences of 55 CPV2 subtypes obtained from isolates in Europe, Asia, and America. Phylogenetic analyses were performed using the maximum-likelihood method (Poisson model) in MEGA X software (v.10.1.8; https://www.megasoftware.net/). A bootstrap value was determined with 1,000 replicates.
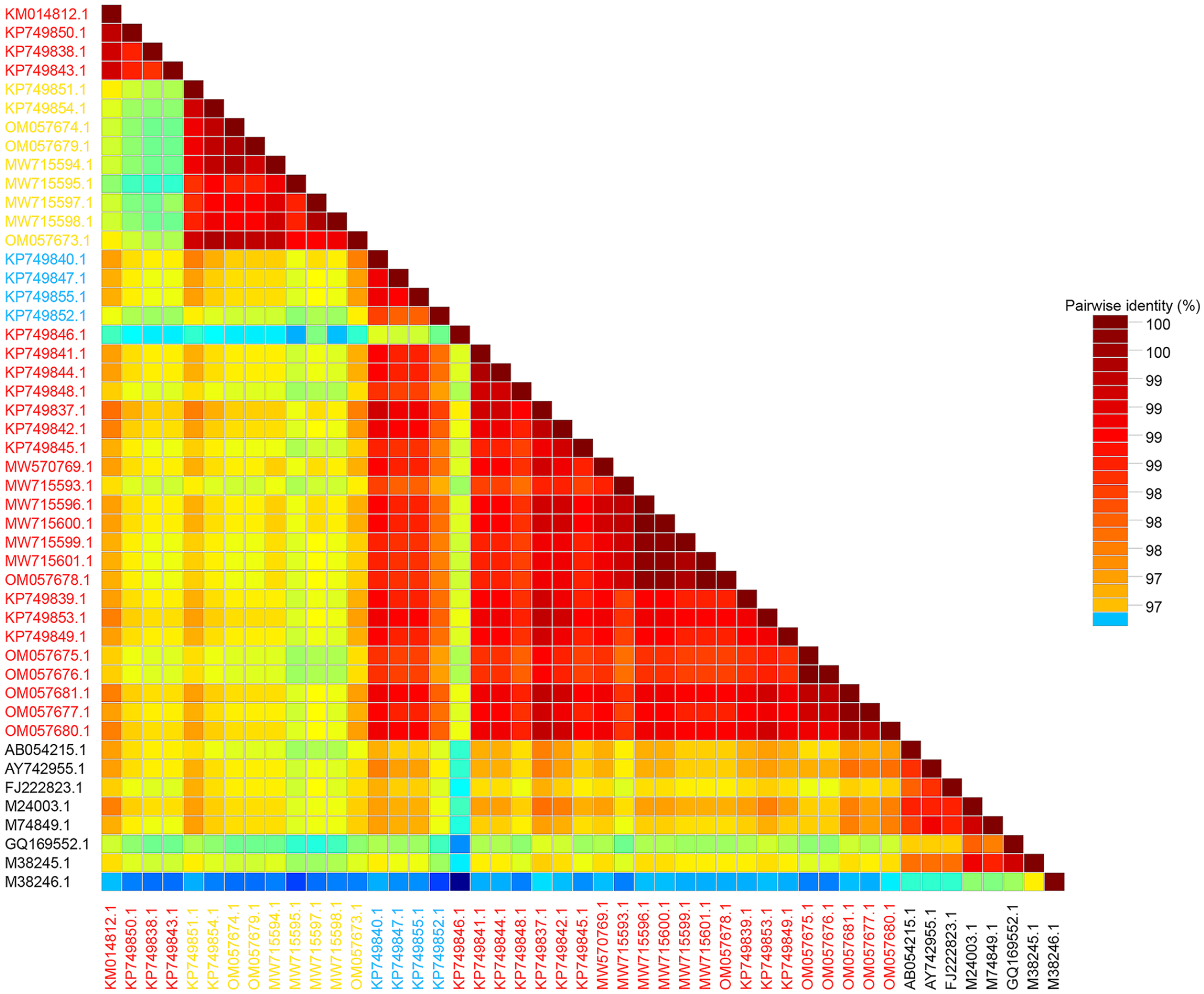
Homology analysis for the investigated canine parvovirus 2 (CPV2)-VP2 sequences and 8 reference strains (CPV2 GenBank AB054215, AY742955, M24003, M74849, GQ169552, and M38245; FPV GenBank M38246). For our 39 sequences, new CPV2a variants are labeled in red, new CPV2b in blue, and CPV2c in yellow.
We determined the genomic diversity of CPV2 strains in the Yanbian region by comprehensive phylogenetic analysis and amino acid mutation analysis, with 97.4–99.8% nucleotide identity and 95.6–99.8% amino acid identity among our 39 samples and the reference strains. The YBYJ sample (GenBank KM014812) was most homologous to the new CPV2a reference strain (GenBank AB054215), with 99.8% identity. By analyzing amino acid mutation sites combined with our 39 sequences, we discovered that our CPV2 strains were distributed in clades A–D and were highly homologous with strains from China, South Korea, and Thailand. We discovered that China’s new CPV2a/2b strains differed significantly from those of other nations. The new CPV2a/2b strain from China was mainly concentrated in clades A and B. In contrast, the new CPV2a/2b strain from other countries were mainly concentrated in clade E. Additionally, the CPV2 vaccine strains in clade F were more evolutionarily distant from the current circulating strain in the Yanbian region (Fig. 3).
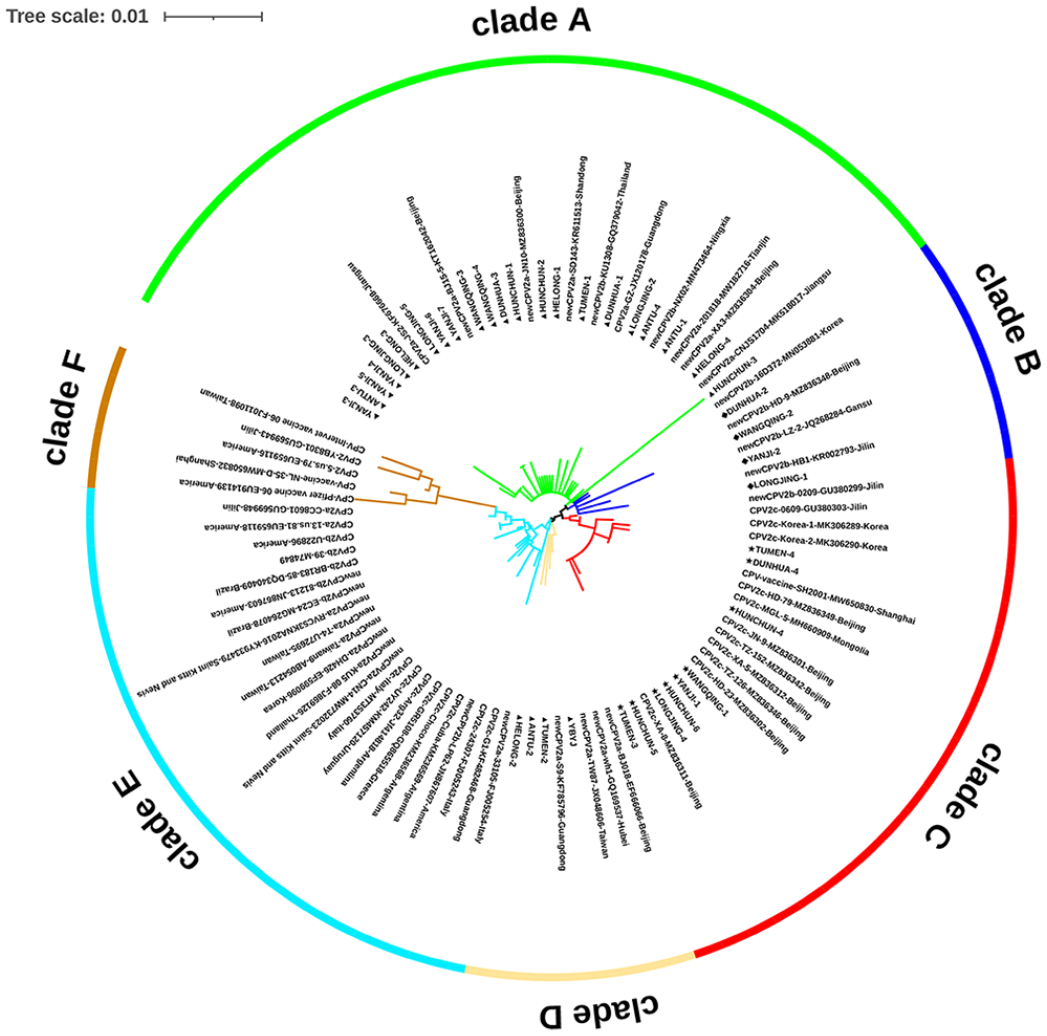
Phylogenetic tree of canine parvovirus 2 (CPV2)-VP2 gene sequences together with the 39 sequences identified in our study performed with the maximum-likelihood method using MEGA 5.0 software with a bootstrap test of 1,000 replicates. The clade A of new CPV2a, clade B of new CPV2b, clade C of CPV2c, clade D of new CPV2a, clade E of CPV2a/2b and new CPV2a/2b from global CPV-VP2 sequences for the derived viral sequences are indicated using the name of the city or country from which the sample was collected, GenBank accessions, and the type of sequences. The symbols used to differentiate the different strains were as follows: triangular icons (▲) represent new CPV2a strains, diamond icons (◆) represent new CPV2b strains, and star icons (★) represent CPV2c strains.
In previous epidemiologic investigations, we found that the new CPV2a and new CPV2b variants predominated in northeast China, 21 and CPV2c was reported rarely; however, here we detected CPV2c in 9 of 39 samples. We found that the new CPV2a strain was the predominant genotype circulating in the Yanbian region, similar to the prevalence of the new CPV2a strain elsewhere in China. Although CPV2c was not the dominant strain in our samples, our detection may indicate an upward trend in infections by the variant. CPV2c has been reported as the dominant strain in other countries.9,13,14
The CPV2c and new CPV2b strains that we detected were characterized by the emergence of novel mutations. Analysis of the phylogenetic tree also showed 98% similarity between these sequences and CPV2 isolated from Jilin Province (GenBank KR002793). Two samples, TUMEN-4 and DUNHUA-4, had 99.7% similarity to the CPV2c strain from Korea (GenBank MK306290). Thus, we suspect that cross-border transmission had occurred. Even though sequence data for CPV2 isolates from North Korea and northeastern Russia were not available, our findings could indirectly indicate the presence of these strains in these 2 countries.
The Ser297Ala substitutions have been identified in the new CPV2a/2b strains 4 ; all 39 sequences in our investigation had this mutation. This substitution has facilitated host range adaptation under selection pressure. 16 The Tyr324Iso mutation was also present in all of our samples and is adjacent to residue 323, a critical amino acid position for host range determination, 13 with mutations occurring frequently in Korea, Thailand, Uruguay, Japan, India, and China.10,17,19,22 To our knowledge, there is no evidence to confirm that residue 324 influences the host range and antigenic properties of CPV2 variants. Therefore, the functional effect of these 2 amino acid residues on antigenic variation is unclear. Additionally, Ser297Ala and Tyr324Ile mutations occurred in all of the sequences in our investigation, meaning that the samples collected in our study were all new CPV2 variants. Researchers have reported that the Gly300Asp and Ala300Asp mutations affect surface-exposed capsid residues 18 ; however, we did not see these substitutions in our study samples.
Additionally, we identified the Phe267Tyr mutation in 16 samples and the Thr440Ala mutation in 14. Among the 39 sequences, Thr440Ala appeared in ~36% of the sequences. Thr440Ala variants are widespread in India, Japan, Korea, Argentina, and China, with some authors hypothesizing that residue 440 is the main antigenic site of the virus and has provided a selective advantage during its evolution. A total of 92% of our CPV2 samples bore the Phe267Tyr mutation, which is highly prevalent in dogs in China. However, residue 267 is not a CPV2 capsid surface site, and it has not been demonstrated if this mutation affects the infectivity of the virus in dogs. Notably, we found that 5 samples were mutated at residue 447, which is not exposed on the viral capsid surface. Mutations at this locus have been reported in Vietnam; however, the relevance of the mutation and its pathogenicity have not been established. 5 Only 2 of our 39 samples had mutations at residue 5. Although most of this genetic drift might be neutral, a fixed-effects likelihood method using Datamonkey (www.datamonkey.org) to calculate the selection pressure for sites showed that 256 and 426 were positive selection sites; 426 is a key site, and whether it plays a role in the evolution of CPV2 needs to be further evaluated.7,12
The vaccine used most commonly in the Yanbian region is the Pfizer vaccine. Compared with the vaccine strains (GenBank EU914139, FJ011098), which were Intervet and Pfizer vaccines, we found that none of our 39 samples contained vaccine strain sequences. High nucleotide substitution rates have resulted in differences between vaccine and field strains, which may have affected the immunologic efficacy of vaccines. VP2 gene substitution could reflect a high background mutation rate, and the differences between the CPV2 genotypes in vaccine and field samples might be a major cause of vaccine failure. The vaccines used in our region against the CPV2 variants are not ideal, and immunized dogs are still at risk. The development of vaccines against the CPV2 variants is imperative. 15
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387221117556 – Supplemental material for Molecular epidemiology of canine parvovirus 2 from 2014, 2019, and 2021 shows CPV2 circulating and CPV2c increasing in Yanbian, China
Supplemental material, sj-pdf-1-vdi-10.1177_10406387221117556 for Molecular epidemiology of canine parvovirus 2 from 2014, 2019, and 2021 shows CPV2 circulating and CPV2c increasing in Yanbian, China by Haoyuan Ma, Xu Gao, Jingfeng Fu, Haowen Xue, Kunru Zhu, Biying Mu, Yanhao Song, Peng Dong and Zhenliang Wang in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported by the Natural Science Foundation (20210101368JC) and “13th 5-Year” Science and Technology Project of Education Department of Jilin Province (JJKH202-00521KJ) of Yanbian University, China.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
