Abstract
The Brachyspira species traditionally associated with swine dysentery and other diarrheal diseases in pigs are Brachyspira hyodysenteriae, Brachyspira pilosicoli, and, to a lesser extent, Brachyspira murdochii. “Brachyspira hampsonii” is a recently proposed novel species that causes clinical disease similar to that caused by B. hyodysenteriae. Matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) systems are increasingly available in veterinary diagnostic laboratories, are less expensive, and are faster than traditional microbiological and molecular methods for identification. Thirty-three isolates associated with Brachyspira species of importance to swine were added to an existing MALDI-TOF MS database library. In total, species included in the library were: B. hyodysenteriae, “B. hampsonii” clades I and II, Brachyspira innocens, Brachyspira intermedia, B. murdochii, and B. pilosicoli. A comparison between MALDI-TOF MS and nox sequencing was completed on 176 field isolates. Of the 176 field isolates, 174 (98.9%) matched species identification by both methods. Thirty field isolates were identified by both methods as “B. hampsonii”. Twenty-seven of the 30 (90%) “B. hampsonii” field isolates matched clade designation in both assays. The nox sequencing identified 26 as “B. hampsonii” clade I and 4 as clade II. Comparatively, MALDI-TOF MS identified 25 of the 30 as “B. hampsonii” clade I and 5 as clade II. The current study indicates MALDI-TOF MS is a reliable tool for the identification of swine Brachyspira species; however, final clade designation for “B. hampsonii” may still require molecular techniques.
Brachyspira spp. commonly recovered from swine feces include the agents of swine dysentery and other diarrheal diseases and species traditionally associated with disease in pigs. Such species include Brachyspira hyodysenteriae, Brachyspira pilosicoli, and, to a lesser extent, Brachyspira murdochii. 11 “Brachyspira hampsonii” is a recently proposed novel species associated with mucohemorrhagic diarrhea in swine, 6 a disease indistinguishable from swine dysentery, which has been experimentally reproduced following infection with representative strains of this pathogen.3,15
Brachyspira are spiral-shaped, Gram-negative, obligate anaerobic bacteria. 9 Common methods used to identify Brachyspira spp. include phenotypic and biochemical analysis, 7 targeted polymerase chain reaction (PCR) assays, and nox sequencing.1,14 Previously, it was discovered that species identification via 16S ribosomal DNA (rDNA) gene sequencing was not as useful as nox sequencing because of the high homology of the 16S rDNA gene sequence between species. 14 However, nox sequencing revealed higher sequence diversity between species and thus greater species discrimination. 14
Matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) bacterial identification systems are increasingly available and being used in veterinary diagnostic laboratories. Some advantages of these systems include their ability to identify bacterial isolates quickly, accurately, and inexpensively.2,16 Another advantage is the ability to add mass spectral profiles (MSP) to a user-defined library increasing the available profiles for comparison and identification. The database library provided with one of the MALDI-TOF MS a systems used in the current study includes only 1 strain of B. murdochii and 2 strains of B. pilosicoli. Previous work has been done to create user-defined libraries which include Brachyspira species5,13; however, profiles of “B. hampsonii” have been omitted up to this point. The present report describes the creation of a user-defined MSP library that includes the major Brachyspira species commonly isolated from swine, including “B. hampsonii”, and the use of this library to rapidly identify field isolates of Brachyspira from swine samples.
The 33 Brachyspira strains to be included in the user-defined library were selected based on phenotypic cultural characteristics followed by species identification using a combination of nox, 16S rDNA, and/or 23S rDNA sequencing. The strains are part of a culture collection maintained at the Iowa State University Veterinary Diagnostic Laboratory (ISU VDL; Ames, Iowa). The collection was started in 1974 and includes historical strains and more recent isolates from cases of swine diarrhea received for diagnostic microbiology. All isolates were cloned 3 times by single colony picks before inclusion in the culture collection. In addition, the isolates were characterized by colony characteristics and molecular methods. At least 5 strains from each Brachyspira species were chosen except for B. pilosicoli and B. intermedia where 4 and 2 strains were used, respectively. The strains are detailed in Table 1. Brachyspira cultures were performed at 42°C in anaerobic jars b utilizing gas generators. c Surface growth from each strain was harvested from trypticase soy agar (TSA) containing 10% bovine blood after 2–6 days of anaerobic incubation into 0.85% physiological saline. One milliliter of the bacterial suspension was transferred to a centrifuge tube and centrifuged (17,968 × g, 2 min) to form a bacterial pellet. The pellet was resuspended in 300 µl of sterile water. Subsequently, 900 µl of 100% ethanol was added to the suspension and mixed thoroughly. The sample was then centrifuged (17,968 × g, 2 min), and the ethanol solution was decanted. The remaining ethanol was allowed to evaporate from the centrifuge tube. The pellet was then resuspended in 50 µl of 70% formic acid. Next, 50 µl of 100% acetonitrile was added to the solution and mixed completely. The suspension was centrifuged a final time (17,968 × g, 2 min). One microliter of the supernatant was then spotted onto the 96-well plate 12 times. Each time a database addition was performed, the MALDI-TOF MS was calibrated to a bacterial test standard according to the manufacturer’s protocol. Thirty-six raw mass spectra were collected from each strain with at least 20 of these spectra compiled according to the manufacturer’s protocol to create a MSP that became available for use in a new user-defined Brachyspira library.
Brachyspira strains used for the creation of the user-defined Brachyspira library and methods by which each strain was characterized.*
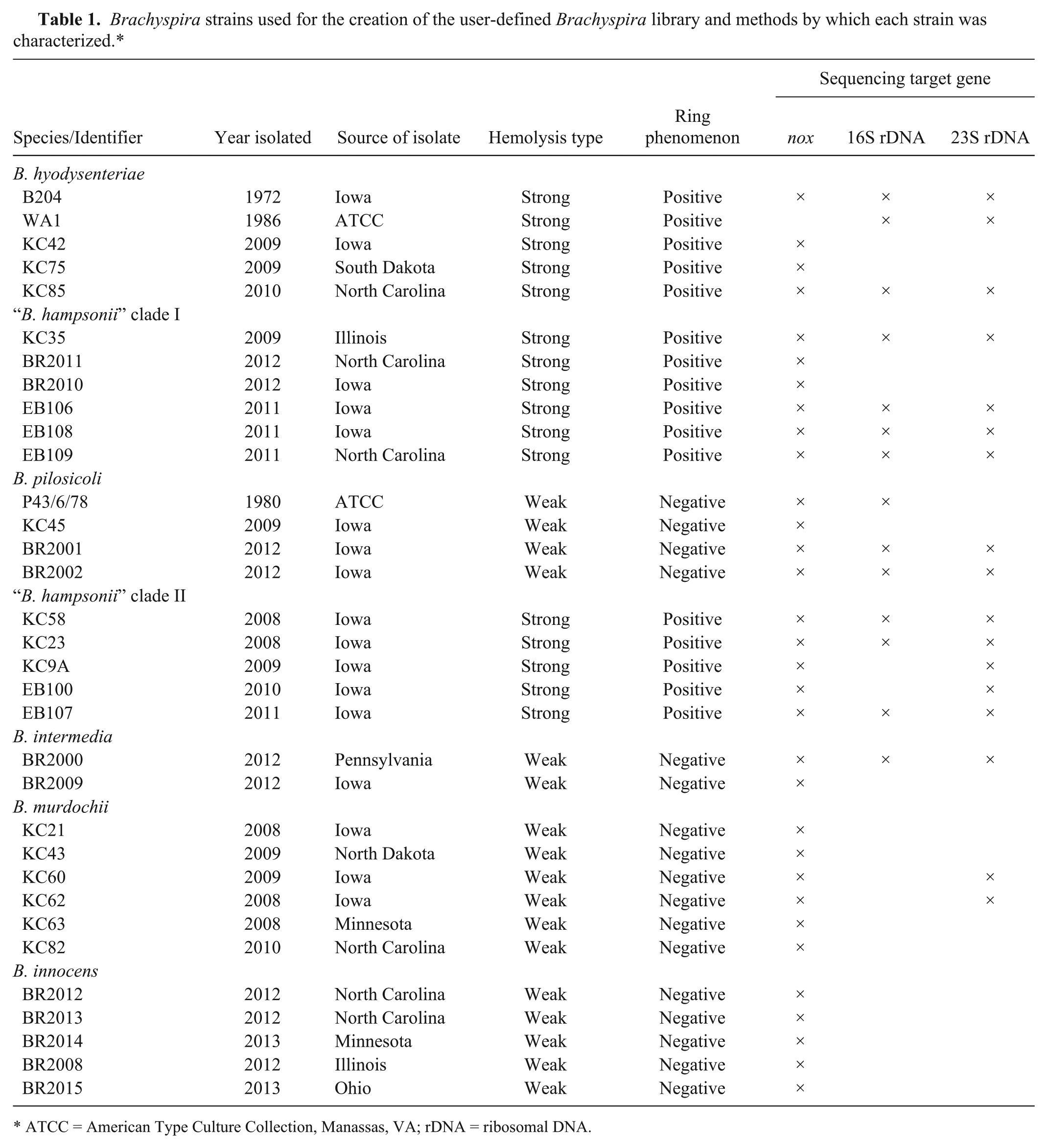
ATCC = American Type Culture Collection, Manassas, VA; rDNA = ribosomal DNA.
After creation of the Brachyspira library, all field isolates obtained at the ISU VDL between October 1, 2012 and December 31, 2013 (176 isolates) from swine samples and representing various Brachyspira species were compared using MALDI-TOF MS and nox sequencing. Isolates from this time frame were all from the United States with the exception of 1 isolate from Canada. Field isolates were cultured anaerobically on TSA plates containing 5% bovine blood, and colistin, vancomycin, and spectinomycin (CVS 10 ) and TSA plates containing 5% bovine blood, and colistin, vancomycin, spectinomycin, spiramycin, and rifampicin (BJ 12 ). Brachyspira growth was subcultured from either CVS or BJ by at least 1 single colony pick for each field isolate. After 2, 4, or 6 days of incubation, cultures of Brachyspira spp. were spotted in duplicate onto a steel plate and allowed to dry. The CVS or BJ agar plates were used to spot onto the steel plate based on the amount of surface growth. One microliter of matrix solution was applied to each spot and allowed to dry. Next, the plate was processed according to the MALDI-TOF MS manufacturer’s protocol for bacterial identification.
The generated spectra from these isolates were then compared against the combined manufacturer’s bacterial library and the Brachyspira library. The resulting output is a score-based bacterial identification ranging from 0 to 3.0. The manufacturer’s guidelines suggest that scores ≥2.0 are considered reliable identifications at the species level and scores ≥1.7 are considered reliable at the genus level. For this evaluation, an isolate with an identification score ≥1.7 was considered acceptable at the species level and was used for nox sequencing comparison. The cutoff value of ≥1.7 was used to test the robustness of the new user-defined Brachyspira library. Multiple attempts of MALDI-TOF MS were performed on some isolates to obtain a score of ≥1.7. Two weakly beta-hemolytic isolates were further characterized by a species-specific PCR that targets portions of the nox gene to detect B. pilosicoli, B. murdochii, B. intermedia, and B. innocens as previously described. 17 Additional molecular techniques were not performed on any other isolates. Sanger sequencing of the nox gene was performed using previously published forward and reverse primers 14 and a DNA analyzer. d Primers amplify an approximate 939 base pair fragment of the central portion of the nox gene. Sequencing of the 16S rDNA was performed using the following forward and reverse primers: F: 5′-TGGAGAGTTTGATCCTGGCT CAG-3′; R: 5′-TACCGCGGCTGCTGGCAC-3′. The primers amplify an approximate 500 nucleotide region at the 5’-end of the 16S rDNA gene. 8 Sequencing of the 23S rDNA was performed as previously described. 4 Used together, all 3 sets of primers were used to amplify and sequence approximately 2,400 nucleotides of the 23S rDNA. Consensus sequences were created and analyzed using a commercial software program. e Sequence identification was determined by alignment comparison with nox sequences available in GenBank from previously identified Brachyspira species. Consensus sequences were also subjected to BLAST analysis (http://blast.ncbi.nlm.nih.gov/Blast.cgi) to compare sequence similarity and to determine percent similarity to the entirety of the GenBank sequence database.
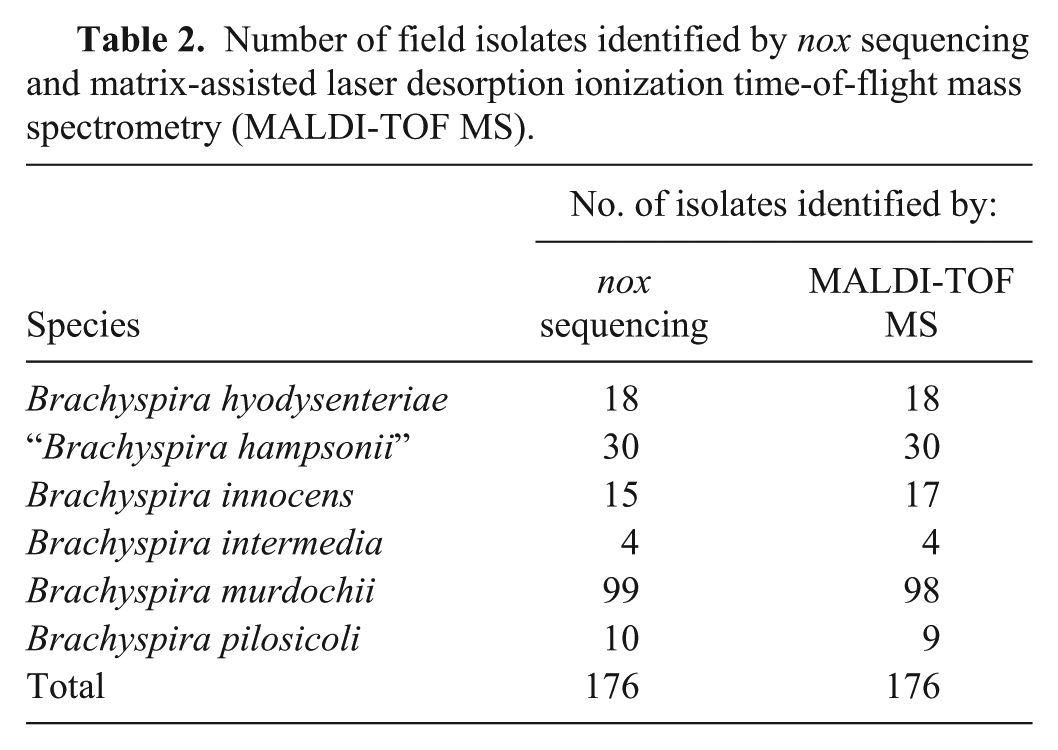
The results of identification using MALDI-TOF MS and nox sequencing performed on 176 Brachyspira field isolates are summarized in Table 2. All 176 field isolates had detectable peaks with scores ≥1.7. Of the 176 field isolates compared, 174 (98.9%) MALDI-TOF MS identification results matched speciation results obtained with nox sequencing. Of the 2 non-matched results, MALDI-TOF MS identified both as B. innocens whereas nox sequencing identified one as B. pilosicoli and the other as B. murdochii. These 2 weakly beta-hemolytic isolates were further characterized by PCR. The results of this additional work indicated that 1 isolate was likely B. pilosicoli, which matched the nox sequencing results and the other isolate was likely B. innocens, which matched the MALDI-TOF MS results. Of the 176 field isolates in the current study, 30 were identified as “B. hampsonii” by MALDI-TOF MS and nox sequencing. Twenty-six field isolates were identified by nox sequencing as “B. hampsonii” clade I, and 4 field isolates were identified by nox sequencing as clade II. Comparatively, 25 field isolates were identified by MALDI-TOF MS as “B. hampsonii” clade I, and 5 field isolates were identified by MALDI-TOF MS as clade II. Overall, of the 30 “B. hampsonii” isolates compared, 27 (90%) matched clade designation in both assays. One isolate was identified as both clades by MALDI-TOF MS and, in this case, the highest score was used for the final clade identification. A phylogenetic analysis was performed on the nox sequence of the “B. hampsonii” field isolates (Fig. 1). Two different groups were formed representing the 2 clades of “B. hampsonii.” In addition to nox sequences, MALDI-TOF MS identities and scores were included in the phylogenetic dendrogram.
Number of field isolates identified by nox sequencing and matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS).
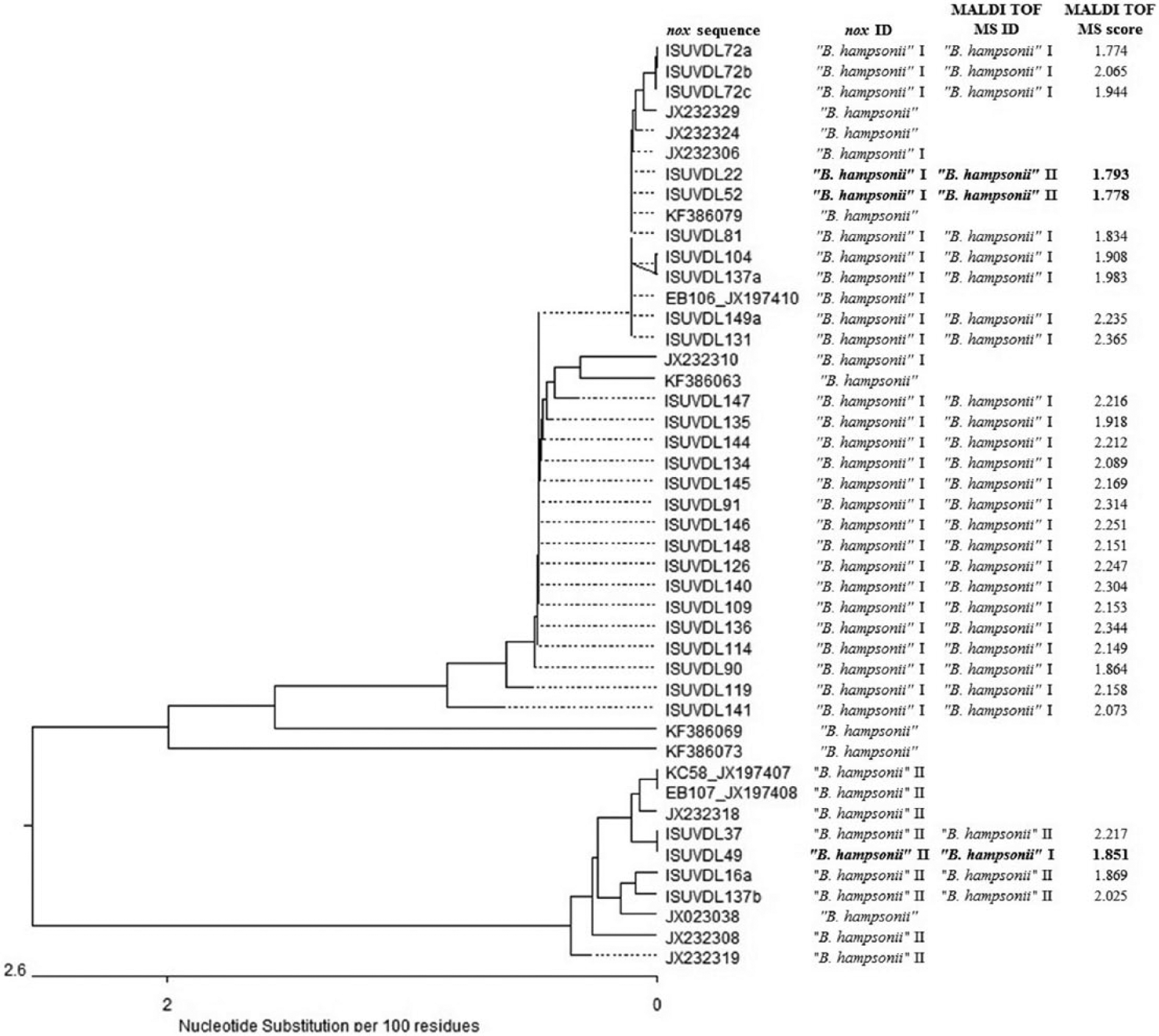


Phylogenetic analysis of the 30 “Brachyspira hampsonii” field isolates. Sequences starting with ISU VDL (Iowa State University Veterinary Diagnostic Laboratory, Ames, Iowa) are the field isolates compared, and other sequences were obtained from GenBank and are identified with accession numbers. The additional columns are the nox sequence ID, matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) ID, and MALDI-TOF MS score. Bolded rows indicate differing clade designation by the 2 methods for the same isolate.
In conclusion, after creation of a Brachyspira library, MALDI-TOF MS identification compared favorably to results obtained by nox sequencing methods. A total of 98.9% of field isolates could be identified using MALDI-TOF MS with the same results as a more complex and expensive nox sequencing method, which has been considered the most reliable method to speciate swine Brachyspira. 14 The results support work presented previously.5,13 In addition, the current study shows that MALDI-TOF MS is capable of differentiating between clades of “B. hampsonii” with a high degree of accuracy. Ninety percent (27/30) of the isolates matched clade designation by nox sequencing indicating that MALDI-TOF MS is a useful screening tool for “B. hampsonii” clade designation. The few anomalous results may be due to the potential for swine feces to contain multiple Brachyspira species. Resolution is possible through extensive and complicated cloning methods and may be warranted in certain clinical presentations. Therefore, the data from the current study shows that MALDI-TOF MS is a fast, cost-effective, reliable, and complete tool for the identification of swine Brachyspira.
Footnotes
a.
MALDI biotyper, Bruker Daltonics, Bremen, Germany.
b.
AnaeroPack system, Mitsubishi Gas Chemical Co. Inc., Japan.
c.
GasPak EZ anaerobe container system, BD Diagnostic Systems, Sparks, MD.
d.
Applied Biosystems 3730xl DNA analyzer, Life Technologies, Carlsbad, CA.
e.
Lasergene, DNASTAR Inc., Madison, WI.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, or publication of this article.
