Abstract
Brucella canis was recovered from dogs that were canine brucellosis suspect by blood culture using a modified lysis method. Organism identity was established by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS). The instrument-provided security library identified the isolates as Brucella species. The isolates were further identified as B. canis with the help of phenotypic and genotypic characteristics. The mass spectral profiles from characterized B. canis isolates, when added to the MALDI-TOF MS standard reference library, allowed successful presumptive identification of B. canis.
Brucella canis is a gram-negative organism responsible for reproductive problems in dogs, such as abortions, stillbirths, epididymitis, orchitis, and sperm abnormalities. 3 Occasionally, the organism can cause human infections and thus is considered a zoonotic pathogen.4,9,17 Diagnosing brucellosis in dogs is problematic because currently available serologic tests are not very sensitive or specific.3,8,20 Culture is considered necessary for a final diagnosis. 1 However, infected dogs often may not be bacteremic because, after infection, the organisms tend to localize in the genital tract and associated lymph nodes. 3 Hence, a single negative blood culture may not be sufficient to rule out the disease in suspect animals. A series of 3 blood samples, collected at least 24 h apart, must be culture negative to ensure that a dog is not infected.3,15
Blood cultures for Brucella are performed by a variety of methods, including the conventional Ruiz–Castaneda method, automated blood culture systems, lysis centrifugation (LC), and clot culture. Among these methods, the LC procedure has been shown to have greater sensitivity and faster recovery. 5 For bacterial identifications, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) has increasingly become the method of choice.10,14,16 MALDI-TOF MS has been successfully used for rapid identification of Brucella, mostly at the genus level 6 and, in a previous study, even at the species level when genotypic characterization was combined with MALDI-TOF. 11 The conventional methods used to identify Brucella isolates to the species level are complex, labor-intensive, and time consuming.2,3 Even with genotypic methods, establishing the identity of Brucella isolates at species and biovar level has not always been completely successful.7,11 We report here the modification of the original LC isolation procedure for culturing of canine blood samples2,5 and the presumptive species-level identification of B. canis using MALDI-TOF MS.
Blood samples from dogs with possible canine brucellosis, including a known exposure history to B. canis–infected animals, were tested. Briefly, 1 mL of citrated blood was transferred to a tube containing 5 mL of brain–heart infusion (BHI) broth and gently mixed. The inoculated BHI broth was then frozen at −20°C overnight to facilitate cell lysis, followed by incubation at 37°C for 18–24 h for enrichment. After incubation, the enriched culture was plated on 2 blood agar plates (BAPs); one plate was incubated at 37°C with 7.5% CO2 and another plate at 37°C without CO2. The plates were observed daily for any growth, and bacterial colonies isolated were subjected to MALDI-TOF MS (MALDI Biotyper, Bruker, Billerica, MA) analysis.
B. canis poses a potential health risk to laboratory workers, if not handled appropriately.4,9,17 A rapid sample preparation method for Brucella isolates compatible with MALDI-TOF MS analysis has been described that involves incubation of Brucella isolates in ethanol for 90 min. 11 However, it has also been reported that incubation of Brucella in ethanol for 10 min did not inactivate Brucella consistently. 12 Therefore, we chose to verify inactivation of B. canis isolates after incubation in ethanol for 10 min and 90 min. For the extraction procedure recommended by the manufacturer (Bruker), inside a biosafety cabinet, a single bacterial colony was transferred from the plate to a 1.5-mL microfuge tube (Eppendorf, Hauppauge, NY) containing 300 μL of molecular biology–grade water. The tube was vortexed, and then 900 μL of ethanol (Sigma-Aldrich, St. Louis, MO) was added to the tube. The bacterial suspension was either centrifuged with no further incubation or was incubated for either 10 min or 90 min in ethanol. The tube was centrifuged at 14,500 × g for 2 min and the supernatant discarded. Tubes were centrifuged for another 2 min and allowed to dry. Subsequently, 10 µL of formic acid (Sigma-Aldrich; 70% v/v) was added to the pellet and mixed, followed with the addition of 10 µL of acetonitrile. The suspension was then centrifuged at 14,500 × g for 2 min. A 1-μL volume of the supernatant was spotted on the target plate and air-dried. Each sample was overlaid with 1 µL of alpha-cyano-4-hydroxycinnamic acid (Bruker) in organic solvent (Sigma-Aldrich; 50% acetonitrile and 2.5% trifluoroacetic acid) and air-dried.
To verify inactivation of the B. canis isolates, after the ethanol incubation step, aliquots from 3 isolates in triplicate were streaked on the BAP immediately after the addition of ethanol, or at 10 min, and at 90 min. No growth was recorded on the BAP after 5 d for both 37°C CO2 and non-CO2 incubations at 10 min and 90 min after the addition of ethanol, but inactivation was inconsistent when there was no incubation (inactivated 8 of 9 times). The findings were contrary to a previous report where 10-min ethanol incubation was found to be inconsistent in inactivating Brucella. 12 Even though minor differences in mass spectral profiles (MSPs) were observed with incubations in ethanol for 90 min, there was no impact on identification scores for Brucella identification (Fig. 1). In the interest of safety and based on a previous report, 11 the 90-min ethanol incubation step was preferred. This safety step does not impact identification and was added because laboratory-acquired infections with Brucella have been reported.4,9,17
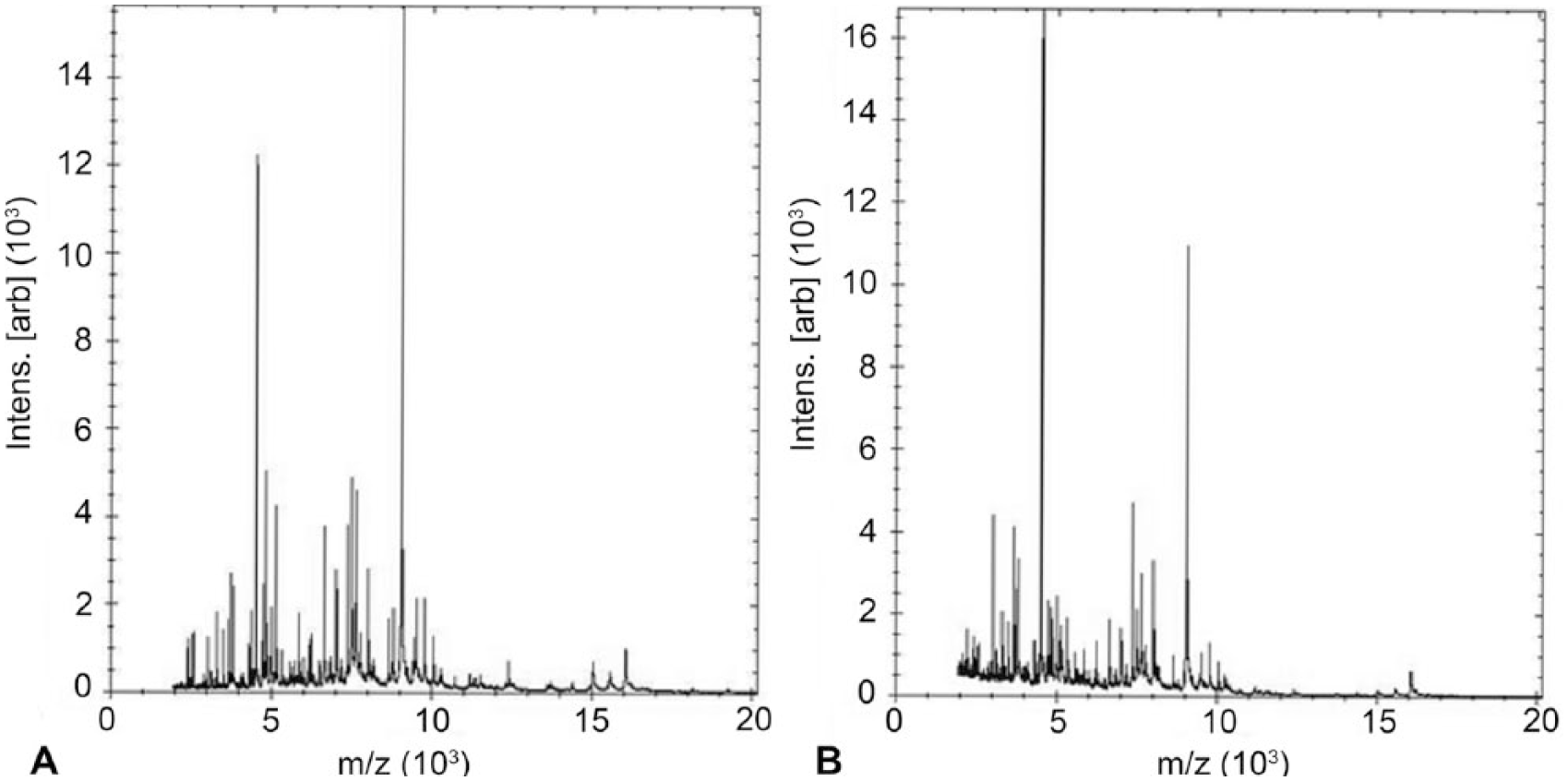
Mass spectral profile of Brucella canis isolate extracted after mixing with ethanol (
When the isolates were subjected to 90 min of formic acid and ethanol extraction followed by analysis with the mass spectrometer (Bruker) using an on-board library (Security-Relevant [SR] library v. 3.3.1.2, Bruker), isolates showed a secure genus-level identity as Brucella, with the highest scores matching the 6 B. melitensis isolates (6767, 6273, 5520, 6073, 5659, and 6074 in the library; score range: 2.21–2.54). This resulted in probable-to-highly probable species-level identification as per the interpretive criteria provided by the manufacturer.
For in-house B. canis library creation, 10 independent spectra were recorded for the 5 well-characterized field isolates,1,7 following the manufacturer’s recommendations (custom MSP and library creation protocol, revision 1, 9/2015, Bruker). Follow-up with 5 suspect B. canis colonies showed the highest MSP match to B. canis in the in-house B. canis library, with a score of 2.61–2.77, and thus highly probable species-level identification. Upon identification of isolates as Brucella species or upon presumptive identification as B. canis with the in-house B. canis library, further confirmation of the isolates was achieved with phenotypic characteristics, growth requirements, and the B. canis species-specific outer membrane protein (Omp)25 gene PCR as described previously. 7 Since 2012, using this culturing and identification procedure but with no ethanol incubation during extraction, 2,979 canine blood samples have been tested, with 2,684 found to be negative, 257 contaminated, and 38 found to be positive and confirmed as B. canis, within 4–5 d. It is only recently that we have considered the addition of the ethanol incubation to our testing protocol for safety considerations.
The current phylogenetic bacterial classification system recognizes B. melitensis as the monospecific type species and other Brucella species and biovars as heterotypic synonyms. However, the current classification also accepts nomenclature based on host range, pathogenicity differences, and biochemical properties. 21 Thus, the well-recognized Brucella species, including B. ovis, B. abortus, B. canis, B. neotomae, and B. suis, are considered nomenspecies.13,18,19
Using the nomenspecies taxonomic criteria, the preferred taxonomic classification for Brucellaceae, 13 a rapid, presumptive identification of B. canis was achieved with MALDI-TOF MS using the in-house reference library. Additional confirmation of isolates as B. canis can be obtained with the help of biochemical reactions 1 and a PCR test. 7
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
