Abstract
This report details 2 outbreaks of dermatophytosis in 2 different mink ranches. On the first farm, only kits were affected, while on the second farm, small numbers of adults were infected. Affected mink were otherwise clinically healthy and in good body condition. Three animals were euthanized and submitted for autopsy. Grossly, mink exhibited locally extensive to coalescing areas of crusting alopecia but no other significant gross lesions in internal organs. Microscopically, skin lesions were characterized by chronic hyperplastic dermatitis with folliculitis, furunculosis, occasional intracorneal pustules, and large numbers of intrafollicular fungal arthrospores and hyphae. The dermatophyte was cultured and identified as Trichophyton equinum based on molecular barcoding of the internal transcribed spacer region of the ribosomal DNA gene.
Dermatophytosis (or ringworm infections) in mammals are common contagious mycoses of the skin, hair, and nails caused by a variety of different keratinophilic fungi belonging to the taxonomic order Onygenales, particularly members of the teleomorphic genus Arthroderma and allied anamorphic states.2,6,8,12 Clinical findings include patchy, often circular areas of alopecia and scaling, with erythematous peripheral margins. 2 Transmission between animals generally occurs via direct contact or by fomites contaminated with hair or scales from infected animals. 2 In farmed mink, ringworm infections are relatively uncommon, and outbreaks on ranches have rarely been reported. In these few case reports, infections were due to 2 different dermatophyte species, Microsporum canis3,7 and Trichophyton mentagrophytes.1,17 Although dermatophyte infections are not fatal, cosmetic damage to pelts can result in significant economic losses to producers. 1 Our study describes 2 outbreaks of dermatophytosis in 2 different mink ranches caused by Trichophyton equinum.
At both ranches, kits and dams were reared in individual cages containing a wooden box. Bedding consisted of straw and wood shavings. Kits were handled at 1, 3, 10, and 21 days after whelping. The diet of the kits consisted primarily of milk from the mother, while the dams, older kits, and juveniles were fed a diet of pork and beef liver, frozen fish, cereal, and some chicken.
In the first ranch, ringworm infection was first detected in 7 litters in 2 different sheds (50–60 kits in total) when the kits were 21 days of age. All were dark color phase mink (black to dark gray). The dams neither exhibited signs of infection nor developed skin lesions later on. All kits from each infected litter were euthanized. According to the ranch records, this was the first time that ringworm was detected. Two affected male kits, weighing 197 and 163 g, were submitted for autopsy (Fig. 1A). Both animals exhibited similar skin lesions consisting of extensive epidermal thickening with lichenification, crusting, and alopecia (Fig. 1A). In 1 mink, cutaneous lesions primarily involved the lateral and ventral thorax while in the other involved front and hind limb, ventral abdomen, and proximal region of the tail. Both minks were in good body condition with abundant fat stores. No significant abnormalities, other than the skin lesions, were noted at autopsy. Samples of lung, heart, digestive tract, liver, kidney, spleen, brain, and skin were fixed in formalin for histopathological examination, and skin samples were submitted for bacteriological and mycological cultures.
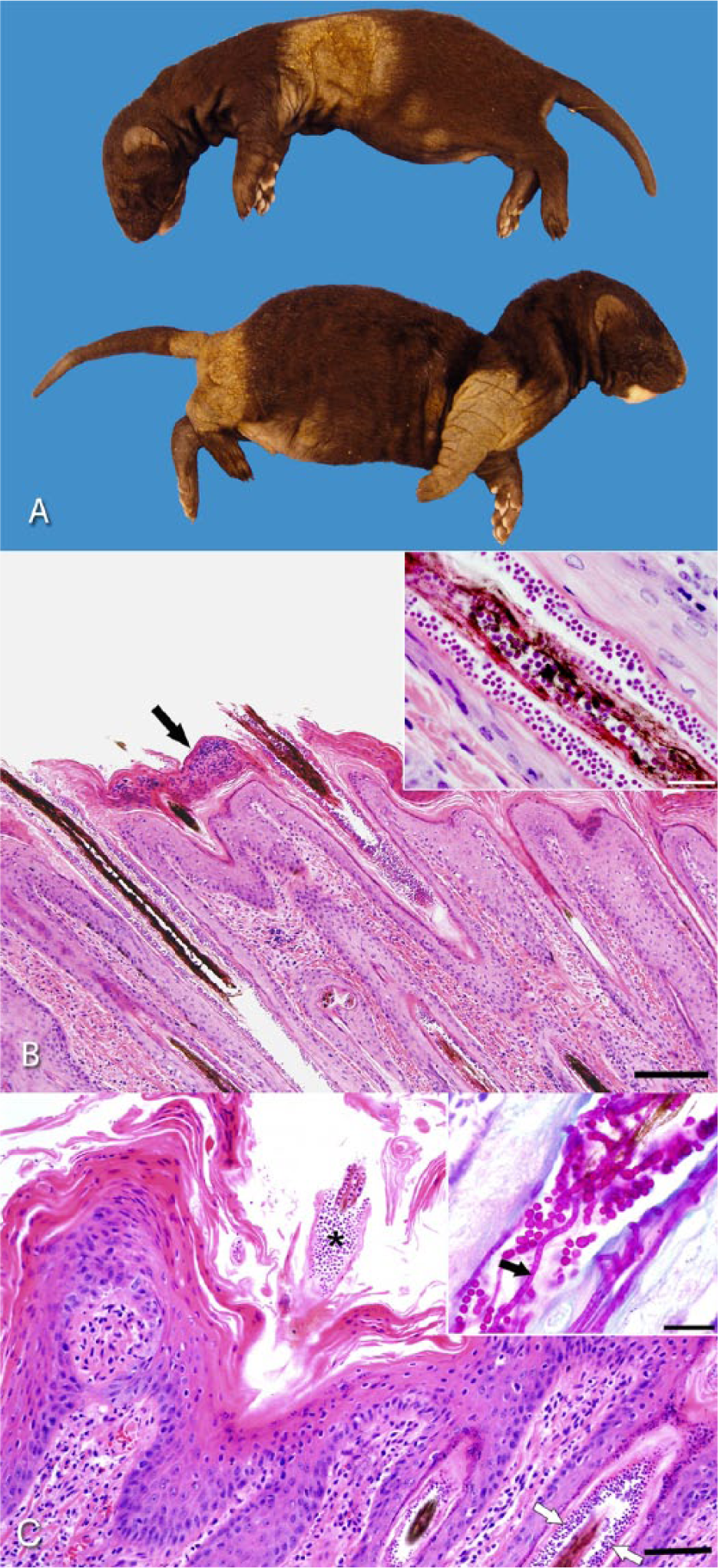
Minks.
On the second ranch, a new line of pastel mink had been purchased in the spring of 2014. One adult, described as having patchy areas of alopecia and scaling, was euthanized at that time. Four more adult females developed similar lesions in September–October of 2014. These were the only 5 animals affected of the 900 housed in the shed. As in ranch 1, there were no previous reported cases of ringworm. However, the farm from which these new mink originated had previously reported a few animals with similar skin lesions, which were presumed to be due to ringworm infection. All 4 infected mink were euthanized, and one of the animals was submitted for autopsy. At autopsy, along the length of the dorsal trunk from approximately the shoulder area extending to the lumbar area, multifocal to coalescing, serpiginous or “track-like” areas of alopecia, scaling, and mild crusting were observed. Skin scrapings from the affected area did not reveal mites. The carcass was in good body condition with abundant fat stores. The spleen was moderately enlarged. Otherwise, no significant abnormalities were noted. Samples of lung, heart, digestive tract, liver, kidney, spleen, brain, and skin were fixed in 10% buffered formalin for histopathological examination, and skin samples were submitted for bacteriological and mycological cultures.
Sections of haired skin from both kits and the adult mink showed similar microscopic lesions. The epidermis was moderately to severely, irregularly hyperplastic and covered by increased amounts of dense ortho- and parakeratotic keratin that contained occasional small intracorneal aggregates of degenerated neutrophils (Fig. 1B). The walls of the hair follicles were similarly thickened. Myriads of densely packed, round, basophilic, 1–2 μm in diameter, arthrospores encircled hair shafts (ectothrix arthrospores; Fig. 1B). Fewer, similar endothrix arthrospores and occasional pale staining endothrix hyphae were present within the center of affected hair shafts (Fig. 1C). Some hair follicles were ruptured with hair fragments surrounded by a dense infiltrate of epithelioid macrophages admixed with few neutrophils. Mild, multifocal infiltrates of macrophages, lymphocytes, plasma cells, neutrophils, and occasional eosinophils were noted in the surrounding dermis. Arthrospores and hyphae are highlighted with periodic acid–Schiff (PAS) and Grocott methenamine silver stains. With PAS staining, hyphae appeared septated, non-branching, and measured ~1.0 μm in width (Fig. 1C). Based on the observed microscopic finding, the skin lesions were diagnosed as moderate, chronic hyperplastic, and pyogranulomatous dermatitis with hyperkeratosis, folliculitis and furunculosis, and numerous intralesional fungi, consistent with dermatophytosis (ringworm). Splenic enlargement was due to moderate extramedullary hematopoiesis, which was interpreted as a subclinical incidental finding.
Routine aerobic bacterial culture of mink liver, lung, and skin tissue samples was performed using standard microbiological procedures.13,15 Samples were plated on Columbia agar with 5% sheep blood, a and MacConkey agar, b and incubated aerobically at 35°C. Cultures were examined after 24- and 48-hr incubation, and bacterial identifications were performed using matrix-assisted laser desorption ionization time-of-flight mass spectrometry and commercial identification software. c No growth was observed from the liver and lung samples of the 2 mink kits. The skin explants from both kits yielded no significant bacterial pathogens, only moderate growth of Staphylococcus sciuri, a coagulase-negative Staphylococcus species (CNS). However, a moderate growth of fungal colonies was also observed on the blood agar plate. Subcultures were made on Sabouraud dextrose agar d and phytone yeast extract agar e plates and incubated aerobically at 25°C. Culture of the adult mink similarly yielded heavy fungal growth and moderate growth of normal bacterial skin flora (CNS and Corynebacterium spp.). A fungal isolate from one of the kits (designated as X10867) and the adult mink (designated as X27229) were retained for mycological identification.
The fungal isolates were subcultured on both Sabouraud dextrose agar and potato dextrose agar f incubated at 22°C, and colony development was monitored over a 3-week period. Resulting colonies were powdery in appearance due to the production of white to cream colored spores and hyaline hyphae and produced a distinct yellow pigmentation observed from the colony reverse. Microscopic examination of the colonies revealed the production of both distinct macroconidia and microconidia (Fig. 2A–C), confirming the identification of the culture as belonging to the genus Trichophyton. Microconidia ranged from spherical to pyriform or oblong in shape, occurring along the sides of the mycelium and aggregated in dense “grape-like” clusters. Macroconidia were clavate with smooth walls and composed of 3–4 cells. Genomic DNA was extracted from the culture, and both the internal transcribed spacer region (ITS; forward ITS1: 5′-TCCGTAGGTGAACCTGCGG, reverse ITS4: 5′-TCCTCCGCTTATTGATATGC) and nuclear large subunit ribosomal DNA (nLSU rDNA; forward NL1: 5′-GCATATCAATAAGCGGAGGAAAAG, reverse NL4: 5′-GGTCCGTGTTTCAAGACGG) genes were amplified by polymerase chain reaction (PCR) 16 and sequenced at a commercial facility. A BLASTn search, performed using sequences of both the ITS1-5.8S-ITS2 gene and the D1D2 region of the nLSU gene against GenBank, returned a high sequence homology (98–100%) with various isolates of genera Trichophyton and Arthroderma (Arthroderma is a teleomorphic genus with several associated anamorphic states, including the genus Trichophyton 6 ). A data set was compiled to include representative ITS rDNA gene sequences for species of both genera and analyzed to infer the phylogenetic placement of isolates X10867 and X27229. The analysis involved 21 sequences and included 648 positions in the final dataset with an overall mean distance calculated as 0.293 with a standard error of 0.128. Isolates X10867 and X27229 formed a well-supported clade (96% bootstrap support) with the neotype strain of T. equinum and the type strain of T. tonsurans (Fig. 3). Sequence alignment using type and neotype barcoding ITS sequences of both T. tonsurans and T. equinum resulted in a 100% match (with the presence of a thymidine residue in position 18 as previously reported 16 ; Fig. 4) confirming the identity of X10867 and X27229 as T. equinum (ITS sequences were submitted to GenBank as KR232950 and KR232951, respectively). Species identification by molecular barcoding was supported by observations of gross colony morphology and micromorphology as the production of yellow pigment in culture and the spore size and shape of both macro- and microconidia fit with the species description of T. equinum. 4
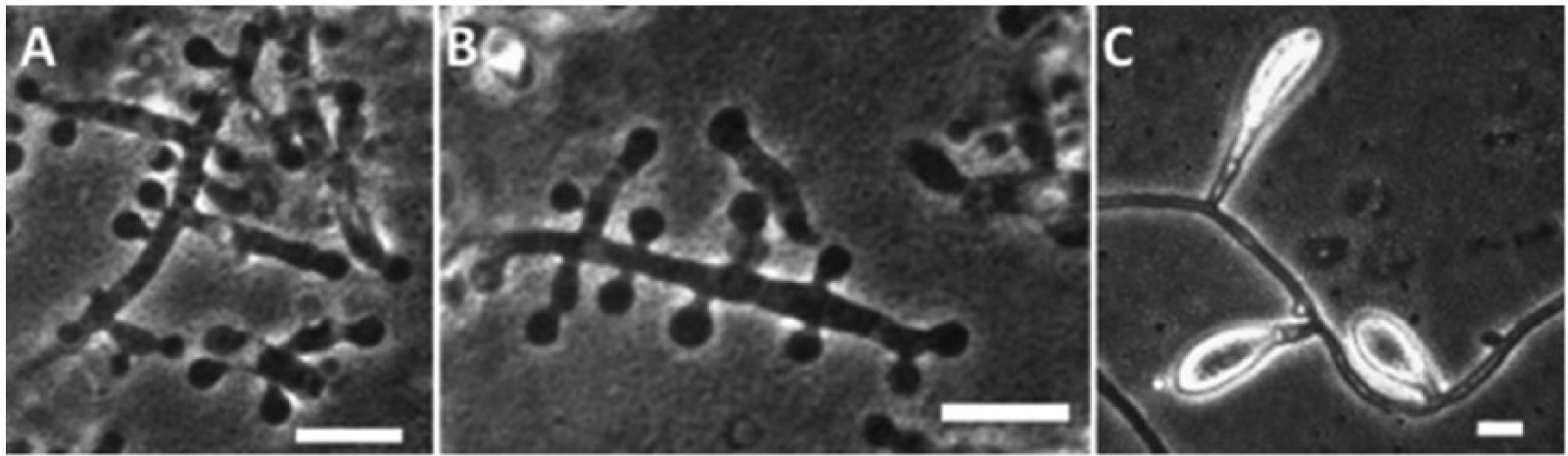
Micromorphology of Trichophyton equinum (isolate X10867) visualized by phase contrast microscopy: microconidia with lateral branching and “grape-like” clusters (
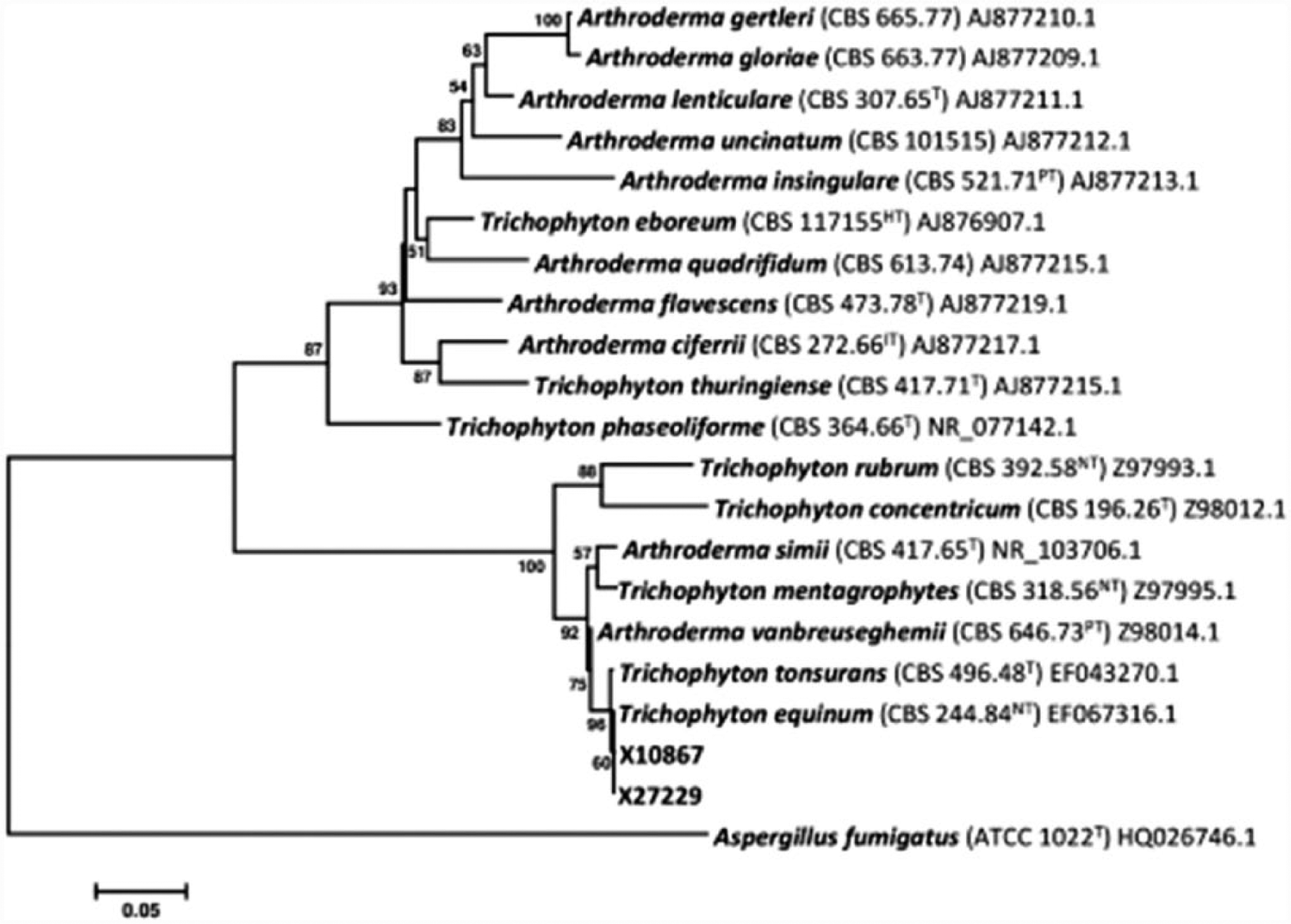

Bootstrap consensus tree inferred from 2,000 replicates using the neighbor-joining method based on Arthroderma and Trichophyton species internal transcribed spacer region (ITS) ribosomal DNA sequences (T = type; NT = neotype; PT = paratype; HT = holotype; IT = isotype). The percentage of replicate trees (>50%) in which the associated taxa clustered together in the bootstrap tests of 2,000 replicates are shown next to the branches. Evolutionary distances were computed using the maximum composite likelihood method and are in the units of the number of base substitutions per site. The tree was rooted with Aspergillus fumigatus (ATCC 1022).
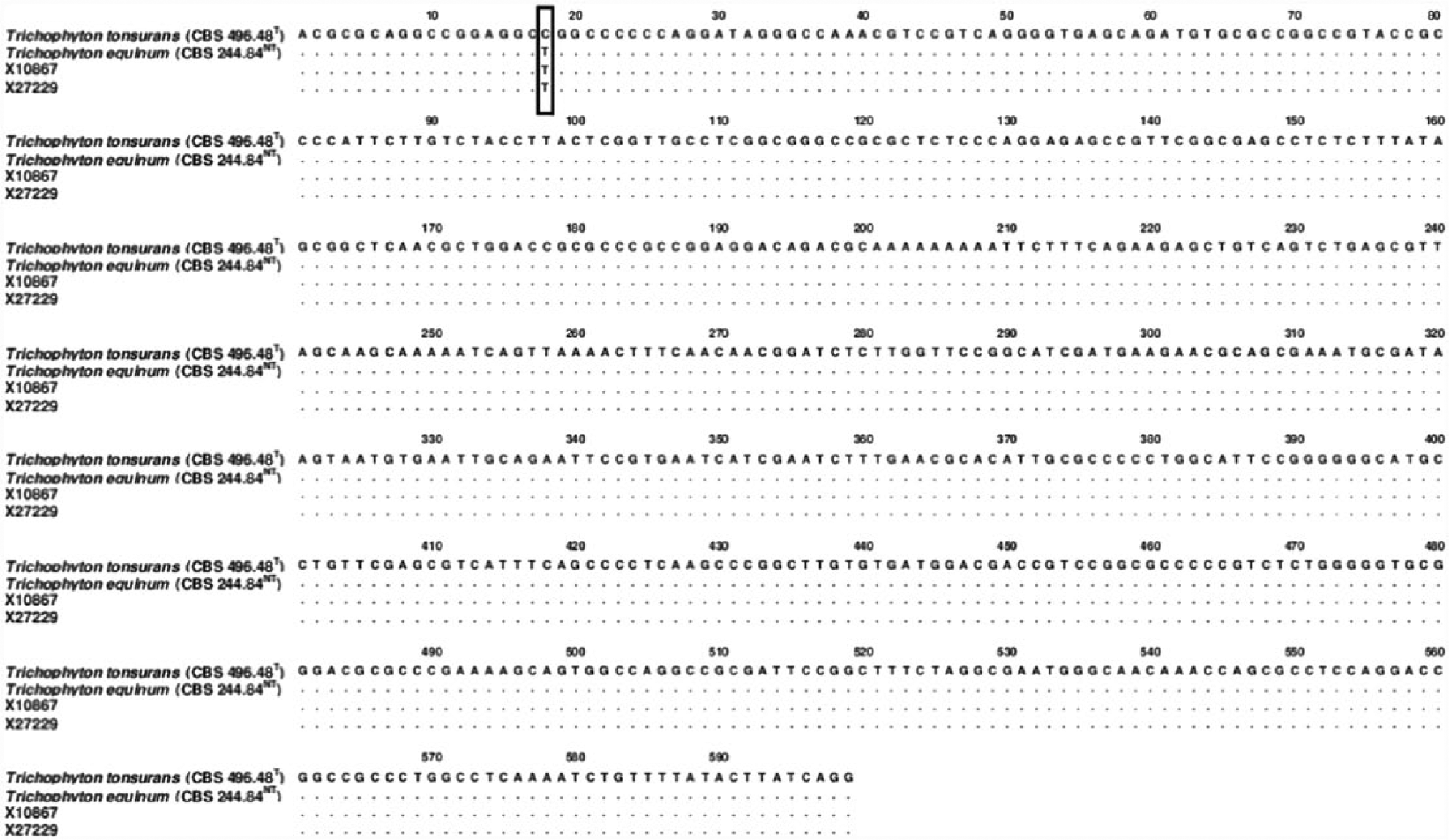

Pairwise alignment of internal transcribed spacer region (ITS) sequences of Trichophyton tonsurans type strain and T. equinum neotype strain with isolates X10867 and X27229 (T = type; NT = neotype). Single nucleotide polymorphism is highlighted with a box.
Close phylogenetic relatedness of T. equinum and T. tonsurans (the anthropophilic agent of endothrix tinea capitis) has previously been demonstrated by molecular phylogeny of the ITS rDNA gene, which at one time was an argument for synonymy of the 2 species. 5 However, similar in phylogenetic relatedness, the synonymy of T. tonsurans and T. equinum has been formally rejected based on sequence comparison using several other genes 6 confirming that T. tonsurans and T. equinum are distinct at the species level. Barcoding for species identification can make highly accurate use of species-specific single nucleotide polymorphisms to distinguish between highly similar phylogenetically related species. 14 Regarding the species T. tonsurans and T. equinum, both species can be readily distinguished from each other based on a single nucleotide polymorphism of the ITS rDNA gene barcodes, a technique that has been robustly evaluated with substantial datasets of clinical isolates of both species.10,16
Preferential host ranges for dermatophytes are based on species preferences for different proteinaceous substrates, where host range adaptations are related to the regulation of particular enzymes involved in the pathogenesis. 9 As the species name implies, T. equinum is a dermatophyte that is distinctly associated with dermatophytosis in horses, with a worldwide distribution. In horses, T. equinum produces visible skin lesions within 7–14 days of initial infection. 11 The nearly exclusive equine host range is believed to occur due to specific nutritional requirements provided by the host, as T. equinum isolates demonstrate accelerated growth on horse hair compared with hair from other mammals (including human). 4 Reports of dermatophycoses caused by T. equinum outside of the equine host range are rare and limited to dogs and cats, 2 as well as multiple cases of zoonosis of humans who have been in direct contact with infected horses.8,12
In dealing with an outbreak of dermatophytosis, environmental disinfection and separation of infected animals from noninfected animals is paramount in order to control and limit the spread of the pathogen.1,2 Systemic antifungal treatment is recommended to shorten the course of the infection and reduce dissemination to other animals. 2 Griseofulvin is the antifungal of choice for the systemic treatment of animal dermatophytosis 2 and has previously proven to be successful in the treatment of an outbreak of T. mentagrophytes in mink. 1 In the current outbreaks, infected animals were euthanized and removed from the population. The origin of the T. equinum inoculum that caused the infection in these mink is an enigma; a possible source of infection might be contaminated straw or wood shavings on which T. equinum might have been growing saprophytically or, in the case of the kits, transmitted from handlers (on day 1 or 3 prior to whelping) possibly exposed to ringworm-infected horses. A horse stable was located near the mink ranch of the first farm; however, there is a biosecurity fence between the 2 buildings, none of the horses in the stable had ringworm lesions, none of the handlers or adult mink had ringworm lesions, and the mink farm bedding was not stored at the horse stable. We cannot rule out the possibility that rodents may have served as a vector; however, T. equinum has never been reported to cause dermatophytosis of rodents. The farm that supplied the pastel mink to the second outbreak farm is the likely source of T. equinum for that case as 1 mink developed skin lesions shortly after purchase. Moreover, that farm had reportedly seen a few mink with similar skin lesions and horses are present on site. The fact that neither outbreak spread rapidly throughout the farm and, in the case of the infected kits, none of the dams developed ringworm, suggests that T. equinum infections of mink can easily be controlled by the removal of infected animals and environmental disinfection of the contaminated area.
Footnotes
a.
Columbia agar, Oxoid Co., Nepean, Ontario, Canada; with 5% defibrinated sheep blood, Alere Inc., Ottawa, Ontario, Canada.
b.
MacConkey agar, Alere Inc., Ottawa, Ontario, Canada.
c.
Microflex LT (MALDI-TOF MS) and MALDI Biotyper RTC 3.0 software, Bruker Daltonics Ltd., Milton, Ontario, Canada.
d.
BD BBL Sabouraud dextrose agar, VWR International, Toronto, Ontario, Canada.
e.
BD BBL phytone yeast extract agar, VWR International, Toronto, Ontario, Canada.
f.
BD Difco potato dextrose agar, VWR International, Toronto, Ontario, Canada.
Authors’ Contributions
D Overy substantially contributed to conception and design of the manuscript. D Overy, F Marron-Lopez, A Muckle, A Bourque, and A Lopez contributed to acquisition, analysis, and interpretation of data. L Lund contributed to analysis and interpretation of data. D MacHattie contributed to acquisition of data. D Overy, A Muckle, A Bourque, and A Lopez drafted the manuscript. All authors critically revised the manuscript, gave final approval, and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
