Abstract
We evaluated the use of MALDI-TOF MS for the identification of 3 major, dairy-associated Prototheca species, namely, Prototheca bovis (formerly P. zopfii genotype 2), P. blaschkeae, and P. ciferrii (formerly P. zopfii genotype 1). The MALDI-TOF MS spectra established for those species were introduced into the reference spectra library of the Bruker Biotyper MALDI-TOF MS analysis software. Next, 31 Prototheca isolates from Holstein cows with mastitis, from herds located in the midwestern area of São Paulo State, Brazil, were subjected to MALDI-TOF MS profiling. MALDI-TOF MS allowed identification of 22 of 27 P. bovis and 3 of 4 P. blaschkeae isolates with scores >2.0, with 5 of 27 P. bovis and 1 of 4 P. blaschkeae isolates identified only to the genus level. With our extended algae database, MALDI-TOF MS can contribute to quick and effective speciation of Prototheca from mastitis cases.
Prototheca spp., which are achlorophyllous yeast-like microalgae distributed ubiquitously in the environment, have become an emerging cause of bovine mastitis around the world, particularly in countries with intensive milk production.2,12 Mastitis is one of the most prevalent diseases of dairy cows, and it affects the dairy industry negatively worldwide. 19 Mastitis caused by Prototheca sp. is nonresponsive to intramammary or systemic treatment with conventional antimicrobial agents, forcing segregation and early culling of cows as the major measures to control protothecal mammary infections.7,8
The revised taxonomy of the Prototheca genus, based on the mitochondrial cytb gene marker, recognizes 14 Prototheca species. 10 Among the dairy-associated species, P. bovis and P. blaschkeae have been identified as the most prevalent etiologic agents of bovine mammary protothecosis; P. ciferrii has been found mainly in the dairy farm environment. 17 P. bovis has been found to be the most pathogenic species, being associated with progressive damage to the mammary gland parenchyma, resulting in apoptosis and alveolar atrophy. 18
Traditionally, identification of Prototheca spp. relies upon phenotypic criteria, such as macro- and micromorphology, and biochemical profiling. Although still in use, the phenotype-based methods do not easily differentiate Prototheca at the species level, and have been increasingly replaced by molecular methods, which provide much higher resolution and throughput and save time.3,12
Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) has been used increasingly for identification of bacterial and fungal pathogens in human and veterinary medicine at the genus- and species-level. 5 However, only a few studies have investigated the use of MALDI-TOF MS to identify Prototheca species.2,6,14 The main limitation for the use of MALDI-TOF MS for Prototheca spp. identification is the absence of entries in the database. Therefore, we aimed to create a MALDI-TOF MS database for the identification, to the species level, of bovine mastitis–causing Prototheca spp. using a published extraction protocol. 13
We included 31 Prototheca spp. isolates recovered from milk samples from Holstein cows with clinical and subclinical mastitis, from herds on 8 farms located in the midwestern area of São Paulo State, Brazil. The culture and identification procedures, the latter based on morphology, were performed according to the National Mastitis Council guidelines. 1 Briefly, a 10-μL loop of milk was streaked onto a 5% bovine blood agar plate. The plates were incubated for 48 h at 37°C in aerobic conditions. Colonies of cream-white or gray-white, pasty appearance were selected and examined microscopically for spherical-to-oval sporangia, with or without endospores.
Suspect Prototheca spp. isolates were streaked on Sabouraud agar and incubated for 24 h at 37°C, and the colonies were used for further analysis. A published ribosomal protein-extraction protocol, 13 with some modifications, was used. Fresh colonies were suspended in 300 μL of sterile molecular-grade water with subsequent addition of 900 μL of absolute ethanol to inactivate the microalgae. The solution was vortexed for 1 min and subjected to 2 cycles of centrifugation (Mikro 200r; Hettich) at 11,700 × g for 2 min. The supernatant was removed in the first cycle, then any remaining supernatant was removed after the second centrifugation. The pellets were allowed to dry at room temperature. A 50-μL aliquot of 70% formic acid was added and used to resuspend the pellet, after which 50 μL of 100% acetonitrile was added. This solution was transferred to a 2-mL tube containing 30 μg of sterile zirconium beads. The suspension was bead-beaten (Loccus; L-Beader 3 Biotechnology) in 5 cycles of 1 min at 4,000 rpm. After allowing the beads to settle, 1 µL of the supernatant was pipetted onto the steel plate (MSP 96 target polished steel; Bruker) in quintuplicate, and then overlaid with 1 μL of matrix solution of α-cyano-4-hydroxy cinnamic acid (Bruker), followed by drying at room temperature.
For database construction in the Biotyper system (Bruker), the type strains of 3 Prototheca species: P. bovis (formerly P. zopfii genotype 1, strain RZI-3), P. ciferrii (formerly P. zopfii genotype 2, strain LZ-5), and P. blaschkeae (strain RZIII-3) were used. The strains were subjected to the same extraction procedure as described above. The ribosomal protein extract from each strain was applied 30 times onto the steel target plate. Afterward, the spectra were captured (flexControl v.3.4; Bruker) and entered into the database (Biotyper 4.1.70; Bruker). The reproducibility of each spectrum was verified using cross-matching with the available information in the database and with the measured spectrum for each strain, as well as blind-coded sample preparation (we used default parameters in the Biotyper 4.1.70 software for smoothing and baseline subtraction of each spectrum).
For the calibration of MALDI-TOF MS, a standard protein solution (Bacterial test standard; Bruker) was used. Spectra were acquired (Microflex LT/SH MALDI-TOF MS; Bruker) via the manual function with flexControl v.3.4 software. The mass range of spectra used for the analysis was 2,000 to 20,000 m/z. The following parameters were configured on the device: ion source 1 set at 20.0 kV; ion source 2 set at 18.2 kV; and lens set at 6.0 kV. For spectra generation, 240 laser shots of each target sample (isolate) were captured. Consequently, the flexControl software generated the spectra through the set of ribosomal protein peaks. Then, the spectra of the 31 Prototheca spp. isolates were compared to the database created in MALDI-TOF MS Biotyper 4.1.70. Scores ≥1.7 and <2.0 were considered reliable for genus identification; scores ≥2.0 were considered reliable for genus and species identification. It is important to note that all MALDI-TOF MS identification results were performed in triplicate.
The MALDI-TOF MS identification of Prototheca isolates was confirmed by PCR. Briefly, all 31 Prototheca spp. isolates were subjected to molecular analysis as described previously. 15 Two protocols were adopted; the first was a genotype-specific PCR protocol used to identify P. bovis, and the second was an assay focused to identify P. blaschkeae. All PCR primers and adopted conditions were performed as described previously. 15 In the first PCR assay, the presence of 2 products (165 and 450 bp) identified an isolate as P. bovis; a single product (450 bp) was suggestive of a non–P. bovis isolate. These non–P. bovis isolates were subjected to the second PCR with the presence of 3 products (126, 213, 450 bp) used to identify an isolate as P. blaschkeae.
For the construction of the local library as the reference spectra, it was necessary to capture 30 spectra from each reference strain (P. blaschkeae: RZIII-3, P. bovis: LZ-5, P. ciferrii: RZI-3; Fig. 1). Spectra of 2–20 kDa were captured successfully by the flexControl software. To carry out the construction of the main spectrum profile (MSP) library, each spectrum obtained was submitted to identification using the commercial library of the Biotyper software. None of the isolates obtained the minimum score for identification (≥1.7). Additionally, there was no matching of the spectra of our library entries of Prototheca spp. compared to the spectrum of P. wickerhamii, which is the only reference present in the Biotyper 4.1.70 database.
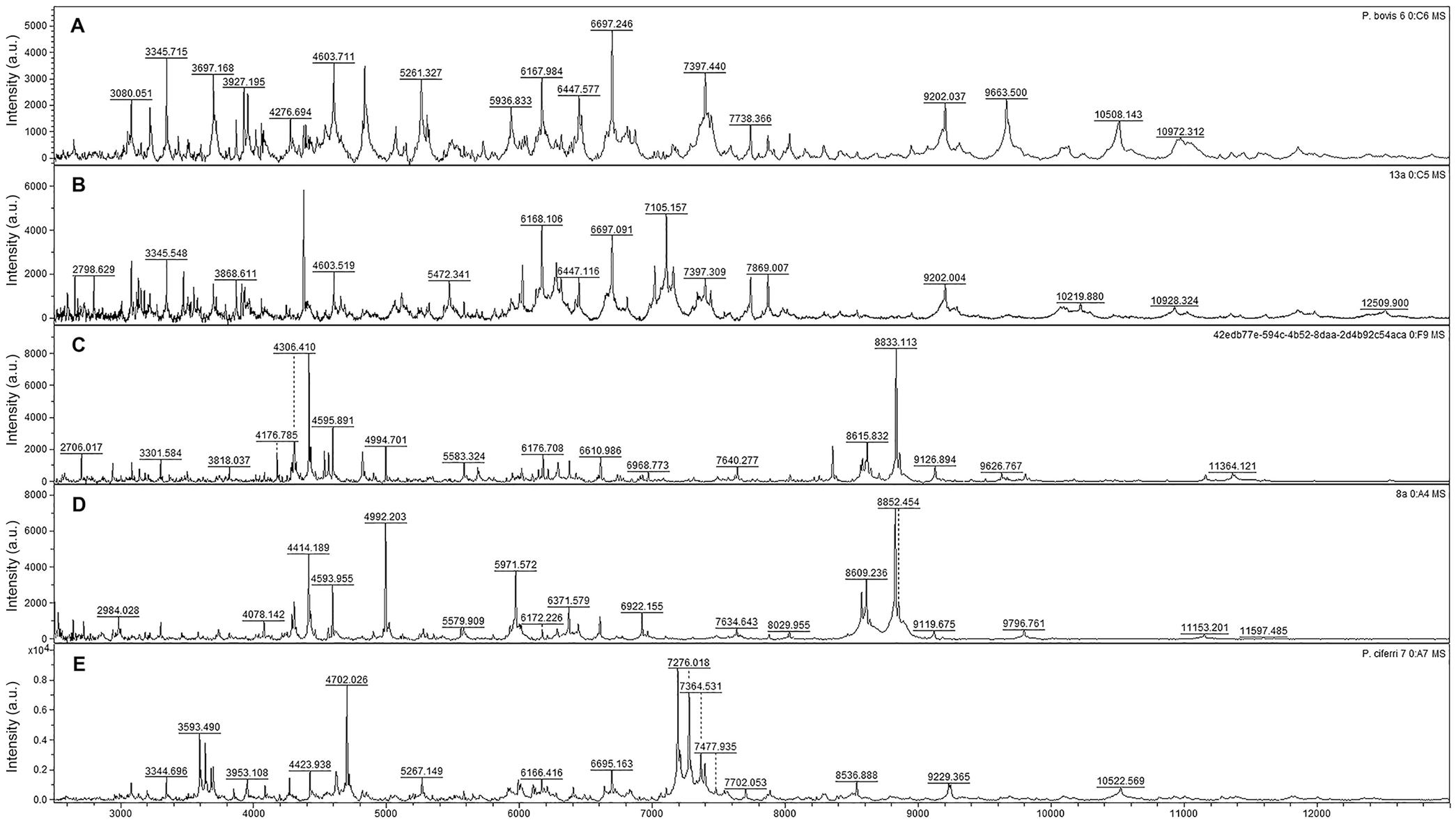
Spectra of
After creation of the database, 27 of 31 isolates submitted to identification were identified as P. bovis; 4 isolates were identified as P. blaschkeae. There were 22 of 27 isolates identified as P. bovis with identification scores ≥2.0, suggesting reliable identification at the species and genus level for P. bovis. The remaining P. bovis isolates produced scores ≥1.7 and <2.0, suggesting that genus-level identification was reliable. Three of 4 P. blaschkeae isolates had an identification score of ≥2.0; the fourth isolate had a score ≥1.7.
In the PCR assays, all Prototheca spp. isolates gave a band at 450 bp, corresponding to the specific fragment of the internal amplification control obtained by Proto 18-4f and Proto 18-4r primers. A total of 27 isolates had a species-specific amplicon of 165 bp (18-4f and PZGT 2/r), corresponding to P. bovis identification. The remaining of 4 isolates were subjected to the second PCR and had the specific amplicons associated with P. blaschkeae of 126 and 213 bp (18-4f, PZGT 3-IK/f, PZGT 3-IK/r).
For optimal analytical performance with the MALDI-TOF MS Biotyper, in-tube, formic acid–acetonitrile (FA-ACN) extraction is recommended by the manufacturer (Bruker). Using this procedure, however, we could not capture spectra of sufficient quality when testing samples obtained from Prototheca isolates, as reported in a previous study. 20 The on-plate extraction protocol, typically used for bacterial identification, 4 also failed when applied to our Prototheca isolates, producing a high level of nonspecific background spectral noise making identification challenging, if not impossible. Some authors have been successful in differentiating Prototheca spp. when using sonication as an additional step for protein extraction.2,14 Extra washing steps, either with 70% ethanol or with pure water, prior to cell disruption, have been recommended as efficient means to remove substances interfering with the spectra collected. 6 All of these modifications were applied to our procedure, yet with no success (data not shown).
With the aim of improving the performance of the extraction procedure, we used homogenization with zirconia beads in a bead-beater apparatus, following the suggestion of a previous study that used zirconia/silica beads in a solution comprising of cell pellet and FA-ACN, agitated by a vortex mixer, but for identification of human-associated species (e.g., P. wickerhamii and P. cutis). 13 There is no entry of Prototheca sp. from mastitis cases in the reference library of Biotyper software 4.1.70. Hence, we incorporated the MSPs of P. blaschkeae RZIII-3, P. bovis LZ-5, and P. ciferrii RZI-3 into the library of Biotyper 4.1.70 software for species-specific identification of Prototheca associated with mastitis. Overall, our library entries of Prototheca spp. allowed the identification of 25 isolates submitted to MALDI-TOF MS analysis, with 22 identified as P. bovis and 3 as P. blaschkeae, with scores ≥2.0. Five isolates of P. bovis and one isolate of P. blaschkeae, all confirmed by PCR, were correctly identified at the genus level by MALDI-TOF MS (score identification ≥1.7 and <2.0). No mastitis-causing isolates were identified as P. ciferrii.
Despite a small sample size, P. bovis was identified as an important causative agent of bovine mammary protothecosis in the enrolled herds, supporting findings of previous studies12,15 that reported the frequent isolation of this species from protothecal mastitis cases. The high prevalence of P. bovis can be attributed to the wide distribution of this microalga in the dairy farm environment, being isolated from diverse sources, such as water, manure, feed, and, particularly, cow bedding. 17 Thus, the risk of infection with P. bovis may stem from poor hygiene, including practices related to milking and treatment of mastitis in cows. 17
Given the small sample size, the finding of P. blaschkeae underlines the potential role of this species in the etiology of bovine mastitis. Although P. blaschkeae has been reported to cause sporadic infections in dairy cows, 15 the organism has also been the cause of an outbreak of mastitis within a herd. 16 The sources of P. blaschkeae infection remain undefined. Further, environmental studies are required given that this organism is distributed abundantly in the dairy farm environment.11,15,17 The absence of P. ciferrii in our sample set is not surprising, because it has not yet been reported in bovine mastitis cases. However, P. ciferrii does exhibit pathogenic potential, as evidenced in a retrospective study of human-derived isolates. 9
The modified extraction method that we used, along with the introduction of an algae database, allowed quick and effective speciation of Prototheca from mastitis cases by MALDI-TOF MS. MALDI-TOF MS identification may enable the adoption of early measures to control and prevent Prototheca mastitis on dairy farms, reducing the economic losses associated with these infections that typically respond poorly to antimicrobial treatment and often result in the culling of affected animals.
Footnotes
Acknowledgements
We thank the “Qualileite” team in the Milk Quality Laboratory at the School of Veterinary Medicine and Animal Science at USP.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was funded by São Paulo Research Foundation (FAPESP) project 2017/50461-5. CAPES (PNPD program) supported a postdoctoral grant for Temporary Assistant Professor Dr. Juliano Leonel Gonçalves.
