Abstract
Metastrongylosis is an infection of the respiratory tract of pigs caused by parasites of the genus Metastrongylus, whose eggs are similar to other Strongylida through light microscopy; species-specific identification can be performed with molecular tools. We explored the species composition and the genetic diversity of Metastrongylus infecting pigs in close contact with humans in impoverished rural communities in the state of Piauí, in northeastern Brazil. Fecal samples (n = 78) were collected for parasitologic tests. Egg morphometry and molecular characterization, using the cytochrome c oxidase subunit 1 (cox1) gene, were performed. For strongyliform eggs, 62 of 78 (80%) pigs were positive and 6 of 99 (6%) eggs had dimensions compatible with Metastrongylus. Of the 37 samples submitted to PCR, 10 were identified as M. salmi. We found 3 M. salmi haplotypes, including 2 new and 1 described previously in Europe. Overall, M. salmi demonstrated lower intraspecific genetic diversity: diversity index (H) ± SD = 0.318 ± 0.164, n = 12, compared with published M. pudendotectus sequences (1.000 ± 0.272, n = 3). To our knowledge, M. salmi DNA sequences have not been published previously from pigs in South America.
Metastrongylosis is caused by helminths of the genus Metastrongylus, found in the airways of wild and domestic pigs.17,19 Pigs become infected by ingesting earthworms that harbor infectious Metastrongylus spp. larvae; in turn, earthworms become infected by ingesting soil contaminated with eggs or first-instar larvae released in pig feces. The infection ranges from subclinical to clinically manifest, including cough, dyspnea, tachypnea, nasal discharge, tremors, and weight loss, which can lead to the pig’s death. The genus Metastrongylus has 6 species: M. apri (syn. M. elongatus), M. salmi, M. pudendotectus, M. confusus, M. asymmetricus, and M. madagascariensis. 6 Mixed infections with different species of this genus can occur. Rare cases in humans have been reported (e.g., in Europe by M. elongatus and in South America by M. salmi).2,14 The zoonotic potential of Metastrongylus infections emphasizes the need for studies using the One Health approach. 21
Globally, the species M. salmi, M. pudendotectus, and M. apri are detected most frequently through the morphologic identification of adult specimens obtained at autopsy. In Brazil, the species detected most frequently is M. salmi, followed by M. pudendotectus and M. apri (Suppl. Table 1).
Brazil has one of the largest swine herds in the world. In low-resource communities, pig farming is practiced extensively or semi-extensively in pigsties located in the peridomestic environment, or with free-range pigs that inhabit the surroundings of houses. In some regions, there is close contact between the residents with animals and contamination of the environment by fecal waste. In general, the prevalence of parasitic diseases in pigs is higher in regions where extensive farming is practiced, causing economic losses for producers. 22
Parasites belonging to different species of the order Strongylida (e.g., Oesophagostomum spp., Trichostrongylus spp., and Metastrongylus spp.) lay eggs that are difficult to distinguish through light microscopy, making specific identification difficult before slaughter. Molecular taxonomy is an alternative means to identify species and analyze the intraspecific genetic variability of these helminths. Information on the genetic diversity of Metastrongylus spp. is scarce in the world and nonexistent in Brazil. Thus, we explored the species composition and the genetic diversity of Metastrongylus spp. infecting pigs raised extensively in rural communities in the state of Piauí, Brazil.
Our fieldwork was conducted in Nossa Senhora de Nazaré (NSN), state of Piauí, northeastern Brazil; the estimated population is 4,890 inhabitants distributed in 146 small and predominantly rural localities. The transition between the Caatinga/Cerrado biomes and Carnauba riparian forest is predominant. The tropical climate is alternately wet and dry. The population lacks basic sanitation, and people live in close contact with domestic animals, such as pigs, goats, and sheep. 15 Extensive pig raising predominates as a source of food and income in rural regions in northeastern Brazil and, in NSN, pigs represent the second largest herd after small ruminants. Pigs are raised loose, without hygiene and organized management, and circulate in the peridomestic environment feeding on what they find, including human feces.
During field visits, fecal samples were collected from pigs after spontaneous defecation on the ground in rudimentary breeding sites or in peri- and intra-households. Samples were stored individually in plastic bags, placed in a container with ice, and sent to the field laboratory for parasitologic examination. Feces were processed using the Ritchie method (centrifugal sedimentation with ethyl acetate) and sucrose flotation. Strongyliform eggs were measured with an ocular micrometer to verify if they were compatible with Metastrongylus spp. (28.5–61.6 µm long, 21–47.5 µm wide). Samples with helminth eggs were kept frozen (−20°C) until DNA was extracted. Our study was approved by the Ethics Committee for the Use of Animals (license LW-21/13 [P-4/13.3]) of the Oswaldo Cruz Institute/Fiocruz.
Genomic DNA was extracted from 37 strongyliform egg–positive samples (DNeasy blood & tissue kit; Qiagen) according to the manufacturer’s instructions. The partial cytochrome c oxidase subunit 1 (cox1) gene was amplified (Platinum Taq DNA polymerase kit; Invitrogen) with a final volume of 50 µL and with a primer cocktail designed for nematode DNA barcoding. 18 PCR conditions were as follows: initial denaturation at 94°C for 5 min, followed by 35 cycles of 94°C for 40 s, 55°C for 40 s, 72°C for 1 min, and a final extension at 72°C for 5 min. PCR products were purified (DNA Illustra GFX PCR and gel band purification kit; GE HealthCare) and subjected to sequencing (BigDye Terminator v.3.1 cycle sequencing kit; Thermo Fisher) in both directions using M13F: 5′-TGTAAAACGACGGCCAGT-3′ (forward) and M13R: 5′-CAGGAAACAGCTATGAC-3′ (reverse) primers. 13 Capillary electrophoresis was performed (3730 automated DNA sequencer; Applied Biosystems).
BioEdit v.7.2.5 (https://bioedit.software.informer.com/) was used to edit the nucleotide sequences. BLASTn (https://www.ncbi.nlm.nih.gov/) was used to verify the similarity of the nucleotides with sequences of nematodes from GenBank. Orthologous sequences (n = 6) were retrieved from GenBank (Suppl. Table 2); sequences with degenerate bases were not included. Ostertagia ostertagi (GenBank AB246108) was used as outgroup. We deposited our sequences in GenBank (MK282827-36).
Phylogenetic inferences were performed using MEGA 7 (v.7.0.20, https://www.megasoftware.net/). The maximum-likelihood (ML) method was applied, and the Tamura–Nei (TN93) substitution model was selected using the Bayesian Information Criterion (BIC) in MEGA 7. The clade stability of the cox1 sequence tag topologies was evaluated using 1,000 bootstrap replicates.
The relationships in the haplotype network were inferred through median-joining using Network v.10.2.0 and DnaSP v.6.1,20 The genetic diversity indexes of Metastrongylus populations were calculated using Arlequin v.5.2.2. 5 The Fst fixation index was performed on all populations using Arlequin to estimate the genetic differentiation among populations with a significance of 1,000 permutations.
We detected strongyliform eggs in 62 of 78 (80%) fecal samples. From the positive samples, the size of 6 of 99 (6%) eggs was compatible with Metastrongylus spp., with mean length of 45.1 µm (SD = 12.3 µm) and average width of 28.6 µm (SD = 7 µm). Of the 37 samples submitted to PCR amplification and cox1 nucleotide sequencing, 10 represented sequences of the genus Metastrongylus. The sequences were characterized as M. salmi. Three different haplotypes were identified. Two haplotypes are previously undescribed, and one of them had 100% similarity to M. salmi isolated from wild boar in France (GenBank MT338559). We constructed a phylogenetic tree (Fig. 1) based on the 10 new sequences from our study and 5 sequences from Metastrongylus spp. available in GenBank.
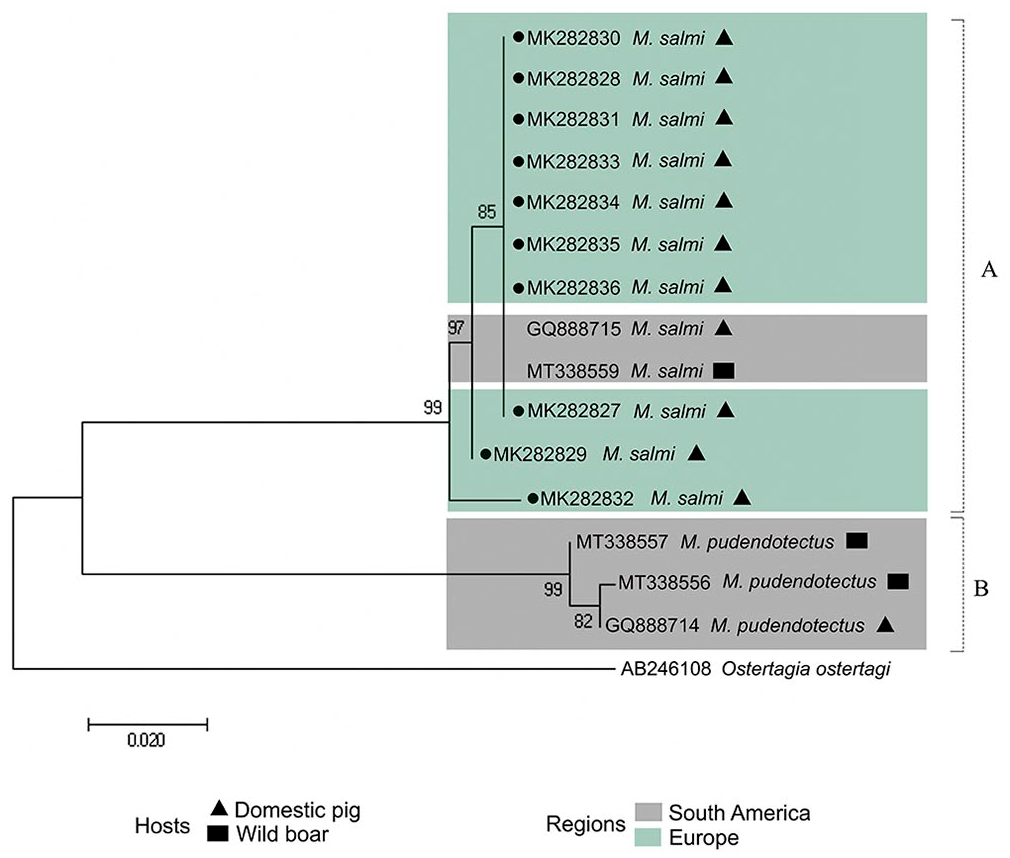
Maximum-likelihood tree constructed using 391-bp cox1 locus of Metastrongylus spp. Ostertagia ostertagi was used as the outgroup. Black dots = samples from our study. GenBank accessions are indicated. Further details of reference strains can be found in Supplementary Table 1.
Analysis of genetic diversity indices showed that there were 2 and 8 polymorphic sites (Table 1; Suppl. Table 3) between the 2 new haplotypes (MK282829 and MK282832) and the GenBank sequences of M. salmi identified in wild and domestic pig samples from Europe. Furthermore, M. salmi had lower intraspecific diversity (H ± SD = 0.318 ± 0.164, n = 12) compared with M. pudendotectus (1.000 ± 0.272, n = 3; Suppl. Table 3). Interspecific diversity was moderate (0.571 ± 0.149, n = 15). The haplotype network was correlated with the ML results, demonstrating a clear division among species with at least 48 mutated positions (Fig. 2). For the species M. salmi, there was a dominant haplotype, with 8 sequences from Brazil (isolated in our study) and 2 from Europe (MT338559 from France, GQ888715 from Estonia). The results of the Fst analysis corroborated the results of the genetic diversity analyses, with interspecific divergence higher than intraspecific divergence (Suppl. Table 4).
Polymorphic sites detected in cox1 locus of Metastrongylus salmi from our study.
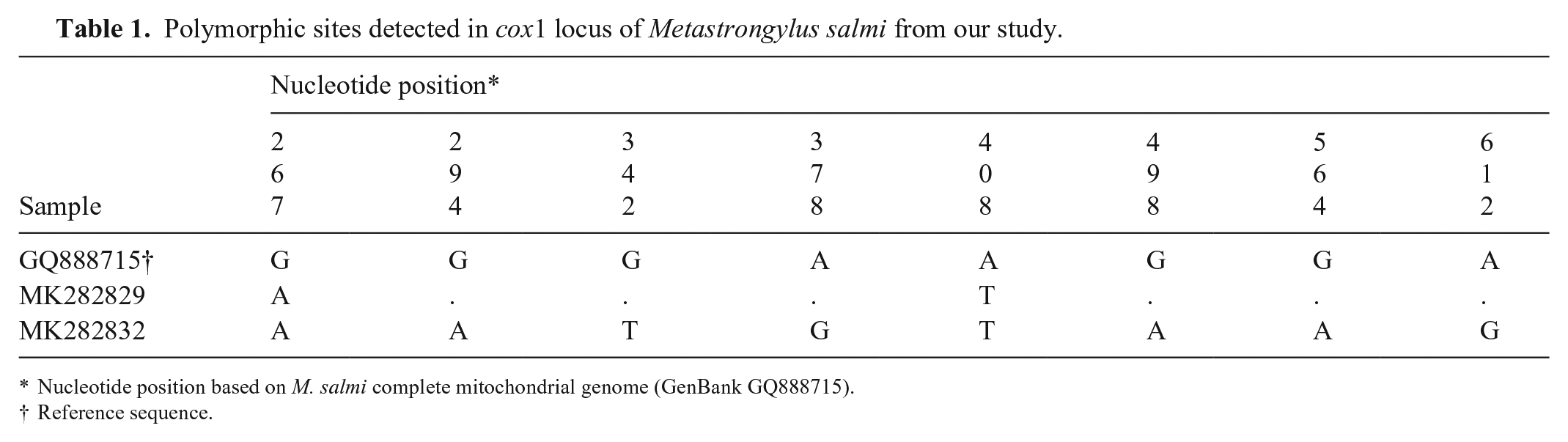

Nucleotide position based on M. salmi complete mitochondrial genome (GenBank GQ888715).
Reference sequence.
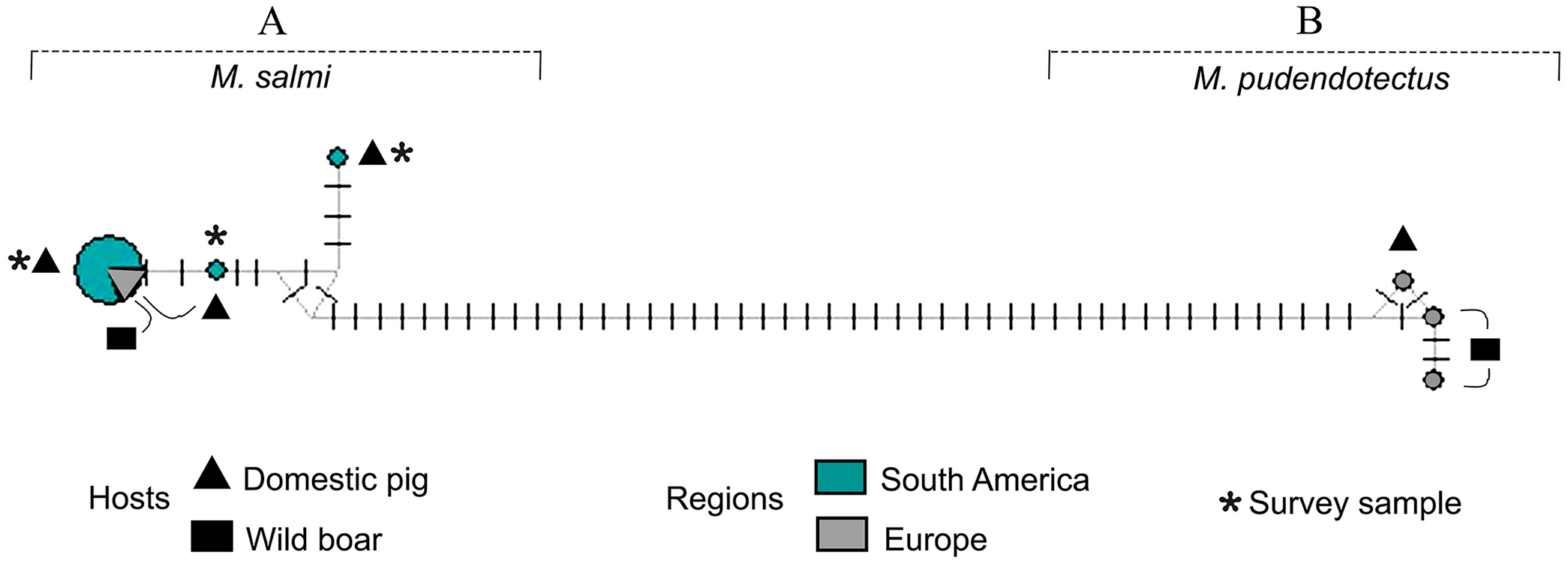
Median-joining network from 391-bp cox1 locus of Metastrongylus spp. Each haplotype is represented by a circle, and the size of each circle is proportional to haplotype frequency.
Our study demonstrated a high positivity rate for strongyliform eggs, including Metastrongylus spp., which is expected in extensively reared animals. 16 Nevertheless, only a small proportion of the eggs that we measured had dimensions compatible with Metastrongylus spp., which suggests the presence of infections by other Strongylida, such as Oesophagostomum sp. and Trichostrongylus sp. Other authors detected the following positivity rates for Metastrongylus spp. eggs: 0.9% (3 of 324), 20% (5 of 25), and 0.9%.4,9,12
Our use of the DNA barcode (cox1 locus) enabled species-specific identification of strongyle eggs in fecal samples of pigs from Piauí; only M. salmi was identified. In other studies, employing this genetic marker, the species M. pudendotectus was identified in domestic pigs in China, and M. pudendotectus and M. salmi were identified in pigs in Estonia and France.7,8,11 However, as a source of DNA, these authors used adult worms or larvae obtained by artificial digestion of tissues, all obtained from animal autopsy. Thus, we demonstrated the effectiveness of using fecal material obtained noninvasively to perform antemortem species-specific identification of these helminths.
Both the phylogenetic tree and the haplotypic network constructed with the cox1 locus revealed a clear distinction between M. salmi and M. pudendotectus. Despite the existence of 2 clusters well defined by species, clusters were not delimited by region or host (domestic and wild pigs). Accordingly, a survey in Vietnam revealed a distant genetic relationship between the group formed by the sequences of M. salmi and M. apri with M. pudendotectus using the cox1 locus and the second internal transcribed spacer (ITS2) of ribosomal DNA. 23 The cox1 locus was considered the most suitable for the species-specific identification of the genus Metastrongylus, given that the various species do not share >88% homology. 8 These authors were not successful in identifying the Metastrongylus species using the 18S locus. We described 2 new M. salmi haplotypes in our study and identified a predominant haplotype. The latter was isolated previously from wild boar in France. 8 Unfortunately, GenBank has few Metastrongylus spp. sequences available, making it difficult to visualize the genetic diversity of this parasite.
Our genetic diversity analyses showed less intraspecific variation for M. salmi compared with M. pudendotectus. Furthermore, interspecific diversity was moderate. Different genetic variability patterns for the 2 species were detected through sequence analysis of ITS2 of the ribosomal DNA and the bands obtained with the random amplified polymorphic DNA assay (RAPD).3,10
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387221092143 – Supplemental material for Molecular detection of Metastrongylus salmi eggs from pigs in low-resource communities in the state of Piauí, northeastern Brazil
Supplemental material, sj-pdf-1-vdi-10.1177_10406387221092143 for Molecular detection of Metastrongylus salmi eggs from pigs in low-resource communities in the state of Piauí, northeastern Brazil by Polyanna A. A. Bacelar, Lauren H. Jaeger, Deiviane A. Calegar, Jéssica P. Santos, Beatriz Coronato-Nunes, Elis R. C. Reis, Márcio N. Bóia, Kerla J. L. Monteiro and Filipe A. Carvalho-Costa in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank the staff of the Municipal Health Department of Nossa Senhora de Nazaré, Piauí, Brazil.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of the present article.
Funding
The authors disclosed the receipt of the following financial support for research, authorship and/or publication of this article: The present study was funded through resources from the regular funds of Instituto Oswaldo Cruz and Fiocruz Piauí.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
